Abstract
Background
Traditional Chinese medicine (TCM) treats diseases in a holistic manner, while TCM formulae are multi-component, multi-target agents at the molecular level. Thus there are many parallels between the key ideas of TCM pharmacology and network pharmacology. These years, TCM network pharmacology has developed as an interdisciplinary of TCM science and network pharmacology, which studies the mechanism of TCM at the molecular level and in the context of biological networks. It provides a new research paradigm that can use modern biomedical science to interpret the mechanism of TCM, which is promising to accelerate the modernization and internationalization of TCM.
Results
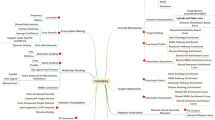
In this paper we introduce state-of-the-art free data sources, web servers and softwares that can be used in the TCM network pharmacology, including databases of TCM, drug targets and diseases, web servers for the prediction of drug targets, and tools for network and functional analysis.
Conclusions
This review could help experimental pharmacologists make better use of the existing data and methods in their study of TCM.

Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Li, S. and Zhang, B. (2013) Traditional Chinese medicine network pharmacology: theory, methodology and application. Chin. J. Nat. Med., 11, 110–120
Zhao, J., Jiang, P. and Zhang, W. (2010) Molecular networks for the study of TCM pharmacology. Brief. Bioinform., 11, 417–430
Li, S., Fan T.-P., Jia, W., Lu, A. and Zhang, W. (2014) Network pharmacology in traditional Chinese medicine, evidence-based complementary and alternative medicine. Article ID 138460 https://doi.org/10.1155/2014/138460
Li, P., Chen, J., Wang, J., Zhou, W., Wang, X., Li, B., Tao, W., Wang, W., Wang, Y. and Yang, L. (2014) Systems pharmacology strategies for drug discovery and combination with applications to cardiovascular diseases. J. Ethnopharmacol., 151, 93–107
Huang, C., Zheng, C., Li, Y., Wang, Y., Lu, A. and Yang, L. (2014) Systems pharmacology in drug discovery and therapeutic insight for herbal medicines. Brief. Bioinform., 15, 710–733
Chen, C. Y.-C. (2011) TCM Database@Taiwan: the world’s largest traditional Chinese medicine database for drug screening in silico. PLoS One, 6, e15939
Chen, X., Zhou, H., Liu, Y. B., Wang, J. F., Li, H., Ung, C. Y., Han, L. Y., Cao, Z. W. and Chen, Y. Z. (2006) Database of traditional Chinese medicine and its application to studies of mechanism and to prescription validation. Br. J. Pharmacol., 149, 1092–1103
Ru, J., Li, P., Wang, J., Zhou, W., Li, B., Huang, C., Li, P., Guo, Z., Tao, W., Yang, Y., et al. (2014) TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J. Cheminform., 6, 13
Xue, R., Fang, Z., Zhang, M., Yi, Z., Wen, C. and Shi, T. (2013) TCMID: traditional Chinese medicine integrative database for herb molecular mechanism analysis. Nucleic Acids Res., 41, D1089–D1095
Li, S., Zhang, B. and Zhang, N. (2011) Network target for screening synergistic drug combinations with application to traditional Chinese medicine. BMC Syst. Biol., 5, S10
Lin, L., Yang, T., Fang, L., Yang, J., Yang, F. and Zhao, J. (2017) Gene gravity-like algorithm for disease gene prediction based on phenotype-specific network. BMC Syst. Biol., 11, 121
Sun, Y., Sheng, Z., Ma, C., Tang, K., Zhu, R., Wu, Z., Shen, R., Feng, J.,Wu, D., Huang, D., et al. (2015) Combining genomic and network characteristics for extended capability in predicting synergistic drugs for cancer. Nat. Commun., 6, 8481
Yang, K., Bai, H., Ouyang, Q., Lai, L. and Tang, C. (2008) Finding multiple target optimal intervention in disease-related molecular network. Mol. Syst. Biol., 4, 228
Fang, H., Wang, Y., Yang T., Ga, Y., Zhang, Y., Liu, R., Zhang, W. and Zhao, J. (2013) Bioinformatics analysis for the antirheumatic effects of Huang-Lian-Jie-Du-Tang from a network perspective. Evid-Based Compl. Alt., Article ID 245357, http://dx.doi.org/10.1155/2013/245357
Le, D. H. and Le, L. (2016) Systems pharmacology: a unified framework for prediction of drug-target interactions. Curr. Pharm. Des., 22, 3569–3575
Fang, H.-Y., Zeng, H.-W., Lin, L.-M., Chen, X., Shen, X.-N., Fu, P., Lv, C., Liu, Q., Liu, R.-H., Zhang, W.-D., et al. (2017) A network-based method for mechanistic investigation of Shexiang Baoxin Pill’s treatment of cardiovascular diseases. Sci. Rep., 7, 43632
Wang, T., Yang, J., Chen, X., Zhao, K., Wang, J., Zhang, Y., Zhao, J. and Ga, Y. (2017) Systems study on the antirheumatic mechanism of Tibetan medicated-bath therapy using Wuwei-Ganlu-Yaoyu-Keli. BioMed Res. Int., 2017, 2320932
Liang, X., Li, H. and Li, S. (2014) A novel network pharmacology approach to analyse traditional herbal formulae: the Liu-Wei-Di-Huang pill as a case study. Mol. Biosyst., 10, 1014–1022
Zhang, W., Tao, Q., Guo, Z., Fu, Y., Chen, X., Shar, P. A., Shahen, M., Zhu, J., Xue, J., Bai, Y., et al. (2016) Systems pharmacology dissection of the integrated treatment for cardiovascular and gastrointestinal disorders by traditional Chinese medicine. Sci. Rep., 6, 32400
Zhou, W., Cheng, X. and Zhang, Y. (2016) Effect of Liuwei Dihuang decoction, a traditional Chinese medicinal prescription, on the neuroendocrine immunomodulation network. Pharmacol. Ther., 162, 170–178
Ye, H., Ye, L., Kang, H., Zhang, D., Tao, L., Tang K., Liu, X., Zhu, R., Liu, Q., Chen, Y. Z. et al. (2011) HIT: linking herbal active ingredients to targets. Nucleic Acids Res., 39 (suppl_1), D1055–D1059
Yu, H., Chen, J., Xu, X., Li, Y., Zhao, H., Fang, Y., Li, X., Zhou, W., Wang, W. and Wang, Y. (2012) A systematic prediction of multiple drug-target interactions from chemical, genomic, and pharmacological data. PLoS One, 7, e37608
Li, Y. H., Yu, C. Y., Li, X. X., Zhang, P., Tang, J., Yang, Q., Fu, T., Zhang, X., Cui, X., Tu, G., et al. (2018) Therapeutic target database update 2018: enriched resource for facilitating bench-toclinic research of targeted therapeutics. Nucleic Acids Res., 46, D1121–D1127
Whirl-Carrillo, M., McDonagh, E. M., Hebert, J. M., Gong, L., Sangkuhl, K., Thorn, C. F., Altman, R. B. and Klein, T. E. (2012) Pharmacogenomics knowledge for personalized medicine. Clin. Pharmacol. Ther., 92, 414–417
Huang, C., Yang, Y., Chen, X., Wang, C., Li, Y., Zheng, C. and Wang, Y. (2017) Large-scale cross-species chemogenomic platform proposes a new drug discovery strategy of veterinary drug from herbal medicines. PLoS One, 12, e0184880
Lee, A. Y., Park, W., Kang, T.-W., Cha, M. H. and Chun, J. M. (2018) Network pharmacology-based prediction of active compounds and molecular targets in Yijin-Tang acting on hyperlipidaemia and atherosclerosis. J. Ethnopharmacol., 221, 151–159
Kuhn, M., von Mering, M., Campillos, M., Jensen, L.J. and Bork, P. (2008) STITCH: interaction networks of chemicals and proteins. Nucleic Acids Res., 36(suppl_1), D684–688
Hamosh, A., Scott, A.F., Amberger, J.S., Bocchini C.A. and McKusick, V.A. (2005) Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders, Nucleic Acids Res., 33(suppl_1), D514–517
Wishart, D. S., Knox, C., Guo, A. C., Shrivastava, S., Hassanali, M., Stothard, P., Chang, Z. and Woolsey, J. (2006) DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res., 34, D668–D672
Mangal, M., Sagar, P., Singh, H., Raghava, G. P. S. and Agarwal, S. M. (2013) NPACT: naturally occurring plant-based anti-cancer compound-activity-target database. Nucleic Acids Res., 41, D1124–D1129
Tao, W., Li, B., Gao, S., Bai, Y., Shar, P. A., Zhang, W., Guo, Z., Sun, K., Fu, Y., Huang, C., et al. (2015) CancerHSP: anticancer herbs database of systems pharmacology. Sci. Rep., 5, 11481
Zeng, X., Zhang, P., He, W., Qin, C., Chen, S., Tao, L., Wang, Y., Tan, Y., Gao, D., Wang, B., et al. (2018) NPASS: natural product activity and species source database for natural product research, discovery and tool development. Nucleic Acids Res., 46, D1217–D1222
Fang, J., Cai, C., Wang, Q., Lin, P., Zhao, Z. and Cheng, F. (2017) Systems pharmacology-based discovery of natural products for precision oncology through targeting cancer mutated genes. CPT Pharmacometrics Syst. Pharmacol., 6, 177–187
Wishart, D. S., Feunang, Y. D., Guo, A. C., Lo, E. J., Marcu, A., Grant, J. R., Sajed, T., Johnson, D., Li, C., Sayeeda, Z., et al. (2018) DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res., 46, D1074–D1082
Bento, A. P., Gaulton, A., Hersey, A., Bellis, L. J., Chambers, J., Davies, M., Krüger, F. A., Light, Y., Mak, L., McGlinchey, S., et al. (2014) The ChEMBL bioactivity database: an update. Nucleic Acids Res., 42, D1083–D1090
Gilson, M. K., Liu, T., Baitaluk, M., Nicola, G., Hwang, L. and Chong, J. (2016) BindingDB in 2015: a public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res., 44, D1045–D1053
Günther, S., Kuhn, M., Dunkel, M., Campillos, M., Senger, C., Petsalaki, E., Ahmed, J., Urdiales, E. G., Gewiess, A., Jensen, L. J., et al. (2008) SuperTarget and Matador: resources for exploring drug-target relationships. Nucleic Acids Res., 36, D919–D922
Kumar, R., Chaudhary, K., Gupta, S., Singh, H., Kumar, S., Gautam, A., Kapoor, P., Raghava, G. P. S. and Cancer, D. R. (2013) CancerDR: cancer drug resistance database. Sci. Rep., 3, 1445
Cotto, K. C., Wagner, A. H., Feng, Y.-Y., Kiwala, S., Coffman, A. C., Spies, G., Wollam, A., Spies, N. C., Griffith, O. L. and Griffith, M. (2018) DGIdb 3.0: a redesign and expansion of the drug-gene interaction database. Nucleic Acids Res., 46, D1068–D1073
Siramshetty, V. B., Eckert, O. A., Gohlke, B.-O., Goede, A., Chen, Q., Devarakonda, P., Preissner, S. and Preissner, R. (2018) SuperDRUG2: a one stop resource for approved/marketed drugs. Nucleic Acids Res., 46, D1137–D1143
Yu, G., Zhang, Y., Ren, W., Dong, L., Li, J., Geng, Y., Zhang, Y., Li, D., Xu, H. and Yang, H. (2016) Network pharmacology-based identification of key pharmacological pathways of Yin-Huang-Qing-Fei capsule acting on chronic bronchitis. Int. J. Chron. Obstruct. Pulmon. Dis., 12, 85–94
Fang, H., Wang, Y., Yang, T., Ga, Y., Zhang, Y., Liu, R., Zhang, W. and Zhao, J. (2013) Bioinformatics analysis for the antirheumatic effects of Huang-Lian-Jie-Du-Tang from a network perspective. Evid. Based Complement. Alternat. Med., 2013, 245357
Zhang, Y., Lin, Y., Zhao, H., Guo, Q., Yan, C. and Lin, N. (2016) Revealing the effects of the herbal pair of Euphorbia kansui and Glycyrrhiza on hepatocellular carcinoma ascites with integrating network target analysis and experimental validation. Int. J. Biol. Sci., 12, 594–606
Okuno, Y., Tamon, A., Yabuuchi, H., Niijima, S., Minowa, Y., Tonomura, K., Kunimoto, R. and Feng, C. (2008) GLIDA: GPCR—ligand database for chemical genomics drug discovery—database and tools update, Nucleic Acids Res., 36(suppl_1), D907–D912
Chen, X., Ji, Z. L. and Chen, Y. Z. (2002) TTD: Therapeutic Target Database. Nucleic Acids Res., 30, 412–415
Davis, A. P., Grondin, C. J., Lennon-Hopkins, K., Saraceni-Richards, C., Sciaky, D., King, B. L., Wiegers, T. C. and Mattingly, C. J. (2015) The Comparative Toxicogenomics Database’s 10th year anniversary: update 2015. Nucleic Acids Res., 43, D914–D920
Kanehisa, M. and Goto, S. (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res., 28, 27–30
Schaefer, C. F., Anthony, K., Krupa, S., Buchoff, J., Day, M., Hannay, T. and Buetow, K. H. (2009) PID: the Pathway Interaction Database. Nucleic Acids Res., 37, D674–D679
Fabregat, A., Sidiropoulos, K., Garapati, P., Gillespie, M., Hausmann, K., Haw, R., Jassal, B., Jupe, S., Korninger, F., McKay, S., et al. (2016) The Reactome pathway Knowledgebase. Nucleic Acids Res., 44, D481–D487
Caspi, R., Billington, R., Ferrer, L., Foerster, H., Fulcher, C. A., Keseler, I. M., Kothari, A., Krummenacker, M., Latendresse, M., Mueller, L. A., et al. (2016) The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res., 44, D471–D480
Roth, B. L., Lopez, E., Patel, S. and Kroeze, W. K. (2000) The multiplicity of serotonin receptors: uselessly diverse molecules or an embarrassment of riches? Neuroscientist, 6, 252–262
Rose, P. W., Prlić, A., Bi, C., Bluhm, W. F., Christie, C. H., Dutta, S., Green, R. K., Goodsell, D. S., Westbrook, J. D., Woo, J., et al. (2015) The RCSB Protein Data Bank: views of structural biology for basic and applied research and education. Nucleic Acids Res., 43, D345–D356
Kim, S., Thiessen, P. A., Bolton, E. E., Chen, J., Fu, G., Gindulyte, A., Han, L., He, J., He, S., Shoemaker, B. A., et al. (2016) PubChem Substance and Compound databases. Nucleic Acids Res., 44, D1202–D1213
Carlson, H. A., Smith, R. D., Damm-Ganamet, K. L., Stuckey, J. A., Ahmed, A., Convery, M. A., Somers, D. O., Kranz, M., Elkins, P. A., Cui, G., et al. (2016) CSAR 2014: a benchmark exercise using unpublished data from pharma. J. Chem. Inf. Model., 56, 1063–1077.
Sterling, T. and Irwin, J. J. (2015) ZINC 15–ligand discovery for everyone. J. Chem. Inf. Model., 55, 2324–2337
Wishart, D. S., Feunang, Y. D., Marcu, A., Guo, A. C., Liang, K., Vázquez-Fresno, R., Sajed, T., Johnson, D., Li, C., Karu, N., et al. (2018) HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res., 46, D608–D617
Liu, Z., Guo, F., Wang, Y., Li, C., Zhang, X., Li, H., Diao, L., Gu, J., Wang, W., Li, D., et al. (2016) BATMAN-TCM: a bioinformatics analysis tool for molecular mechanism of traditional Chinese medicine. Sci. Rep., 6, 21146
Wang, X., Shen, Y., Wang, S., Li, S., Zhang, W., Liu, X., Lai, L., Pei, J. and Li, H. (2017) PharmMapper 2017 update: a web server for potential drug target identification with a comprehensive target pharmacophore database. Nucleic Acids Res., 45, W356–W360
Luo, H., Chen, J., Shi, L., Mikailov, M., Zhu, H., Wang, K., He, L., and Yang, L. (2011) DRAR-CPI: a server for identifying drug repositioning potential and adverse drug reactions via the chemical–protein interactome, Nucleic Acids Res., 39(suppl_2), W492–W498
Pereira, A. S. P., Bester, M. J. and Apostolides, Z. (2017) Exploring the anti-proliferative activity of Pelargonium sidoides DC with in silico target identification and network pharmacology. Mol. Divers., 21, 809–820
Wei, J., Zhang, Y., Jia, Q., Liu, M., Li, D., Zhang, Y., Song, L., Hu, Y., Xian, M., Yang, H., et al. (2016) Systematic investigation of transcription factors critical in the protection against cerebral ischemia by Danhong injection. Sci. Rep., 6, 29823
Nickel, J., Gohlke, B.-O., Erehman, J., Banerjee, P., Rong, W. W., Goede, A., Dunkel, M. and Preissner, R. (2014) SuperPred: update on drug classification and target prediction. Nucleic Acids Res., 42, W26–W31
Gfeller, D., Grosdidier, A., Wirth, M., Daina, A., Michielin, O. and Zoete, V. (2014) SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res., 42, W32–W38
Yao, Z.-J., Dong, J., Che, Y.-J., Zhu, M.-F., Wen, M., Wang, N.-N., Wang, S., Lu, A.-P. and Cao, D.-S. (2016) TargetNet: a web service for predicting potential drug-target interaction profiling via multitarget SAR models. J. Comput. Aided Mol. Des., 30, 413–424
Hsin, K.-Y., Matsuoka, Y., Asai, Y., Kamiyoshi, K., Watanabe, T., Kawaoka, Y. and Kitano, H. (2016) systemsDock: a web server for network pharmacology-based prediction and analysis. Nucleic Acids Res., 44, W507–W513
Hsin, K.-Y., Ghosh, S. and Kitano, H. (2013) Combining machine learning systems and multiple docking simulation packages to improve docking prediction reliability for network pharmacology. PLoS One, 8, e83922
Zsoldos, Z., Reid, D., Simon, A., Sadjad, B. S. and Johnson, A. P. (2006) eHiTS: an innovative approach to the docking and scoring function problems. Curr. Protein Pept. Sci., 7, 421–435.
Piñero, J., Bravo, À., Queralt-Rosinach, N., Gutiérrez-Sacristán, A., Deu-Pons, J., Centeno, E., García-García, J., Sanz, F. and Furlong, L. I. (2017) DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res., 45, D833–D839
Davis, A. P., Grondin, C. J., Johnson, R. J., Sciaky, D., King, B. L., McMorran, R., Wiegers, J., Wiegers, T. C. and Mattingly, C. J. (2017) The Comparative Toxicogenomics Database: update 2017. Nucleic Acids Res., 45, D972–D978
Apweiler, R., Bairoch, A., Wu, C. H., Barker, W. C., Boeckmann, B., Ferro, S., Gasteiger, E., Huang, H., Lopez, R., Magrane, M., et al. (2004) UniProt: the Universal Protein knowledgebase. Nucleic Acids Res., 32, D115–D119
Landrum, M. J., Lee, J. M., Benson, M., Brown, G., Chao, C., Chitipiralla, S., Gu, B., Hart, J., Hoffman, D., Hoover, J., et al. (2016) ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res., 44, D862–D868
Aymé, S. and Schmidtke, J. (2007) Networking for rare diseases: a necessity for Europe. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz, 50, 1477–1483, in German
MacArthur, J., Bowler, E., Cerezo, M., Gil, L., Hall, P., Hastings, E., Junkins, H., McMahon, A., Milano, A., Morales, J., et al. (2017) The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res., 45, D896–D901
Becker, K. G., Barnes, K. C., Bright, T. J. and Wang, S. A. (2004) The Genetic Association Database. Nature Genet., 36 431–432
Blake, J. A., Richardson, J. E., Bult, C. J., Kadin, J. A. and Eppig, J. T. (2003) MGD: the Mouse Genome Database. Nucleic Acids Res., 31, 193–195
Twigger, S., Lu, J., Shimoyama, M., Chen, D., Pasko, D., Long, H., Ginster, J., Chen, C.-F., Nigam, R., Kwitek, A., et al. (2002) Rat Genome Database (RGD): mapping disease onto the genome. Nucleic Acids Res., 30, 125–128
Gutiérrez-Sacristán, A., Grosdidier, S., Valverde, O., Torrens, M., Bravo, À., Piñero, J., Sanz, F. and Furlong, L. I. (2015) PsyGeNET: a knowledge platform on psychiatric disorders and their genes. Bioinformatics, 31, 3075–3077
Robinson, P. N., Köhler, S., Bauer, S., Seelow, D., Horn, D. and Mundlos, S. (2008) The Human Phenotype Ontology: a tool for annotating and analyzing human hereditary disease. Am. J. Hum. Genet., 83, 610–615
Bundschus, M., Dejori, M., Stetter, M., Tresp, V. and Kriegel, H.-P. (2008) Extraction of semantic biomedical relations from text using conditional random fields. BMC Bioinformatics, 9, 207
Bravo, À., Piñero, J., Queralt-Rosinach, N., Rautschka, M. and Furlong, L. I. (2015) Extraction of relations between genes and diseases from text and large-scale data analysis: implications for translational research. BMC Bioinformatics, 16, 55
Rappaport, N., Twik, M., Plaschkes, I., Nudel, R., Iny Stein, T., Levitt, J., Gershoni, M., Morrey, C. P., Safran, M. and Lancet, D. (2017) MalaCards: an amalgamated human disease compendium with diverse clinical and genetic annotation and structured search. Nucleic Acids Res., 45, D877–D887
Roberta, A. (2007) GeneTests: integrating genetic services into patient care. Am. J. Hum. Genet., 81, 658–659
Pletscher-Frankild, S., Pallejà, A., Tsafou, K., Binder, J. X. and Jensen, L. J. (2015) DISEASES: text mining and data integration of disease-gene associations. Methods, 74, 83–89
Allende, R. A. (2009) Accelerating searches of research grants and scientific literature with novoseekSM. Nat. Methods, 6, 394
Safran, M., Dalah, I., Alexander, J., Rosen, N., Iny Stein, T., Shmoish, M., Nativ, N., Bahir, I., Doniger, T., Krug, H., et al. (2010) GeneCards Version 3: the human gene integrator. Database (Oxford), 2010, baq020
Kim, J., So, S., Lee, H.-J., Park, J. C., Kim, J. J. and Lee, H. (2013) DigSee: disease gene search engine with evidence sentences (version cancer). Nucleic Acids Res., 41, W510–W517
Zhang, Y., Bai, M., Zhang, B., Liu, C., Guo, Q., Sun, Y.,Wang, D., Wang, C., Jiang, Y., Lin, N., et al. (2015) Uncovering pharmacological mechanisms of Wu-tou decoction acting on rheumatoid arthritis through systems approaches: drug-target prediction, network analysis and experimental validation. Sci. Rep., 5, 9463
Huang, W., Sherman, B. T. and Lempicki, R. A. (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc., 4, 44–57
Subramanian, A., Narayan, R., Corsello, S. M., Peck, D. D., Natoli, T. E., Lu, X., Gould, J., Davis, J. F., Tubelli, A. A. and Asiedu, J. K. (2017) A next generation connectivity map: L1000 platform and the first 1,000,000 profiles, Cell 171, 1437–1452. e17
Lamb, J., Crawford, E. D., Peck, D., Modell, J. W., Blat, I. C., Wrobel, M. J., Lerner, J., Brunet, J.-P., Subramanian, A. and Ross, K.N. (2006) The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313,1929–1935
Wen, Z., Wang, Z., Wang, S., Ravula, R., Yang, L., Xu, J., Wang, C., Zuo, Z., Chow, M. S., Shi, L., et al. (2011) Discovery of molecular mechanisms of traditional Chinese medicinal formula Si-Wu-Tang using gene expression microarray and connectivity map. PLoS One, 6, e18278
Lv, C., Wu, X., Wang, X., Su, J., Zeng, H., Zhao, J., Lin, S., Liu, R., Li, H., Li, X., et al. (2017) The gene expression profiles in response to 102 traditional Chinese medicine (TCM) components: a general template for research on TCMs. Sci. Rep., 7, 352
Yoo, M., Shin, J., Kim, H., Kim, J., Kang, J. and Tan, A. C. (2018) Exploring the molecular mechanisms of traditional Chinese medicine components using gene expression signatures and connectivity map. Comput. Methods Programs Biomed.
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., Amin, N., Schwikowski, B. and Ideker, T. (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res., 13, 2498–2504
Vennix, P. P., Kuijpers, W., Tonnaer, E. L., Peters, T. A. and Ramaekers, F. C. (1990) Cytokeratins in induced epidermoid formations and cholesteatoma lesions. Arch. Otolaryngol. Head Neck Surg., 116, 560–565
Ravasz, E., Somera, A. L., Mongru, D. A., Oltvai, Z. N. and Barabási, A.-L. (2002) Hierarchical organization of modularity in metabolic networks. Science 297,1551–1555
Padmanabhan, K., Wang, K. and Samatova, N. F. (2012) Functional annotation of hierarchical modularity. PLoS One, 7, e33744
Kim, H. U., Ryu, J. Y., Lee, J. O. and Lee, S. Y. (2015) A systems approach to traditional oriental medicine. Nat. Biotechnol., 33, 264–268
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos. 81520108030, 21472238, 61372194 and 81260672), Professor of Chang Jiang Scholars Program, Shanghai Engineering Research Center for the Preparation of Bioactive Natural Products (No. 16DZ2280200), the Scientific Foundation of Shanghai China (Nos. 13401900103 and 13401900101), the National Key Research and Development Program of China (No. 2017YFC1700200), the Natural Science Foundation of Chongqing (No. cstc2018jcyjAX0090) and Chongqing Education Reform Project of Graduate (No. yjg152017). The funders had no role in study design, data collection, analysis, decision to publish and preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Author summary: TCM network pharmacology studies the therapeutic mechanism of TCM formulae from a systems perspective and at the molecular level. Years of research in related fields has developed many databases and tools that are useful for the study of TCM network pharmacology. In this paper, we introduce some of such free resources.
Rights and permissions
About this article
Cite this article
Zhao, J., Yang, J., Tian, S. et al. A survey of web resources and tools for the study of TCM network pharmacology. Quant Biol 7, 17–29 (2019). https://doi.org/10.1007/s40484-019-0167-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-019-0167-8