Abstract
Plants are continuously exposed to a wide range of pathogens, including parasitic nematodes. Being sessile, plant have developed elaborate defense mechanisms that allow them to recognize potential invading pathogens and to initiate successful defenses. Plant parasitic nematodes are ubiquitous, tiny and microscopic animals, which can attack nearly all parts of the plant and globally cause crop loss of $100 billion annually. The co-evolution of plants and plant-parasitic nematodes with respect to resistance and parasitism has resulted in remarkable adaptations of the host and parasite life cycles. We review here the recent knowledge acquired on the molecular players that help in successful parasitism within the host plant roots. We then also discuss the molecular players and mechanisms underlying plant resistance against these parasitic nematodes. Understanding the molecular basis of plant-nematode interactions will be a way forward to design environmental friendly control strategies to target harmful pathogen.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nematodes are the members of phylum Nematoda represented by more than 25,000 species of roundworms. They constitute one of the most diverse groups of animals distributed in all kinds of habitats and globally they cause crop loss of $100 billion annually (Trudgill and Block 2001). Most of the nematode species are free-living that feed on microorganisms, but others are parasites of insects, animals or plants. Plant parasitic nematodes (PPNs) are tiny and microscopic animals, which can attack nearly all parts of the plant. Majority of PPNs are root parasites, either migratory or sedentary and/or they can be either ectoparasities or endoparasities. For migratory ectoparasitic nematodes, the interactions with plant are limited as they derive nutrients from epidermal cells of plant. For sedentary endoparasitic nematodes the interactions with plant are complex as they modify a few vascular cells to form a specialized feeding site.
PPNs are well-equipped for successful parasitism. They have evolved special structures for efficient plant penetration, root cell modification and withdrawal of nutrients for their growth, development and reproduction. Adaptations for parasitism includes: (1) stylet- a hollow, needle-like structure present at the anterior end. It is used to puncture host cell membrane to invade and to inject effector molecules into the host cells. During sedentary stages, stylet also helps in withdrawing nutrients from feeding sites. (2) Esophageal glands- three esophageal glands, two sub-ventral glands and one dorsal gland secrete effector molecules, which are crucial for penetration, migration, establishment and maintenance of feeding sites. (3) Chemosensory organ- two amphids, which help nematode to migrate towards the host root.
Two of the most damaging groups of sedentary nematodes are cyst nematodes (CNs) and root-knot nematodes (RKNs). CNs have narrow host range whereas RKNs are polyphagous, having a broad host range. Among CNs, majority of crop losses are inflicted by Heterodera and Globodera species. There are four most ubiquitous and polyphagous RKN spp. viz, Meloidogyne incognita, M. javanica and M. arenaria (apomictic) and M.hapla (automictic) which account for most of the damage to agricultural crops. Both CNs and RKNs spend most part of their life-cycle inside the roots of higher plants and complete their life-cycle within 4-6 weeks (Vanholme et al. 2004). We focus mainly on data from these two nematodes to briefly describe the molecular aspects of plant-nematode interactions in the following sections.
Parasitic life cycle of sedentary endoparasitic nematodes
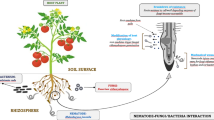
The infective exophytic second stage juveniles (J2) hatch from the eggs (J1) in the soil. Amphids help J2 s to reach their appropriate host by sensing the chemo-attractants present in root exudates (Grundler et al. 1991; Teillet et al. 2013). The main task of J2 is to reach the host tissue before their lipid reserves are depleted. For penetration, J2 use mechanical force of stylet to pierce the cell wall and secrete a variety of cell wall degrading enzymes that loosen the cell wall structure throughout penetration and migration phases. They migrate through cortex either intracellularly (CNs) or intercellularly (RKNs) towards the vascular bundle. Once nematodes reach the vascular bundle, they select a few xylem parenchymal cells and reprogram them to form specialised nematode feeding sites (NFS). After establishment of permanent feeding sites, nematode becomes sedentary and undergoes various moults. J2 develops into J3 and J4 and then into an adult (Fig. 1). An adult male is long, motile, vermiform in shape and migrate out of the plant, whereas an adult female is globose, pear-shaped with a small beak, which helps in ingestion of food. In case of cyst nematodes, females reproduce sexually to produce large number of eggs. When the female dies, the body wall forms a protective covering for eggs, called cyst. These cysts are pushed out of the roots and are visible on the surface of roots. They are, therefore, known as cyst nematodes. On the other hand, RKN females reproduce parthenogetically and produce a large number of eggs, which are enclosed in a gelatinous matrix, called egg masses. These egg masses are visible on root knots as they are pushed out of the roots while female remains inside the root. Upon hatching eggs again form motile infective second stage juveniles (J2 s) to start a new infective cycle (Abad and Williamson 2010).
The lifecycle of Plant-parasitic nematode showing cyst nematode and root-knot nematode. Root-knot nematodes form giant cells in host root as nematode feeding site and reproduce through parthenogenesis to from egg masses. However, cyst nematodes form syncytium as nematode feeding site and undergo sexual reproduction to produce cyst (photo courtesy: http://plantsci.missouri.edu/mitchumlab)
NFS are metabolically active, large, multinucleate cells with dense granular cytoplasm which serve as the sole source of food for developing nematode. They are called syncytia in case of CNs and giant cells (GCs) in case of RKNs. A syncytium is formed by endoreduplication of nuclei and may incorporate >200 cells through dissolution of adjacent cell walls with subsequent fusion of cytoplasm of neighboring cells. GCs are formed by stimulation of karyokinesis without cytokinesis of only 5–7 vascular parenchymal cells. The cortical tissue surrounding GCs exhibit hyperplasia and hypertrophism leading to the formation of characteristic galls, called root-knots.
Nematodes have evolved to interactively communicate with the host so that it becomes suitable for growth and reproduction. In the past, studies on nematode behaviour, morphology, physiology, genetics and lifecycle have been highly informative. In recent years, however, extensive efforts have been made to understand the molecular basis of plant-nematode interactions.
Molecular determinants of parasitism
During infection, parasitic nematodes secrete biochemical compounds (proteins, hormones, etc.) or the so called ‘effector molecules’ that are important for parasitism. Therefore, identification and characterization of effector molecules involved in disease development is critical for designing biotechnological strategies using nematode targets to control them. Further, these effector molecules interact with plant proteins and manipulate them according to their own benefit. Plant gene expression is modified to form NFS and subsequent feeding on plant cytoplasm leads to developmental progression in nematode from J2 to adult. Therefore, annotation and characterization of nematode responsive plant genes is equally essential for understanding the dynamics of disease development. Another aspect to be considered in disease development process is the specific interactions between the effectors and their corresponding host plant proteins.
Nematode parasitism genes
Sedentary endoparasitic nematodes rely on effector molecules to successfully parasitize their host. These effector molecules modulate host response to their own benefit so that they can live inside the host tissue. Various functions attributed to nematode effector molecules include: modification of plant cell walls, suppression of plant defense responses, alteration in the auxin and polyamine signaling, mimicking plant molecules to by-pass defense responses and regulating stress signaling. Since parasitic stages are inside the host tissue, it is difficult to assess them; therefore, in earlier studies, nematode secretions were used for identification of effector molecules. Majority of effector molecules are produced in esophageal glands (two sub-ventral glands and one dorsal gland) while others are synthesized in hypodermis and amphids. Most successful technique in deciphering effector molecules from PPNs have been the protein-based approach where secretions from esophageal glands are analyzed directly from pre-parasitic and parasitic stages of nematodes. One of the major drawbacks of protein-based studies is insufficient availability of protein of interest and only a limited number of proteins could be analyzed in one experiment. Hence, researchers switched to sequence-based approaches. Among sequence-based approaches, EST (expressed sequence tags) databases were explored to reveal potential effector candidates on the basis of their similarity to already identified parasitism genes (Jacob and Mitreva 2011). The detailed analysis of gene expression of a particular gene during parasitic stages has been by far the most successful technique to understand parasitism process but more recent next generation sequencing technology have made it possible to analysis genome-wide expression profile of all the cells of an organism. Genome datasets are now available for some of the PPN species and for many PPN species genome projects are in progress. Analysis of whole-genome and transcriptomes of curated PPNs have revealed a set of parasitism genes from different life stages such pre-parasitic J2, parasitic J2, J3/J4 and adult female (Nicol et al. 2012; Haegeman et al. 2013; Cotton et al. 2014; Rutter et al. 2014a; Petitot et al. 2016). Such genome-wide analysis of gene expression has identified numerous parasitism genes along with novel genes whose functions are still unknown. Following sections describes a few nematode parasitism genes based on their function and a list of functionally characterized nematode effectors with their respective roles is summarized in Table 1.
Cell wall degrading and modifying enzymes
The first structural barrier for parasitic nematodes is plant cell wall, which needs softening via mixture of enzymes secreted from PPNs. Nematode stylet secretions contain various cell wall degrading and modifying enzymes such as, β-1,4-endoglucanases, xylanases, pectate lyases, expansins and polygalacturonases, which break the integrity of plant cell wall through hydrolytic breakdown of cellulose, hemicellose, pectin etc. present in the cell wall. Protein tagging with fluorescent proteins has been used to localize these secretions inside the host tissue. β-1,4-endoglucanase was the first nematode effector molecule to be localized inside the host roots. It was first cloned from Heterodera glycines (Hewezi and Baum 2013) and was the first cellulase gene to be reported in animals. Nematode cellulases and other cell wall degrading enzymes show great similarity with that of bacterial and fungal cell wall degrading enzymes suggesting their probable acquisition by horizontal gene transfer (Baum et al. 2007). Moreover, Heterodera schachtii cellulose binding protein (CBP) was shown to interact with Arabidopsis pectin methyl esterase 3 and thus reduces levels of methylesterified pectins in the cell walls. As a result, the plant becomes susceptible to nematode (Hewezi and Baum 2013). On the other hand, expansins break non-covalent bonds between the cell wall fibrils and thus loosens its structure.
Suppression of plant defense response
Once inside the root tissue, nematode has to deal with plant defenses including reactive oxygen species (ROS), cell wall thickening and secondary metabolites. On the other hand, nematodes deploy various mechanisms to cope with such defense responses and host defense responses are subsequently suppressed for successful parasitism. An active role in plant defense suppression has been proposed for esophageal gland specific chorismate mutase enzyme. Chorismate mutase (CM) enzyme was first cloned using differential screening of cDNA libraries from the excised nematode pharyngeal glands and nematode tail region. Chorismate is the end product of plant shikimate pathway and nematode CM acts on chorismate substrate and catabolize it to prephenate, which subsequently reduces the synthesis of flavonoids, salicylic acid, phytoalexins and auxin (Curtis 2007; Baum et al. 2007). Another effector protein, FAR-1, belongs to fatty acid and retinol-binding family. Nematode surface protein FAR-1 has ability to bind to precursors of jasmonic acid signaling pathway and thus actively suppresses plant defense responses (Hassan et al. 2010). Peroxidase proteins presumably are involved in detoxification of ROS. Peroxidase genes have been reported from hypodermis of potato cyst nematode and proteins were reported to be localized on body surface that helps them to cope with ROS and suppress defense responses (Baum et al. 2007). Another gene encoding glutathione-S-transferase (GSTs) have been identified in endophytic J3 nematodes through suppression subtractive hybridization studies of exophytic J2 and endophytic J3 RKN. GSTs are likely to play a role in suppressing plant defense responses at plasma membrane and cell walls during early and later stages of parasitism (Dubreuil et al. 2007).
Effector proteins that could alter plant developmental gene expression
Feeding site formation inside the host tissue requires re-programming of plant cell cycle genes as well as the genes involved in transport of nutrients. A few nematode effectors are known to produce proteins with nuclear localization signals that possibly alter plant gene expression. Nematode effectors that trigger endoreduplication in nematode feeding sites or acytokinetic mitosis in giant cells are yet to be verified. However, cyclin D3, which is involved in G1 to S phase transition during cell cycle, was reported in GCs and indicates that plant cells have re-entered the cell cycle for formation of multi-nucleate GCs (Zimmet et al. 1997; de Almeida Engler et al. 1999, 2015). Several other effectors have also been shown to play a role in cell cycle regulation, which includes a CDC48-like protein, a ubiquitin and a SKP1-like protein (de Almeida Engler and Gheysen 2013). SKP-1 (S-phase kinase-associated protein-1) is a member of SCF (Skp, Cullin, F-box) ubiquitin ligase protein complex, which is known to be involved in cell cycle progression. A homologue of SKP-1 was identified from cyst nematodes and may be involved in inducing multiple S-phases without cytokinesis which leads to development of syncitia. Another group of effectors, ubiquitin extension proteins, have also been reported from cyst nematodes that play role in nuclear transport and cell cycle regulation. Ubiquitination is the process where the ubiquitin targets the protein to proteasome complex for degradation of proteins (Hassan et al. 2010). RanBPM (Ran-binding protein in microtubule organizing center) has diverse cellular functions such as modulation of proteins, regulation of transcription activity, regulation of cell cycle and neurological functions. RanBPMs were reported from potato cyst nematodes that might play role in cell cycle regulation during syncytium development, however exact mechanism of action of RanBPM in regulation of cell cycle is yet to be identified (Baum et al. 2007). Various effector molecules have also been shown to interact with plant hormones (such as auxin, cytokinin and ethylene) which interfere with plant hormone signaling and balance (Caillaud et al. 2008). An effector, Hs19C07, identified from cyst nematode Heterodera schachtii have been shown to interact with LAX3 auxin influx protein, thus increases auxin influx during initial syncytium development (Hassan et al. 2010).
Small bioactive peptides and effectors with uncertain function
Small bioactive peptides play significant role in plant growth and development. For instance, CLAVATA-like peptides have been shown to regulate cell differentiation in root and shoot meristem. The CLAVATA3 (CLV3)/Endosperm surrounding region (ESR) (CLE) peptides function as signaling molecules in regulation of various physiological and developmental processes particularly in meristem differentiation. Interestingly, CLE peptides have been reported from nematode secretions both in CNs and RKNs that mimic plant CLE peptides but their receptors in plant are unknown. 16-D-10 from RKN is a 13 amino acid peptide that has sequence similarity to CLE peptide and probably interacts with SCARECROW-like transcription factors inside the plant during giant cell formation (Hassan et al. 2010). Through suppression subtractive hybridization, Dubreuil et al. (2007) identified genes encoding a predicted cathepsin B-like cysteine proteases that is thought to help nematode in digesting nutrients in intestine and putatively act as effector of plant parasitism. A gene encoding a predicted galectin (galactose-binding lectin) which expressed exclusively in the intestine and esophageal glands have also been identified in the same study from RKNs. However, the role of galectins in plant parasitism is yet to be identified.
Effectors that modulate calcium concentrations in plant cells
Calcium ions are generally involved in signaling pathways, cell adhesion, plant defense responses, cell death by apoptosis and molecular chaperon activity in endoplastic reticulum. Annexins are calcium dependent phospholipid binding proteins that protects the plant cell against stress like drought, oxidative and osmotic. Homologues of annexins have been reported from dorsal esophageal glands of Heterodera glycines and their interaction with plant genes are important for initiation of syncycial cell. Annexin-like genes had also been identified from C. elegans and Globodera pallida, however, their function is not established as they do not contain signal peptide sequences, which is required for secretion (Patel et al. 2010). Calreticulins are calcium binding proteins and modulates concentration of calcium during signaling. RKN calreticulin is involved in maintaining intercellular trafficking in giant cells and cell cycle regulation during giant cell formation (Baum et al. 2007; Patel et al. 2010).
Nematode responsive plant genes
At each level of plant nematode interaction, nematode responsive plant genes are as important as nematode effector molecules. Plant genes responding to nematode effectors will determine the aggressiveness of nematode. Nematode alters the expression of plant cytoskeleton genes, cell cycle genes, defense responses and hormone signaling genes that allows the formation of nematode feeding site. Sedentary nematodes select certain plant cells to initiate nematode feeding site which respond to effector molecules and make them nutrient sinks. This signal exchange between plants and nematodes is very significant to understand the mechanism of disease development. Still the functional roles of many nematode responsive plant genes in feeding site formation remain elusive. Studying the transcriptional changes during plant-nematode interactions has increased our knowledge in understanding the function of plant response to nematode infection. Various molecular approaches have been used to study plant responses like differential display, promoter–reporter gene fusions, promoter-trap strategies, RNA blotting, protein immunolocalization, in situ hybridization and differential library screening (Abad and Williamson 2010).
Initial studies were conducted using differential screening of cDNA libraries from infected and uninfected roots. The first differentially expressed gene identified using this method was a giant cell-induced gene, TobRB7, from tobacco infected roots. Since nematode infection is not synchronous, so the initial studies were restricted to late infection stages (Escobar et al. 2011). Various molecular approaches like differential screening and subtraction of cDNAs, differential display, promoter-β-glucuronidase (GUS) fusions, mRNA in situ hybridization, (in situ) reverse-transcription polymerase chain reaction (RT-PCR) have been used to compare differential gene expression in nematode feeding sites and corresponding uninfected root regions (Gheysen and Fenoll. 2002). Recent advances in technology such as laser capture microdissection (LCM) and microaspiration, DNA microarrays and more recently RNA sequencing will help to understand plant-nematode interactions at a deeper depth.
There are microarray-based transcriptome studies in Arabidopsis (Hammes et al. 2005; Jammes et al. 2005; Fuller et al. 2007a; Barcala et al. 2010), tomato (Bar-Or et al. 2005; Schaff et al. 2007; Portillo et al. 2013), a resistant soybean line (Ibrahim et al. 2011) and RKN tolerant egg plant Solanum torvum (Bagnaresi et al. 2013). More recently, advances in high-throughput profiling approaches such as RNA-Seq, gives us a global picture of host response and directly identifies both conserved and novel transcripts along with their abundance. Transcriptome profiling of Aegilops variabilis was done to study plant response and defense against cereal cyst nematode using mRNA-Seq (Illumina HiSeq™ 2000 instrument) of infected and uninfected roots (Xu et al. 2012). The study provided a platform to explore the genome of resistant wheat plant and discover novel genes involved in plant defense which may subsequently be used for plant improvement programs. To date, only a few studies have utilized NGS technique to investigate differential host gene expression patterns during PPN-host interaction including rice galls and GCs (Kyndt et al. 2012; Ji et al. 2013), resistant soybean roots (Beneventi et al. 2013), resistant and susceptible alfalfa cultivars (Postnikova et al. 2015) and common bean roots (Santini et al. 2016). Following sections describes the plant genes involved in parasitism and few nematode responsive plant genes are summarized in Table 2.
Pathogenesis-related (PR) genes
The damage caused by nematodes induces plant defense responses which are subsequently silenced by pathogen for successful parasitism. Genes encoding reactive oxygen species (ROS) was up-regulated during root-knot nematode invasion (Caillaud et al. 2008) while gene encoding gibberellin (GA) 2-oxidase-like protein was up-regulated throughout disease development in both RKN- and CN-infected roots. GA 2-oxidase enzyme is responsible for deactivation of gibberellin thus inhibiting plant elongation during parasitic process (Bar-or et al. 2005). A recent study on rice and root-knot nematode interaction revealed the role of brassinosteroids in pathogenicity process. Pathogen takes over the plant brassinosteroid pathway, which suppresses the salicylic acid (SA)- and gibberellin (GA)-mediated signal transduction pathways (Nahar et al. 2013). Genes encoding two WRKY transcription factors were up-regulated during disease development and are known to repress pathogenesis related genes, including peroxidases (Bar-or et al. 2005).
Hormone associated genes
The plant hormone, auxin plays significant role in plant development particularly organogenesis and cell expansion. High auxin concentrations are required to initiate development of a new organ and auxin transporters and auxin signaling molecules are induced during early stages of feeding site development. AUX1 and AUX4/LAX3 (putative auxin transporters) and DR5 (auxin responsive promoter element) were induced at early stages of giant cell formation, which causes local, transient increase in auxin levels (Caillaud et al. 2008). Genes encoding cytokinin-response factors were up-regulated during early stages of giant cell formation suggesting the role of cytokinin for giant cell initiation (Caillaud et al. 2008). No production or over-production of ethylene reduces the attractiveness of root-knot nematodes towards the roots suggesting the role of ethylene for initial attraction of nematodes towards the root system (Fudali et al. 2013).
Membrane transport proteins
The transport across plasma membrane is important for a constant supply of nutrients inside the developing nematode feeding site. Genes involved in cellular transport were differentially regulated in galls and syncytia. Three of the Arabidopsis aquaporin genes were upregulated (one NOD26-like intrinsic protein and 2 plasma memebrane PIPs) whereas 7 (3 PIPs and 4 tonoplastic aquaporins TIPs) were downregulated in galls (Jammes et al. 2005). Two amino acid transporters, AAP3 and AAP6, were up-regulated during RKN infestation in Arabidopsis roots (Marella et al. 2013).
Cytoskeleton and cell cycle genes
A characteristic feature of feeding site is cell wall remodeling or its expansion. Plant cell wall degrading or modifying enzymes (CWD/MEs) are also involved in feeding cell formation. After initiation of feeding cells, expression of genes encoding CWD/MEs are reduced in nematodes and expression of genes encoding CWD/MEs are induced in plants for allowing cell wall remodelling during formation of feeding cells (reviewed in Sobczak et al. 2011). Additionally, genes involved in cytoskeleton reorganization were differentially regulated in PPN-infected host roots. For instance, membrane anchored formins (AtFH6 and AtFH10), involved in actin nucleation, were upregulated during RKN infection (Jammes et al. 2005).
Molecular basis of resistance
When plant encounters a pathogen, multiple layers of defense responses are activated. The first layer of plant immune system, called basal defense responses or pattern-triggered immunity (PTI), is activated by the interaction of pattern recognition receptors (PRRs; present on the plant cell surface) with pathogen-associated molecular patterns (PAMPs; conserved pathogen molecules). PTI aim at inhibiting the growth of pathogen and restrict the infection. Virulent pathogens overcome PTI and alter the host physiology by secreting the effector molecules and evoke effector-triggered susceptibility (ETS). Plants have evolved multiple resistance (R) genes that either directly or indirectly recognize specific effectors [avirulence (Avr) proteins]. This R/Avr interaction activates a stronger defense response referred to as effector-triggered immunity (ETI) (Jones and Dangl 2006). Two interacting, but distinct plant defense responses, PTI and ETI, defend the host against attacking pathogen and pest. ETI-based resistance has been shown against nematodes, but PTI-based resistance has not yet been reported.
Host resistance genes
The largest class of R-genes encodes proteins that contain a central nucleotide-binding (NB) and a C-terminal leucine-rich repeat (LRR) domain. The N-terminal of NB-LRRs is varied structurally and presence of LRR domain is a typical hallmark of most of R-genes. High variability of LRR domain is predicted to be related to recognition of diverse pathogen elicitors and likely to be originated from point-mutations combined with positive selection (Jones and Dangl 2006). Proteins carrying N-terminal domain sharing homology to the Toll and human interleukin-1 receptor (TIR) domain are called TIR-NB-LRRs or TNLS. Other Non-TIR NB-LRR members of protein contain a predicted coiled coil region (CC), sometimes extended by a DNA binding domain such as BEAF/DREAF Zinc finger domain (BED) or a solanaceous domains (SD) are referred to as CC-NB-LRRs or CNLs.
Nematodes are important pathogens of many crop plants and host resistance is a highly desirable approach to control (Williamson and Kumar 2006). Several nematode-R-genes (Nem-R-genes) have been characterized genetically and a handful have been cloned belonging to the class of a single dominant genes (also called major resistance genes). While many of the Nem-R-genes are canonical immune receptors resembling those that confer resistance to other plant pathogen, the recent isolation and functional characterization of two major quantitative trait loci Rhg1 and Rhg4 contributing to resistance to the soybean cyst nematode do not fit this pattern and reveal that the resistance against PPNs can be achieved by mechanisms other than effector triggered immunity. To date all the cloned Nem-R-genes are summarized in Table 3.
Cloned Nem-R-genes in plants
The first nematode R-gene to be cloned was Hs1Pro−1 gene from wild sugar beets. The Hs1 Pro−1 gene confers resistance against sugar beet cyst nematode, H. schachtii. This gene encodes for a protein that lacks LRR domain and its precise role in conferring resistance is still elusive. Other R-genes identified from crop plants include Mi-1 of tomato, Hero A of tomato, Gpa2 of potato, Gro1-4 of potato, Ma of plum and rhg genes (rhg1 & rhg 4) of soybean. The Mi1.2 gene of tomato confers resistance to several root-knot nematode species and acts when infective juvenile tries to form a feeding site. A hypersensitive reaction is activated in the region of plant where a feeding structure is being induced, leading to cell death surrounding the invaded J2. In contrast, another tomato Nem-R-gene, Hero A, which confers resistance to potato cyst nematode, acts after feeding cells have developed by inciting an HR in the cells immediately surrounding the developing feeding structure. Interestingly, two rhg genes cloned from soybean show atypical properties as compared to other R-genes (Cook et al. 2012; Liu et al. 2012). Resistance is conferred by rhg1 gene is due to the presence of tandem repeats containing several different genes whereas rhg4 gene encodes for a serine hydroxymethyl transferase. However, detailed mechanisms of resistance are yet unknown.
Nematode Avr effector genes
Substantial progress has been made to identify and clone Avr genes from bacteria and fungi, but only a few have been identified from PPNs. The cognate effectors (i.e., Avirulence genes) for the Mi1.2 and Hero A have not yet been identified. However, the Avirulence gene recognized by another Nem-R-gene, Gpa2, has been identified as the GpRBP1 gene from the potato cyst nematode Globodera pallida (Fuller et al. 2007b). This protein is part of a large family of secreted proteins called SPRYSECs (Sacco et al. 2009; Postma et al. 2012), some of which are known to suppress host defense responses (Quentin et al. 2013). Recently, members of a conserved family of nematode pheromones (ascarosides) have been identified as nematode PAMPs (Manosalva et al. 2015). Low concentrations of a major ascaroside, ascr#18, activate plant immune responses and increases resistance against the pathogen. Another gene encoding a putative RKN avirulence (MAP-1) protein, has previously been described based on differential expression between near-isogenic virulent and avirulent lines of M. incognita (Semblat et al. 2001; Castagnone-Sereno et al. 2009). More recently, members of MAP-1 gene family have been shown to possess conserved CLE-like motifs but the number and arrangements of repeats are different (Rutter et al. 2014b). The gene silencing of Mi-Cg1 of M. incognita converts the avirulent species to become virulent, which suggest that this gene is required for Mi-1.2 gene to confer resistance. However, the protein encoded by Mi-Cg1 gene does not contain a signal peptide sequence, which is a hallmark of secretory proteins that might interact with the host plant proteins. It is speculated that Mi-Cg1 gene is involved in regulting the expression of another effector gene, which is recognized by Mi-1.2 gene (Gleason et al. 2008).
MicroRNAs in plant-nematode interactions
The role of microRNAs (miRNA) in regulating the expression of genes that are involved in plant-nematode interactions have already been demonstrated (Jones-Rhoades et al. 2006; Hewezi and Baum 2015). miRNAs are 20–22 nucleotide long non-coding RNA molecules generated by dicer-like protein activity (Reinhart et al. 2002). They are transcribed from MIR genes by their own promoters and RNA pol II to form single strand of RNA precursor (pri-miRNA) in the nucleus (Lee et al. 2004). Pri-miRNAs form stem-loop hairpin structure (precursor, pre-miRNA) and are then cleaved by DCL proteins in the cytoplasm to generate miRNA duplex (Bartel 2004). A mature strand of this duplex is incorporated into RNA-induced silencing complex (RISC) and suppresses the expression of homologous mRNA targets by cleavage mediated through argonaute (AGO1) or by translational inhibition.
Host miRNAs have been shown to play a role in regulating the expression of genes involved in nematode parasitism (Hewezi and Baum 2015; Cabrera et al. 2016). For example, 19 miRNA families were differentially regulated at different time points of cyst nematode (Heterodera schachtii) infection in Arabidopsis indicating the role of miRNAs in host-nematode interaction (Hewezi et al. 2008). Majority of the miRNAs during early stage of infection were downregulated whereas at later stage, 7 out of 16 miRNAs were upregulated, 5 were downregulated and four remain unchanged. Recently, a study on host-RKN interaction during early stages of disease development suggested the role of miRNAs during RKN parasitism (Cabrera et al. 2016). For instance, downregulation of miRNAs in galls at early infection stage during Arabidopsis-M. javanica interaction was observed (Cabrera et al. 2016). The most abundant targets of miRNAs are transcription factors that are deregulated in response to nematode infection (Hewezi et al. 2012; Cabrera et al. 2016). Also, miR396 regulates growth regulating factors (GRFs), miR171 regulates scarecrow-like transcription factors (SCL), miR156 regulates squamosa promoter-binding protein transcription factor (SBP), miR159 regulates myleoblastosis (MYB) family of transcription factor, miR166 regulated homeobox (HB), miR319 regulates Teosinte branched 1/Cycloidea/Proliferating cell factor 1 (TCP) and miR390/TAS3 regulates Auxin responsive factors (ARFs) during host-nematode interactions. These transcription factors have been shown to play important roles in regulating key genes involved in plant-nematode interaction. The regulatory role of miR396/GRFs during transition of syncytium formation and maintenance phase has been demonstrated during nematode infection (Hewezi et al. 2012). These GRFs regulate various biological processes involved in plant defenses and disease resistance in plants. Zhao et al. (2015) reported that miR319/TCP module acts as a regulator of jasmonic acid levels in tomato upon M. incognita infection and thereby affects the nature of host resistance. Recently, functional role of miR390/TAS3 in regulating ARFs in early gall development was studied during Arabidopsis-M. javanica interaction (Cabrera et al. 2016). Further work on the extent of involvement of miRNA in plant-nematode interactions will provide a novel insight into gene regulation during nematode parasitism.
General conclusions and future perspectives
PPNs cause extensive damage and substantial yield loss as other biotic constraints but difficulties in recognizing nematode threats lead to ignorance of damage caused due to these pathogens. A wide range of nematode management strategies is being employed to manage PPN impact on economic losses in crop plants. These strategies include (1) use of nematicides (2) cultural practices and (3) production of nematode resistant varieties. Use of nematicides is restricted to limit environmental harm. Cultural practices generally involve crop rotation but with a drawback that cyst nematodes remain dormant for many years and root knot nematodes are polyphagous. Due to inadequacy of these measures, host resistance is considered to be the most effective which can be either natural or can be produced by transferring resistant gene from wild species to cultivated one through conventional breeding. But this involves crossing and selection, which is time consuming and limits the gene pool. Although transgenic transfer of nematode resistance genes within a closely related species has been successful, this technique has limited success in producing nematode resistance in distantly related species (reviewed in Williamson and Kumar 2006). Understanding molecular aspects of plant-nematode interactions hold wide application and relevance in present context of society. Knowledge of molecular determinants in host-pathogen interaction will be a way forward to design environmental friendly controls to target harmful pathogen. This will help in reducing economic loss to agriculture incurred by pathogen.
References
Abad, P., & Williamson, V. M. (2010). Plant nematode interaction: a sophisticated dialogue. In C. Escobar & C. Fenoll (Eds.), Advances in botanical research (pp. 147–192). Amsterdam: Elsevier.
Bagnaresi, P., Sala, T., Irdani, T., Scotto, C., Lamontanara, A., Beretta, M., et al. (2013). Solanum torvum responses to the root-knot nematode Meloidogyne incognita. BMC Genomics, 540, 1–21.
Barcala, M., Garcia, A., Cabrera, J., Casson, S., Lindsey, K., Favery, B., et al. (2010). Early transcriptomic events in microdissected Arabidopsis nematode-induced giant cells. The Plant Journal, 61, 698–712.
Bar-Or, C., Kapulnik, Y., & Koltai, H. (2005). A broad characterization of the transcriptional profile of the compatible tomato response to the plant parasitic root knot nematode Meloidogyne javanica. European Journal of Plant Pathology, 111, 181–192.
Bartel, D. P. (2004). MicroRNAs: Genomics, biogenesis, mechanism, and function genomics: The miRNA genes. Cell, 116, 281–297.
Baum, T. J., Hussey, R. S., & Davis, E. L. (2007). Root-knot and cyst nematode parasitism genes: the molecular basis of plant parasitism. In J. K. Setlow (Ed.), Genetic engineering (pp. 17–43). New York: Springer.
Beneventi, M. A., da Silva, O. B., de Sá, M. E. L., Firmino, A. A. P., de Amorim, R. M. S., Albuquerque, E. V. S., et al. (2013). Transcription profile of soybean-root-knot nematode interaction reveals a key role of phythormones in the resistance reaction. BMC Genomics, 322, 1–17.
Cabrera, J., Barcala, M., García, A., Rio-Machín, A., Medina, C., Jaubert-Possamai, S., et al. (2016). Differentially expressed small RNAs in Arabidopsis galls formed by Meloidogyne javanica: A functional role for miR390 and its TAS3-derived tasiRNAs. New Phytologist, 209, 1625–1640.
Cai, D., Kleine, M., Kifle, S., Harloff, H. J., Sandal, N. N., Marcker, K. A., et al. (1997). Positional cloning of a gene for nematode resistance in sugar beet. Science, 275, 832–834.
Caillaud, M. C., Dubreuil, G., Quentin, M., Barbeoch, L. P., Lecomte, P., Engler, J. A., et al. (2008). Root-knot nematodes manipulate plant cell functions during a compatible interaction. Journal of Plant Physiology, 165, 104–113.
Castagnone-Sereno, P., Semblat, J. P., & Castagnone, C. (2009). Modular architecture and evolution of the map-1 gene family in the root-knot nematode Meloidogyne incognita. Molecular Genetics and Genomics, 282, 547–554.
Cook, D. E., Lee, T. G., Guo, X., Melito, S., Wang, K., Bayless, A. M., et al. (2012). Copy number variation of multiple genes at Rhg1 mediates nematode resistance in soybean. Science, 338, 1206–1209.
Cotton, J. A., Lilley, C. J., Jones, L. M., Kikuchi, T., Reid, A., Thorpe, P., et al. (2014). The genome and life-stage specific transcriptomes of Globodera pallida elucidate key aspects of plant parasitism by a cyst nematode. Genome Biology, 15, 1–17.
Curtis, R. H. C. (2007). Plant parasitic nematode proteins and the host-parasite interaction. Briefings in Functional Genomics and Proteomics, 6, 50–58.
de Almeida Engler, J. A., & Gheysen, G. (2013). Nematode-induced endoreduplication in plant host cells: Why and How? The American Phytopathological Society, 26, 17–24.
de Almeida Engler, J., de Vleesschauwer, V., Burssens, S., Celenza, J. L., Jr., Inzé, D., van Montagu, M., et al. (1999). Molecular markers and cell cycle inhibitors show the importance of cell cycle progression in nematode-induced galls and syncytia. Plant Cell, 11, 793–807.
de Almeida Engler, J., Vieira, P., Rodiuc, N., de Sa, M. F. G., & Engler, G. (2015). The plant cell cycle machinery: usurped and modulated by plant-parasitic nematodes. In C. Escobar & C. Fenoll (Eds.), Advances in botanical research (pp. 91–118). Amsterdam: Elsevier.
Dubreuil, G., Magliano, M., Deleury, E., Abad, P., & Rosso, M. N. (2007). Transcriptome analysis of root-knot nematode functions induced in the early stages of parasitism. New Phytologist, 176, 426–436.
Ernst, K., Kumar, A., Kriseleit, D., Kloos, D. U., Phillips, M. S., & Ganal, M. W. (2002). The broad-spectrum potato cyst nematode resistance gene (Hero) from tomato is the only member of a large gene family of NBS-LRR genes with an unusual amino acid repeat in the LRR region. Plant Journal, 31, 127–136.
Escobar, C., Brown, S., & Mitchum, M. G. (2011). Transcriptomic and proteomic analysis of the plant response to nematode infection. In J. Jones, et al. (Eds.), Genomics and molecular genetics of plant-nematode interactions (pp. 157–173). Netherlands: Springer.
Fudali, S. L., Wang, C., & Williamson, V. M. (2013). Ethylene signaling pathway modulates attractiveness of host roots to the root-knot nematode Meloidogyne hapla. Molecular Plant-Microbe Interaction, 26, 75–86.
Fuller, V. L., Lilley, C. J., Atkinson, H. J., & Urwin, P. E. (2007a). Differential gene expression in Arabidopsis following infection by plant-parasitic nematodes Meloidogyne incognita and Heterodera schachtii. Molecular Plant Pathology, 8, 595–609.
Fuller, V. L., Lilley, C. J., & Urwin, P. E. (2007b). Nematode resistance. The New phytologist, 180, 27–44.
Gheysen, G., & Fenoll, C. (2002). Gene expression in nematode feeding sites. Annual Review of Phytopathology, 40, 191–219.
Gleason, C. A., Liu, Q. L., & Williamson V. M. (2008). Silencing a candidate nematode effector gene corresponding to the tomato resistance gene Mi-1 leads to acquisition of virulence. Molecular Plant Microbe Interactions, 21, 576–585.
Grundler, F., Schnibbe, L., & Wyss, U. (1991). In vitro studies on behavior of 2nd stage juveniles Heterodera schachtii (nematode, heteroderidae) in response to plant-roots exudates. Parasitology, 103, 149–155.
Haegeman, A., Bauters, L., Kyndt, T., Rahman, M. M., & Gheysen, G. (2013). Identification of candidate effector genes in the transcriptome of the rice root knot nematode Meloidogyne graminicola. Molecular Plant Pathology, 14, 379–390.
Hammes, U. Z., Schachtman, D. P., Berg, R. H., Nielsel, E., Koch, W., McIntyre, L. M., et al. (2005). Nematode-induced changes of transporter gene expression in Arabidopsis roots. The American Phytopathological Society, 18, 1247–1257.
Hassan, S., Behm, C. A., & Mathesius, U. (2010). Effectors of plant parasitic nematodes that re-program root cell development. Functional Plant Biology, 37, 933–942.
Hewezi, T., & Baum, T. J. (2013). Manipulation of plant cells by cyst and root-knot nematode effectors. Molecular Plant Microbe Interactions, 26, 9–16.
Hewezi, T., & Baum, T. J. (2015). Gene silencing in nematode feeding sites. In C. Escobar & C. Fenoll (Eds.), Advances in botanical research (pp. 221–239). Amsterdam: Elsevier.
Hewezi, T., Howe, P., Maier, T. R., & Baum, T. J. (2008). Arabidopsis small RNAs and their targets during cyst nematode parasitism. Molecular Plant Microbe Interactions, 21, 1622–1634.
Hewezi, T., Maier, T. R., Nettleton, D., & Baum, T. J. (2012). The Arabidopsis MicroRNA396-GRF1/GRF3 regulatory module acts as a developmental regulator in the reprogramming of root cells during cyst nematode infection. Plant Physiology, 159, 321–335.
Ibrahim, H. M., Hosseini, P., Alkharouf, N. W., Hussein, E. H., Abd El-Kader, Y., Aly, M. A., et al. (2011). Analysis of gene expression in soybean (Glycine max) roots in response to the root-knot nematode Meloidogyne incognita using microarrays and KEGG pathways. BMC Genomics, 12, 1–16.
Jablonska, B., Ammiraju, J. S. S., Bhattarai, K. K., Mantelin, S., de Ilarduya, O. M., Roberts, P. A., et al. (2007). The Mi-9 gene from Solanum arcanum conferring heat-stable resistance to root-knot nematodes is a homolog of Mi-1. Plant Physiology, 143, 1044–1054.
Jacob, J., & Mitreva, M. (2011). Transcriptomes of plant-parasitic nematodes. In J. Jones (Ed.), Genomics and molecular genetics of plant-nematode Interaction (pp. 119–138). Berlin: Springer.
Jammes, F., Lecomte, P., Engler, J. A., Bitton, F., Magniette, M. L. M., Renou, J. P., et al. (2005). Genome-wide expression profiling of the host response to root-knot nematode infection in Arabidopsis. The Plant Journal, 44, 447–458.
Ji, H., Gheysen, G., Denil, S., Lindsey, K., Topping, J. F., Nahar, K., et al. (2013). Transcriptional analysis through RNA sequencing of giant cells induced by Meloidogyne graminicola in rice roots. Journal of Experimental Botany, 64, 3885–3898.
Jones, J. D. G., & Dangl, J. L. (2006). The Plant Immune System. Nature, 444, 323–329.
Jones-Rhoades, M. W., Bartel, D. P., & Bartel, B. (2006). MicroRNAs and their regulatory roles in plants. Annual Review of Plant Biology, 57, 19–53.
Kyndt, T., Denil, S., Haegeman, A., Trooskens, G., Bauters, L., Van Criekinge, W., et al. (2012). Transcriptional reprogramming by root knot and migratory nematode infection in rice. New Phytologist, 196, 887–900.
Lee, Y., Kim, M., Han, J., Yeom, K., Lee, S., Baek, S. H., et al. (2004). MicroRNA genes are transcribed by RNA polymerase II. EMBO Journal, 23, 4051–4060.
Liu, S., Kandoth, P. K., Warren, S. D., Yeckel, G., Heinz, R., Alden, J., et al. (2012). A soybean cyst nematode resistance gene points to a new mechanism of plant resistance to pathogens. Nature, 492, 256–260.
Manosalva, P., Manohar, M., Von Reuss, S. H., Chen, S., Koch, A., Kaplan, F., et al. (2015). Conserved nematode signalling molecules elicit plant defenses and pathogen resistance. Nature Communications, 6, 7795. doi:10.1038/ncomms8795.
Marella, H. H., Nielsen, E., Schachtman, D. P., & Taylor, C. G. (2013). The amino acid permeases AAP3 and AAP6 are involved in root-knot nematode parasitism of Arabidopsis. Molecular Plant-Microbe Interactions, 26, 44–54.
Milligan, S. B., Bodeau, J., Yaghoobi, J., Kaloshian, I., Zabel, P., & Williamson, V. M. (1998). The root knot nematode resistance gene Mi from tomato is a member of the leucine zipper, nucleotide binding, leucine-rich repeat family of plant genes. Plant Cell, 10, 1307–1319.
Murray, S. L., Ingle, R. A., Petersen, L. N., & Denby, K. J. (2007). Basal resistance against Pseudomonas syringae in Arabidopsis involves WRKY53 and a protein with homology to a nematode resistance protein. Molecular Plant-Microbe Interactions, 20, 1431–1438.
Nahar, K., Kyndt, T., Hause, B., Hofte, M., & Gheysen, G. (2013). Brassinosteroids suppress rice defense against root-knot nematodes through antagonism with the jasmonate pathway. Molecular Plant Microbe Interactions, 26, 106–115.
Nicol, P., Gill, R., Nyarko, J. F., & Jones, G. K. (2012). De novo analysis and functional classification of the transcriptome of the root lesion nematode, Pratylenchus thornei, after 454 GS FLX sequencing. International Journal for Parasitology, 42, 225–237.
Paal, J., Henselewski, H., Muth, J., Meksem, K., Menéndez, C. M., Salamini, F., et al. (2004). Molecular cloning of the potato Gro1-4 gene conferring resistance to pathotype Ro1 of the root cyst nematode Globodera rostochiensis, based on a candidate gene approach. The Plant Journal, 38, 285–297.
Patel, N., Hamamouch, N., Li, C., Hewezi, T., Hussey, R. S., Baum, T. J., et al. (2010). A nematode effector protein similar to annexins in host plants. Journal of Experimental Botany, 61, 235–248.
Petitot, A. S., Dereeper, A., Agbessi, M., da Silva, C., Guy, J., Ardisson, M., et al. (2016). Dual RNA-Seq reveals Meloidogyne graminicola transcriptome and candidate effectors during the interaction with rice plants. Molecular Plant Pathology, 17, 860–874.
Portillo, M., Cabrera, J., Lindsey, K., Topping, J., Andres, M. F., Emiliozzi, M., Oliveros, J. C., Gracia-Casado, G., Solano, R., Koltai, H., Resnick, N., Fenoll, C., & Escobar, C. (2013). Distinct and conserved transcriptomic changes during nematode-induced giant cell development in tomato compared with Arabidopsis: a functional role for gene repression. New Phytologist, 197, 1276–1290.
Postma, W. J., Slootweg, E. J., Rehman, S., Finkers-Tomczak, A., Tytgat, T. O. G., van Gelderen, K., Lozano-Torres, J. L., Roosien, J., Pomp, R., van Schaik, C., Bakker, J., Goverse, A., Smant, G., Postma, W. J., Slootweg, E. J., Rehman, S., Finkers-Tomczak, A., Tytgat, T. O. G., van Gelderen, K., Lozano-Torres, J. L., Roosien, J., Pomp, R., van Schaik, C., Bakker, J., Goverse, A., & Smant, G. (2012). The effector SPRYSEC-19 of globodera rostochiensis suppresses CC-NB-LRR-mediated disease resistance in plants. Plant Physiology, 160(2), 944–954.
Postnikova, O. A., Hult, M., Shao, J., Skantar, A., & Nemchinov, L. G. (2015). Transcriptome analysis of resistant and susceptible alfalfa cultivars infected with root-knot nematode Meloidogyne incognita. PLoS ONE, 10, e0118269. doi:10.1371/journal.pone.0118269.
Quentin, M., Abad, P., & Favery, B. (2013). Plant parasitic nematode effectors target host defense and nuclear functions to establish feeding cells. Frontier in Plant Science, 4, 53. doi:10.3389/fpls.2013.00053.
Reinhart, B. J., Weinstein, E. G., Rhoades, M. W., Bartel, B., & Bartel, D. P. (2002). MicroRNAs in plants. Genes and Development, 16, 1616–1626.
Rutter, W. B., Hewezi, T., Abubucker, S., Maier, T. R., Huang, G., Mitreva, M., Hussey, R. S. & Baum, T. J. (2014a) Mining novel effector proteins from the oesophageal gland cells of Meloidogyne incognita. Molecular Plant Microbe Interactions, 27, 965–974.
Rutter, W. B., Hewezi, T., Maier, T. R., Mitchum, M. G., Davis, E. L., Hussey, R. S., & Baum, T. J. (2014b). Members of the Meloidogyne avirulence protein family contain multiple plant ligand-like motifs. Nematology, 104, 875–885.
Sacco, M. A., et al. (2009). The cyst nematode SPRYSEC protein RBP-1 elicits Gpa2- and RanGAP2-dependent plant cell death. PLoS Pathology, 5, e1000564. doi:10.1371/journal.ppat.1000564.
Santini, L., Munhoz, C. D. F., Bonfim, M. F., Jr., Brandão, M. M., Inomoto, M. M., & Vieira, M. L. C. (2016). Host transcriptional profiling at early and later stages of the compatible interaction between Phaseolus vulgaris and meloidogyne incognita. Phytopathology, 106, 282–294.
Schaff, J. E., Nielsen, D. M., Smith, C. P., Scholl, E. H., & Bird, D. M. (2007). Comprehensive transcriptome profiling in tomato reveals a role for glycosyltransferase in Mi-mediated nematode resistance. Plant Physiology, 144, 1079–1092.
Semblat, J. P., Rosso, M. N., Hussey, R. S., Abad, P., & Castagnone-Sereno, P. (2001). Molecular cloning of a cDNA encoding an amphid-secreted putative avirulence protein from the root-knot nematode Meloidogyne incognita. Molecular Plant Microbe Interactions, 14, 72–79.
Sobczak, M., Avrova, A., Jupowicz, J., Phillips, M. S., Ernst, K., & Kumar, A. (2005). Characterization of susceptibility and resistance responses to potato cyst nematode (Globodera spp.) infection of tomato lines in the absence and presence of the broad-spectrum nematode resistance Hero gene. Molecular Plant-Microbe Interactions, 18, 158–168.
Sobczak, M., Fudali, S., & Wieczorek, K. (2011). Cell wall modifications induced by nematodes. In J. Jones, G. Gheysen & C. Fenoll (Eds.), Genomics and molecular genetics of plant-nematode interactions (pp. 395–422). Netherlands: Springer.
Teillet, A., Dybal, K., Kerry, B. R., Miller, A. J., Curtis, R. H. C., & Hedden, P. (2013). Transcriptional changes of the root–knot nematode Meloidogyne incognita in response to Arabidopsis thaliana root signals. PLoS ONE, 8, e61259. doi:10.1371/journal.pone.0061259.
Trudgill, D. L., & Block, V. C. (2001). Apomictic, polyphagous root-knot nematodes, exceptionally successful and damaging biotrophic root pathogens. Annual Review of Phytopathology, 39, 53–77.
van der Vossen, E. A., Der Voort, V., Rouppe, J. N., Kanyuka, K., Bendahmane, A., Sandbrink, H., et al. (2000). Homologues of a single resistance-gene cluster in potato confer resistance to distinct pathogens: A virus and a nematode. The Plant Journal, 23, 567–576.
Vanholme, B., Meutter, J. D., Tytgat, T., Montagu, M. V., Coomans, A., & Gheysen, G. (2004). Secretions of plant—parasitic nematodes: A molecular update. Gene, 332, 13–27.
Williamson, V. M., & Kumar, A. (2006). Nematode resistance in plants: The battle underground. Trends in Genetics, 22, 396–403.
Xu, D. L., Long, H., Liang, J. J., Zhang, J., Chen, X., Li, J. L., et al. (2012). De novo assembly and characterization of the root transcriptome of Aegilops variabilis during an interaction with the cereal cyst nematode. BMC Genomics, 13, 1–9.
Zhao, W., Li, Z., Fan, J., Hu, C., Yang, R., Qi, X., et al. (2015). Identification of jasmonic acid-associated microRNAs and characterization of the regulatory roles of the miR319/TCP4 module under root-knot nematode stress in tomato. Journal of Experimental Botany, 66, 4653–4667.
Zimmet, J. M., Ladd, D., Jackson, C. W., Stenberg, P. E., & Ravid, K. (1997). A role for cyclin D3 in the endomitotic cell cycle. Molecular and Cellular Biology, 17, 7248–7259.
Acknowledgements
We thank Delhi University R & D grant.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shukla, N., Kaur, P. & Kumar, A. Molecular aspects of plant-nematode interactions. Ind J Plant Physiol. 21, 477–488 (2016). https://doi.org/10.1007/s40502-016-0263-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-016-0263-y