Abstract
Geological events and glacial history have all induced habitat fragmentation, which has long been recognized as a major threat to the survival of many species. However, fragmentation has differential effects on population genetic patterns of individual species, such as loss of genetic diversity, genetic differentiation enhance, inbreeding increase, allelic deletion, and no effect. Calocedrus macrolepis Kurz is an endangered ancient relictual and fragmented conifer. In this study, the genetic diversity and structure were analyzed to shed light on the factors that determine their contemporary genetic patterns and to provide optimum strategies for future conservation. The genetic diversity within and among 14 extant populations of C. macrolepis was analyzed using nine microsatellite markers. The genetic diversity (H E = 0.636), genetic differentiation (F ST = 0.163), and gene flow (Nm = 1.496) were disclosed, respectively. No significant correlation was detected by the Mantel test between genetic and geographical distances among pair-wise comparisons of populations (r = 0.197, P = 0.103). Fourteen populations could be generally assigned to two separate groups, and significant asymmetrical migration among populations within the regional groups was revealed. Both Quaternary glaciations and neotectonic movements seem to be associated with a long history of population contraction and fragmentation of C. macrolepis, enhancing its genetic drift and population divergence. The results indicate that long-term habitat fragmentation could be responsible for the genetic structure observed. In situ conservation strategies should be designed, especially small and isolated populations. Also, special attention should be given to populations LY and DZ because of their private alleles, as well as to population CJ isolated from the mainland.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Habitat fragmentation threatens the survival of many species due to the formation of smaller and discrete plant populations that eventually become more spatially isolated (Fahrig and Merriam 1994; Young et al. 1996; Frankham et al. 2002). If populations are separated over successive generations, their heterozygosity decreases because of greater random genetic drift and inbreeding (Rumball et al. 1994). This produces a series of negative effects, e.g., the accumulation of deleterious recessive alleles, lower fecundity in individual plants, higher seed/seedling mortality, and slower individual growth rates, thereby reducing fitness and perhaps leading to local extinction (Lande 1988; Young et al. 1996; Newman and Pilson 1997; Aguilar et al. 2008).
Also, certain life-history traits in plants may confer different levels of vulnerabilities to fragmentation effects. For example, the onset of genetic influences from fragmentation is frequently delayed in long-lived woody species (Torres-Díaz et al. 2007). For long-lived woody species, two alternative hypotheses might explain why they can counteract negative inputs (Van Geert et al. 2008). First, although the habitats for some species might become fragmented, their populations are still interconnected through pollen flow and/or seed dispersal, which can counteract genetic erosion (Ellstrand 1992; Kwak et al. 1998; Newman and Tallmon 2001). Second, for some species with relatively long-life cycles of up to a few hundred even over 1000 years, the long lifespan of individual plants reflects a historic situation in which habitat fragmentation is less expansive (Ellstrand and Elam 1993; Oostermeijer et al. 1994; Sork et al. 1999; Honnay et al. 2005). However, evidence from both theoretical (Hanski and Ovaskainen 2002) and empirical (Lindborg and Eriksson 2004) indicates that a woody species can become extinct after 50–100 years of habitat fragmentation. Thus, when making conservation decisions about endangered species, it is of the utmost importance to understand the genetic variation and structure of the species in order to elucidate the factors that determine the genetic architecture of plant populations (Milligan et al. 1994; Schaal et al. 1998; Avise 2000). Research on the influence of habitat fragmentation has largely concentrated on assessing the genetic consequences of recent events associated with human activity (Mona et al. 2014). For example, Fagus sylvatica L. is a wind-pollinated tree, which dominates forest over large regions of Europe. The habitat fragmentation induced by historic deforestation in this species has led to genetic bottlenecks and the disruption of the species’ breeding system (Alistair and Josep 2006). In Scotland, deforestation caused chronic habitat fragmentation on Sorbus aucuparia Poir., which led to the reduced pollen-mediated gene flow (Bacles et al. 2004). English yew (Taxus baccata L.) is a Tertiary relict, whose genetic variation was highly structured at the local scale, while only a low proportion of the observed variation was attributed to regional differences. The patterns probably reflect the combined effects of Quaternary climatic changes and recent impact of human activities (González-Martínez et al. 2010). Although the effects of long-term fragmentation on genetic structure have been reported, but the role that historical processes play in affecting population genetic structure is still unclear.
C. macrolepis (Chinese Incense-cedar) is a relictual, long-lived woody plant. It is native to southwestern China [southeast of the Yunnan-Guizhou Plateau (YGP)] and the adjacent regions of Myanmar and Vietnam (Fu and Yu 1999; Farjon 2005). The genus includes four species, which have a typical eastern Asia–western North American disjunct pattern of distribution and represent an ancient lineage (Averyanov et al. 2008; Chen et al. 2009). It is an endangered ancient relictual and fragmented conifer, and is a heliophile and subtropical plant. It is monoecious with unisexual flowers on the same branches. It belongs to wind-pollinated plant and contains light-weight seeds that have unequal wings (Liao et al. 2014; Wang et al. 2004). Based on fossil evidence from the European Tertiary, C. macrolepis was formerly widespread in temperate regions (Kracek 1999), and sediments also exist from the Asiatic Oligocene to Pliocene (WGCPC 1978; Liu et al. 1996). Because of its valuable wood, it has been threatened by over-harvesting in the past century and its habitat has also been severely degraded ecologically. Its wild populations are dramatically shrinking, and an overall decline continues throughout its entire area of distribution (Liao et al. 2014; Liu et al. 2010). According to criteria from the 2001 International Union for the Conservation of Nature and Natural Resources Red List (Farjon 2001), C. macrolepis is classified as a vulnerable species and is listed as an endangered species in China (Fu 1995) and Vietnam (WCMC 1997).
Although few microsatellite characterizations have been made of its genetic population structure and diversity, recently markers have been developed (Liao et al. 2010). In this study, we conducted genetic diversity and structure analyses of this species by microsatellite. Our objectives were to examine: (1) the degree of genetic diversity within and among remnant populations, (2) significant genetic differentiation among our study populations, (3) factors that determine their genetic patterns, and (4) optimum strategies for future conservation.
Materials and methods
Population descriptions and sample collection
In China, Calocedrus macrolepis is distributed on sites within Yunnan and southern Guizhou Provinces, at elevations ranging from 300 to 2000 m. This species also can occur sporadically in western Guangxi and Hainan Provinces. As a medium-sized, subtropical evergreen, trees can be 15–25 m tall, with trunks as large as 60–80 cm in diameter at breast height (dbh). Remnant populations are generally fragmented and highly disturbed because of heavy deforestation. We analyzed 14 isolated populations that represent those found throughout China. These included eight from Yunnan Province, three from Guizhou Province, two from Guangxi Province, and one from Hainan province (Fig. 1; Table 1). For each population, Needle samples from 17 to 34 mature individuals (dbh > 10 cm) were randomly collected. After collection, samples were dried in silica gel and stored at 4 °C prior to DNA extractions.
DNA extraction, microsatellite amplification, and scoring
Genomic DNA samples were extracted from dry leaf tissue, which was grounded in liquid nitrogen according to CTAB methodology (Doyle and Doyle 1987). Nuclear DNA was PCR-amplified by microsatellite primer. Nine microsatellite primers (Cm2–Cm4, Cm6–Cm8, and Cm10–Cm12) (Liao et al. 2010) that showed good repetition and distinct polymorphism were selected for this study. Amplification conditions were as described by Liao et al. (2010). PCR products were separated on an 8 % denaturing polyacrylamide gel, using a 20 bp DNA ladder molecular size standard (TaKaRa, China) and silver staining.
Genetic diversity, Hardy–Weinberg equilibrium, and linkage disequilibrium
To estimate overall genetic diversity, we used GenAlEx, Version 6 (Peakall and Smouse 2006) to calculate the following measures: mean number of alleles per locus (Na), average number of effective alleles (Ne), observed heterozygosity (H O) and expected heterozygosity (H E) for each locus and population, as well as private alleles (Np), and the inbreeding coefficient (F IS) for each population. The presence of null allele was tested via MICRO-CHECKER 2.2 (Van Oosterhout et al. 2004). We applied the Markov chain method for estimating the probability of significant deviation from the Hardy–Weinberg equilibrium (HWE) and also used default parameters to perform pair-wise tests for linkage disequilibrium (LD), using the online version of GENEPOP (2013). Sequential Bonferroni corrections were made to determine significance levels for all tests at P < 0.05 (Rice 1989).
Genetic differentiation and assignment analyses
Overall and pair-wise values for genetic differentiation (F ST) were generated by the AMOVA routine in GenAlEx program, Version 6. Significance of the differentiation between pairs of populations was tested using permutation procedures (1000 replicates). The average effective number of migrants exchanged between populations in each generation, i.e., gene flow (Nm), was estimated from F ST.
Cluster analysis
Unbiased genetic distances (Nei 1978) were computed with PopGene software, Version 1.32 (Yeh et al. 1997). A dendrogram based on these resulting coefficients of genetic distance was constructed using the unweighted pair-group method with arithmetic mean (UPGMA). This distance matrix was analyzed with NTSYSpc, Version 2.11c (Rohlf 1997). Cluster analysis (UPGMA) and dendrogram construction were performed with the SHAN program in NTSYSpc to determine the genetic distance among the 14 populations. A simple Mantel test was conducted with TFPGA 1.3 (Miller 1997) to assess the correlation between the Nei’s unbiased genetic distance and the geographic distance that separated populations.
Genetic structure
Bayesian clustering was used for inferring the population structure by STRUCTURE v2.3.4 (Pritchard et al. 2000). The each level of (K = 1–20) was tested and determined by a no-admixture model with 106 iterations and 105 burn-in steps. Ten independent runs and tests for each K were performed to ensure consistent results. Different clustering scenarios were evaluated using the mean log probability [L(K)] and the change in log probability, ΔL(K) (Evanno et al. 2005).
Long-term migration and effective population size
The MIGRATE-N 3.6.5 (2013) was employed to estimate long-term migration rates and effective population sizes. Analyses assumed a Brownian motion model which represents a continuous approximation to the stepwise mutation model for microsatellite evolution (Kimura and Ohta 1978). Long-term migration rates, M, 4N e generations in the past based on a coalescent approach, were estimated using the maximum-likelihood mode. MIGRATE-N tests all possible combinations of symmetrical or no migration between each regional population group and between populations within a region with the likelihood ratio test. MIGRATE-N estimates the long-term effective population size (N e) of each population group as parameter θ (that is, 4N e μ, where μ is the mutation rate per site). MIGRATE-N also estimates the bi-directional M among population groups and among populations within a region (M = m/μ, where m is the immigration rate per generation) (Hsieh et al. 2013). The Markov Chain settings consisted of ten short and three long chains per run, with an increment of 20. For short and long chains, 500 and 1000 genealogies were respectively recorded, with a burn-in of 1000.
Results
Genetic diversity
A total of 70 alleles at nine microsatellite loci were revealed across 291 individuals of 14 natural populations for C. macrolepis. Allele numbers ranged from four at locus Cm7 to 13 at locus Cm12, with a mean value of 7.8 per locus. Mean gene diversity ranged from H E = 0.497 at locus Cm7 to H E = 0.748 at locus Cm10, H O values ranged from 0.534 at locus Cm8 to 0.869 at locus Cm3. Null alleles were detected at a negligible frequency in two loci (Cm11 and Cm12). The average variation for intrapopulation microsatellites was Na = 4.7 (3.6–5.9), Ne = 3.2 (2.5–4.0), H E = 0.636 (0.563–0.701), and H O = 0.712 (0.467–0.844). Four private alleles (Np) were detected in two populations—DZ (1; 2.5 %) and LY (3; 5–25 %) (Table 2). The level of inbreeding for each population was between −0.311 and 0.365, and significant inbreeding coefficients (F) were detected in the LY population (F = 0.365) (Table 2). When all loci were combined, nine populations deviated from the HWE at P < 0.05 (Table 2). Significant linkage disequilibrium between SSR loci within each population was detected for 31 out of 485 comparisons; only four remained significant after the Bonferroni correction (P ≤ 0.05), suggesting that the loci used in this study were independent of each other.
Population genetics differentiation and structure
The global genetic differentiation coefficient (F ST) was 0.163 (Table 3). Pair-wise F ST estimates ranged from 0.060 (LS vs. JH) to 0.245 (CN vs. LY) (Table 4). Historical gene flow (Nm) between pairs of populations averaged Nm = 1.562 and ranged from 0.770 (between CN and LY) to 3.895 (between JH and LS) (Table 3). No significant correlation was detected by the Mantel test between genetic and geographical distances among pair-wise comparisons (r = 0.197, P = 0.103). The relationships among these 14 populations were clustered based on Nei’s genetic distances (Fig. 2). Population CJ from Hainan province was clustered with YM and DZ from Yunnan and Guizhou provinces, respectively. The easternmost population, LS, was grouped with the westernmost population, TC. Guangxi populations HJ and LY were clustered into another group.
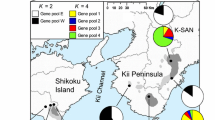
Based on nine loci, a global analysis of pooled data from all populations was implemented by Structure program. The results showed that individuals of the 14 C. macrolepis populations could be generally assigned to two separate groups (western and southern) (Fig. 3) according to changes in the log probability (ΔL(K); Fig. 4). Western group included populations Yimen (YM), Tengchong (TC), Changning (CN), Leishan (LS), Jianhe (JH), Lincang (LC), Mojiang (MJ), Puer (PE), Shiping (SP), and Yuanjiang (YJ). Southern group comprised populations Changjiang (CJ), Huanjiang (HJ), Danzhai (DZ), and Leyie (LY). According to historical migration rates (M) calculated by MIGRATE-N, there is no significant asymmetrical migration among the two regional groups. However, significant asymmetrical migration among populations within the western and southern regional groups according to non-overlapping 95 % CIs was revealed (Table 5; Fig. 5, respectively).
Analysis with STRUCTURE for K = 2. Each color represents a different cluster. Black lines separate populations. See Table 1 for population codes
Graphical representation of results of migrations estimated using MIGRATE-N among the four regional populations. Maximum-likehood estimates and extreme value in each locus (in parentheses) of the long-term migration rate (M) and mutation-scaled effective population size (θ) using the MIGRATE-N program. Arrows represent the direction of gene flow among sites. Dotted lines means no value
Discussion
Genetic diversity
Compared with previously reported genetic patterns for C. macrolepis that were based on inter simple sequence repeat (ISSR) analysis (mean H E = 0.1114, overall F ST = 0.042) (Wang et al. 2004), we found much greater variation and higher differentiation in our 14 remnant natural populations (mean H E = 0.636, overall F ST = 0.163). One potential explanation for this discrepancy is that only five populations were sampled in that earlier study, which may have led to somewhat biased estimates of genetic diversity. Another possibility is that techniques utilizing dominant molecular markers, such as ISSR, AFLP (amplified fragment length polymorphisms), or RAPD (rapid amplification of polymorphic DNA), provide limited resolution when examining interpopulation differentiation and gene flow among populations. By contrast, microsatellite markers that rely on specific primers for a certain species, and which are present in all species genomes, are highly polymorphic, codominant, and widespread across the genome (Rallo et al. 2000; Sefc et al. 2000). Therefore, they can be used to discriminate between heterozygotes and homozygotes, and can truly reveal the genetic diversity within a species.
Compared with other plants, most conifers show a relatively high level (H E = 0.151) of variation within populations and a low level (F ST = 0.073) of differentiation among populations, as measured by isozyme markers (Hamrick et al. 1992; Matusova 1995). However, values for genetic diversity vary among types of markers, making it necessary to compare diversities in different species based on the same molecular markers. Here, we found that diversity (H E = 0.636) was similar with the mean values (H E = 0.65) reported from studies of other regional species such as Larix occidentalis Nutt. (H E = 0.58) (Khasa et al. 2006) Fitzroya cupressoides I. M. Johnst. (H E = 0.55) (Allnutt et al. 1999) and long-lived perennial species (H E = 0.68) (Nybom 2004) such as Araucaria araucana (Molina) K. Koch (H E = 0.69) (Bekessy et al. 2002), Quercus lobata Née (H E = 0.78), Juniperus oxycedrus Hochst. (H E = 0.72) (Boratyński et al. 2014), and Quercus douglasii Hook. & Arn. (H E = 0.88) (Craft et al. 2002). However, the diversity was much lower than that estimated with SSR markers for five pinaceae species with broad distribution (H E = 0.734–0.815, mean = 0.79) (Mariette et al. 2001; Acheré et al. 2005; Maghuly et al. 2006; Mehes et al. 2009) and was higher than the cupressaceae species with narrow distribution, such as Pilgerodendron uviferum (D. Don) Florín (H E = 0.035) (Premoli et al. 2001), F. cupressoides (H E = 0.077) (Premoli et al. 2000), and Podocarpus nubigena Lindl. (H E = 0.218) (Quiroga and Premoli 2010).
Genetic structure and gene flow
Because of greater random genetic drift and inbreeding as well as reduced gene flow, habitat fragmentation is expected to erode genetic variability and increase the interpopulation genetic divergence of plant populations (Young et al. 1996; Sork et al. 1999; Lowe et al. 2005; Aguilar et al. 2008). However, many tree species are probably resilient to habitat fragmentation for one or two generations after it occurs because they already contain high diversity (Bacles et al. 2004, 2005; Yao et al. 2007). In this study, we calculated that the degree of population differentiation was 0.163, based on SSR loci. This indicated that 16 % of the total genetic diversity could be attributed to differences among populations. Our F ST value for C. macrolepis was relatively higher than the level of genetic differentiation observed in other conifer species when examined with microsatellite markers. These include Pinus monticola, F ST = 0.057 and P. strobus, F ST = 0.084 (Mehes et al. 2009); Picea abies (L.) H. Karst., F ST = 0.009 (Acheré et al. 2005); P. abies, F ST = 0.005 (Maghuly et al. 2006); Pinus pinaster, F ST = 0.111 (Mariette et al. 2001); J. oxycedrus, F ST = 0.160 (Boratyński et al. 2014); J. cedrus, F ST = 0.08 (Rumeu et al. 2014); P. uviferum, F ST = 0.159 (Premoli et al. 2002); F. cupressoides, F ST = 0.078 (Premoli et al. 2001) and F. cupressoides, F ST = 0.14 (Allnutt et al. 1999). However, our findings were similar to those for other relictual conifer species in China (Wang et al. 1996; Ge et al. 1998; Tang et al. 2008). The level of gene flow for C. macrolepis (Nm = 1.562) suggests that the genetic diversity in C. macrolepis is low, and the probability of population differentiation has increased due to the fragmentation of habitat. Similar results were found for populations with narrow and fragmented distributions (Larix lyallii Parl., Nm = 1.4, (Khasa et al. 2006) and P. pinceana, Nm = 1.39, (Ledig et al. 2001). C. macrolepis is wind-pollinated, monoecious, and long-lived, and its small, light-weight seeds have two sub-apical, unequal wings (Wang et al. 2004). These biological features generally play a significant role in enhancing gene flow and decreasing genetic differentiation among populations. The genetic differentiation coefficient (F ST) represents the extent of gene differentiation among populations. Generally, 0.05–0.15 of F ST indicates the relatively middle level of gene differentiation, and 0.15–0.25 of F ST indicates the relatively high level of gene differentiation (Balloux and Lugon-Moulin 2002). Nevertheless, we found relatively high population differentiation (F ST = 0.163) and low gene flow.
From the UPGMA, populations HJ and LY are clearly separated from the rest, also MJ, YJ, and PE form a clear cluster, marginal populations TC, LS, JH, CN, and LC form another cluster, which is supported by Structure results. A reasonable hypothesis for the modern distribution patterns is that because of their suitability as subtropical climates, regions that include the southeastern portion of the YGP on the southeastern edge of the Qinghai-Tibetan Plateau may have been a Pleistocene refugium for C. macrolepis after the Quaternary glaciations. Concurrently, because of neotectonic movement since earlier in the Quaternary, complex and various climates and topography have appeared, and various microclimates have been shaped in the YGP. C. macrolepis is a heliophile and subtropical plant and after the Quaternary glaciations, it was maintained only in areas with subtropical microclimates. The genetic patterns presented here possibly reflect a scenario of glacial refugees (may be in lowland areas) with posterior recolonization of highlands when the temperature raised. Populations LS, JH, and DZ show great genetic diversity, also population DZ and LY present private alleles. In addition these populations have the lowest elevations, therefore these populations could be considered may be as glacial refugees. These deductions accorded with the results of Migrate analysis. Because the evolutionary rate of conifers are very slow (Petit and Hampe 2006; Savolainen and Pyhäjärvi 2007), these results implied that the genetic patterns within C. macrolepis were caused mainly by a long history of population isolation.
Management implications and recommendations for future research
Genetic diversity plays an important role in determining the survival and adaptability of a species. The loss of such diversity is often associated with reduced fitness. Maintaining genetic variation is a major objective when developing most programs for species conservation (Avise and Hamrick 1996). In the present study, although the majority of genetic variation by C. macrolepis resided within (84 %) rather than between populations, its diversity was lower and its differentiation was much greater than is found with other conifers that become broadly distributed due to population fragmentation. Nine populations in 14 sampled populations deviated from HW equilibrium. All of the results indicate that populations of C. macrolepis are declining. Moreover, their fragmentation may have been shaped by long-term historical events as well as independent evolutionary processes, such as neotectonic movement, glaciation, and glacial retreat.
Hence, to capture the greatest within-species genetic variation, decision-makers must include as many populations as possible in their programs for ex situ conservation or breeding. Because all extant populations are small and isolated, in situ conservation strategies should be designed to preserve large areas in order to minimize the loss of diversity that may result from future genetic drift. From our example, special attention should be given to Populations LY and DZ because of their private alleles, as well as to Population CJ in Hainan province because of its isolation from the mainland. The negative consequences of habitat fragmentation, such as genetic erosion and increased genetic divergence among populations, may be delayed in long-lived perennial plant species. This is because mature, established individuals may still reflect a historic situation with less-severe habitat fragmentation (Sork et al. 1999; Tomimatsu and Ohara 2003; Galeuchet et al. 2005; Van Geert et al. 2008). To determine the extent to which fragmentation affects the genetic variation and structure of C. macrolepis among generations, we must compare diversity and differentiation between adult and seedling populations, and integrate simultaneous investigations of demography, reproductive biology, and genetics.
References
Acheré V, Favre JM, Besnard G, Jeandroz S (2005) Genomic organization of molecular differentiation in Norway spruce (Picea abies). Mol Ecol 14:3191–3201
Aguilar R, Quesada M, Ashworth L, Herrerias-Diego Y, Lobo J (2008) Genetic consequences of habitat fragmentation in plant populations: susceptible signals in plant traits and methodological approaches. Mol Ecol 17:5177–5188
Alistair SJ, Josep P (2006) Genetic effects of chronic habitat fragmentation in a wind-pollinated tree. P Natl Acad Sci USA 103:8096–8100
Allnutt TR, Newton AC, Lara A, Premoli A, Armesto JJ, Vergara R, Gardner M (1999) Genetic variation in Fitzroya cupressoides (alerce), a threatened South American conifer. Mol Ecol 8:975–987
Averyanov LV, Hiep NT, Loc PK (2008) The Genus Calocedrus (Cupressaceae) in the Flora of Vietnam. Taiwania 53:11–22
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Avise JC, Hamrick JL (1996) Conservation genetics: case histories from nature. Chapman & Hall, New York
Bacles CFE, Lowe AJ, Ennos RA (2004) Genetic effects of chronic habitat fragmentation on tree species: the case of Sorbus aucuparia in a deforested Scottish landscape. Mol Ecol 13:573–584
Bacles CFE, Burczyk J, Lowe AJ, Ennos RA (2005) Historical and contemporary mating patterns in remnant populations of the forest tree Fraxinus excelsior L. Evolution 59:979–990
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165
Bekessy SA, Allnutt TR, Premoli AC, Lara A, Ennos RA, Burgman MA, Cortes M, Newton AC (2002) Genetic variation in the vulnerable and endemic Monkey Puzzle tree, detected using RAPDs. Heredity 88:243–249
Boratyński A, Wachowiak W, Dering M, Boratyńska K, Sękiewicz K, Sobierajska K, Jasińska AK, Klimko M, Montserrat JM, Romo A, Ok T, Didukh Y (2014) The biogeography and genetic relationship of J. oxycedrus and related taxa from the Mediterranean and Macaronesian regions. Bot J Linn Soc 174:637–653
Chen CH, Huang JP, Tsai CC, Chaw SM (2009) Phylogeny of Calocedrus (Cupressaceae), an eastern Asian and western North American disjunct gymnosperm genus, inferred from nuclear ribosomal nrITS sequences. Bot Stud 50:425–433
Craft KJ, Ashley MV, Koenig WD (2002) Limited hybridization between Quercus lobata and Quercus douglasii (Fagaceae) in a mixed stand in central coastal California. Am J Bot 89:1792–1798
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem bull 19:11–15
Ellstrand NC (1992) Gene flow by pollen: implications for plant conservation genetics. Oikos 63:77–86
Ellstrand NC, Elam DR (1993) Population genetic consequences of small population size: implications for plant conservation. Annu Rev Ecol Syst 24:217–242
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fahrig L, Merriam G (1994) Conservation of fragmented populations. Conserv Biol 8:50–59
Farjon A (2001) World checklist and bibliography of conifers. The Royal Botanic Gardens Press, Kew
Farjon A (2005) A monograph of Cupressaceae and Sciadopitys. The Royal Botanic Gardens Press, Kew
Frankham R, Briscoe DA, Ballou JD (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Fu LG (1995) China plant red data book. Science Press, Beijing
Fu LG, Yu YF (1999) Cupressaceae. In: Raven PH, Wu CY (eds) Flora of China. Science Press, Missouri Botanical Garden Press, Beijing, St. Louis
Galeuchet DJ, Perret C, Fischer M (2005) Microsatellite variation and structure of 28 populations of the common wetland plant, Lychnis floscuculi L., in a fragmented landscape. Mol Ecol 14:991–1000
Ge S, Hong D, Wang H, Liu Z, Zhang C (1998) Population genetic structure and conservation of an endangered conifer, Cathaya argyrophylla (Pinaceae). Int J Plant Sci 159:351–357
GENEPOP (2013) Genepop - population genetics software. http://genepop.curtin.edu.au/
González-Martínez SC, Dubreuil M, Riba M, Vendramin GG, Sebastiani F, Mayol M (2010) Spatial genetic structure of Taxus baccata L. in the western Mediterranean Basin: past and present limits to gene movement over a broad geographic scale. Mol Phylogenet Evol 55:805–815
Hamrick JL, Godt MJW, Sherman-Broyles SL (1992) Factors influencing levels of genetic diversity in woody plant species. New Forest 6:95–124
Hanski I, Ovaskainen O (2002) Extinction debt at extinction threshold. Conserv Biol 16:666–673
Honnay O, Jacquemyn H, Bossuyt B, Hermy M (2005) Forest fragmentation effects on patch occupancy and population viability of herbaceous plant species. New Phytol 166:723–736
Hsieh YC, Chung JD, Wang CN, Chang CT, Chen CY, Hwang SY (2013) Historical connectivity, contemporary isolation and local adaptation in a widespread but discontinuously distributed species endemic to Taiwan, Rhododendron oldhamii (Ericaceae). Heredity 111:147–156
Khasa DP, Jaramillo-Correa JP, Jaquish B, Bousquet J (2006) Contrasting microsatellite variation between subalpine and western larch, two closely related species with different distribution patterns. Mol Ecol 15:3907–3918
Kimura M, Ohta T (1978) Stepwise mutation model and distribution of allelic frequencies in a finite population. P Natl Acad Sci USA 75:2868–2872
Kracek Z (1999) An ancient Calocedrus (Cupressaceae) from the European Tertiary. Flora: Morphol Geobot, Oekophysiol 194:237–248
Kwak MM, Velterop O, van Andel J (1998) Pollen and gene flow in fragmented habitats. Appl Veg Sci 1:37–54
Lande R (1988) Genetics and demography in biological conservation. Science 241:1455–1460
Ledig FT, Capó-Arteaga MA, Hodgskiss PD, Sbay H, Flores-López C, Thompson Conkle M, Bermejo-Velázquez B (2001) Genetic diversity and the mating system of a rare Mexican piñon, Pinus pinceana, and a comparison with Pinus maximartinezii (Pinaceae). Am J Bot 88:1977–1987
Liao SX, Mi XJ, Liu AZ, Kun L, Yang ZY (2010) Isolation and characterization of polymorphic microsatellite markers in Calocedrus macrolepis Kurz (Cupressaceae). HortScience 45:169–171
Liao SX, Cui K, Wan YM, Zhou ZS, Li ZH, Cui YZ (2014) Reproductive biology of the endangered cypress Calocedrus macrolepis. Nord J Bot 32:98–105
Lindborg R, Eriksson O (2004) Historical landscape connectivity affects present plant species diversity. Ecology 85:1840–1845
Liu YS, Guo SX, Ferguson DK (1996) Catalogue of cenozoic megafossil plants in China. Palaeontogr Abt B 238:141–179
Liu FY, Li K, Liao SX, Cui YZ (2010) Interspecific competition, population structure and growth dynamics of endangered Calocedrus macrolepis. Sci Silvae Sin 46:23–28
Lowe AJ, Boshier D, Ward M, Bacles CFE, Navarro C (2005) Genetic resource impacts of habitat loss and degradation; reconciling empirical evidence and predicted theory for neotropical trees. Heredity 95:255–273
Maghuly F, Pinsker W, Praznik W, Fluch S (2006) Genetic diversity in managed subpopulations of Norway spruce [Picea abies (L.) Karst.]. Forest Ecol Manag 222:266–271
Mariette S, Chagne D, Lezier C, Pastuszka P, Raffin A, Plomion C, Kremer A (2001) Genetic diversity within and among Pinus pinaster populations: comparison between AFLP and microsatellite markers. Heredity 86:469–479
Matusova R (1995) Genetic variation in five populations of silver fir (Abies alba Mill.) in Slovakia. Biologia 50:53–59
Mehes M, Nkongolo KK, Michael P (2009) Assessing genetic diversity and structure of fragmented populations of eastern white pine (Pinus strobus) and western white pine (P. monticola) for conservation management. J Plant Ecol 2:143–151
MIGRATE-N 3.6.5 (2013) Estimation of population sizes and gene flow using the coalescent Software. Current version 3.6.5. http://popgen.sc.fsu.edu/Migrate-n.html
Miller MP (1997) Tools for population genetic analysis (TFPGA), version 1.3: a windows program for the analysis of allozyme and molecular population genetics data. http://www.markgeneticsoftware.net/
Milligan BG, Leebens-Mack J, Strand AE (1994) Conservation genetics: beyond the maintenance of marker diversity. Mol Ecol 3:423–435
Mona S, Ray N, Arenas M, Excoffier L (2014) Genetic consequences of habitat fragmentation during a range expansion. Heredity 112:291–299
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Newman D, Pilson D (1997) Increased probability of extinction due to decreased genetic effective population size: experimental populations of Clarkia pulchella. Evolution 512:354–362
Newman D, Tallmon DA (2001) Experimental evidence for beneficial fitness effects of gene flow in recently isolated populations. Conserv Biol 15:1054–1063
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
Oostermeijer J, Van’t Veer R, Den Nijs J (1994) Population structure of the rare, long-lived perennial Gentiana pneumonanthe in relation to vegetation and management in the Netherlands. J Appl Ecol 31:428–438
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Petit RJ, Hampe A (2006) Some evolutionary consequences of being a tree. Annu Rev Ecol Evol Syst 37:187–214
Premoli AC, Kitzberger T, Veblen TT (2000) Isozyme variation and recent biogeographical history of the long-lived conifer Fitzroya cupressoides. J Biogeogr 27:251–260
Premoli AC, Souto CP, Allnut TR, Newton AC (2001) Effects of population disjunction on isozyme variation in the widespread Pilgerodendron uviferum. Heredity 87:337–343
Premoli AC, Souto CP, Rovere AE, Allnut TR, Newton AC (2002) Patterns of isozyme variation as indicators of biogeographic history in Pilgerodendron uviferum (D. Don) Florín. Divers Distrib 8:57–66
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Quiroga MP, Premoli AC (2010) Genetic structure of Podocarpus nubigena (Podocarpaceae) provides evidence of Quaternary and ancient historical events. Palaeogeogr Palaeoclimatol 285:186–193
Rallo P, Dorado G, Martín A (2000) Development of simple sequence repeats (SSRs) in olive tree (Olea europaea L.). Theor Appl Genet 101:984–989
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rohlf FJ (1997) NTSYS-pc: numerical taxonomy and multivariate analysis system. Version 2.11c. Exeter Software, Setauket
Rumball W, Franklin IR, Franklin R, Sheldon BL (1994) Decline in heterozygosity under full-sib and double first-cousin inbreeding in Drosophia melanogaster. Genetics 136:1039–1049
Rumeu B, Vargas P, Jaén-molina R, Nogales M, Caujapé-castells J (2014) Phylogeography and genetic structure of the threatened Canarian Juniperus cedrus (cupressaceae). Bot J Linn Soc 175:376–394
Savolainen O, Pyhäjärvi T (2007) Genomic diversity in forest trees. Curr Opin Plant Biol 10:162–167
Schaal B, Hayworth D, Olsen K, Rauscher J, Smith W (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474
Sefc K, Lopes M, Mendonça D, Santos MRD, Machado L, Machado ADC (2000) Identification of microsatellite loci in olive(Olea europaea) and their characterization in Italian and Iberian olive trees. Mol Ecol 9:1171–1173
Sork VL, Nason J, Campbell DR, Fernandez JF (1999) Landscape approaches to historical and contemporary gene flow in plants. Trends Ecol Evol 14:219–223
Tang S, Dai W, Li M, Zhang Y, Geng Y, Wang L, Zhong Y (2008) Genetic diversity of relictual and endangered plant Abies ziyuanensis (Pinaceae) revealed by AFLP and SSR markers. Genetica 133:21–30
Tomimatsu H, Ohara M (2003) Genetic diversity and local population structure of fragmented populations of Trillium camschatcense (Trilliaceae). Biol Conserv 109:249–258
Torres-Díaz C, Ruiz E, González F, Fuentes G, Cavieres LA (2007) Genetic diversity in Nothofagus alessandrii (Fagaceae), an endangered endemic tree species of the Coastal Maulino Forest of Central Chile. Ann Bot 100:75–82
Van Geert A, Van Rossum F, Triest L (2008) Genetic diversity in adult and seedling populations of Primula vulgaris in a fragmented agricultural landscape. Conserv Genet 9:845–853
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Wang X, Zou Y, Zhang D, Hong D (1996) RAPD analysis of genetic diversity of Cathaya argyrophylla. Sci China (Serial C) 26:54–59
Wang DL, Li ZC, Hao G, Chiang TY, Ge XJ (2004) Genetic diversity of Calocedrus macrolepis (Cupressaceae) in southwestern China. Biochem Syst Ecol 32:797–807
WCMC (1997) The Socialist Republic of Vietnam Appendix 5-threatened plant species. http://www.wcmc.org.uk/infoserv/countryp/vietnam/app5.html
WGCPC (The Writing Group of Cenozoic Plants of China) (1978) Cenozoic plants from China. In: WGCPC (ed) Fossil plants of China. Science Press, Beijing, p 15
Yao X, Ye Q, Kang M, Huang H (2007) Microsatellites analysis reveals interpopulation differentiation and gene flow in endangered tree Changiostyrax dolichocarpa (Styracaceae) with fragmented distribution in central China. New Phytol 176:472–480
Yeh FC, Yang RC, Boyle T, Ye ZH, Mao JX (1997) POPGENE, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Centre, University of Alberta, Canada
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Acknowledgments
The authors would like to thank Mr. Priscilla Anne Licht for his help in language editing of the manuscript, and Hongtao Li, Longqian Xiao and Rong Huang for their assistance and advice.
Conflict of interest
None declared.
Funding
This work was supported by the Forestry Science and Technology Extension Project (Grant Number 2015[35]), the Forest Commonweal Foundation of Chinese State Forestry Administration (Grant Number 200704029) and the Knowledge Innovation Program of the Chinese Academy of Sciences (Grant Number KSCX2-YW-G-035-1).
Author information
Authors and Affiliations
Corresponding author
Additional information
Shengxi Liao and Kai Cui have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Liao, S., Cui, K., Tian, B. et al. The effect of long-term historical habitat fragmentation on genetic diversity of the relictual conifer Calocedrus macrolepis (Cupressaceae) in China. Braz. J. Bot 38, 567–577 (2015). https://doi.org/10.1007/s40415-015-0168-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-015-0168-4