Abstract
Purpose of Review
This review summarizes recent studies on the molecular mechanisms of RNA-binding proteins (RBPs) that control neurological functions and pathogenesis in various neurodevelopmental and neurodegenerative diseases, including autism spectrum disorders, schizophrenia, Alzheimer’s disease, amyotrophic lateral sclerosis, frontotemporal dementia, and spinocerebellar ataxia.
Recent Findings
RBPs are critical players that regulate every step of posttranscriptional modifications of gene expression. Recent genome-wide approaches revealed that many proteins associate with RNA, but do not contain any known RNA-binding motifs. Additionally, many causal and risk genes of neurodevelopmental and neurodegenerative diseases are RBPs. Development of high-throughput sequencing methods has mapped out the fingerprints of RBPs on transcripts and provided unprecedented potential to discover new mechanisms of neurological diseases. Insights into how RBPs modulate neural development are important for designing effective therapies for numerous neurodevelopmental and neurodegenerative diseases.
Summary
RBPs have diverse mechanisms for modulating RNA processing and, thereby, controlling neurogenesis. Understanding the role of disease-associated RBPs in neurogenesis is vital for developing novel treatments for neurological diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
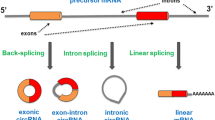
Neurodevelopment requires the functions of different proteins at different developmental stages. It involves diverse transcriptional and posttranscriptional events, such as RNA transportation, alternative splicing, stabilization, degradation, and translation. RNA-binding proteins (RBPs) are critical in regulating these processes (Fig. 1). Dysfunction of RBPs in the early stages of neural system development may affect neuronal migration, synaptic plasticity, and behavioral functions, which eventually lead to neurodevelopmental and neurodegenerative diseases [1].
Autism spectrum disorders (ASD) and schizophrenia (SCZ) are two major neurodevelopmental disorders with strong genetic components. Multiple risk genes for ASD and SCZ encode RBPs. RNA processing defects are observed in several neurodegenerative diseases, including amyotrophic lateral sclerosis (ALS), frontotemporal dementia (FTD), spinocerebellar ataxia (SCA), and Alzheimer’s disease (AD) [2,3,4,5,6,7,8,9,10]. Thus, we will focus on their diverse functions in regulating RNAs as a variety of etiologies of neurodevelopmental and neurodegenerative diseases.
RBPs in ASDs
FMR1
The neurodevelopmental disorder, fragile X syndrome (FXS), features intellectual deficits and autistic behaviors [11]. FXS is the well-known leading monogenic cause of ASD [12]. The increased trinucleotide repeats at the 5′ untranslated region of FMR1 lead to a decrease of fragile X mental retardation protein (FMRP). Local translation at the synapse is important for neuronal plasticity [13]. As an RBP, FMRP modulates mRNA localization and local translation at the synapse [14]. It represses polypeptide elongation of protein synthesis by stalling polyribosomes [15]. Loss-of-function of FMR1 significantly affects local protein translation and impairs plasticity. In addition, FMR1 serves as a sequence- and context-dependent N6-methyladenosine (m6A) reader, indicating that the m6A modification regulates mRNA stability [16].
The FXS mouse (Fmr1−/y) shows hyperactive ERK and mTOR signaling. Chronic metformin treatment selectively downregulates the ERK and mTOR signaling pathway and rescues core autistic phenotypes in this mouse model [12]. Metformin treatment also corrects the phenotypes of increased spine density and exaggerated metabotropic glutamate receptor (mGluR)-dependent long-term depression (LTD), providing an exciting drug target for FXS. Using translating ribosome affinity purification (TRAP) and RNA-seq, excessively translating mRNAs were identified in CA1 pyramidal neurons of the Fmr1−/y mouse model [17]. The muscarinic acetylcholine receptor 4 (M4) is excessively translated and consequently suppresses mGluR-induced LTD of synaptic transmission. VU0152100, a positive allosteric M4 modulator, enhances the cholinergic effects on M4, which significantly reduce the audiogenic seizures [17], suggesting a potential pathway to reverse FXS-associated phenotypes.
ELAV-Like RBPs
Neuronal ELAV-like RBPs are involved in several neurological disorders [17]. The combination of crosslinking-immunoprecipitation and RNAseq (CLIP-seq) has identified more than 8000 targets that bind to ELAV-like RBPs. They regulate splicing and abundance of bound RNAs [8]. Knockdown of ELAVL2 in primary human neurons alters mRNA alternative splicing, including RBFOX1 and FMR1, which are well-known ASD risk genes [18]. The CUGBP ELAV-like family member 4 (CELF4) gene encodes an RBP. Haploinsufficiency of CELF4 was found in ASD patients [19]. Similarly, CELF4 mutant mice show complex seizure disorders [20]. CELF4 regulates excitatory neurotransmission by stabilizing mRNA and supporting synaptic local translation [21]. It regulates about 30% of potential ASD risk genes at pre- and postsynaptic sites.
RBFOX1
RBFOX proteins are a family of RNA-binding proteins that contain a single high-affinity RNA recognition domain. RBFOX proteins bind to UGCAUG motifs to regulate RNA processing in neurons, muscle, and heart [22]. RBFOX1 has been associated with neurodevelopmental disorders, such as ASD and epilepsy [23, 24]. Alternative splicing of RBFOX1 generates different protein isoforms localized to either the nucleus or cytoplasm. The nuclear isoform regulates mRNA splicing. Downregulation of nuclear RBFOX1 delays neuronal migration in the brains of embryonic day 14.5 mice [25]. CLIP-seq shows that targets of Rbfox1 in mouse brains enriched in regulating brain development and ASD risk genes [26]. Unlike nuclear RBFOX1, cytoplasmic RBFOX1 predominantly regulates mRNA stability and translation. CLIP-seq results at single-nucleotide resolution indicate that cytoplasmic RBFOX1 binds to 3′-UTR of mRNA targets and increases their abundance [23]. The Rbfox1-bound genes control synaptic activity and calcium signaling.
Exon Junction Complex
The exon junction complex (EJC) is an RNA-binding protein complex that controls pre-mRNA splicing, maturation, translation, and nonsense-mediated mRNA decay (NMD) [27]. The core protein components are eIF4AIII, MAGOH, RBM8A, and BTZ [28, 29]. Pre-mRNA splicing plays an important role in the development of the central neural system, and multiple NMD factors are known to associate with neurodevelopmental disorders [30].
RBM8A is highly expressed in the neural progenitor cells (NPCs) of the subventricular zone at embryonic brain. Downregulation of RBM8A at embryonic day 13 promotes the neuronal migration in the neocortex and decreases the proliferation of NPCs. Upregulation of RBM8A suppresses the neuronal migration and increases the NPC dividing [31]. Consistently, haploinsufficiency of eIF4AIII, MAGOH, and RBM8A in NPCs of the dorsal telencephalon reduces cortical area and volume of mouse brains [32,33,34]. Conditional deletion of TP53 in NPCs reverses the microcephaly phenotype in the embryonic state [32]. In adult mice, abnormal RBM8A expression in the dentate gyrus leads to anxiety behaviors [35].
NMD is an mRNA surveillance mechanism that eliminates mRNAs containing premature termination codons. UPF1, UPF2, UPF3A, and UPF3B are required for activation of NMD in eukaryotes [36]. UPF3B mutations were identified in patients with ASD, SCZ, or attention-deficit hyperactivity disorder [37,38,39,40]. Knockdown of UPF3B increases proliferation of NPCs and decreases primary axon growth. UPF3B-null mice have fewer dendritic spines and less neural activity. The mutant UPF3B mice are deficient in the prepulse inhibition and fear learning tests [41]. Interestingly, UPF3A, an assumed redundant paralog of UPF3B, has an opposing function against the NMD pathway [42]. In the nervous system, besides regulating NMD, UPF1 facilitates mRNA transport and local translation for synaptic plasticity [43]. Downregulation of UPF1 decreases MAP1B mRNA in neurites. STAU2, another RBP, physically interacts with UPF1 to regulate mRNA transportation. Thus, RBPs controlling NMD are important in protecting neuronal development from inaccurate mRNA splicing and incorrect synaptic development.
Methyl-DNA-Binding Proteins
Epigenetic regulators, such as MECP2 and DNA methyl-transferases (DNMTs), contain a well-characterized methyl-DNA-binding domain (MBD) and modulate DNA methylation. Abnormal copies of MECP2 affect neuronal development and lead to neurodevelopmental disorders. MECP2 deletion causes Rett’s syndrome [44], whereas MECP2 duplication leads to similar autistic behaviors and intellectual disability. Conditionally, overexpressed MECP2 in a mouse model can be behaviorally corrected by removing one MECP2 allele or using antisense oligonucleotides to silence MECP2 [45], suggesting that gene dosage is important for the brain development. Intriguingly, some MBD proteins and DNMTs, including MECP2, interact with RNA and form an RNA protein complex [46]. The RNA-binding motifs in these proteins are different from their MBDs. These data suggest that MBD-containing proteins and DNMTs associate with RNAs to participate in DNA methylation [47]. MBD1 and MECP2 dysfunction affect adult neurogenesis and the hippocampal functions [48,49,50].
Other RBPs in ASDs
Heterogeneous nuclear ribonucleoproteins (hnRNPs) are a family of RBPs that control variant transcriptional and translational events. Abnormalities of hnRNPs are associated with different neurological diseases and cancers [51]. Missense mutation of HNRNPH2, which localizes in the X chromosome, associates with autistic behaviors and ataxia in females [52]. HNRNPU deletion was reported in patients with infantile spasms, seizures, and brain malformation [53, 54].
The growth cone responds to axonal guidance cues to reach a targeted region and form synapses. This process involves local mRNA translation to generate rapidly appropriate responses. The RBPs, hnRNPK and PCBP1, associate with mRNA and Mena (ENAH), an actin-regulatory protein, to form an RNP complex [55]. CLIP data show that Mena-bound mRNAs modulate axon guidance. DYRK1A (dual specificity tyrosine phosphorylation regulated kinase 1A), a Down syndrome and ASD risk gene, is guided by the Mena complex regulating the synaptic local translation.
Janus kinase and microtubule-interacting protein 1 (JAKMIP1) is an RBP that is involved in RNP translation. JAKMIP1 is highly expressed in glutamatergic neurons in developmental brains [56]. Differential expression of JAKMIP1 has been observed in ASD patients [57]. Protein interactome analysis indicates that JAKMIP1’s binding partners participate in translational regulation [58]. Jakmip1 knockout mice show autistic behaviors, such as social deficits, repetitive behavior, and impaired vocalization [58]. Translation initiation factors, such as EIF4E, regulate local translation in the synapse. De novo mutations of EIF4E have been associated with autistic behaviors [59].
RBPs in SCZ
Disrupted in Schizophrenia 1
SCZ is a devastating mental disorder affected by genetic risks. Disrupted in Schizophrenia 1 (DISC1) was first identified in a big Scottish family with high incidences of mental diseases [60]. The chromosomal translocation within the DISC1 gene locus is associated with SCZ [61]. DISC1 associates with many proteins and is important in neurogenesis and neural plasticity [62, 63]. It modulates the Wnt pathway and NPC proliferation and neuronal migration/differentiation by inhibiting GSK3β activity [64,65,66,67,68]. Interactome screens indicate that DISC1 physically interacts with several RBPs [69]. Its targets involve RNA-transporting granules and synaptic plasticity, such as the ITPR1 gene.
ZNF804A
ZNF804A was the first SCZ risk gene reaching genome-wide signifcance in a genome-wide association study (GWAS) [70]. Several follow-up GWASs replicated that result and also confirmed the association of rs1344706 with SCZ in different populations [71,72,73,74,75,76,77,78,79]. In addition to common variants, the SGENE-plus consortium reported rare copy number variants (CNVs) at the ZNF804A locus in psychotic patients, including a deletion in a SCZ patient, a deletion in a patient with an anxiety disorder, and a duplication in a BD patient [73], but none in controls. Interestingly, chromosome duplication, deletion, inversion, and translocation at the ZNF804A locus were found in patients with autism [80, 81] and developmental delay [82, 83]. Consistent with the reproducible genetic association of risk SNP in ZNF804A with SCZ, neuroimaging and neuropsychological studies provide mounting evidence that ZNF804A risk allele modulates human brain structures and functions [84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110].
ZNF804A contains a zinc-finger domain that shows both DNA- and RNA-binding ability. Although it has been proposed to function as a transcription factor [111], using RNA immunoprecipitation sequencing (RIP) and interactome analysis, we found that ZNF804A binds to RNAs [112]. ZNF804A is highly expressed in the prenatal central nerve system. Knockdown of ZNF804A affects translation, as well as neural migration toward the neocortex in mouse embryonic stage [112]. Suppression of ZNF804A expression in human NPCs and primary rat cortical neurons reduces neurite formation and dendritic spine formation [113], supporting the important function of ZNF804A in brain functions.
Quaking
Quaking (QKI) is a member of the signal transduction and activation of RNA (STAR) protein family and the HNRNPK homology (KH)-type family [114]. QKI binds to its downstream mRNAs carrying a conserved QKI response element (QRE). QKI regulates several RNA processes, including alternative splicing, micro-RNA processing, and mRNA stabilization and translation. Differential splicing of QKI mRNA produces several isoforms. Decreased QKI-7 and QKI-7b were observed in 55 SCZ patients [114]. These isoforms are regulated by HNRNPC1/C2 [115]. QKI isoforms are also expressed in astrocytes. Downregulation of QKI-7 in astrocytes decreases glial fibrillary acidic protein (GFAP) expression. A typical antipsychotic medication, haloperidol, increases QKI-7 and GFAP expression, suggesting that QKI-7 coordinates with GFAP to regulate the function of astrocytes [116]. This study proposes a new potential link of RNA processing with SCZ.
A recent GWAS identified 108 genetic loci associated with SCZ [78]. SCZ shares many risk genes with intellectual disability and ASDs. Interestingly, de novo mutations in SCZ enriched in glutamatergic postsynaptic proteins related to activity-regulated cytoskeleton-associated protein (ARC) and N-methyl-D-aspartate receptor (NMDAR) complexes. Many transcripts of these complexes are targets of FMRP [117]. Interestingly, FMRP is also a substrate of glycogen synthase kinase 3β (GSK3β) [118]. Inhibition of GSK3β showed antipsychotic effects and mood stabilization.
RBPs in ALS/FTD
ALS and FTD are two diseases with similar symptoms and pathogenesis [119]. Recent evidence indicates that disrupting RNA homeostasis is a leading cause of ALS/FTD [120]. Mutations in several RBPs have been identified in ALS/FTD patients.
STAU1
Staufen 1 (STAU1) is an RNA-binding protein responsible for RNA transportation, localization, translation, and the ribonucleoprotein formation [121]. Gershoni-Emek et al. observed altered localization of synaptic STAU1 in an ALS SOD1G93A mouse model, probably due to interrupted retrograde transportation from the synapses [122].
TDP-43
TAR DNA-binding protein 43 (TDP-43, also called TARDBP) is a causal gene for ALS [123]. It encodes an RBP involved in transcription, mRNA splicing, stability, and transportation [124]. Cytoplasmic mislocalization and decreased nuclear expression are associated with TDP-43 mutations. Both mislocalization and downregulation of expression contribute to the cellular toxicity [125, 126]. Though the protein has been associated with ALS for almost two decades, until recently, studies revealed that an oligomeric form of TDP-43 is functional in the nucleus, and this state is essential for its role in RNA metabolism [127]. Additionally, TDP-43 has been reported to regulate ER-mitochondria communication and Ca2+ homeostasis, probably through regulating GSK3β signaling pathway [128]. Using motor neurons derived from human-induced pluripotent stem (iPS) cells, Alami et al. reported that anterograde axonal transportation of mRNAs was reduced in ALS-causing mutations of TDP-43 [129].
Fused in Sarcoma
Fused in sarcoma (FUS), another ALS/FTD risk gene, shares many common pathophysiological characteristics with TDP-43 [123]. They are both involved in RNA processing and neuronal development [130]. Specifically, FUS binds to nascent mRNA and modifies alternative splicing [130]. A recent microarray study suggested that these two proteins share some downstream targets, including splicing and expression regulation [131]. In fibroblasts derived from an ALS patient, FUS formed nuclear aggregates [132]. Patel et al. reported that FUS undergoes a dynamic liquid-like phase transition in vivo, and converted to aggregates upon aging. This transition to aggregates is accelerated with the disease-related mutations in prion-like domains [133]. This could be the mechanism for other age-related diseases involving proteins carrying prion-like domains. Later in the same year, Murakami and colleagues reported a similar phenotype that mutant FUS generates irreversible hydrogels from liquid droplets [134]. ALS-associated FUS mutations often show higher cytoplasmic expression than wild-type controls [135], and the nuclear-localization of FUS does not seem to be required for protein aggregation and neuronal toxicity [136].
When ALS-associated FUS mutant aggregates, other ALS-associated RNA-binding proteins are sequestered in the same complex, including SMN1, hnRNPA1, hnRNPA2, and STAU-1 [134, 137]. Using an in vitro fluorescent molecule tracking assay, Murakami et al. reported that irreversible FUS aggregates trap cargo RNPs and lead to cellular toxicity. Especially among the affected proteins, SMN1 is the major causal gene for spinal muscular atrophy (SMA). Sun et al. reported that the interaction of FUS with SMN protein is increased in ALS-associated FUS mutants [138]. At the same time, the mutated FUS has reduced interaction with U1-snRNP, through which it affects global mRNA splicing [138, 139]. FUS also regulates translation. Yasuda et al. observed that FUS promotes translation preferentially within cell protrusions, and the translation process is not halted by FUS-positive granules [140]. Fragmentation of the Golgi apparatus has been observed in ALS [141, 142], and FUS disease mutation induces Golgi fragmentation [143]. Although FUS is associated with both ALS and FTD, Suárez-Calvet and colleagues identified a difference: mono-methylated arginines occur exclusively in FTLD-FUS mutations, which makes the protein to bind tightly to the nuclear import receptor, transportin-1, but not in ALS-FUS mutations [144].
C9ORF72
The hexanucleotide “GGGGCC” repeat expansion in the noncoding region of the C9ORF72 gene is another common genetic cause for ALS/FTD. Donnelly et al. reported an alteration in gene expression profiles and sequestration of a GGGGC binding RBP, ADARB2, in nuclear aggregates of patient-derived iPS cells with C9ORF72 repeats [145]. Other ALS-associated RBPs, including FUS, TDP-43, and HNRNPA1, were not identified in the same complexes [145]. However, another study did find inclusion of other RBPs, including SF2, SC35, and HNRNPH. HNRNPH was identified as a binding partner to the hexanucleotide expansion [146]. Aggregation results in increased sensitivity to glutamate toxicity [145] and enhanced apoptosis [146]. Besides interrupting RNA processing, ATM-mediated DNA repair was disrupted by the expansion, suggesting another possible cause of neurodegeneration by repeat expansion [147]. Unbiased yeast screening revealed that karyopherins and other nucleocytoplasmic transport proteins are involved in the cytotoxicity [148]. Several potential drug targets have been reported that might help prevent neurodegeneration-associated deficits, including inhibiting SRSF1-dependent nuclear export of C9ORF72 repeat transcripts [149] and antisense intervention [145].
GLE1
Recent exome screening studies linked mutations in GLE1 with ALS [150]. GLE1 expresses two isoforms in human cells, GLE1A and GLE1B [151]. GLE1A regulates translation and is localized to stress granules (SG) upon stress and regulates SG assembly and disassembly [152]. GLE1B is an mRNA export factor associated with nuclear pore complex [151, 153]. GLE1 mutations lead to dysregulation at nuclear pore complexes and cause human lethal congenital contracture syndrome-1 (LCCS1) [154, 155]. In zebrafish, Gle1 knockout results in defective Schwann cell development [156]. The ALS-linked GLE1 allele was reported to encode a protein that has the function of both GLE1A and GLE1B, which may affect the normal regulatory roles of GLE1 [157].
RBPs in SCA
SCAs are a group of over 35 progressive neurodegenerative diseases. At least six of them (SCA1, SCA2, SCA3, SCA6, SCA7, and SCA17) are caused by a polyglutamine expansion encoded by CAG repeats [158]. Two of the risk proteins, polyglutamine expansion within the ataxin-1 protein (ATXN1) and ATXN2, causing SCA1 and 2, respectively, have been identified as RBPs.
ATXN1
The ATXN1 mutation causes SCA1 [159]. The binding ability of this RBP is disrupted by the expanded polyglutamine [160]. ATXN1 regulates neuronal proliferation in vitro and in vivo [161, 162]. By modulating the GSK3β-mTOR pathway, ATXN1 regulates energy homeostasis in the mouse cerebellum [163]. On the other hand, the RAS-MAPK-MSK1 pathway modulates ATXN1 protein level and its associated toxicity [164]. Meanwhile, ATXN1 expression level is negatively regulated by PUMILIO1 (PUM1), which is also an RBP, by modulating ATXN1 RNA stability [165]. By increasing the expression of ATXN1, Pumilio1 haploinsufficiency leads to SCA1-like neurodegeneration [165]. Transcriptome profiles of the cerebellum in ATXN1 transgenic mice at several ages and genotypes revealed that upregulation of cholecystokinin (CCK) may play a protective role against Purkinje cell death [166]. Intracellular expression of HMGB1 prolongs lifespan in mutant ATXN1 knock-in mice by repairing mitochondrial DNA damage, which may serve as a potential treatment for SCA1 [167].
ATXN2
Abnormal polyglutamate expansion in ATXN2 results in SCA2 [168, 169] and ALS [170]. ATXN2 regulates metabolism through modulating the mTOR pathway [171,172,173]. It belongs to the like-Sm (LSm) protein family that regulates multiple aspects of RNA metabolism [174]. It directly interacts with poly(A)-binding protein, cytoplasmic 1 (PABPC1), and possibly regulates translation and mRNA stability partly through its binding partners [175, 176]. Meanwhile, PAR-CLIP revealed that ATXN2 also targets genes in a PABPC1-independent manner and helps to maintain mRNA stability, including genes involved in posttranscriptional processes and metabolic processes [177]. For example, TDP-43 is targeted by ATXN2 [177]. Crossing Ataxin-2 knockout mice to TDP-43 transgenic mice showed drastic reduction in TDP-43 aggregation [178]. In cell culture, ATXN2 carrying an intermediate length of polyglutamate repeats promotes mutant FUS translocation and stimulates Golgi fragmentation and apoptosis caused by mutant FUS [143]. The same ATXN2 mutant also enhances neuronal toxicity caused by C9ORF72 depletion [179]. Mitochondria dysfunction is associated with ALS [180]. ATXN2 was identified as a transcriptional regulator upstream of PINK1, a key regulator for mitochondrial stress response [181]. By positively regulating the translation of a circadian rhythm gene, Period (PER), ATXN2 modulates circadian cycle in Drosophila [182, 183]. Antisense oligonucleotides (ASO) of Ataxin-2 effectively improve motor functions in several SCA mouse models, suggesting ASO as a potential treatment for ATXN2-associated human neurodegenerative diseases [178, 184].
RNA Metabolism in AD
The neurofibrillary tangle caused by microtubule-associated protein, tau, is a well-known pathological feature of AD. Though there is still a controversy on whether tau is an RBP or not, it has been reported decades ago that RNA facilitates the formation of paired helical filaments of tau [185]. A study using a Tau-knock-out mouse model suggests that RNA-integrity in neurons is affected under heat shock with tau deficiency [186]. Recently, using PAR-iCLIP, Zhang et al. found that the major associated RNAs of tau are tRNAs [187]. Intriguingly, mixing tau and RNA in vitro created dynamic liquid droplets, which may be the mechanism underlying tau pathology [187]. Besides direct association with RNA, tau interacts with RBPs. Vanderweyde et al. reported that TIA-1 modulates tau pathology, and synergistically, they promote neuronal death [188]. Besides tau, other AD-associated proteins have been suggested to be regulated by RNA. Faghihi et al. reported that the concentration of β-secretase 1 (BACE1) antisense RNA is elevated in both postmortem human brains and Tg19959 mouse [189]. The elevation was associated with BACE1 mRNA stability and Aβ accumulation [189]. Knockdown of this RNA transcript induces neuronal differentiation [190].
Conclusion
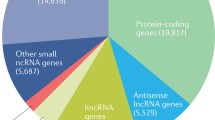
In the human genome of ~ 20,000 protein coding genes, about 7.5% directly associate with RNA and regulate different RNA processes. Many RBPs are implicated in human diseases and only a small fraction of RBPs are summarized in this review (Table 1). The rapid development of next-generation sequencing-based methods, such as RIP- and CLIP-based methods, ribosome profiling, in vivo RNA secondary structure profiling, and small and long RNA-seq, will help define the regulation of each RBP in the whole RNA network. However, many details of RNA regulatory mechanisms and their disease relevance remain to be determined. These studies will reveal novel targets and pathways that potentially facilitate new therapeutic development.
References
Bryant CD, Yazdani N. RNA-binding proteins, neural development and the addictions. Genes Brain Behav. 2016;15(1):169–86. https://doi.org/10.1111/gbb.12273.
Oswald F, Kloble P, Ruland A, Rosenkranz D, Hinz B, Butter F, et al. The FOXP2-driven network in developmental disorders and neurodegeneration. Front Cell Neurosci. 2017;11:212. https://doi.org/10.3389/fncel.2017.00212.
Maziuk B, Ballance HI, Wolozin B. Dysregulation of RNA binding protein aggregation in neurodegenerative disorders. Front Mol Neurosci. 2017;10:89. https://doi.org/10.3389/fnmol.2017.00089.
Barker HV, Niblock M, Lee YB, Shaw CE, Gallo JM. RNA misprocessing in C9orf72-linked neurodegeneration. Front Cell Neurosci. 2017;11:195. https://doi.org/10.3389/fncel.2017.00195.
Alkallas R, Fish L, Goodarzi H, Najafabadi HS. Inference of RNA decay rate from transcriptional profiling highlights the regulatory programs of Alzheimer’s disease. Nat Commun. 2017;8(1):909. https://doi.org/10.1038/s41467-017-00867-z.
Khermesh K, D'Erchia AM, Barak M, Annese A, Wachtel C, Levanon EY, et al. Reduced levels of protein recoding by A-to-I RNA editing in Alzheimer’s disease. RNA. 2016;22(2):290–302. https://doi.org/10.1261/rna.054627.115.
Brunello CA, Yan X, Huttunen HJ, Scheckel C, Drapeau E, Frias MA, et al. Internalized Tau sensitizes cells to stress by promoting formation and stability of stress granules. Sci Rep. 2016;6:30498. https://doi.org/10.1038/srep30498.
Scheckel C, Drapeau E, Frias MA, Park CY, Fak J, Zucker-Scharff I, et al. Regulatory consequences of neuronal ELAV-like protein binding to coding and non-coding RNAs in human brain. elife. 2016;5:e10421. https://doi.org/10.7554/eLife.10421.
Peretti D, Bastide A, Radford H, Verity N, Molloy C, Martin MG, et al. RBM3 mediates structural plasticity and protective effects of cooling in neurodegeneration. Nature. 2015;518(7538):236–9. https://doi.org/10.1038/nature14142.
Goodwin M, Mohan A, Batra R, Lee KY, Charizanis K, Fernandez Gomez FJ, et al. MBNL sequestration by toxic RNAs and RNA misprocessing in the myotonic dystrophy brain. Cell Rep. 2015;12(7):1159–68. https://doi.org/10.1016/j.celrep.2015.07.029.
Davis JK, Broadie K. Multifarious functions of the fragile X mental retardation protein. Trends Genet. 2017;33(10):703–14. https://doi.org/10.1016/j.tig.2017.07.008.
Gantois I, Khoutorsky A, Popic J, Aguilar-Valles A, Freemantle E, Cao R, et al. Metformin ameliorates core deficits in a mouse model of fragile X syndrome. Nat Med. 2017;23(6):674–7. https://doi.org/10.1038/nm.4335.
Donlin-Asp PG, Rossoll W, Bassell GJ. Spatially and temporally regulating translation via mRNA-binding proteins in cellular and neuronal function. FEBS Lett. 2017;591(11):1508–25. https://doi.org/10.1002/1873-3468.12621.
El Fatimy R, Davidovic L, Tremblay S, Jaglin X, Dury A, Robert C, et al. Tracking the fragile X mental retardation protein in a highly ordered neuronal ribonucleoparticles population: a link between stalled polyribosomes and RNA granules. PLoS Genet. 2016;12(7):e1006192. https://doi.org/10.1371/journal.pgen.1006192.
Richter JD, Bassell GJ, Klann E. Dysregulation and restoration of translational homeostasis in fragile X syndrome. Nat Rev Neurosci. 2015;16(10):595–605. https://doi.org/10.1038/nrn4001.
Edupuganti RR, Geiger S, Lindeboom RGH, Shi H, Hsu PJ, Lu Z, et al. N6-methyladenosine (m6A) recruits and repels proteins to regulate mRNA homeostasis. Nat Struct Mol Biol. 2017;24(10):870–8. https://doi.org/10.1038/nsmb.3462.
Hinney A, Albayrak Ö, Antel J, Volckmar A-L, Sims R, Chapman J, et al. Genetic variation at the CELF1 (CUGBP, elav-like family member 1 gene) locus is genome-wide associated with Alzheimer’s disease and obesity. Am J Med Genet B Neuropsychiatr Genet. 2014;165(4):283–93. https://doi.org/10.1002/ajmg.b.32234.
Berto S, Usui N, Konopka G, Fogel BL. ELAVL2-regulated transcriptional and splicing networks in human neurons link neurodevelopment and autism. Hum Mol Genet. 2016;25(12):2451–64. https://doi.org/10.1093/hmg/ddw110.
Barone R, Fichera M, De Grandi M, Battaglia M, Lo Faro V, Mattina T, et al. Familial 18q12.2 deletion supports the role of RNA-binding protein CELF4 in autism spectrum disorders. Am J Med Genet A. 2017;173(6):1649–55. https://doi.org/10.1002/ajmg.a.38205.
Wagnon JL, Mahaffey CL, Sun W, Yang Y, Chao HT, Frankel WN. Etiology of a genetically complex seizure disorder in Celf4 mutant mice. Genes Brain Behav. 2011;10(7):765–77. https://doi.org/10.1111/j.1601-183X.2011.00717.x.
Wagnon JL, Briese M, Sun W, Mahaffey CL, Curk T, Rot G, et al. CELF4 regulates translation and local abundance of a vast set of mRNAs, including genes associated with regulation of synaptic function. PLoS Genet. 2012;8(11):e1003067. https://doi.org/10.1371/journal.pgen.1003067.
Conboy JG. Developmental regulation of RNA processing by Rbfox proteins. Wiley interdisciplinary reviews. RNA. 2017;8(2):e1398. https://doi.org/10.1002/wrna.1398.
Lee JA, Damianov A, Lin CH, Fontes M, Parikshak NN, Anderson ES, et al. Cytoplasmic Rbfox1 regulates the expression of synaptic and autism-related genes. Neuron. 2016;89(1):113–28. https://doi.org/10.1016/j.neuron.2015.11.025.
Lal D, Pernhorst K, Klein KM, Reif P, Tozzi R, Toliat MR, et al. Extending the phenotypic spectrum of RBFOX1 deletions: sporadic focal epilepsy. Epilepsia. 2015;56(9):e129–33. https://doi.org/10.1111/epi.13076.
Hamada N, Ito H, Nishijo T, Iwamoto I, Morishita R, Tabata H, et al. Essential role of the nuclear isoform of RBFOX1, a candidate gene for autism spectrum disorders, in the brain development. Sci Rep. 2016;6:30805. https://doi.org/10.1038/srep30805.
Weyn-Vanhentenryck SM, Mele A, Yan Q, Sun S, Farny N, Zhang Z, et al. HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Rep. 2014;6(6):1139–52. https://doi.org/10.1016/j.celrep.2014.02.005.
Xiong HY, Alipanahi B, Lee LJ, Bretschneider H, Merico D, Yuen RK, et al. RNA splicing. The human splicing code reveals new insights into the genetic determinants of disease. Science. 2015;347(6218):1254806. https://doi.org/10.1126/science.1254806.
Bono F, Ebert J, Lorentzen E, Conti E. The crystal structure of the exon junction complex reveals how it maintains a stable grip on mRNA. Cell. 2006;126(4):713–25. https://doi.org/10.1016/j.cell.2006.08.006.
Andersen CB, Ballut L, Johansen JS, Chamieh H, Nielsen KH, Oliveira CL, et al. Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science. 2006;313(5795):1968–72. https://doi.org/10.1126/science.1131981.
Nguyen LS, Kim HG, Rosenfeld JA, Shen Y, Gusella JF, Lacassie Y, et al. Contribution of copy number variants involving nonsense-mediated mRNA decay pathway genes to neuro-developmental disorders. Hum Mol Genet. 2013;22(9):1816–25. https://doi.org/10.1093/hmg/ddt035.
Zou D, McSweeney C, Sebastian A, Reynolds DJ, Dong F, Zhou Y, et al. A critical role of RBM8a in proliferation and differentiation of embryonic neural progenitors. Neural Dev. 2015;10:18. https://doi.org/10.1186/s13064-015-0045-7.
Mao H, McMahon JJ, Tsai YH, Wang Z, Silver DL. Haploinsufficiency for Core exon junction complex components disrupts embryonic neurogenesis and causes p53-mediated microcephaly. PLoS Genet. 2016;12(9):e1006282. https://doi.org/10.1371/journal.pgen.1006282.
Mao H, Pilaz LJ, McMahon JJ, Golzio C, Wu D, Shi L, et al. Rbm8a haploinsufficiency disrupts embryonic cortical development resulting in microcephaly. J Neurosci Off J Soc Neurosci. 2015;35(18):7003–18. https://doi.org/10.1523/JNEUROSCI.0018-15.2015.
Silver DL, Watkins-Chow DE, Schreck KC, Pierfelice TJ, Larson DM, Burnetti AJ, et al. The exon junction complex component Magoh controls brain size by regulating neural stem cell division. Nat Neurosci. 2010;13(5):551–8. https://doi.org/10.1038/nn.2527.
Alachkar A, Jiang D, Harrison M, Zhou Y, Chen G, Mao Y. An EJC factor RBM8a regulates anxiety behaviors. Curr Mol Med. 2013;13(6):887–99.
He F, Jacobson A. Nonsense-mediated mRNA decay: degradation of defective transcripts is only part of the story. Annu Rev Genet. 2015;49:339–66. https://doi.org/10.1146/annurev-genet-112414-054639.
Jolly LA, Homan CC, Jacob R, Barry S, Gecz J. The UPF3B gene, implicated in intellectual disability, autism, ADHD and childhood onset schizophrenia regulates neural progenitor cell behaviour and neuronal outgrowth. Hum Mol Genet. 2013;22(23):4673–87. https://doi.org/10.1093/hmg/ddt315.
Tarpey PS, Raymond FL, Nguyen LS, Rodriguez J, Hackett A, Vandeleur L, et al. Mutations in UPF3B, a member of the nonsense-mediated mRNA decay complex, cause syndromic and nonsyndromic mental retardation. Nat Genet. 2007;39(9):1127–33. https://doi.org/10.1038/ng2100.
Addington AM, Gauthier J, Piton A, Hamdan FF, Raymond A, Gogtay N, et al. A novel frameshift mutation in UPF3B identified in brothers affected with childhood onset schizophrenia and autism spectrum disorders. Mol Psychiatry. 2011;16(3):238–9. https://doi.org/10.1038/mp.2010.59.
Laumonnier F, Shoubridge C, Antar C, Nguyen LS, Van Esch H, Kleefstra T, et al. Mutations of the UPF3B gene, which encodes a protein widely expressed in neurons, are associated with nonspecific mental retardation with or without autism. Mol Psychiatry. 2010;15(7):767–76. https://doi.org/10.1038/mp.2009.14.
Huang L, Shum EY, Jones SH, Lou CH, Dumdie J, Kim H, et al. A Upf3b-mutant mouse model with behavioral and neurogenesis defects. Mol Psychiatry. 2017; https://doi.org/10.1038/mp.2017.173.
Shum EY, Jones SH, Shao A, Dumdie J, Krause MD, Chan WK, et al. The antagonistic gene paralogs Upf3a and Upf3b govern nonsense-mediated RNA decay. Cell. 2016;165(2):382–95. https://doi.org/10.1016/j.cell.2016.02.046.
Graber TE, Freemantle E, Anadolu MN, Hebert-Seropian S, MacAdam RL, Shin U, et al. UPF1 governs synaptic plasticity through association with a STAU2 RNA granule. J Neurosci Off J Soc Neurosci. 2017;37(38):9116–31. https://doi.org/10.1523/JNEUROSCI.0088-17.2017.
Mellios N, Feldman DA, Sheridan SD, Ip JPK, Kwok S, Amoah SK, et al. MeCP2-regulated miRNAs control early human neurogenesis through differential effects on ERK and AKT signaling. Mol Psychiatry. 2017;23:1051–65. https://doi.org/10.1038/mp.2017.86.
Sztainberg Y, Chen HM, Swann JW, Hao S, Tang B, Wu Z, et al. Reversal of phenotypes in MECP2 duplication mice using genetic rescue or antisense oligonucleotides. Nature. 2015;528(7580):123–6. https://doi.org/10.1038/nature16159.
Khan AW, Ziemann M, Rafehi H, Maxwell S, Ciccotosto GD, El-Osta A. MeCP2 interacts with chromosomal microRNAs in brain. Epigenetics. 2017;12(12):1028–37. https://doi.org/10.1080/15592294.2017.1391429.
Jeffery L, Nakielny S. Components of the DNA methylation system of chromatin control are RNA-binding proteins. J Biol Chem. 2004;279(47):49479–87. https://doi.org/10.1074/jbc.M409070200.
Jobe EM, Gao Y, Eisinger BE, Mladucky JK, Giuliani CC, Kelnhofer LE, et al. Methyl-CpG-binding protein MBD1 regulates neuronal lineage commitment through maintaining adult neural stem cell identity. J Neurosci. 2017;37(3):523–36. https://doi.org/10.1523/jneurosci.1075-16.2016.
Liu C, Teng Z-Q, Santistevan NJ, Szulwach KE, Guo W, Jin P, et al. Epigenetic regulation of miR-184 by MBD1 governs neural stem cell proliferation and differentiation. Cell Stem Cell. 2010;6(5):433–44. https://doi.org/10.1016/j.stem.2010.02.017.
Li H, Zhong X, Chau KF, Santistevan NJ, Guo W, Kong G, et al. Cell cycle-linked MeCP2 phosphorylation modulates adult neurogenesis involving the notch signalling pathway. Nat Commun. 2014;5:5601. https://doi.org/10.1038/ncomms6601. https://www.nature.com/articles/ncomms6601#supplementary-information
Geuens T, Bouhy D, Timmerman V. The hnRNP family: insights into their role in health and disease. Hum Genet. 2016;135(8):851–67. https://doi.org/10.1007/s00439-016-1683-5.
Bain JM, Cho MT, Telegrafi A, Wilson A, Brooks S, Botti C, et al. Variants in HNRNPH2 on the X chromosome are associated with a neurodevelopmental disorder in females. Am J Hum Genet. 2016;99(3):728–34. https://doi.org/10.1016/j.ajhg.2016.06.028.
Du X, An Y, Yu L, Liu R, Qin Y, Guo X, et al. A genomic copy number variant analysis implicates the MBD5 and HNRNPU genes in Chinese children with infantile spasms and expands the clinical spectrum of 2q23.1 deletion. BMC Med Genet. 2014;15:62. https://doi.org/10.1186/1471-2350-15-62.
Poot M, Kas MJ. Antisense may make sense of 1q44 deletions, seizures, and HNRNPU. Am J Med Genet A. 2013;161A(4):910–2. https://doi.org/10.1002/ajmg.a.35770.
Vidaki M, Drees F, Saxena T, Lanslots E, Taliaferro MJ, Tatarakis A, et al. A requirement for Mena, an actin regulator, in local mRNA translation in developing neurons. Neuron. 2017;95(3):608–22 e5. https://doi.org/10.1016/j.neuron.2017.06.048.
Vidal RL, Valenzuela JI, Lujan R, Couve A. Cellular and subcellular localization of Marlin-1 in the brain. BMC Neurosci. 2009;10:37. https://doi.org/10.1186/1471-2202-10-37.
Nishimura Y, Martin CL, Vazquez-Lopez A, Spence SJ, Alvarez-Retuerto AI, Sigman M, et al. Genome-wide expression profiling of lymphoblastoid cell lines distinguishes different forms of autism and reveals shared pathways. Hum Mol Genet. 2007;16(14):1682–98. https://doi.org/10.1093/hmg/ddm116.
Berg JM, Lee C, Chen L, Galvan L, Cepeda C, Chen JY, et al. JAKMIP1, a novel regulator of neuronal translation, modulates synaptic function and autistic-like behaviors in mouse. Neuron. 2015;88(6):1173–91. https://doi.org/10.1016/j.neuron.2015.10.031.
Neves-Pereira M, Muller B, Massie D, Williams JH, O'Brien PC, Hughes A, et al. Deregulation of EIF4E: a novel mechanism for autism. J Med Genet. 2009;46(11):759–65. https://doi.org/10.1136/jmg.2009.066852.
Millar JK, Christie S, Semple CA, Porteous DJ. Chromosomal location and genomic structure of the human translin-associated factor X gene (TRAX; TSNAX) revealed by intergenic splicing to DISC1, a gene disrupted by a translocation segregating with schizophrenia. Genomics. 2000;67(1):69–77.
Blackwood DH, Fordyce A, Walker MT, St Clair DM, Porteous DJ, Muir WJ. Schizophrenia and affective disorders—cosegregation with a translocation at chromosome 1q42 that directly disrupts brain-expressed genes: clinical and P300 findings in a family. Am J Hum Genet. 2001;69(2):428–33.
Yates D. Psychiatric disorders: multiple pathways to DISC1-related disease? Nat Rev Neurosci. 2011;13(1):4–5. https://doi.org/10.1038/nrn3166.
Brandon NJ, Sawa A. Linking neurodevelopmental and synaptic theories of mental illness through DISC1. Nat Rev Neurosci. 2011;12(12):707–22. https://doi.org/10.1038/nrn3120.
Mao Y, Ge X, Frank CL, Madison JM, Koehler AN, Doud MK, et al. Disrupted in schizophrenia 1 regulates neuronal progenitor proliferation via modulation of GSK3beta/beta-catenin signaling. Cell. 2009;136(6):1017–31. https://doi.org/10.1016/j.cell.2008.12.044.
Singh KK, Ge X, Mao Y, Drane L, Meletis K, Samuels BA, et al. Dixdc1 is a critical regulator of DISC1 and embryonic cortical development. Neuron. 2010;67(1):33–48. https://doi.org/10.1016/j.neuron.2010.06.002.
Ishizuka K, Kamiya A, Oh EC, Kanki H, Seshadri S, Robinson JF, et al. DISC1-dependent switch from progenitor proliferation to migration in the developing cortex. Nature. 2011;473(7345):92–6. https://doi.org/10.1038/nature09859.
Srikanth P, Han K, Callahan DG, Makovkina E, Muratore CR, Lalli MA, et al. Genomic DISC1 disruption in hiPSCs alters Wnt signaling and neural cell fate. Cell Rep. 2015;12(9):1414–29. https://doi.org/10.1016/j.celrep.2015.07.061.
De Rienzo G, Bishop JA, Mao Y, Pan L, Ma TP, Moens CB, et al. Disc1 regulates both beta-catenin-mediated and noncanonical Wnt signaling during vertebrate embryogenesis. FASEB J. 2011;25(12):4184–97. https://doi.org/10.1096/fj.11-186239.
Tsuboi D, Kuroda K, Tanaka M, Namba T, Iizuka Y, Taya S, et al. Disrupted-in-schizophrenia 1 regulates transport of ITPR1 mRNA for synaptic plasticity. Nat Neurosci. 2015;18(5):698–707. https://doi.org/10.1038/nn.3984.
O'Donovan MC, Craddock N, Norton N, Williams H, Peirce T, Moskvina V, et al. Identification of loci associated with schizophrenia by genome-wide association and follow-up. Nat Genet. 2008;40(9):1053–5. https://doi.org/10.1038/ng.201.
Zhang R, Lu SM, Qiu C, Liu XG, Gao CG, Guo TW, et al. Population-based and family-based association studies of ZNF804A locus and schizophrenia. Mol Psychiatry. 2011;16(4):360–1.
Williams HJ, Norton N, Dwyer S, Moskvina V, Nikolov I, Carroll L, et al. Fine mapping of ZNF804A and genome-wide significant evidence for its involvement in schizophrenia and bipolar disorder. Mol Psychiatry. 2011;16(4):429–41. https://doi.org/10.1038/mp.2010.36.
Steinberg S, Mors O, Borglum AD, Gustafsson O, Werge T, Mortensen PB, et al. Expanding the range of ZNF804A variants conferring risk of psychosis. Mol Psychiatry. 2011;16(1):59–66.
Riley B, Thiselton D, Maher BS, Bigdeli T, Wormley B, McMichael GO, et al. Replication of association between schizophrenia and ZNF804A in the Irish Case-Control Study of Schizophrenia sample. Mol Psychiatry. 2010;15(1):29–37.
Schwab SG, Kusumawardhani A, Dai N, Qin W, Wildenauer MDB, Agiananda F, et al. Association of rs1344706 in the ZNF804A gene with schizophrenia in a case/control sample from Indonesia. Schizophr Res. 2013;147(1):46–52. https://doi.org/10.1016/j.schres.2013.03.022.
Xiao X, Luo XJ, Chang H, Liu Z, Li M. Evaluation of European schizophrenia GWAS loci in Asian populations via comprehensive meta-analyses. Mol Neurobiol. 2017;54(6):4071–80. https://doi.org/10.1007/s12035-016-9990-3.
Huang L, Ohi K, Chang H, Yu H, Wu L, Yue W, et al. A comprehensive meta-analysis of ZNF804A SNPs in the risk of schizophrenia among Asian populations. Am J Med Genet B Neuropsychiatr Genet. 2016;171b(3):437–46. https://doi.org/10.1002/ajmg.b.32425.
Consortium SWGotPG. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511(7510):421–7. https://doi.org/10.1038/nature13595.
Ripke S, O’Dushlaine C, Chambert K, Moran JL, Kähler AK, Akterin S, et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet. 2013;45(10):1150–9.
Griswold AJ, Ma D, Cukier HN, Nations LD, Schmidt MA, Chung RH, et al. Evaluation of copy number variations reveals novel candidate genes in autism spectrum disorder-associated pathways. Hum Mol Genet. 2012;21(15):3513–23. https://doi.org/10.1093/hmg/dds164.
Anitha A, Thanseem I, Nakamura K, Vasu MM, Yamada K, Ueki T, et al. Zinc finger protein 804A (ZNF804A) and verbal deficits in individuals with autism. J Psychiatry Neurosci. 2014;39(5):294–303.
Talkowski ME, Rosenfeld JA, Blumenthal I, Pillalamarri V, Chiang C, Heilbut A, et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell. 2012;149(3):525–37. https://doi.org/10.1016/j.cell.2012.03.028.
Blake J, Riddell A, Theiss S, Gonzalez AP, Haase B, Jauch A, et al. Sequencing of a patient with balanced chromosome abnormalities and neurodevelopmental disease identifies disruption of multiple high risk loci by structural variation. PLoS One. 2014;9(3):e90894. https://doi.org/10.1371/journal.pone.0090894.
Guella I, Sequeira A, Rollins B, Morgan L, Myers RM, Watson SJ, et al. Evidence of allelic imbalance in the schizophrenia susceptibility gene ZNF804A in human dorsolateral prefrontal cortex. Schizophr Res. 2014;152(1):111–6. https://doi.org/10.1016/j.schres.2013.11.021.
Wei Q, Kang Z, Diao F, Shan B, Li L, Zheng L, et al. Association of the ZNF804A gene polymorphism rs1344706 with white matter density changes in Chinese schizophrenia. Prog Neuro-Psychopharmacol Biol Psychiatry. 2012;36(1):122–7. https://doi.org/10.1016/j.pnpbp.2011.08.021.
Wassink TH, Epping EA, Rudd D, Axelsen M, Ziebell S, Fleming FW, et al. Influence of ZNF804a on brain structure volumes and symptom severity in individuals with schizophrenia. Arch Gen Psychiatry. 2012;69(9):885–92.
Rasetti R, Sambataro F, Chen Q, Callicott JH, Mattay VS, Weinberger DR. Altered cortical network dynamics: a potential intermediate phenotype for schizophrenia and association with ZNF804A. Arch Gen Psychiatry. 2011;68(12):1207–17. https://doi.org/10.1001/archgenpsychiatry.2011.103.
Paulus FM, Krach S, Bedenbender J, Pyka M, Sommer J, Krug A, et al. Partial support for ZNF804A genotype-dependent alterations in prefrontal connectivity. Hum Brain Mapp. 2013;34(2):304–13. https://doi.org/10.1002/hbm.21434.
Esslinger C, Walter H, Kirsch P, Erk S, Schnell K, Arnold C, et al. Neural mechanisms of a genome-wide supported psychosis variant. Science. 2009;324(5927):605.
Esslinger C, Kirsch P, Haddad L, Mier D, Sauer C, Erk S, et al. Cognitive state and connectivity effects of the genome-wide significant psychosis variant in ZNF804A. NeuroImage. 2011;54(3):2514–23. https://doi.org/10.1016/j.neuroimage.2010.10.012.
Del Re EC, Bergen SE, Mesholam-Gately RI, Niznikiewicz MA, Goldstein JM, Woo TU, et al. Analysis of schizophrenia-related genes and electrophysiological measures reveals ZNF804A association with amplitude of P300b elicited by novel sounds. Transl Psychiatry. 2013;4:e346. https://doi.org/10.1038/tp.2013.117.
Hashimoto R, Ohi K, Yasuda Y, Fukumoto M, Iwase M, Iike N, et al. The impact of a genome-wide supported psychosis variant in the ZNF804A gene on memory function in schizophrenia. Am J Med Genet B Neuropsychiatr Genet. 2010;153b(8):1459–64. https://doi.org/10.1002/ajmg.b.31123.
Walters JT, Corvin A, Owen MJ, Williams H, Dragovic M, Quinn EM, et al. Psychosis susceptibility gene ZNF804A and cognitive performance in schizophrenia. Arch Gen Psychiatry. 2010;67(7):692–700. https://doi.org/10.1001/archgenpsychiatry.2010.81.
Balog Z, Kiss I, Keri S. ZNF804A may be associated with executive control of attention. Genes Brain Behav. 2011;10(2):223–7. https://doi.org/10.1111/j.1601-183X.2010.00657.x.
Donohoe G, Rose E, Frodl T, Morris D, Spoletini I, Adriano F, et al. ZNF804A risk allele is associated with relatively intact gray matter volume in patients with schizophrenia. NeuroImage. 2011;54(3):2132–7. https://doi.org/10.1016/j.neuroimage.2010.09.089.
Chen M, Xu Z, Zhai J, Bao X, Zhang Q, Gu H, et al. Evidence of IQ-modulated association between ZNF804A gene polymorphism and cognitive function in schizophrenia patients. Neuropsychopharmacology. 2012;37(7):1572–8. https://doi.org/10.1038/npp.2012.1.
Kuswanto CN, Woon PS, Zheng XB, Qiu A, Sitoh YY, Chan YH, et al. Genome-wide supported psychosis risk variant in ZNF804A gene and impact on cortico-limbic WM integrity in schizophrenia. Am J Med Genet B Neuropsychiatr Genet. 2012;159b(3):255–62. https://doi.org/10.1002/ajmg.b.32032.
Mossner R, Schuhmacher A, Wagner M, Lennertz L, Steinbrecher A, Quednow BB, et al. The schizophrenia risk gene ZNF804A influences the antipsychotic response of positive schizophrenia symptoms. Eur Arch Psychiatry Clin Neurosci. 2012;262(3):193–7. https://doi.org/10.1007/s00406-011-0235-1.
Sprooten E, McIntosh AM, Lawrie SM, Hall J, Sussmann JE, Dahmen N, et al. An investigation of a genomewide supported psychosis variant in ZNF804A and white matter integrity in the human brain. Magn Reson Imaging. 2012;30(10):1373–80. https://doi.org/10.1016/j.mri.2012.05.013.
Del Re EC, Bergen SE, Mesholam-Gately RI, Niznikiewicz MA, Goldstein JM, Woo TU, et al. Analysis of schizophrenia-related genes and electrophysiological measures reveals ZNF804A association with amplitude of P300b elicited by novel sounds. Transl Psychiatry. 2014;4:e346. https://doi.org/10.1038/tp.2013.117.
Ikuta T, Peters BD, Guha S, John M, Karlsgodt KH, Lencz T, et al. A schizophrenia risk gene, ZNF804A, is associated with brain white matter microstructure. Schizophr Res. 2014;155(1–3):15–20. https://doi.org/10.1016/j.schres.2014.03.001.
Mohnke S, Erk S, Schnell K, Schutz C, Romanczuk-Seiferth N, Grimm O, et al. Further evidence for the impact of a genome-wide-supported psychosis risk variant in ZNF804A on the theory of mind network. Neuropsychopharmacology. 2014;39(5):1196–205. https://doi.org/10.1038/npp.2013.321.
Nicodemus KK, Hargreaves A, Morris D, Anney R, Gill M, Corvin A, et al. Variability in working memory performance explained by epistasis vs polygenic scores in the ZNF804A pathway. JAMA Psychiatry. 2014;71(7):778–85. https://doi.org/10.1001/jamapsychiatry.2014.528.
Cousijn H, Tunbridge EM, Rolinski M, Wallis G, Colclough GL, Woolrich MW, et al. Modulation of hippocampal theta and hippocampal-prefrontal cortex function by a schizophrenia risk gene. Hum Brain Mapp. 2015;36(6):2387–95. https://doi.org/10.1002/hbm.22778.
Wickramasinghe A, Tulloch AD, Hayes RD, Chang CK, Broadbent M, Di Forti M, et al. Associations between the schizophrenia susceptibility gene ZNF804A and clinical outcomes in psychosis. Transl Psychiatry. 2015;5:e698. https://doi.org/10.1038/tp.2015.198.
Mallas EJ, Carletti F, Chaddock CA, Woolley J, Picchioni MM, Shergill SS, et al. Genome-wide discovered psychosis-risk gene ZNF804A impacts on white matter microstructure in health, schizophrenia and bipolar disorder. PeerJ. 2016;4:e1570. https://doi.org/10.7717/peerj.1570.
Mallas E, Carletti F, Chaddock CA, Shergill S, Woolley J, Picchioni MM, et al. The impact of CACNA1C gene, and its epistasis with ZNF804A, on white matter microstructure in health, schizophrenia and bipolar disorder(1). Genes Brain Behav. 2017;16(4):479–88. https://doi.org/10.1111/gbb.12355.
Squarcina L, Houenou J, Altamura AC, Soares J, Brambilla P. Association of increased genotypes risk for bipolar disorder with brain white matter integrity investigated with tract-based spatial statistics: special section on “Translational and Neuroscience Studies in Affective Disorders”. Section editor, Maria Nobile MD, PhD. This section of JAD focuses on the relevance of translational and neuroscience studies in providing a better understanding of the neural basis of affective disorders. The main aim is to briefly summarise relevant research findings in clinical neuroscience with particular regards to specific innovative topics in mood and anxiety disorders. J Affect Disord. 2017;221:312–7. https://doi.org/10.1016/j.jad.2017.06.031.
Tecelao D, Mendes A, Martins D, Bramon E, Toulopoulou T, Kravariti E, et al. The impact of psychosis genome-wide associated ZNF804A variation on verbal fluency connectivity. J Psychiatr Res. 2017;98:17–21. https://doi.org/10.1016/j.jpsychires.2017.12.005.
Xu Q, Xiong Y, Yuan C, Liu F, Zhao F, Shen J, et al. ZNF804A rs1344706 interacts with COMT rs4680 to affect prefrontal volume in healthy adults. Brain Imaging Behav. 2017;12:13–9. https://doi.org/10.1007/s11682-016-9671-x.
Girgenti MJ, LoTurco JJ, Maher BJ. ZNF804a regulates expression of the schizophrenia-associated genes PRSS16, COMT, PDE4B, and DRD2. PLoS One. 2012;7(2):e32404.
Zhou Y, Dong F, Lanz TA, Reinhart V, Li M, Liu L, et al. Interactome analysis reveals ZNF804A, a schizophrenia risk gene, as a novel component of protein translational machinery critical for embryonic neurodevelopment. Mol Psychiatry. 2017;23:952–62. https://doi.org/10.1038/mp.2017.166.
Deans PJM, Raval P, Sellers KJ, Gatford NJF, Halai S, Duarte RRR, et al. Psychosis risk candidate ZNF804A localizes to synapses and regulates neurite formation and dendritic spine structure. Biol Psychiatry. 2017;82(1):49–61. https://doi.org/10.1016/j.biopsych.2016.08.038.
Darbelli L, Richard S. Emerging functions of the quaking RNA-binding proteins and link to human diseases. Wiley interdisciplinary reviews. RNA. 2016;7(3):399–412. https://doi.org/10.1002/wrna.1344.
Iwata K, Matsuzaki H, Manabe T, Mori N. Altering the expression balance of hnRNP C1 and C2 changes the expression of myelination-related genes. Psychiatry Res. 2011;190(2–3):364–6. https://doi.org/10.1016/j.psychres.2011.05.043.
Radomska KJ, Halvardson J, Reinius B, Lindholm Carlstrom E, Emilsson L, Feuk L, et al. RNA-binding protein QKI regulates glial fibrillary acidic protein expression in human astrocytes. Hum Mol Genet. 2013;22(7):1373–82. https://doi.org/10.1093/hmg/dds553.
Fromer M, Pocklington AJ, Kavanagh DH, Williams HJ, Dwyer S, Gormley P, et al. De novo mutations in schizophrenia implicate synaptic networks. Nature. 2014;506(7487):179–84. https://doi.org/10.1038/nature12929.
Del'Guidice T, Latapy C, Rampino A, Khlghatyan J, Lemasson M, Gelao B, et al. FXR1P is a GSK3beta substrate regulating mood and emotion processing. Proc Natl Acad Sci U S A. 2015;112(33):E4610–9. https://doi.org/10.1073/pnas.1506491112.
Ferrari R, Kapogiannis D, Huey ED, Momeni P. FTD and ALS: a tale of two diseases. Curr Alzheimer Res. 2011;8(3):273–94.
Weishaupt JH, Hyman T, Dikic I. Common molecular pathways in amyotrophic lateral sclerosis and frontotemporal dementia. Trends Mol Med. 2016;22(9):769–83. https://doi.org/10.1016/j.molmed.2016.07.005.
Heraud-Farlow JE, Kiebler MA. The multifunctional Staufen proteins: conserved roles from neurogenesis to synaptic plasticity. Trends Neurosci. 2014;37(9):470–9. https://doi.org/10.1016/j.tins.2014.05.009.
Gershoni-Emek N, Mazza A, Chein M, Gradus-Pery T, Xiang X, Li KW, et al. Proteomic analysis of dynein-interacting proteins in amyotrophic lateral sclerosis synaptosomes reveals alterations in the RNA-binding protein Staufen1. Mol Cell Proteomics. 2016;15(2):506–22. https://doi.org/10.1074/mcp.M115.049965.
Kwiatkowski TJ, Bosco DA, LeClerc AL, Tamrazian E, Vanderburg CR, Russ C, et al. Mutations in the <em>FUS/TLS</em> gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science. 2009;323(5918):1205–8. https://doi.org/10.1126/science.1166066.
Lee EB, Lee VM, Trojanowski JQ. Gains or losses: molecular mechanisms of TDP43-mediated neurodegeneration. Nat Rev Neurosci. 2011;13(1):38–50. https://doi.org/10.1038/nrn3121.
Winton MJ, Igaz LM, Wong MM, Kwong LK, Trojanowski JQ, Lee VM. Disturbance of nuclear and cytoplasmic TAR DNA-binding protein (TDP-43) induces disease-like redistribution, sequestration, and aggregate formation. J Biol Chem. 2008;283(19):13302–9. https://doi.org/10.1074/jbc.M800342200.
Ayala YM, Zago P, D'Ambrogio A, Xu YF, Petrucelli L, Buratti E, et al. Structural determinants of the cellular localization and shuttling of TDP-43. J Cell Sci. 2008;121(Pt 22):3778–85. https://doi.org/10.1242/jcs.038950.
Afroz T, Hock EM, Ernst P, Foglieni C, Jambeau M, Gilhespy LAB, et al. Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation. Nat Commun. 2017;8(1):45. https://doi.org/10.1038/s41467-017-00062-0.
Stoica R, De Vos KJ, Paillusson S, Mueller S, Sancho RM, Lau KF, et al. ER-mitochondria associations are regulated by the VAPB-PTPIP51 interaction and are disrupted by ALS/FTD-associated TDP-43. Nat Commun. 2014;5:3996. https://doi.org/10.1038/ncomms4996.
Alami NH, Smith RB, Carrasco MA, Williams LA, Winborn CS, Han SSW, et al. Axonal transport of TDP-43 mRNA granules is impaired by ALS-causing mutations. Neuron. 2014;81(3):536–43. https://doi.org/10.1016/j.neuron.2013.12.018.
Rogelj B, Easton LE, Bogu GK, Stanton LW, Rot G, Curk T, et al. Widespread binding of FUS along nascent RNA regulates alternative splicing in the brain. Sci Rep. 2012;2:603. https://doi.org/10.1038/srep00603.
Honda D, Ishigaki S, Iguchi Y, Fujioka Y, Udagawa T, Masuda A, et al. The ALS/FTLD-related RNA-binding proteins TDP-43 and FUS have common downstream RNA targets in cortical neurons. FEBS Open Bio. 2013;4:1–10. https://doi.org/10.1016/j.fob.2013.11.001.
Schwartz JC, Podell ER, Han SS, Berry JD, Eggan KC, Cech TR. FUS is sequestered in nuclear aggregates in ALS patient fibroblasts. Mol Biol Cell. 2014;25(17):2571–8. https://doi.org/10.1091/mbc.E14-05-1007.
Patel A, Lee HO, Jawerth L, Maharana S, Jahnel M, Hein MY, et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell. 2015;162(5):1066–77. https://doi.org/10.1016/j.cell.2015.07.047.
Murakami T, Qamar S, Lin JQ, Schierle GS, Rees E, Miyashita A, et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron. 2015;88(4):678–90. https://doi.org/10.1016/j.neuron.2015.10.030.
Dormann D, Rodde R, Edbauer D, Bentmann E, Fischer I, Hruscha A, et al. ALS-associated fused in sarcoma (FUS) mutations disrupt transportin-mediated nuclear import. EMBO J. 2010;29(16):2841–57. https://doi.org/10.1038/emboj.2010.143.
Shiihashi G, Ito D, Yagi T, Nihei Y, Ebine T, Suzuki N. Mislocated FUS is sufficient for gain-of-toxic-function amyotrophic lateral sclerosis phenotypes in mice. Brain. 2016;139(Pt 9):2380–94. https://doi.org/10.1093/brain/aww161.
Takanashi K, Yamaguchi A. Aggregation of ALS-linked FUS mutant sequesters RNA binding proteins and impairs RNA granules formation. Biochem Biophys Res Commun. 2014;452(3):600–7. https://doi.org/10.1016/j.bbrc.2014.08.115.
Sun S, Ling SC, Qiu J, Albuquerque CP, Zhou Y, Tokunaga S, et al. ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP. Nat Commun. 2015;6:6171. https://doi.org/10.1038/ncomms7171.
Qiu H, Lee S, Shang Y, Wang WY, Au KF, Kamiya S, et al. ALS-associated mutation FUS-R521C causes DNA damage and RNA splicing defects. J Clin Invest. 2014;124(3):981–99. https://doi.org/10.1172/JCI72723.
Yasuda K, Zhang H, Loiselle D, Haystead T, Macara IG, Mili S. The RNA-binding protein Fus directs translation of localized mRNAs in APC-RNP granules. J Cell Biol. 2013;203(5):737–46. https://doi.org/10.1083/jcb.201306058.
Gonatas NK, Stieber A, Mourelatos Z, Chen Y, Gonatas JO, Appel SH, et al. Fragmentation of the Golgi apparatus of motor neurons in amyotrophic lateral sclerosis. Am J Pathol. 1992;140(3):731–7.
Mourelatos Z, Adler H, Hirano A, Donnenfeld H, Gonatas JO, Gonatas NK. Fragmentation of the Golgi apparatus of motor neurons in amyotrophic lateral sclerosis revealed by organelle-specific antibodies. Proc Natl Acad Sci U S A. 1990;87(11):4393–5.
Farg MA, Soo KY, Warraich ST, Sundaramoorthy V, Blair IP, Atkin JD. Ataxin-2 interacts with FUS and intermediate-length polyglutamine expansions enhance FUS-related pathology in amyotrophic lateral sclerosis. Hum Mol Genet. 2013;22(4):717–28. https://doi.org/10.1093/hmg/dds479.
Suarez-Calvet M, Neumann M, Arzberger T, Abou-Ajram C, Funk E, Hartmann H, et al. Monomethylated and unmethylated FUS exhibit increased binding to transportin and distinguish FTLD-FUS from ALS-FUS. Acta Neuropathol. 2016;131(4):587–604. https://doi.org/10.1007/s00401-016-1544-2.
Donnelly CJ, Zhang PW, Pham JT, Haeusler AR, Mistry NA, Vidensky S, et al. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron. 2013;80(2):415–28. https://doi.org/10.1016/j.neuron.2013.10.015.
Lee YB, Chen HJ, Peres JN, Gomez-Deza J, Attig J, Stalekar M, et al. Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep. 2013;5(5):1178–86. https://doi.org/10.1016/j.celrep.2013.10.049.
Walker C, Herranz-Martin S, Karyka E, Liao C, Lewis K, Elsayed W, et al. C9orf72 expansion disrupts ATM-mediated chromosomal break repair. Nat Neurosci. 2017;20(9):1225–35. https://doi.org/10.1038/nn.4604.
Jovicic A, Mertens J, Boeynaems S, Bogaert E, Chai N, Yamada SB, et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat Neurosci. 2015;18(9):1226–9. https://doi.org/10.1038/nn.4085.
Hautbergue GM, Castelli LM, Ferraiuolo L, Sanchez-Martinez A, Cooper-Knock J, Higginbottom A, et al. SRSF1-dependent nuclear export inhibition of C9ORF72 repeat transcripts prevents neurodegeneration and associated motor deficits. Nat Commun. 2017;8:16063. https://doi.org/10.1038/ncomms16063.
Kaneb HM, Folkmann AW, Belzil VV, Jao LE, Leblond CS, Girard SL, et al. Deleterious mutations in the essential mRNA metabolism factor, hGle1, in amyotrophic lateral sclerosis. Hum Mol Genet. 2015;24(5):1363–73. https://doi.org/10.1093/hmg/ddu545.
Kendirgi F, Barry DM, Griffis ER, Powers MA, Wente SR. An essential role for hGle1 nucleocytoplasmic shuttling in mRNA export. J Cell Biol. 2003;160(7):1029–40. https://doi.org/10.1083/jcb.200211081.
Aditi, Folkmann AW, Wente SR. Cytoplasmic hGle1A regulates stress granules by modulation of translation. Mol Biol Cell. 2015;26(8):1476–90. https://doi.org/10.1091/mbc.E14-11-1523.
Kendirgi F, Rexer DJ, Alcazar-Roman AR, Onishko HM, Wente SR. Interaction between the shuttling mRNA export factor Gle1 and the nucleoporin hCG1: a conserved mechanism in the export of Hsp70 mRNA. Mol Biol Cell. 2005;16(9):4304–15. https://doi.org/10.1091/mbc.E04-11-0998.
Nousiainen HO, Kestila M, Pakkasjarvi N, Honkala H, Kuure S, Tallila J, et al. Mutations in mRNA export mediator GLE1 result in a fetal motoneuron disease. Nat Genet. 2008;40(2):155–7. https://doi.org/10.1038/ng.2007.65.
Folkmann AW, Collier SE, Zhan X, Aditi, Ohi MD, Wente SR. Gle1 functions during mRNA export in an oligomeric complex that is altered in human disease. Cell. 2013;155(3):582–93. https://doi.org/10.1016/j.cell.2013.09.023.
Seytanoglu A, Alsomali NI, Valori CF, McGown A, Kim HR, Ning K, et al. Deficiency in the mRNA export mediator Gle1 impairs Schwann cell development in the zebrafish embryo. Neuroscience. 2016;322:287–97. https://doi.org/10.1016/j.neuroscience.2016.02.039.
Aditi GL, Dawson TR, Wente SR. An amyotrophic lateral sclerosis-linked mutation in GLE1 alters the cellular pool of human Gle1 functional isoforms. Adv Biol Regul. 2016;62:25–36. https://doi.org/10.1016/j.jbior.2015.11.001.
Orr HT. Cell biology of spinocerebellar ataxia. J Cell Biol. 2012;197(2):167–77. https://doi.org/10.1083/jcb.201105092.
Orr HT, Chung MY, Banfi S, Kwiatkowski TJ Jr, Servadio A, Beaudet AL, et al. Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nat Genet. 1993;4(3):221–6. https://doi.org/10.1038/ng0793-221.
Yue S, Serra HG, Zoghbi HY, Orr HT. The spinocerebellar ataxia type 1 protein, ataxin-1, has RNA-binding activity that is inversely affected by the length of its polyglutamine tract. Hum Mol Genet. 2001;10(1):25–30.
Asher M, Johnson A, Zecevic B, Pease D, Cvetanovic M. Ataxin-1 regulates proliferation of hippocampal neural precursors. Neuroscience. 2016;322:54–65. https://doi.org/10.1016/j.neuroscience.2016.02.011.
Cvetanovic M, Hu YS, Opal P. Mutant ataxin-1 inhibits neural progenitor cell proliferation in SCA1. Cerebellum. 2017;16(2):340–7. https://doi.org/10.1007/s12311-016-0794-9.
Sanchez I, Balague E, Matilla-Duenas A. Ataxin-1 regulates the cerebellar bioenergetics proteome through the GSK3beta-mTOR pathway which is altered in Spinocerebellar ataxia type 1 (SCA1). Hum Mol Genet. 2016;25(18):4021–40. https://doi.org/10.1093/hmg/ddw242.
Park J, Al-Ramahi I, Tan Q, Mollema N, Diaz-Garcia JR, Gallego-Flores T, et al. RAS-MAPK-MSK1 pathway modulates ataxin 1 protein levels and toxicity in SCA1. Nature. 2013;498(7454):325–31. https://doi.org/10.1038/nature12204.
Gennarino VA, Singh RK, White JJ, De Maio A, Han K, Kim JY, et al. Pumilio1 haploinsufficiency leads to SCA1-like neurodegeneration by increasing wild-type ataxin1 levels. Cell. 2015;160(6):1087–98. https://doi.org/10.1016/j.cell.2015.02.012.
Ingram M, Wozniak EAL, Duvick L, Yang R, Bergmann P, Carson R, et al. Cerebellar transcriptome profiles of ATXN1 transgenic mice reveal SCA1 disease progression and protection pathways. Neuron. 2016;89(6):1194–207. https://doi.org/10.1016/j.neuron.2016.02.011.
Ito H, Fujita K, Tagawa K, Chen X, Homma H, Sasabe T, et al. HMGB1 facilitates repair of mitochondrial DNA damage and extends the lifespan of mutant ataxin-1 knock-in mice. EMBO Mol Med. 2015;7(1):78–101. https://doi.org/10.15252/emmm.201404392.
Pulst SM, Nechiporuk A, Nechiporuk T, Gispert S, Chen XN, Lopes-Cendes I, et al. Moderate expansion of a normally biallelic trinucleotide repeat in spinocerebellar ataxia type 2. Nat Genet. 1996;14(3):269–76. https://doi.org/10.1038/ng1196-269.
Sanpei K, Takano H, Igarashi S, Sato T, Oyake M, Sasaki H, et al. Identification of the spinocerebellar ataxia type 2 gene using a direct identification of repeat expansion and cloning technique, DIRECT. Nat Genet. 1996;14(3):277–84. https://doi.org/10.1038/ng1196-277.
Elden AC, Kim HJ, Hart MP, Chen-Plotkin AS, Johnson BS, Fang X, et al. Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature. 2010;466(7310):1069–75. https://doi.org/10.1038/nature09320.
Lastres-Becker I, Nonis D, Eich F, Klinkenberg M, Gorospe M, Kotter P, et al. Mammalian ataxin-2 modulates translation control at the pre-initiation complex via PI3K/mTOR and is induced by starvation. Biochim Biophys Acta. 2016;1862(9):1558–69. https://doi.org/10.1016/j.bbadis.2016.05.017.
Bar DZ, Charar C, Dorfman J, Yadid T, Tafforeau L, Lafontaine DL, et al. Cell size and fat content of dietary-restricted Caenorhabditis elegans are regulated by ATX-2, an mTOR repressor. Proc Natl Acad Sci U S A. 2016;113(32):E4620–9. https://doi.org/10.1073/pnas.1512156113.
Meierhofer D, Halbach M, Sen NE, Gispert S, Auburger G. Ataxin-2 (Atxn2)-knock-out mice show branched chain amino acids and fatty acids pathway alterations. Mol Cell Proteomics. 2016;15(5):1728–39. https://doi.org/10.1074/mcp.M115.056770.
Wilusz CJ, Wilusz J. Eukaryotic Lsm proteins: lessons from bacteria. Nat Struct Mol Biol. 2005;12(12):1031–6. https://doi.org/10.1038/nsmb1037.
Kozlov G, Safaee N, Rosenauer A, Gehring K. Structural basis of binding of P-body-associated proteins GW182 and ataxin-2 by the Mlle domain of poly(A)-binding protein. J Biol Chem. 2010;285(18):13599–606. https://doi.org/10.1074/jbc.M109.089540.
Fittschen M, Lastres-Becker I, Halbach MV, Damrath E, Gispert S, Azizov M, et al. Genetic ablation of ataxin-2 increases several global translation factors in their transcript abundance but decreases translation rate. Neurogenetics. 2015;16(3):181–92. https://doi.org/10.1007/s10048-015-0441-5.
Yokoshi M, Li Q, Yamamoto M, Okada H, Suzuki Y, Kawahara Y. Direct binding of Ataxin-2 to distinct elements in 3′ UTRs promotes mRNA stability and protein expression. Mol Cell. 2014;55(2):186–98. https://doi.org/10.1016/j.molcel.2014.05.022.
Becker LA, Huang B, Bieri G, Ma R, Knowles DA, Jafar-Nejad P, et al. Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature. 2017;544(7650):367–71. https://doi.org/10.1038/nature22038.
Sellier C, Campanari ML, Julie Corbier C, Gaucherot A, Kolb-Cheynel I, Oulad-Abdelghani M, et al. Loss of C9ORF72 impairs autophagy and synergizes with polyQ Ataxin-2 to induce motor neuron dysfunction and cell death. EMBO J. 2016;35(12):1276–97. https://doi.org/10.15252/embj.201593350.
Palomo GM, Manfredi G. Exploring new pathways of neurodegeneration in ALS: the role of mitochondria quality control. Brain Res. 2015;1607:36–46. https://doi.org/10.1016/j.brainres.2014.09.065.
Sen NE, Drost J, Gispert S, Torres-Odio S, Damrath E, Klinkenberg M, et al. Search for SCA2 blood RNA biomarkers highlights Ataxin-2 as strong modifier of the mitochondrial factor PINK1 levels. Neurobiol Dis. 2016;96:115–26. https://doi.org/10.1016/j.nbd.2016.09.002.
Zhang Y, Ling J, Yuan C, Dubruille R, Emery P. A role for Drosophila ATX2 in activation of PER translation and circadian behavior. Science. 2013;340(6134):879–82. https://doi.org/10.1126/science.1234746.
Lim C, Allada R. ATAXIN-2 activates PERIOD translation to sustain circadian rhythms in Drosophila. Science. 2013;340(6134):875–9. https://doi.org/10.1126/science.1234785.
Scoles DR, Meera P, Schneider MD, Paul S, Dansithong W, Figueroa KP, et al. Antisense oligonucleotide therapy for spinocerebellar ataxia type 2. Nature. 2017;544(7650):362–6. https://doi.org/10.1038/nature22044.
Kampers T, Friedhoff P, Biernat J, Mandelkow EM, Mandelkow E. RNA stimulates aggregation of microtubule-associated protein tau into Alzheimer-like paired helical filaments. FEBS Lett. 1996;399(3):344–9.
Violet M, Delattre L, Tardivel M, Sultan A, Chauderlier A, Caillierez R, et al. A major role for tau in neuronal DNA and RNA protection in vivo under physiological and hyperthermic conditions. Front Cell Neurosci. 2014;8:84. https://doi.org/10.3389/fncel.2014.00084.
Zhang X, Lin Y, Eschmann NA, Zhou H, Rauch JN, Hernandez I, et al. RNA stores tau reversibly in complex coacervates. PLoS Biol. 2017;15(7):e2002183. https://doi.org/10.1371/journal.pbio.2002183.
Vanderweyde T, Apicco DJ, Youmans-Kidder K, Ash PEA, Cook C, Lummertz da Rocha E, et al. Interaction of tau with the RNA-binding protein TIA1 regulates tau pathophysiology and toxicity. Cell Rep. 2016;15(7):1455–66. https://doi.org/10.1016/j.celrep.2016.04.045.
Faghihi MA, Modarresi F, Khalil AM, Wood DE, Sahagan BG, Morgan TE, et al. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat Med. 2008;14(7):723–30. https://doi.org/10.1038/nm1784.
Modarresi F, Faghihi MA, Patel NS, Sahagan BG, Wahlestedt C, Lopez-Toledano MA. Knockdown of BACE1-AS nonprotein-coding transcript modulates beta-amyloid-related hippocampal neurogenesis. Int J Alzheimers Dis. 2011;2011:929042–11. https://doi.org/10.4061/2011/929042.
Thomson SR, Seo SS, Barnes SA, Louros SR, Muscas M, Dando O, et al. Cell-type-specific translation profiling reveals a novel strategy for treating fragile X syndrome. Neuron. 2017;95(3):550–63 e5. https://doi.org/10.1016/j.neuron.2017.07.013.
Funding
We thank Young Investigator Grant of NARSAD and Scientist Development Grant of American Heart Association for Yingwei Mao. This work is supported by NIH 1R21MH108983.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors confirm that this article content has no conflicts of interest.
Additional information
This article is part of the Topical Collection on Neurogenesis and Disease
Rights and permissions
About this article
Cite this article
Zhou, Y., Dong, F. & Mao, Y. Control of CNS Functions by RNA-Binding Proteins in Neurological Diseases. Curr Pharmacol Rep 4, 301–313 (2018). https://doi.org/10.1007/s40495-018-0140-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40495-018-0140-7