Abstract
Background
E2F1 protein, a major effector of the Rb/E2F pathway plays a central role in regulating cell-fate decisions involved in proliferation, apoptosis, and differentiation. Its expression is highly dynamic and tightly modulated through a combination of transcriptional, translational and posttranslational controls. However, the mechanisms by which its expression and activity can promote different cellular outcomes remain to be fully elucidated. To better document E2F1 expression in live cells, we have engineered a series of fluorescent E2F1 protein reporters that quantitatively capture E2F1 protein dynamics.
Methods
Reporter constructs, under the control of the mouse or human E2F1 proximal promoter, were designed to express an E2F1-Venus fusion protein incapable of binding DNA. In addition, constructs either included or excluded the 3′ untranslated region (3′UTR) of the E2F1 gene. These constructs were introduced into fibroblasts and epithelial cells, and expression of the fusion reporter protein was validated and quantified in single cells using live imaging.
Results
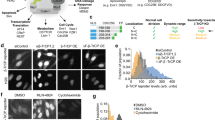
In all cases, expression of the reporter protein effectively recapitulated the behavior of E2F1 under various conditions, including cell cycle progression and genotoxic stress. No or little fluorescent signal of the reporter was detected in G0, but as the cycle progressed, expression of the reporter protein steadily increased in the nucleus, peaking a few hours before cell division, but declining to baseline 2–3 h prior to the onset of mitosis. The absence of the E2F1 3′UTR in the constructs led to considerably higher steady-state levels of the fusion protein, which although normally regulated, exhibited a slightly less complex dynamic profile during the cell cycle or genotoxic stress. Lastly, the presence or absence of Rb failed to impact the overall detection and levels of the reporter proteins.
Conclusions
Our validated E2F1 protein reporters complement nicely other reporters of the Rb/E2F pathway and provide a unique tool to follow the complex dynamics of E2F1 expression in real time in single cells.

Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Fischer, M. and Müller, G. A. (2017) Cell cycle transcription control: DREAM/MuvB and RB-E2F complexes. Crit. Rev. Biochem. Mol. Biol., 52, 638–662
Bertoli, C., Skotheim, J. M. and de Bruin, R. A. M. (2013) Control of cell cycle transcription during G1 and S phases. Nat. Rev. Mol. Cell Biol., 14, 518–528
Ren, B., Cam, H., Takahashi, Y., Volkert, T., Terragni, J., Young, R. A. and Dynlacht, B. D. (2002) E2F integrates cell cycle progression with DNA repair, replication, and G(2)/M checkpoints. Genes Dev., 16, 245–256
van den Heuvel, S. and Dyson, N. J. (2008) Conserved functions of the pRB and E2F families. Nat. Rev. Mol. Cell Biol., 9, 713–724
Henley, S. A. and Dick, F. A. (2012) The retinoblastoma family of proteins and their regulatory functions in the mammalian cell division cycle. Cell Div., 7, 10
Sherr, C. J. and Roberts, J. M. (2004) Living with or without cyclins and cyclin-dependent kinases. Genes Dev., 18, 2699–2711
Chen, H.-Z., Tsai, S.-Y. and Leone, G. (2009) Emerging roles of E2Fs in cancer: an exit from cell cycle control. Nat. Rev. Cancer, 9, 785–797
Dong, P., Maddali, M. V., Srimani, J. K., Thélot, F., Nevins, J. R., Mathey-Prevot, B. and You, L. (2014) Division of labour between Myc and G1 cyclins in cell cycle commitment and pace control. Nat. Commun., 5, 4750
Dong, P., Zhang, C., Parker, B.-T., You, L. and Mathey-Prevot, B. (2018) Cyclin D/CDK4/6 activity controls G1 length in mammalian cells. PLoS One, 13, e0185637
Dimova, D. K. and Dyson, N. J. (2005) The E2F transcriptional network: old acquaintances with new faces. Oncogene, 24, 2810–2826
Dimri, G. P., Itahana, K. and Acosta, M. (2000). Regulation of a senescence checkpoint response by the E2F1 transcription factor and p14ARF tumor suppressor. Mol. Cell. Biol
Cuitiño, M. C., Pécot, T., Sun, D., Kladney, R., Okano-Uchida, T., Shinde, N., Saeed, R., Perez-Castro, A. J., Webb, A., Liu, T., et al. (2019) Two distinct E2F transcriptional modules drive cell cycles and differentiation. Cell Reports, 27, 3547–3560.e5
O’Donnell, K. A., Wentzel, E. A., Zeller, K. I., Dang, C. V. and Mendell, J. T. (2005) c-Myc-regulated microRNAs modulate E2F1 expression. Nature, 435, 839–843
Pickering, M. T., Stadler, B. M. and Kowalik, T. F. (2009) miR-17 and miR-20a temper an E2F1-induced G1 checkpoint to regulate cell cycle progression. Oncogene, 28, 140–145
Iaquinta, P. J. and Lees, J. A. (2007) Life and death decisions by the E2F transcription factors. Curr. Opin. Cell Biol., 19, 649–657
Poppy Roworth, A., Ghari, F. and La Thangue, N. B. (2015) To live or let die–complexity within the E2F1 pathway. Mol. Cell. Oncol., 2, e970480
Zheng, N., Fraenkel, E., Pabo, C. O. and Pavletich, N. P. (1999) Structural basis of DNA recognition by the heterodimeric cell cycle transcription factor E2F-DP. Genes Dev., 13, 666–674
Munro, S., Carr, S. M. and La Thangue, N. B. (2012) Diversity within the pRb pathway: is there a code of conduct?. Oncogene, 31, 4343–4352
Kozak, M. (1987) An analysis of 5′-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res., 15, 8125–8148
Wong, J. V., Dong, P., Nevins, J. R., Mathey-Prevot, B. and You, L. (2011) Network calisthenics: control of E2F dynamics in cell cycle entry. Cell Cycle, 10, 3086–3094
Lin, W. C., Lin, F. T. and Nevins, J. R. (2001) Selective induction of E2F1 in response to DNA damage, mediated by ATMdependent phosphorylation. Genes Dev., 15, 1833–1844
Magae, J., Wu, C. L., Illenye, S., Harlow, E. and Heintz, N. H. (1996) Nuclear localization of DP and E2F transcription factors by heterodimeric partners and retinoblastoma protein family members. J. Cell. Sci., 109, 1717–1726
Allen, K. E., de la Luna, S., Kerkhoven, R. M., Bernards, R. and La Thangue, N. B. (1997) Distinct mechanisms of nuclear accumulation regulate the functional consequence of E2F transcription factors. J. Cell. Sci., 110, 2819–2831
Ivanova, I. A., Vespa, A. and Dagnino, L. (2007) A novel mechanism of E2F1 regulation via nucleocytoplasmic shuttling: determinants of nuclear import and export. Cell Cycle, 6, 2186–2195
Hofmann, F., Martelli, F., Livingston, D. M. and Wang, Z. (1996) The retinoblastoma gene product protects E2F-1 from degradation by the ubiquitin-proteasome pathway. Genes Dev., 10, 2949–2959
Campanero, M. R. and Flemington, E. K. (1997) Regulation of E2F through ubiquitin-proteasome-dependent degradation: stabilization by the pRB tumor suppressor protein. Proc. Natl. Acad. Sci. USA, 94, 2221–2226
Blanchet, E., Annicotte, J.-S., Lagarrigue, S., Aguilar, V., Clapé, C., Chavey, C., Fritz, V., Casas, F., Apparailly, F., Auwerx, J., et al. (2011) E2F transcription factor-1 regulates oxidative metabolism. Nat. Cell Biol., 13, 1146–1152
Shats, I., Deng, M., Davidovich, A., Zhang, C., Kwon, J. S., Manandhar, D., Gordân, R., Yao, G. and You, L. (2017) Expression level is a key determinant of E2F1-mediated cell fate. Cell Death Differ., 24, 626–637
Yu, W., Chan-On, W., Teo, M., Ong, C. K., Cutcutache, I., Allen, G. E., Wong, B., Myint, S. S., Lim, K. H., Voorhoeve, P. M., et al. (2011) First somatic mutation of E2F1 in a critical DNA binding residue discovered in well-differentiated papillary mesothelioma of the peritoneum. Genome Biol., 12, R96
Helin, K., Lees, J. A., Vidal, M., Dyson, N., Harlow, E. and Fattaey, A. (1992) A cDNA encoding a pRB-binding protein with properties of the transcription factor E2F. Cell, 70, 337–350
Hanahan, D. and Weinberg, R. A. (2011) Hallmarks of cancer: the next generation. Cell, 144, 646–674
Morita, S., Kojima, T. and Kitamura, T. (2000) Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther., 7, 1063–1066
Sanjana, N. E., Shalem, O. and Zhang, F. (2014) Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods, 11, 783–784
Shalem, O., Sanjana, N. E., Hartenian, E., Shi, X., Scott, D. A., Mikkelson, T., Heckl, D., Ebert, B. L., Root, D. E., Doench, J. G., et al. (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science, 343, 84–87
Hayflick, L. (1965) The limited in vitro lifetime of human diploid cell strains. Exp. Cell Res., 37, 614–636
Acknowledgements
We thank Y. Gao for his help with time-lapse and confocal microscopy at the Duke Light Microscopy Core Facility. This research was supported by a grant from NIH (1R01-GM106107) (L.Y and B.M-P.), and funds from the School of Medicine at Duke University (B.M-P).
Author information
Authors and Affiliations
Contributions
B.M-P., P.D., G.Y. and L.Y. developed the concept of the paper. B.M.-P. designed the research approach and performed experiments with the help of P.D., B-T.P., C.I., C.H. B.M-P. analyzed the data; B.M-P., P.D, G.Y. and L.Y. interpreted the results and wrote the manuscript with contributions from C.I.
Corresponding author
Ethics declarations
The authors Bernard Mathey-Prevot, Bao-Tran Parker, Carolyn Im, Cierra Hong, Peng Dong, Guang Yao and Lingchong You declare that they have no conflict of interests.
All procedures performed in studies were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Author summary: Cell proliferation in response to growth signals is regulated through the action of the Rb/E2F circuit, which includes the E2F1 protein as one of its effectors. E2F1 expression is highly dynamic and changes in E2F1 protein can promote different decisions based on a complex balance with other effectors. To document how E2F1 levels change in single cells under different experimental conditions, we have engineered fluorescent protein reporters that serve as proxies for endogenous E2F1expression. Here we report the design of these reporters and show by live imaging and quantitative analysis in single cells that they faithfully recapitulate E2F1 behavior.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Mathey-Prevot, B., Parker, BT., Im, C. et al. Quantifying E2F1 protein dynamics in single cells. Quant Biol 8, 20–30 (2020). https://doi.org/10.1007/s40484-019-0193-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-019-0193-6