Abstract
The signalling pathways in human cells mostly rely on protein–protein interactions (PPI) for their function. Such a PPI site in 3 Phosphoinositide dependent Kinase-1 (PDK1) is targeted to design the small molecule modulators. Based on the hotspot residues in its PPI site, a pharmacophore with seven different features was developed and screened against 2.9 million lead like compounds in Zinc database. A phthalazine derivative was identified as a potent allosteric inhibitor through virtual screening, molecular docking and 100 ns dynamics simulations. The modified hit possessed hydrogen bonds with Lys115, Arg131, Thr148, Glu150 as well as pi–pi stacking interactions with Phe157 which are the key residues in the PIF pocket of PDK1. Comparison between the free energy profiles by metadynamics simulation with the presence and absence of the modified ligand (MH) in the binding pocket indicates that the binding of MH enhances the hinge motion making PDK1 to adopt open conformation also and stabilizes the fluctuation of the end-to-end distance in αB helix of PDK1. The modified hit compound was synthesized, characterized and found to be cytotoxic to cancerous cells that are rich in PDK1 expression. These results propose that MH can serve as a new scaffold template for the design of novel drugs targeting PDK1 as well as promising allosteric regulator of PDK1 targeting its protein–protein interface.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Kinase inhibitors occupied the vast space in the cancer drug discovery. It is reviewed that around 53 kinase inhibitors are approved by FDA and over 200 potential inhibitors are in different phases of clinical trials (Gagic et al. 2020). PDK1 is one such well known cancer target kinase which plays multiple roles in cell signalling pathways such as PI3K/Akt, Ras/MAPK, and Myc (Emmanouilidi and Falasca 2017; Engel et al. 2006). Though many orthosteric and allosteric inhibitors of PDK1 such as UCN-01(Komander et al. 2003), substituted thieno[3,2-c]pyridine-7-carboxamides, indolinones(Medina et al. 2010), pyridinonyl-PDK1inhibitors, N-phenylpyrimidin-2-amines, 4-heterocycloalkyl-2-aminopyrimidines, diazepinones to tetracyclic imidazo phenanthrenones imidazo[4,5-c]quinolones, pyrrole derivatives, quinazolines, celecoxib derivatives, 4-aryl-7-azaindoles, 3,5-diaryl-7-azaindoles, pyrrolo[2,3-d]pyrimidines (Bai et al. 2015; Qin et al. 2023), pyrazole[1,5-α]pyrimidines, triazolo [1,5-α]pyrimidines, pyrazolyl benzimidazoles, indazoles, dibenzo[c,f][2,7]-naphthyridines, 3-hydroxy anthranilic acid have identified so far, except GSK-2334470 (Najafov et al. 2012; Zhang et al. 2018) none has succeeded the clinical trials. Also, there are no FDA approved drugs in the market against PDK1. Thus, the discovery of specific and potent inhibitors for PDK1 remains open and motivates the authors to work on. Phosphorylation is responsible for transmitting the signals through pathway proteins. PDK1 recruits other proteins with its PIF site as it lacks the hydrophobic motif like other Serine/Threonine kinases. Disrupting the recruitment by a small molecule inhibiting the protein–protein interaction is becoming popular with the name of PPI (protein–protein interaction site inhibitors) drug discovery (Wang et al. 2021).
The development of drugs against protein–protein interaction site in kinases is still in its earlier stage due to the difficulties such as flattened surface, deeper cavities, larger surface area etc. Designing small molecules to inhibit the protein–protein interaction site still requires more discoveries. Though it is difficult, potent small molecular inhibitors were developed either by directly targeting the PPI site or by allosterically from the ATP site. The PDK1 has single allosteric site which is the PDK1 interacting fragment (PIF) pocket (Fig. S1). But, there are recent works on bidirectional allosteric and reverse allosteric mechanism in PDK1 in which the kinase is regulated by a small molecule binding at the ATP site and produces a conformational change at protein–protein interaction site (Leroux and Biondi 2020). The protein–protein interaction site in PDK1 is the PIF-binding pocket where the kinase recruits and activates at least 23 AGC kinases including S6 Kinase, protein kinase C and serum and glucocorticoid-induced protein kinase (SGK), etc., for phosphorylation. PDK1 kinase domain consists of ATP site and PIF pocket which are ligand binding sites. The PIF pocket is wrapped up by Lys115, Ile118, Ile119, Val124, Val127 and Leu155. In the same structure, the residues Gly89, Glu90, Gly91, Lys111, Ser92, Val143, Leu159, Ser 160, Tyr161, Ala162 and Glu166 forms the ATP binding site. It is always challenging to develop potent PIF pocket binding ligands which disrupt the protein -protein interactions and also that freely diffuse into cells. It does not need to be competent with ATP which is at very high intra cellular concentrations. This type of PPI inhibition will lead to more specific inhibitor with minimum off-target pharmacology. The in silico tools will help in rational drug design and modulating the stability of the protein kinase interactions and protein complexes. Pharmacophore mapping with hotspots, virtual library screening, molecular modelling and dynamic simulations are being used in the drug discovery process and have yielded experimentally confirmed hits for many protein targets (Sarvagalla et al. 2016). It was reported that for proteins with multiple interacting partners, the key binding elements reside in the protein–protein interface adjacent to the concave hot region (Guo et al. 2014).
As a well-known cancer target, PDK1 has two domains, Kinase domain and PH domain. The kinase domain is with two lobes with a cleft between the two has ATP binding site. The small compounds binding at this site may have off target effects because of the conserved residues in this region of similar kinases. Also, from the recent communication, it is observed that the compounds binding at ATP site also induce some conformational changes at the PIF pocket which will in turn inhibits/enhance the activity of the protein (Arencibia et al. 2017; Schulze et al. 2016). Either small molecule allosteric modulator or PPI inhibitor or orthosteric inhibitor, no anticancer compounds functioning against PDK1 are approved drugs in the market still. From the recent advances in the PPI drug discovery, it is always interesting to look for specific small molecule activator/regulator for PDK1 through computational dynamics and biophysical characterisations. In an attempt to identify novel PDK1 inhibitor scaffolds, we pursued the combined molecular modelling approach with its structure dynamics activity relationship (SDAR) study by utilizing the pharmacophore mapping, virtual screening, molecular docking and atomistic simulations. The cytotoxic activity of the identified compound was further studied by SPR assay against cancer cell lines which were rich in expressing mRNA of PDK1 and the workflow (Fig. 1) utilized is represented below.
Experimental
Pharmacophore development and validation
The pharmacophore can be developed computationally from the structures of n number of ligands or a single ligand structure known as ligand-based method (Anant et al. 2021; Asati et al. 2020; Bhattacharya et al. 2021, 2022; Rathore et al. 2022; Saha et al. 2023). The development of pharmacophores from the active residues of the target protein or from different conformers of protein is commonly known as structure -based pharmacophore. In this study, the structure-based pharmacophore was aimed from the important residues in the active site region of the target protein. The hotspot residues in the protein–protein interaction site were picked up with the residues involved in hydrogen bonding with PIFtide substrate that are experimentally identified important residues in protein–protein interaction and hotspots prediction by hotpoint (Beekman et al. 2019; Cukuroglu et al. 2014). Utilizing the pharmacophore hypothesis development tool in Schrodinger, a seven featured pharmacophore model was developed based on the spatial coordinates of the hotspot residues. In order to validate the pharmacophore model, the developed hypothesis was screened against a set of compounds which includes 84 decoys generated from the Phase database and 4 actives taken from the literature which were experimentally identified as most active against PDK1. The statistical parameter, area under Receiver Operation Characteristics (ROC) was utilized to validate the model. The ligand screening was then applied with the prepared Zinc database ligands against developed pharmacophore model. It was set that the hits must match minimum 4 features and partial matches as preferred.
Ligands and protein preparation
The three-dimensional coordinates for the ligands were downloaded from Zinc database (Sterling and Irwin 2015) that contains over 230 million compounds ready to dock, in 3D formats. The filters like molecular weight less than 500, in-stock compounds which could be easy to buy earlier, clean and lead like compounds were applied in tranches to choose a subset of 2.9 million compounds. Their structures were optimized with OPLS2005 force field by Ligprep in Schrodinger (2015). The optimized structures were taken for pharmacophore matching.
The three-dimensional protein coordinates were downloaded from the protein data bank with PDB id: 4RRV. The missing side chains were built using Prime in Maestro. The hydrogens were added and the restrained energy minimization was done with OPLS2005 force field. The grid was generated with enclosing box set at the centroid of the residues which are at the interacting site of PPI region. The grid box was instructed to dock ligands with length less than or equal to 20 Å without any constraints.
Virtual screening and molecular docking
Virtual screening workflow in Maestro was utilized for this purpose. No filters were chosen as the ligands were lead like. The ligands were initially screened for pharmacophoric feature matches with the settings including the partial match and with the requirement of minimum six matches out of seven features and to give out the top thousand hits. The molecular docking was performed with Glide HTVS at first and the top ten percent hits were then redocked by Glide Simple precision (SP). The obtained top ten percent hits from SP were subjected to Glide extra precision mode (XP) whose poses were post processed with Prime MM-GBSA. The hits were ranked based on their gscores and MM-GBSA energy values. During the docking calculations, the protein was treated as rigid and ligands as flexible to determine their best conformations in the binding site of the protein. The protocol is summarized in the flowchart below (Fig. 2).
Molecular dynamics simulations
The best pose of the docked protein–ligand system with the modified hit was subjected to metadynamics simulations. Metadynamics simulation was performed with Desmond software developed by D.E. Shaw Research (Singh et al. 2012). The molecular system was initially set up with predefined TIP3P water molecules constrained in the orthorhombic box of 10 Å in each side and its volume was minimized according to the size of the complex. The system was neutralized by adding 0.15 M NaCl and Na+ /Cl− ions were added to neutralize the system and the model system was relaxed with the default parameters. The fully neutralized system was subjected to energy minimization to eliminate any unfavourable interactions due to model building and to adjust the initial structure to the OPLS2005 force field before production MD runs. The protein ligand system was simulated as NPT ensemble for 100 ns at 300 K with 1.0 atmospheric pressure by recording the trajectory at the interval of 100 ps which generated around 1000 snapshots of the motion of the system. Noose–Hoover chain Thermostat and Martyna–Tobias–Klein barostat was applied and the resulted trajectories were analysed with simulations interaction diagram in Maestro.
General procedure for synthesis of modified hit
The compound 4-Oxo-3, 4-dihydrophthalazine-1-carboxylic acid (400 mg, 1.0 eq) and 2-(lH-pyrrolo[2,3-b] pyridine-3-yl) ethanamine (422 mg, 1.25 eq) was taken in dimethyl formamide (4.0 mL, 10 vol) and cooled to 0–10 °C. HATU (878 mg, 1.1 eq) was added and stirred for 30 min. Triethyl amine (0.44 mL, 1.5 eq) was added to the reaction mass. The reaction mass temperature was raised to ambient temperature and stirred for 24 h (Scheme 1). After completion of reaction quench the reaction mass with purified water (20 mL) and filtered the solid. The solid was slurried with ethyl acetate (20 mL) and followed by methanol (20 mL) to obtain compound (7) (200 mg). The detailed scheme and synthesis of compounds 2 and 6 are given in supporting information Scheme S1–S2. The corresponding 1H NMR spectra are also included therein (Figs. S3–S8).
1H NMR (400 M Hz, DMSO-d6) δppm = 2.948–2.982 (t, J = 6.4, 2H), 3.344(s), 3.579–3.595 (d, J = 6.4, 1H), 6.999–7.029 (q, J = 4.8, 1H), 7.318 (s, 1H), 7.855–7.909 (q, J = 7, 3H), 7.997–8.016 (d, J = 7.6, 1H), 8.165–8.175 (d, J = 4, 2H), 8.251–8.269 (d, J = 7.2, 1H), 8.376–8.394 (d, J = 7.2, 1H), 8.720 (s), 11.366 (s, 1H), 12.934 (s,1H) 13C NMR (200 M Hz, DMSO-d6) δppm = 163.55, 159.64, 148.65, 142.42, 139.68, 133.69, 131.88, 127.73, 127.70, 126.68, 126.56, 125.79, 123.32, 119.55, 114.87, 110.74, 38.89 (Figs. S9–S10) HR-MS m/z: 333.344 calculated for C17H15O2N5 and found 334.200(+ MS, 2.3min).
SRB assay
The SRB assay is one of the more sensitive and reliable assays to determine the cytotoxic activity of the compounds (Rajini et al. 2017; Skehan et al. 1990). The cell lines were grown in appropriate medium containing 10% fetal bovine serum and 2 mM L-glutamine. For present screening experiment, 5000 cells/well were inoculated into 96 well microtiter plates in 100 µL. After cell inoculation, the microtiter plates were incubated at 37 °C, 5% CO2, 95% air and 100% relative humidity for 24 h prior to addition of experimental drugs.
Experimental compound was solubilized in DMSO solvent at 100 mg/ml and diluted to 1 mg/ml using water and stored frozen prior to use. At the time of drug addition, an aliquote of frozen concentrate (1 mg/ml) was thawed and diluted to 100 μg/ml, 200 μg/ml, 400 μg/ml and 800 μg/ml with complete medium containing test article. Aliquots of 10 µl of these different drug dilutions were added to the appropriate microtiter wells already containing 90 µl of medium, resulting in the required final drug concentrations, i.e., 10 μg/ml, 20 μg/ml, 40 μg/ml, 80 μg/ml.
After compound addition, plates were incubated at standard conditions for 48 h and assay was terminated by the addition of cold TCA. Cells were fixed in situ by the gentle addition of 50 µl of cold 30% (w/v) TCA (final concentration, 10% TCA) and incubated for 60 min at 4 °C. The supernatant was discarded; the plates were washed five times with tap water and air dried. Sulforhodamine B (SRB) solution (50 µl) at 0.4% (w/v) in 1% acetic acid was added to each of the wells, and plates were incubated for 20 min at room temperature. After staining, unbound dye was recovered and the residual dye was removed by washing five times with 1% acetic acid. The plates were air dried. Bound stain was subsequently eluted with 10 mM trizma base, and the absorbance was read on a plate reader at a wavelength of 540 nm with 690 nm reference wavelength.
Percent growth was calculated on a plate-by-plate basis for test wells relative to control wells. Percent Growth was expressed as the ratio of average absorbance of the test well to the average absorbance of the control wells × 100. Using the six absorbance measurements [time zero (Tz), control growth (C), and test growth in the presence of drug at the four concentration levels (Ti)], the percentage growth was calculated at each of the drug concentration levels. Percentage growth inhibition was calculated as: [Ti/C] × 100%.
Results and discussion
Identification of hit compound
The crystal structure of PDK1 with PIFtide, a well specific substrate is available in the protein data bank (PDB id: 4RRV) and this peptide-protein interaction site is considered for lead identification. The amino acid residues interacting with PIFtide that are crucial for the binding without which will fail to form that particular interaction are termed as “hotspots”. Hotspots can be predicted using experimental techniques such as Alanine mutation scanning and many computational methods including molecular simulations, machine learning algorithms and based on energy functions (Rettenmaier et al. 2014). In this study, hotspots were selected from the experimentally reported residues (Biondi et al. 2002) as well as residues predicted by the program ‘Hotpoint’ based on machine learning algorithms. In this way, the amino acid residues chosen as hotspots are Val127, Arg131, Leu155, Phe157 and Lys76, Tyr146, Phe147, Thr148, Phe149, Gln150 as warm spots in the PIF pocket. Based on the coordinates of atoms of hotspot residues and their chemical environment along with their energies, a Pharmacophore hypothesis was developed with ADDDNRR as seven features (Fig. 3) by using Phase in Schrodinger.
The hypothesis was then screened against 80 decoys generated from the Phase database (Fig. S2) in Maestro and 4 active PIF pocket binders shown in Fig. 4 reported in the recent work. The hypothesis was validated by an enrichment factor (ROC). ROC (Receiver operation characteristics) is a statistical method which tells about the efficiency of the created model against picking up the actives from a pool of non-actives (Marondedze et al. 2020). The ROC curve obtained for the hypothesis (Fig. 5) in which the curve pointing on the left side of the diagonal of false positives proved the hypothesis to be good with area under ROC curve score of 0.82 (Meharena et al. 2016). The developed Pharmacophore has predicted the most active compound PS210 without any false positives. The verified hypothesis can thus be employed to search for any matching small molecules from Zinc database.
Pharmacophore matching was done against lead like, clean compounds with the Ligand and database screening in Maestro holding the conditions that the ligand must match at least six features out of seven including the partial matching of pharmacophoric sites to give out top thousand hits. The hits were ranked according to their fitness scores and the topper has a fitness score of 0.6. The top 1000 compounds are further subjected to high-throughput virtual screen with the grid generated using the PDK1 structural coordinates and the top ten percent performers was then fed to SP docking. The resulting 11 compounds were subjected to extended XP docking in the HTVS workflow. A hit was identified with least gscore of -6.058 and the ligands whose gscore > 4.0 (Fig. 6) with their fitness score are reported in Table 1.
Hit compound and the binding site analysis
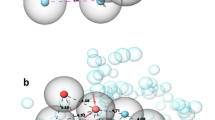
The identified hit (Fig. 6, first compound) consists of phthalazine and indole system connected by a peptide bonded aliphatic chain. In view of its binding with the PPI site of PDK1, it was found that the phthalazine ring system perfectly occupied the first cleft in the PIF pocket and the indole ring occupied the second cleft as shown in Fig. 7c. The amide linkage stabilizes the occupation of the molecule in the clefts. Another interesting factor is that the amide linkage which is essential for ligand binding was found to be present in almost all the identified compounds. The extra precision docking results proved the existence of hydrogen bond between Phe149 and –NH in the indole ring of the compound. The nitrogen atom in the amide linkage donates a proton to Glu150 and Pi-Pi stacking between the benzene ring of Phe157 and the indole moiety also stabilizes the protein–ligand complex (Fig. 7b).
a The alignment of top hit compound (ZINC000009387255) with the hypothesis developed. 5 partially matched sites out of seven can be seen. b The identified hit ineracts with PDK1 with two hydrogen bonds (purple arrows) and a pi–pi interaction (green arrow). c Comparison of PIFtide and the hit compound in the PPI site of PDK1 (Pdb id:4RRV). d Comparison of the hit compound with PS210 in the PIF pocket
When compared to the PIFtide, strong substrate of PDK1, the identified hit has positioned well with its ring system in the PIF pocket’s cleft 1 identical to the phenyl ring of PHE 14 and to that of PHE 17 in cleft 2 (Fig. 7c). Also similar to PS210 (an active PIF binder, (Busschots et al. 2012)), the hit has positioned well into the clefts. These views predicted the compound as good binder to PDK1 PIF binding pocket. In order to improve the number of hydrogen bonds, it is also attempted with slightly modified basicity of the indole ring to pyrrolopyridine in the hit compound which shows improved hydrogen bonds with PDK1 (Fig. 8 and Table T1) together with a hydrogen bond, linking the modified pyrrolopyridine with hotspot residues Arg 131 and Phe 157.
Metadynamics simulations and free energy analysis
To analyse the changes in the dynamics of PDK1 upon binding of the ligand, Metadynamics simulations of the two systems, PDK1/ATP and PDK1/ATP/designed ligand were carried out using DESMOND package. Metadynamics is a free energy perturbation method which enhances sampling of the underlying free energy space by biasing against previously-visited values of user-specified collective variables. The two collective variables used in the dynamics study are the end-to-end distance of αB helix (residues 115–125) and the distance between the Gly-rich loop (residues 90–94) and Asp 205 in order to visualize the hinge motion of PDK1 (Schulze et al. 2016). The root mean squared deviation (RMSD) of the Cα atoms of PDK1/ATP + MH and PDK1/ATP complexes was calculated against their initial structure and the results are shown in Fig. 9a. The highest RMSD value in PDK1/ATP + MH of 2.6 which is lower to 3.3 in PDK1/ATP structure implicates that the structure gets stabilized (Fig. 9a) around 40 ns of simulation upon ligand binding. The notably smaller deviation in RMSF values of the Cα atoms in PDK1/ATP + MH (Fig. 9b) could indicate the more proper orientation of T-loop which is more fluctuated in case of the enzyme without MH. In addition, the fluctuations in the PIF pocket lining of helices αB and αC are found to be reduced in PDK1/ATP + MH due to the direct contact with Lys115 in helix αB and Arg131 in helix αC.
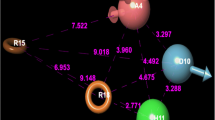
To understand the conformational changes in PDK1, 1000 frames of motion were analysed with free energy landscape projected along the variable describing the length of αB helix and the distance between the Gly-rich loop (residues 90–94) and Asp 205, which describes the kinase hinge motion. The ligand bound structure in solvent environment were observed to enhance the hinge motion when compared to structure with ATP. The two minima obtained, indicates the presence of populated structures in both the open and closed conformations of PDK1. The hinge motion is described by the presence and absence of salt bridges in PDK1 structure particularly the salt bridge between Lys111 and Glu130 which is the hall mark for the closed active form of PDK1. It is found that the dynamic structures are well populated in the first minima with energy value of − 10.42 kcal/mol (Fig. 10) were in closed form having the hall mark salt bridge. The structures found in the second minima with least energy value of − 8.92 kcal/mol were found to be in open form without the Lys–Glu salt bridge. These results indicate that the binding of ligand in the PIF pocket disturbed the hinge region allosterically and stabilizes the complex in open inactive form in addition to the closed active conformation. Equilibrium between these two conformations might exist. The two different conformations of αB helix were observed in both the cases. The conformational changes in αB and αC helices were not altered by the binding of MH thus stabilizing the PIF pocket. This information leads to conclude that the binding of MH with PDK1 stabilizes PIF pocket preventing the PPI interaction similar to PIFtide and allosterically hinder the hinge region and making the protein inactive.
Biological activity
The cytotoxic activity for the synthesized compound was assessed by SRB assay against MCF7 (breast), SiHa (cervical) and Vero (normal) cell lines. The compound showed moderate activity in MCF7 cell lines with GI50 of 64.6 µg/ml when compared with Adriamycin (Figs. 11, S11), the positive control used in the assay. For SiHa cells, the GI50 value was calculated as > 80 µg/ml. The compound was not toxic to normal vero cells. Being an activator, the identified hit compound may enhance the PDK1 activity and inhibit the protein–protein interaction which is more profound in certain cancer cell lines and which is the reason for no activity in SiHa cancer cells. This permits the authors to conclude that the hit can be taken through the modulator development pipeline for PDK1 whose downstream signalling pathway has to be experimentally studied.
Conclusions
A seven featured hotspot-based pharmacophore model was developed and validated for PIF pocket of PDK1. The virtual screening of the pharmacophore model was carried out against 2.9 million Zinc database compounds. The molecular docking of top hits resulted in a novel phthalazine derivative. The metadynamics simulations of the modified hit predicted the molecule binds in the PIF pocket with binding energy of − 10.42 kcal/mol. The allostery analysis by free energy landscape further confirmed that the ligand hindered the hinge region of PDK1 by binding in the PIF pocket and stabilizing the protein in open inactive conformation. The proposed allosteric inhibitor was synthesized, characterized and checked for its cytotoxic GI50 against the cancerous cell lines that are rich in PDK1 expression as well as normal cell line. The identified hit was toxic to breast cancer cell lines at the concentration of 60 µg/ml. Thus, the identified phthalazine analogue is proposed as a good starting point towards the allosteric drug design of PDK1 and the methodology and findings can be implemented for future drug design of PPI inhibitors to other similar drug targets.
References
Anant A, Ali A, Ali A, Gupta GD, Asati V (2021) A Computational approach to discover potential quinazoline derivatives against CDK4/6 kinase. J Mol Struct 1245:131079
Arencibia JM, Fröhner W, Krupa M, Pastor-Flores D, Merker P, Oellerich T et al (2017) An allosteric inhibitor scaffold targeting the PIF-pocket of atypical protein kinase C isoforms. ACS Chem Biol 12(2):564–573
Asati V, Agarwal S, Mishra M, Das R, Kashaw SK (2020) Structural prediction of novel pyrazolo-pyrimidine derivatives against PIM-1 kinase: In-silico drug design studies. J Mol Struct 1217:128375
Bai LY, Chiu CF, Kapuriya NP, Shieh TM, Tsai YC, Wu CY et al (2015) BX795, a TBK1 inhibitor, exhibits antitumor activity in human oral squamous cell carcinoma through apoptosis induction and mitotic phase arrest. Eur J Pharmacol [internet] 769:287–296
Beekman AM, Cominetti MMD, Walpole SJ, Prabhu S, O’Connell MA, Angulo J et al (2019) Identification of selective protein-protein interaction inhibitors using efficient: in silico peptide-directed ligand design. Chem Sci 10(16):4502–4508
Bhattacharya S, Asati V, Mishra M, Das R, Kashaw V, Kashaw SK (2021) Integrated computational approach on sodium-glucose co-transporter 2 (SGLT2) Inhibitors for the development of novel antidiabetic agents. J Mol Struct 1227:129511
Bhattacharya S, Asati V, Ali A, Ali A, Gupta GD (2022) In-silico studies for the development of novel RET inhibitors for cancer treatment. J Mol Struct 1251:132040
Biondi RM, Komander D, Thomas CC, Lizcano JM, Deak M, Alessi DR et al (2002) High resolution crystal structure of the human PDK1 catalytic domain defines the regulatory phosphopeptide docking site. EMBO J 21(16):4219–4228
Busschots K, Lopez-Garcia LA, Lammi C, Stroba A, Zeuzem S, Piiper A et al (2012) Substrate-selective inhibition of protein kinase PDK1 by small compounds that bind to the PIF-pocket allosteric docking site. Chem Biol 19(9):1152–1163
Cukuroglu E, Engin HB, Gursoy A, Keskin O (2014) Hot spots in protein-protein interfaces: towards drug discovery. Prog Biophys Mol Biol Internet. 116(2–3):165–173. https://doi.org/10.1016/j.pbiomolbio.2014.06.003
Emmanouilidi A, Falasca M (2017) Targeting PDK1 for chemosensitization of cancer cells. Cancers (basel). 9:140
Engel M, Hindie V, Lopez-Garcia LA, Stroba A, Schaeffer F, Adrian I et al (2006) Allosteric activation of the protein kinase PDK1 with low molecular weight compounds. EMBO J 25(23):5469–5480
Gagic Z, Ruzic D, Djokovic N, Djikic T, Nikolic K (2020) In silico methods for design of kinase inhibitors as anticancer drugs. Front Chem 7(January):1–25
Guo W, Wisniewski JA, Ji H (2014) Hot spot-based design of small-molecule inhibitors for protein-protein interactions. Bioorg Med Chem Lett [internet]. 24(11):2546–2554. https://doi.org/10.1016/j.bmcl.2014.03.095
Komander D, Kular GS, Bain J, Elliott M, Alessi DR, Van ADMF (2003) Structural basis for UCN-01 (7-hydroxystaurosporine) specificity and PDK1 (3-phosphoinositide-dependent protein kinase-1) inhibition. Biochem J 375:255–262
Leroux AE, Biondi RM (2020) Renaissance of allostery to disrupt protein kinase interactions. Trends Biochem Sci Internet 45(1):27–41. https://doi.org/10.1016/j.tibs.2019.09.007
Marondedze EF, Govender KK, Govender PP (2020) Ligand-based pharmacophore modelling and virtual screening for the identification of amyloid-beta diagnostic molecules. J Mol Graph Model [internet]. 101:107711. https://doi.org/10.1016/j.jmgm.2020.107711
Medina JR, Blackledge CW, Heerding DA, Campobasso N, Ward P, Briand J et al (2010) Aminoindazole PDK1 inhibitors: a case study in fragment-based drug discovery. ACS Med Chem Lett 1(8):439–442
Meharena HS, Fan X, Ahuja LG, Keshwani MM, McClendon CL, Chen AM et al (2016) Decoding the interactions regulating the active state mechanics of eukaryotic protein kinases. PLoS Biol 14(11):e2000217
Najafov A, Shpiro N, Alessi DR (2012) Akt is efficiently activated by PIF-pocket- and PtdIns(3,4,5) P3-dependent mechanisms leading to resistance to PDK1 inhibitors. Biochem J [internet]. 448(2):285–295. https://doi.org/10.1042/BJ20121287
Qin J, Cao M, Hu X, Tan W, Ma B, Cao Y et al (2023) Dual inhibitors of ASK1 and PDK1 kinases: design, synthesis, molecular docking and mechanism studies of N-benzyl pyridine-2-one containing derivatives as anti-fibrotic agents. Eur J Med Chem 247:115057
Rajini A, Nookaraju M, Reddy IAK, Venkatathri N (2017) Synthesis, characterization, antimicrobial and cytotoxicity studies of a novel titanium dodecylamino phosphate. J Saudi Chem Soc [internet]. 21:S77-85. https://doi.org/10.1016/j.jscs.2013.10.005
Rathore A, Asati V, Mishra M, Das R, Kashaw V, Kashaw SK (2022) Computational approaches for the design of novel dopamine D2 and serotonin 5-HT2A receptor dual antagonist towards schizophrenia. In Silico Pharmacol [internet]. https://doi.org/10.1007/s40203-022-00121-5
Rettenmaier TJ, Sadowsky JD, Thomsen ND, Chen SC, Doak AK, Arkin MR et al (2014) A small-molecule mimic of a peptide docking motif inhibits the protein kinase PDK1. Proc Natl Acad Sci Internet. 111(52):18590–18595. https://doi.org/10.1073/pnas.1415365112
Saha M, Gupta S, Dhiman S, Asati V, Ali A, Ali A (2023) Field and atom-based 3D-QSAR models of chromone (1-benzopyran-4-one) derivatives as MAO inhibitors. J Biomol Struct Dyn [internet]. https://doi.org/10.1080/07391102.2023.2166122
Sarvagalla S, Cheung CHA, Tsai JY, Hsieh HP, Coumar MS (2016) Disruption of protein-protein interactions: hot spot detection, structure-based virtual screening and: in vitro testing for the anti-cancer drug target-survivin. RSC Adv [internet]. 6(38):31947–31959. https://doi.org/10.1039/C5RA22927H
Schrodinger Release 2021–3: Bioluminate, Schrodinger, LLC Newyork, NY,2021. New York: NY: LLC; 2015
Schulze JO, Saladino G, Busschots K, Neimanis S, Süß E, Odadzic D et al (2016) Bidirectional allosteric communication between the ATP-binding site and the regulatory PIF pocket in PDK1 protein kinase. Cell Chem Biol 23(10):1193–1205
Singh KD, Kirubakaran P, Nagarajan S, Sakkiah S, Muthusamy K, Velmurgan D et al (2012) Homology modeling, molecular dynamics, e-pharmacophore mapping and docking study of Chikungunya virus nsP2 protease. J Mol Model 1:39–51
Skehan P, Storeng R, Scudiero D, Monks A, McMahon J, Vistica D et al (1990) New colorimetric cytotoxicity assay for. J Natl Cancer Inst 82(13):1107–1112
Sterling T, Irwin JJ (2015) ZINC 15—ligand discovery for everyone. J Chem Inf Model 55(11):2324–2337
Wang X, Ni D, Liu Y, Lu S (2021) Rational design of peptide-based inhibitors disrupting protein-protein interactions. Front Chem [internet] 9:682675
Zhang J, Yang C, Zhou F, Chen X (2018) PDK1 inhibitor GSK2334470 synergizes with proteasome inhibitor MG-132 in multiple myeloma cells by inhibiting full AKT activity and increasing nuclear accumulation of the PTEN protein. Oncol Rep 39(6):2951–2959
Acknowledgements
The Senior Research Associateship from CSIR, Govt. of India by Scientists’ pool scheme is greatly acknowledged by KNV. The cell cytotoxicity assays from ACTREC, Mumbai, India is also acknowledged by the authors.
Author information
Authors and Affiliations
Contributions
VKN: idea, analysis, manuscript writing SB: synthesis of the compound and characterization, EKP: idea, results and Manuscript evaluation.
Corresponding author
Ethics declarations
Conflict of interest
The authors state that “There are no conflicts to declare”.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kailasam Natesan, V., Balaraman, S. & KuppannaGounder Pitchaimuthu, E. Insilico design of an allosteric modulator targeting the protein–protein interaction site of 3 Phosphoinositide dependent Kinase-1: design, synthesis and biological activity. In Silico Pharmacol. 11, 26 (2023). https://doi.org/10.1007/s40203-023-00160-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40203-023-00160-6