Abstract
Background
Our understanding of post-transcriptional gene regulation has increased exponentially with the development of robust methods to define protein-RNA interactions across the transcriptome. In this review, we highlight the evolution and successful applications of crosslinking and immunoprecipitation (CLIP) methods to interrogate protein-RNA interactions in a transcriptome-wide manner.
Results
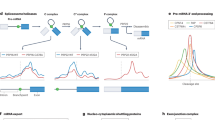
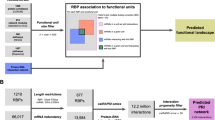
Here, we survey the vast array of in vitro and in vivo approaches used to identify protein-RNA interactions, including but not limited to electrophoretic mobility shift assays, systematic evolution of ligands by exponential enrichment (SELEX), and RIP-seq. We particularly emphasize the advancement of CLIP technologies, and detail protocol improvements and computational tools used to analyze the output data. Importantly, we discuss how profiling protein-RNA interactions can delineate biological functions including splicing regulation, alternative polyadenylation, cytoplasmic decay substrates, and miRNA targets.
Conclusions
In summary, this review summarizes the benefits of characterizing RNA-protein networks to further understand the regulation of gene expression and disease pathogenesis. Our review comments on how future CLIP technologies can be adapted to address outstanding questions related to many aspects of RNA metabolism and further advance our understanding of RNA biology.

Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Linder, B., Fischer, U. and Gehring, N. H. (2015) mRNA metabolism and neuronal disease. FEBS Lett., 589, 1598–1606
Brais, B., Bouchard, J. P., Xie, Y. G., Rochefort, D. L., Chrétien, N., Tomé, F. M., Lafrentére, R. G., Rommens, J. M., Uyama, E., Nohira, O., et al. (1998) Short GCG expansions in the PABP2 gene cause oculopharyngeal muscular dystrophy. Nat. Genet., 18, 164–167
Sreedharan, J., Blair, I. P., Tripathi, V. B., Hu, X., Vance, C., Rogelj, B., Ackerley, S., Durnall, J. C., Williams, K. L., Buratti, E., et al. (2008) TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science, 319, 1668–1672
Bentley, D. L. (2014) Coupling mRNA processing with transcription in time and space. Nat. Rev. Genet., 15, 163–175
Baltz, A. G., Munschauer, M., Schwanhäusser, B., Vasile, A., Murakawa, Y., Schueler, M., Youngs, N., Penfold-Brown, D., Drew, K., Milek, M., et al. (2012) The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell, 46, 674–690
Gerstberger, S., Hafner, M. and Tuschl, T. (2014) A census of human RNA-binding proteins. Nat. Rev. Genet., 15, 829–845
Dreyfuss, G., Kim, V. N. and Kataoka, N. (2002) Messenger-RNAbinding proteins and the messages they carry. Nat. Rev. Mol. Cell Biol., 3, 195–205
Setzer, D. R. (1999) Measuring equilibrium and kinetic constants using gel retardation assays. Methods Mol. Biol., 118, 115–128
Katsamba, P. S., Park, S. and Laird-Offringa, I. A. (2002) Kinetic studies of RNA-protein interactions using surface plasmon resonance. Methods, 26, 95–104
Hook, B., Bernstein, D., Zhang, B. and Wickens, M. (2005) RNAprotein interactions in the yeast three-hybrid system: affinity, sensitivity, and enhanced library screening. RNA, 11, 227–233
Ellington, A. D. and Szostak, J. W. (1990) In vitro selection of RNA molecules that bind specific ligands. Nature, 346, 818–822
Tuerk, C. and Gold, L. (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science, 249, 505–510
Tenenbaum, S. A., Carson, C. C., Lager, P. J. and Keene, J. D. (2000) Identifying mRNA subsets in messenger ribonucleoprotein complexes by using cDNA arrays. Proc. Natl. Acad. Sci. USA, 97, 14085–14090
Keene, J. D., Komisarow, J. M. and Friedersdorf, M. B. (2006) RIP-chip: the isolation and identification of mRNAs, microRNAs and protein components of ribonucleoprotein complexes from cell extracts. Nat. Protoc., 1, 302–307
Mili, S. and Steitz, J. A. (2004) Evidence for reassociation of RNA-binding proteins after cell lysis: implications for the interpretation of immunoprecipitation analyses. RNA, 10, 1692–1694
Dreyfuss, G., Adam, S. A. and Choi, Y. D. (1984) Physical change in cytoplasmic messenger ribonucleoproteins in cells treated with inhibitors of mRNA transcription. Mol. Cell. Biol., 4, 415–423
Ule, J., Jensen, K. B., Ruggiu, M., Mele, A., Ule, A. and Darnell, R. B. (2003) CLIP identifies Nova-regulated RNA networks in the brain. Science, 302, 1212–1215
Sugimoto, Y., Vigilante, A., Darbo, E., Zirra, A., Militti, C., D’Ambrogio, A., Luscombe, N. M. and Ule, J. (2015) hiCLIP reveals the in vivo atlas of mRNA secondary structures recognized by Staufen 1. Nature, 519, 491–494
Kudla, G., Granneman, S., Hahn, D., Beggs, J. D. and Tollervey, D. (2011) Cross-linking, ligation, and sequencing of hybrids reveals RNA-RNA interactions in yeast. Proc. Natl. Acad. Sci. USA, 108, 10010–10015
Zarnegar, B. J., Flynn, R. A., Shen, Y., Do, B. T., Chang, H. Y. and Khavari, P. A. (2016) irCLIP platform for efficient characterization of protein-RNA interactions. Nat. Methods, 13, 489–492
Singh, G., Ricci, E. P. and Moore, M. J. (2014) RIPiT-Seq: a highthroughput approach for footprinting RNA:protein complexes. Methods, 65, 320–332
McMahon, A. C., Rahman, R., Jin, H., Shen, J. L., Fieldsend, A., Luo, W. and Rosbash, M. (2016) TRIBE: hijacking an RNAediting enzyme to identify cell-specific targets of RNA-binding proteins. Cell, 165, 742–753
Kargapolova, Y., Levin, M., Lackner, K. and Danckwardt, S. (2017) sCLIP—an integrated platform to study RNA-protein interactomes in biomedical research: identification of CSTF2tau in alternative processing of small nuclear RNAs. Nucleic Acids Res., 45, 6074–6086
Rosenberg, M., Blum, R., Kesner, B., Maier, V. K., Szanto, A. and Lee, J. T. (2017) Denaturing CLIP, dCLIP, pipeline identifies discrete RNA footprints on chromatin-associated proteins and reveals that CBX7 targets 3′ UTRs to regulate mRNA expression. Cell Syst., 5, 368–385
Brugiolo, M., Botti, V., Liu, N., Müller-McNicoll, M. and Neugebauer, K. M. (2017) Fractionation iCLIP detects persistent SR protein binding to conserved, retained introns in chromatin, nucleoplasm and cytoplasm. Nucleic Acids Res., 45, 10452–10465
Licatalosi, D. D., Mele, A., Fak, J. J., Ule, J., Kayikci, M., Chi, S. W., Clark, T. A., Schweitzer, A. C., Blume, J. E., Wang, X., et al. (2008) HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature, 456, 464–469
Darnell, R. B. (2010) HITS-CLIP: panoramic views of protein-RNA regulation in living cells. Wiley Interdiscip. Rev. RNA, 1, 266–286
Hafner, M., Landthaler, M., Burger, L., Khorshid, M., Hausser, J., Berninger, P., Rothballer, A., Ascano, M. Jr, Jungkamp, A. C., Munschauer, M., et al. (2010) Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell, 141, 129–141
Kishore, S., Jaskiewicz, L., Burger, L., Hausser, J., Khorshid, M. and Zavolan, M. (2011) A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins. Nat. Methods, 8, 559–564
König, J., Zarnack, K., Rot, G., Curk, T., Kayikci, M., Zupan, B., Turner, D. J., Luscombe, N. M. and Ule, J. (2010) iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol., 17, 909–915
Urlaub, H., Hartmuth, K. and Lührmann, R. (2002) A two-tracked approach to analyze RNA-protein crosslinking sites in native, nonlabeled small nuclear ribonucleoprotein particles. Methods, 26, 170–181
Sugimoto, Y., König, J., Hussain, S., Zupan, B., Curk, T., Frye, M. and Ule, J. (2012) Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions. Genome Biol., 13, R67
Marchese, D., de Groot, N. S., Lorenzo Gotor, N., Livi, C. M. and Tartaglia, G. G. (2016) Advances in the characterization of RNAbinding proteins. Wiley Interdiscip. Rev. RNA, 7, 793–810
Van Nostrand, E. L., Pratt, G. A., Shishkin, A. A., Gelboin-Burkhart, C., Fang, M. Y., Sundararaman, B., Blue, S. M., Nguyen, T. B., Surka, C., Elkins, K., et al. (2016) Robust transcriptomewide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods, 13, 508–514
Björling, E. and Uhlén, M. (2008) Antibodypedia, a portal for sharing antibody and antigen validation data. Mol. Cell. Proteomics, 7, 2028–2037
de Boer, E., Rodriguez, P., Bonte, E., Krijgsveld, J., Katsantoni, E., Heck, A., Grosveld, F. and Strouboulis, J. (2003) Efficient biotinylation and single-step purification of tagged transcription factors in mammalian cells and transgenic mice. Proc. Natl. Acad. Sci. USA, 100, 7480–7485
Gerace, E. and Moazed, D. (2015) Affinity purification of protein complexes using TAP tags. Methods Enzymol., 559, 37–52
Nelles, D. A., Fang, M. Y., O’Connell, M. R., Xu, J. L., Markmiller, S. J., Doudna, J. A. and Yeo, G. W. (2016) Programmable RNA tracking in live cells with CRISPR/Cas9. Cell, 165, 488–496
Wheeler, E. C., Van Nostrand, E. L. and Yeo, G. W. (2018) Advances and challenges in the detection of transcriptome-wide protein-RNA interactions. Wiley Interdiscip. Rev. RNA, 9, e1436
Maragkakis, M., Alexiou, P., Nakaya, T. and Mourelatos, Z. (2016) CLIPSeqTools — a novel bioinformatics CLIP-seq analysis suite. RNA, 22, 1–9
Kucukural, A., Özadam, H., Singh, G., Moore, M. J. and Cenik, C. (2013) ASPeak: an abundance sensitive peak detection algorithm for RIP-seq. Bioinformatics, 29, 2485–2486
Khorshid, M., Rodak, C. and Zavolan, M. (2011) CLIPZ: a database and analysis environment for experimentally determined binding sites of RNA-binding proteins. Nucleic Acids Res., 39, D245–D252
Chen, B., Yun, J., Kim, M. S., Mendell, J. T. and Xie, Y. (2014) PIPE-CLIP: a comprehensive online tool for CLIP-seq data analysis. Genome Biol., 15, R18
Uren, P. J., Bahrami-Samani, E., Burns, S. C., Qiao, M., Karginov, F. V., Hodges, E., Hannon, G. J., Sanford, J. R., Penalva, L. O. and Smith, A. D. (2012) Site identification in high-throughput RNAprotein interaction data. Bioinformatics, 28, 3013–3020
Wang, T., Chen, B., Kim, M., Xie, Y. and Xiao, G. (2014) A model-based approach to identify binding sites in CLIP-seq data. PLoS One, 9, e93248
Althammer, S., González-Vallinas, J., Ballaré, C., Beato, M. and Eyras, E. (2011) Pyicos: a versatile toolkit for the analysis of highthroughput sequencing data. Bioinformatics, 27, 3333–3340
Lovci, M. T., Ghanem, D., Marr, H., Arnold, J., Gee, S., Parra, M., Liang, T. Y., Stark, T. J., Gehman, L. T., Hoon, S., et al. (2013) Rbfox proteins regulate alternative mRNA splicing through evolutionarily conserved RNA bridges. Nat. Struct. Mol. Biol., 20, 1434–1442
Webb, S., Hector, R. D., Kudla, G. and Granneman, S. (2014) PAR-CLIP data indicate that Nrd1-Nab3-dependent transcription termination regulates expression of hundreds of protein coding genes in yeast. Genome Biol., 15, R8
Corcoran, D. L., Georgiev, S., Mukherjee, N., Gottwein, E., Skalsky, R. L., Keene, J. D. and Ohler, U. (2011) PARalyzer: definition of RNA binding sites from PAR-CLIP short-read sequence data. Genome Biol., 12, R79
Bailey, T. L., Johnson, J., Grant, C. E. and Noble, W. S. (2015) The MEME Suite. Nucleic Acids Res., 43, W39–W49
Heinz, S., Benner, C., Spann, N., Bertolino, E., Lin, Y. C., Laslo, P., Cheng, J. X., Murre, C., Singh, H. and Glass, C. K. (2010) Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell, 38, 576–589
Weyn-Vanhentenryck, S. M. and Zhang, C. (2016) mCarts: genome-wide prediction of clustered sequence motifs as binding sites for RNA-binding proteins. Methods Mol. Biol., 1421, 215–226
Kulakovskiy, I. V., Boeva, V. A., Favorov, A. V. and Makeev, V. J. (2010) Deep and wide digging for binding motifs in ChIP-seq data. Bioinformatics, 26, 2622–2623
Zhang, C. and Darnell, R. B. (2011) Mapping in vivo protein-RNA interactions at single-nucleotide resolution from HITS-CLIP data. Nat. Biotechnol., 29, 607–614
Bahrami-Samani, E., Penalva, L. O., Smith, A. D. and Uren, P. J. (2015) Leveraging cross-link modification events in CLIP-seq for motif discovery. Nucleic Acids Res., 43, 95–103
Ray, D., Kazan, H., Chan, E. T., Castillo, L. P., Chaudhry, S., Talukder, S., Blencowe, B. J., Morris, Q. and Hughes, T. R. (2009) Rapid and systematic analysis of the RNA recognition specificities of RNA-binding proteins. Nat. Biotechnol., 27, 667–670
Lambert, N., Robertson, A., Jangi, M., McGeary, S., Sharp, P. A. and Burge, C. B. (2014) RNA Bind-n-Seq: quantitative assessment of the sequence and structural binding specificity of RNA binding proteins. Mol. Cell, 54, 887–900
Liu, H. X., Zhang, M. and Krainer, A. R. (1998) Identification of functional exonic splicing enhancer motifs recognized by individual SR proteins. Genes Dev., 12, 1998–2012
Johnson, J. M., Castle, J., Garrett-Engele, P., Kan, Z., Loerch, P. M., Armour, C. D., Santos, R., Schadt, E. E., Stoughton, R. and Shoemaker, D. D. (2003) Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science, 302, 2141–2144
Ule, J., Ule, A., Spencer, J., Williams, A., Hu, J. S., Cline, M., Wang, H., Clark, T., Fraser, C., Ruggiu, M., et al. (2005) Nova regulates brain-specific splicing to shape the synapse. Nat. Genet., 37, 844–852
Sugnet, C. W., Srinivasan, K., Clark, T. A., O’Brien, G., Cline, M. S., Wang, H., Williams, A., Kulp, D., Blume, J. E., Haussler, D., et al. (2006) Unusual intron conservation near tissue-regulated exons found by splicing microarrays. PLoS Comput. Biol., 2, e4
Boutz, P. L., Stoilov, P., Li, Q., Lin, C. H., Chawla, G., Ostrow, K., Shiue, L., Ares, M. Jr and Black, D. L. (2007) A posttranscriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev., 21, 1636–1652
Licatalosi, D. D., Yano, M., Fak, J. J., Mele, A., Grabinski, S. E., Zhang, C. and Darnell, R. B. (2012) Ptbp2 represses adult-specific splicing to regulate the generation of neuronal precursors in the embryonic brain. Genes Dev., 26, 1626–1642
Hannigan, M. M., Zagore, L. L. and Licatalosi, D. D. (2017) Ptbp2 controls an alternative splicing network required for cell communication during spermatogenesis. Cell Reports, 19, 2598–2612
Wang, E. T., Cody, N. A., Jog, S., Biancolella, M., Wang, T. T., Treacy, D. J., Luo, S., Schroth, G. P., Housman, D. E., Reddy, S., et al. (2012) Transcriptome-wide regulation of pre-mRNA splicing and mRNA localization by muscleblind proteins. Cell, 150, 710–724
Charizanis, K., Lee, K. Y., Batra, R., Goodwin, M., Zhang, C., Yuan, Y., Shiue, L., Cline, M., Scotti, M. M., Xia, G., et al. (2012) Muscleblind-like 2-mediated alternative splicing in the developing brain and dysregulation in myotonic dystrophy. Neuron, 75, 437–450
Änkö, M. L., Müller-McNicoll, M., Brandl, H., Curk, T., Gorup, C., Henry, I., Ule, J. and Neugebauer, K. M. (2012) The RNAbinding landscapes of two SR proteins reveal unique functions and binding to diverse RNA classes. Genome Biol., 13, R17
Pandit, S., Zhou, Y., Shiue, L., Coutinho-Mansfield, G., Li, H., Qiu, J., Huang, J., Yeo, G. W., Ares, M. Jr and Fu, X. D. (2013) Genome-wide analysis reveals SR protein cooperation and competition in regulated splicing. Mol. Cell, 50, 223–235
Weyn-Vanhentenryck, S. M., Mele, A., Yan, Q., Sun, S., Farny, N., Zhang, Z., Xue, C., Herre, M., Silver, P. A., Zhang, M. Q., et al. (2014) HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Reports, 6, 1139–1152
Damianov, A., Ying, Y., Lin, C. H., Lee, J. A., Tran, D., Vashisht, A. A., Bahrami-Samani, E., Xing, Y., Martin, K. C., Wohlschlegel, J. A., et al. (2016) Rbfox proteins regulate splicing as part of a large multiprotein complex LASR. Cell, 165, 606–619
Wang, E. T., Ward, A. J., Cherone, J. M., Giudice, J., Wang, T. T., Treacy, D. J., Lambert, N. J., Freese, P., Saxena, T., Cooper, T. A., et al. (2015) Antagonistic regulation of mRNA expression and splicing by CELF and MBNL proteins. Genome Res., 25, 858–871
Kapeli, K., Pratt, G. A., Vu, A. Q., Hutt, K. R., Martinez, F. J., Sundararaman, B., Batra, R., Freese, P., Lambert, N. J., Huelga, S. C., et al. (2016) Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses. Nat. Commun., 7, 12143
Rothrock, C. R., House, A. E. and Lynch, K. W. (2005) HnRNP L represses exon splicing via a regulated exonic splicing silencer. EMBO J., 24, 2792–2802
Motta-Mena, L. B., Heyd, F. and Lynch, K. W. (2010) Contextdependent regulatory mechanism of the splicing factor hnRNP L. Mol. Cell, 37, 223–234
Zhang, C., Frias, M. A., Mele, A., Ruggiu, M., Eom, T., Marney, C. B., Wang, H., Licatalosi, D. D., Fak, J. J. and Darnell, R. B. (2010) Integrative modeling defines the Nova splicing-regulatory network and its combinatorial controls. Science, 329, 439–443
Yang, Y., Park, J. W., Bebee, T. W., Warzecha, C. C., Guo, Y., Shang, X., Xing, Y. and Carstens, R. P. (2016) Determination of a comprehensive alternative splicing regulatory network and combinatorial regulation by key factors during the epithelial-tomesenchymal transition. Mol. Cell. Biol., 36, 1704–1719
Dittmar, K. A., Jiang, P., Park, J. W., Amirikian, K., Wan, J., Shen, S., Xing, Y. and Carstens, R. P. (2012) Genome-wide determination of a broad ESRP-regulated posttranscriptional network by high-throughput sequencing. Mol. Cell. Biol., 32, 1468–1482
Batra, R., Charizanis, K., Manchanda, M., Mohan, A., Li, M., Finn, D. J., Goodwin, M., Zhang, C., Sobczak, K., Thornton, C. A., et al. (2014) Loss of MBNL leads to disruption of developmentally regulated alternative polyadenylation in RNA-mediated disease. Mol. Cell, 56, 311–322
Masuda, A., Takeda, J., Okuno, T., Okamoto, T., Ohkawara, B., Ito, M., Ishigaki, S., Sobue, G. and Ohno, K. (2015) Positionspecific binding of FUS to nascent RNA regulates mRNA length. Genes Dev., 29, 1045–1057
Rot, G., Wang, Z., Huppertz, I., Modic, M., Lence, T., Hallegger, M., Haberman, N., Curk, T., von Mering, C. and Ule, J. (2017) High-resolution RNA maps suggest common principles of splicing and polyadenylation regulation by TDP-43. Cell Reports, 19, 1056–1067
Hurt, J. A., Robertson, A. D. and Burge, C. B. (2013) Global analyses of UPF1 binding and function reveal expanded scope of nonsense-mediated mRNA decay. Genome Res., 23, 1636–1650
Masuda, A., Andersen, H. S., Doktor, T. K., Okamoto, T., Ito, M., Andresen, B. S. and Ohno, K. (2012) CUGBP1 and MBNL1 preferentially bind to 3′ UTRs and facilitate mRNA decay. Sci. Rep., 2, 209
Moore, M. J., Zhang, C., Gantman, E. C., Mele, A., Darnell, J. C. and Darnell, R. B. (2014) Mapping Argonaute and conventional RNA-binding protein interactions with RNA at single-nucleotide resolution using HITS-CLIP and CIMS analysis. Nat. Protoc., 9, 263–293
Helwak, A., Kudla, G., Dudnakova, T. and Tollervey, D. (2013) Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell, 153, 654–665
Majoros, W. H., Lekprasert, P., Mukherjee, N., Skalsky, R. L., Corcoran, D. L., Cullen, B. R. and Ohler, U. (2013) MicroRNA target site identification by integrating sequence and binding information. Nat. Methods, 10, 630–633
Reczko, M., Maragkakis, M., Alexiou, P., Grosse, I. and Hatzigeorgiou, A. G. (2012) Functional microRNA targets in protein coding sequences. Bioinformatics, 28, 771–776
Liu, C., Mallick, B., Long, D., Rennie, W. A., Wolenc, A., Carmack, C. S. and Ding, Y. (2013) CLIP-based prediction of mammalian microRNA binding sites. Nucleic Acids Res., 41, e138
Khorshid, M., Hausser, J., Zavolan, M. and van Nimwegen, E. (2013) A biophysical miRNA-mRNA interaction model infers canonical and noncanonical targets. Nat. Methods, 10, 253–255
Lu, Y. and Leslie, C. S. (2016) Learning to predict miRNA-mRNA interactions from AGO CLIP sequencing and CLASH data. PLoS Comput. Biol., 12, e1005026
Zhu, Y., Wang, X., Forouzmand, E., Jeong, J., Qiao, F., Sowd, G. A., Engelman, A. N., Xie, X., Hertel, K. J. and Shi, Y. (2018) Molecular mechanisms for CFIm-mediated regulation of mRNA alternative polyadenylation. Mol. Cell, 69, 62–74
Simon, M. D. (2013) Capture hybridization analysis of RNA targets (CHART). Curr. Protoc. Mol. Biol. 101, 21.25.1–21.25.16
Chu, C., Quinn, J. and Chang, H. Y. (2012) Chromatin isolation by RNA purification (ChIRP). J. Vis. Exp., 61, 3912
McHugh, C. A., Chen, C. K., Chow, A., Surka, C. F., Tran, C., McDonel, P., Pandya-Jones, A., Blanco, M., Burghard, C., Moradian, A., et al. (2015) The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature, 521, 232–236
Castello, A., Horos, R., Strein, C., Fischer, B., Eichelbaum, K., Steinmetz, L. M., Krijgsveld, J. and Hentze, M. W. (2016) Comprehensive identification of RNA-binding proteins by RNA interactome capture. Methods Mol. Biol., 1358, 131–139
Bao, X., Guo, X., Yin, M., Tariq, M., Lai, Y., Kanwal, S., Zhou, J., Li, N., Lv, Y., Pulido-Quetglas, C., et al. (2018) Capturing the interactome of newly transcribed RNA. Nat. Methods, 15, 213–220
Hwang, H. W., Park, C. Y., Goodarzi, H., Fak, J. J., Mele, A., Moore, M. J., Saito, Y. and Darnell, R. B. (2016) PAPERCLIP identifies microRNA targets and a role of CstF64/64tau in promoting non-canonical poly(A) site usage. Cell Reports, 15, 423–435
Acknowledgements
This work was supported by funds from the National Institutes of Health to LLZ (T32 GM08056) and DDL (R01 GM107331), USA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Author summary: Rapidly advancing sequencing methodologies continue to improve our understanding of gene regulation and disease pathogenesis. An emerging area of interest is the contribution of RNA regulation in development and disease. In this review, we highlight techniques widely used to interrogate RNA regulatory networks that fine tune gene expression and how they impact cell biology.
Rights and permissions
About this article
Cite this article
Hannigan, M.M., Zagore, L.L. & Licatalosi, D.D. Mapping transcriptome-wide protein-RNA interactions to elucidate RNA regulatory programs. Quant Biol 6, 228–238 (2018). https://doi.org/10.1007/s40484-018-0145-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-018-0145-6