Abstract
Triple-negative breast cancer (TNBC) is a lethal and aggressive breast cancer subtype. It is characterized by the deficient expression of the three main receptors implicated in breast cancers, making it unresponsive to hormone therapy. Hence, an existing need to develop a targeted molecular therapy for TNBC. The PI3K/AKT/mTOR signaling pathway mediates critical cellular processes, including cell proliferation, survival, and angiogenesis. It is activated in approximately 10–21% of TNBCs, emphasizing the importance of this intracellular target in TNBC treatment. AKT is a prominent driver of the PI3K/AKT/mTOR pathway, validating it as a promising therapeutic target. Dysphania ambrosioides is an important ingredient of Nigeria’s traditional herbal recipe for cancer treatment. Thus, our present study explores its anticancer properties through a structure-based virtual screening of 25 biologically active compounds domiciled in the plant. Interestingly, our molecular docking study identified several potent inhibitors of AKT 1 and 2 isoforms from D. ambrosioides. However, cynaroside and epicatechin gallate having a binding energy of − 9.9 and − 10.2 kcal/mol for AKT 1 and 2, respectively, demonstrate considerable drug-likeness than the reference drug (capivasertib), whose respective binding strengths for AKT 1 and 2 are − 9.5 and − 8.4 kcal/mol. Lastly, the molecular dynamics simulation experiment showed that the simulated complex systems of the best hits exhibit structural stability throughout the 50 ns run. Together, our computational modeling analysis suggests that these compounds could emerge as efficacious drug candidates in the treatment of TNBC. Nevertheless, further experimental, translational, and clinical research is required to establish an empirical clinical application.
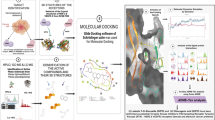
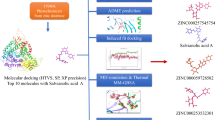
Graphical Abstract
A structure-based virtual screening and simulation of Dysphania ambrosioides phytochemicals in the active pocket of AKT 1 and 2 isoforms

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BC) is one of the most common and prevalent malignancies in women worldwide, with over 2 million cases reported in 2020 (Koo et al. 2017; Łukasiewicz et al. 2021; Sung et al. 2021). Even though numerous studies are underway to progress the diagnosis and give a significant treatment (Siegel et al. 2022, 2023). Nonetheless, it is expected to rise as risk factors increase owing to modernization and economization (Ciriello and Magnani 2021; Kumar Prusty et al. 2020; Sung et al. 2021). Based on molecular markers, BC is divided into three subtypes: hormone receptor-positive/erythroblastic oncogene B-2 (ERBB2) negative, ERBB2 positive, and triple-negative (TN) (Sherman et al. 2022; Waks and Winer 2019). Because of the absence or inexpression of all three primary breast cell receptors [Progesterone Receptor (PR), Estrogen Receptor (ER), and Human Epidermal Growth Factor Receptor 2 (HER2)], TNBC is the most aggressive cancer subtype (Mahmoud et al. 2022; Yin et al. 2020). TNBC is also known for its strong metastatic potential, invasiveness, and poor prognosis (Yin et al. 2020). Hence, it has a more aggressive clinical course than other breast cancer subtypes.
Based on recent studies, TNBC has been further categorized into six subtypes using molecular pathology, gene expression profiling, and histopathology (Faje et al. 2018; Gottifredi 2020). And these include luminal androgen receptor (LAR), mesenchymal stem-like (MSL), basal-like 1 (BL1), basal-like 2 (BL2), immunomodulatory (IM), and mesenchymal (M), each exhibiting unique gene expression patterns and mutations (Mahmoud et al. 2022; Nazmy et al. 2021). The molecular and genetic profiling and clinicopathological studies of TNBC have proved its heterogeneity (Zakaria et al. 2018). Nevertheless, TP53, BRCA1/2, PIK3CA, AKT, PTEN, EGFR, FGFR1, VEGFRA, AR, and BCL2 have all been identified as TNBC genetic markers (Rasul et al. 2022; Sporikova et al. 2018). These genes have mutations that affect their main functions, such as genome integrity, DNA repair, apoptosis, survival, proliferation, differentiation, angiogenesis, and metastasis through changes in expression, amplification, overexpression, deletion, or inactivation (Sporikova et al. 2018).
However, certain driver genes, such as PIK3CA and AKT, have been discovered in several TNBC subtypes (BL1, BL2, MSL, LAR, and M) and may serve as prospective therapeutic targets (Marra et al. 2020). Although biomarker-driven, multi-omics technology and immune checkpoint-based therapeutics have been investigated with promise but with poor success (Anders et al. 2016; Marra et al. 2020). Thus, as a key driver gene in TNBC, the AKT gene contributes significantly to the PI3K/AKT/mTOR-mediating pathway, resulting in cell survival, proliferation, and tumor formation. The PI3K/AKT/mTOR pathway is frequently dysregulated in TNBC (Nazmy et al. 2021). Hyperactivation of AKT and mTOR is linked to a poor prognosis in TNBC patients, highlighting the PI3K/AKT/mTOR pathway as an appealing therapeutic target (Nazmy et al. 2021; Sporikova et al. 2018). While adhering to concepts and new results on TNBC, evidence about gene changes such as PIK3CA and AKT has offered a good hotspot for TNBC targeted therapy (Aine et al. 2021).
AKT (Protein Kinase B) governs cellular activities downstream of PI3K, such as cell proliferation, survival, and metabolism (Basu and Lambring 2021; Hinz and Jücker 2019) (Fig. 1). There are three AKT isoforms: AKT1 (PKBα), AKT2 (PKBβ), and AKT3 (PKBγ), all of which have 80% structural similarity but perform different activities (Manning and Toker 2017). Breast tumors have been found to exhibit AKT isoform amplification or high levels of mRNA expression (Turner et al. 2015). Furthermore, systemic studies have indicated unique roles for each isoform; in animal models, AKT1 knockdown decreased lung metastases, but AKT2 deletion prevented mammary tumorigenesis and metastasis (Chen et al. 2020). CyclinD1 levels in MDA MB 231 cells were reduced by nanobodies targeting the C-terminal regulatory hydrophobic motif site (HM) (Merckaert et al. 2021). AKT1, AKT2, and AKT3 genes are found on chromosomes 14q32 (Basu and Lambring 2021; Staal et al. 1988), 19q13 (Cheng et al. 1992), and 1q44 (Risso et al. 2015) respectively. They each have three domains: an N-terminal pleckstrin homology domain (PH), a central catalytic domain (CD), and a regulatory hydrophobic motif site at the C-terminus. AKT1 amplification or high mRNA expression levels are the least common among all AKT isoforms in breast cancer (Turner et al. 2015). However, they are related to a poor prognosis in basal-like 2 TNBCs (Wang et al. 2019). AKT1 participates in PI3K/AKT signaling via importation. AKT1 also promotes cell proliferation by altering proteins such as p21, p27, and cyclin D1 and inhibiting apoptosis via regulating protein 53 (p53) (Hinz and Jücker 2019).
Furthermore, AKT2 amplification has been identified in 3% of breast cancer cases (Bellacosa et al. 1995; Turner et al. 2015). AKT2 promotes breast cancer cell motility and invasion through modulating -integrins, epithelial-mesenchymal transition (EMT) proteins, and F-actin. AKT2 deletion has also been linked to glucose homeostasis by reducing glucose absorption into the cell, leading to an energy shortage for cell metabolism (Hinz and Jücker 2019; Merckaert et al. 2021). In breast cancer, however, AKT3 is the most elevated AKT isoform (Turner et al. 2015). A recurring fusion of MAGI3 (membrane-associated guanylate kinase, WW, and PDZ domain containing 3)-AKT3 was shown to activate AKT pathways in TNBCs constitutively and could only be dispersed or disrupted by utilizing an ATP-competitive AKT inhibitor (Banerji et al. 2012). However, systemic research on AKT3 knockdown found no substantial effect (Chen et al. 2020). Though it has been examined for its involvement in lowering CyclinD1 levels in MDA MB 231 cells (Merckaert et al. 2021), its overall function in TNBC is unknown.
The AKT signaling pathway involves PIP3 binding to AKT’s PH domain, which attaches it to the cellular membrane and allows it to be phosphorylated and activated. Thorough AKT activation necessitates the phosphorylation of two highly conserved sites, a conformational shift and Thr308/Thr309 phosphorylation by PDK1. The mTORC2 complex is responsible for phosphorylating Ser473/Ser474 in the hydrophobic motif, which is required for effective AKT activation. Other Ser473/Ser474 phosphorylation pathways include PDK1, integrin-linked kinase, and autophosphorylation (Khan et al. 2019; LoRusso 2016; Manning and Toker 2017; Unger and van Golen 2009). Once thoroughly activated, AKT translocates inside the cell to the cytoplasm, nucleus, mitochondria, and other organelles. NF-κB transcription factors, glycogen synthase kinase (GSK) 3β, BAD, FOXO1, MDM2, human caspase 9, mTOR, Raf, p21, and BRCA-1 are all phosphorylated by AKT using its kinase domain. These substrates play a role in cancer-related phenotypes. As a result, the AKT signaling pathway is critical in regulating cell survival, proliferation, and growth, and abnormal activation of this pathway is implicated in TNBC development (Hoxhaj and Manning 2020; Khan et al. 2019; LoRusso 2016; Manning and Toker 2017; Unger and van Golen 2009).
Due to high rates of pathway aberrations, such as the loss of negative regulators like PTEN, the PI3K/AKT/mTOR signaling pathway is frequently activated in TNBC (Davies et al. 2012; Gener et al. 2019). AKT inhibitors like ipatasertib (GDC-0068), ONC201, and capivasertib (AZD5363) have demonstrated preclinical activity in TNBC models and have been linked to sensitivity in tumors with PI3K/AKT pathway activation (Davies et al. 2012; Greer and Lipkowitz 2015; Khan et al. 2019; Lin et al. 2013; Stein et al. 2017). Therapeutic inhibition of AKT, PI3K, and mTOR has also been found to activate feedback loops that may restrict efficacy and enhance acquired resistance to single-agent receptor tyrosine kinase inhibition (Chandarlapaty et al. 2011; O’Reilly et al. 2006; Rodrik-Outmezguine et al. 2011). In breast cancer, for example, dual suppression of P13K/AKT and DNA damage checkpoint pathways or tumors with AKT mutations or amplification has been reported (Lin et al. 2013; LoRusso 2016; Millis et al. 2015; Skladanowski et al. 2007).
Currently, there are no standardized FDA (Food and Drug Administration)-approved AKT-targeting regimens, and viable medicines are still in the preclinical stages. As a result, TNBC researchers have been investigating new therapy options, such as targeted medicines, to enhance TNBC patient health outcomes. However, developing a directed treatment regimen requires a deeper knowledge of TNBC’s molecular and driver gene characteristics, such as protein kinase B (AKT). Dysphania ambrosioides, originally Chenopodium ambrosioides, are medicinal plants with important pharmacological properties. Previous studies have extensively researched its phytochemistry, ethnopharmacology, and geographical range (da Silva et al. 2021; Zohra et al. 2018). In addition, its hydroethanolic and chloroform extracts have been reported to exhibit myorelaxant, antispasmodic, and antioxidant action (Kandsi et al. 2021). Interestingly, studies have revealed that Dysphania ambrosioides contains several phytochemical components, including catechin gallate (epicatechin gallate), amentoflavone, luteolin-7-O-glucoside (cynaroside), gallocatechin/epigallocatechin gallate, arbutin, caffeic acid, ferulic acid, hesperetin, isorhamnetin-3-O-rutinoside, and isorhamnetin-7-O-rutinoside (Kandsi et al. 2021). These components are active to varied degrees and have been linked to antioxidant, anti-inflammatory, anti-diabetic, and anti-cancer activities (Degenhardt et al. 2016; Kandsi et al. 2021; Zohra et al. 2018). Howbeit, the presence, and combination of these components in Dysphania ambrosioides may contribute to the plant’s medicinal potential. With a UMi (use-mention index) of 0.129, Mephors et al. (2017) ranked Dysphania ambrosioides as one of the most frequently used herbal recipes for cancer treatment by Traditional Medical Practitioners in the Southwestern part of Nigeria.
The scientific pursuit of chemicals domiciled in plants with antitumor activity is central to the greener approach of revolutionizing cancer treatment. Dysphania ambrosioides have demonstrated pharmacological properties that could be harnessed to develop new medications. Its cytotoxic, anti-proliferative, and apoptosis-inducing actions on cancer cells have been reported by (Tauchen et al. 2019; Wang et al. 2021; Zohra et al. 2018). Similarly, its ethanolic extract has been shown to exhibit substantial cytotoxic action in RAJI and K562 tumor cell lines (Degenhardt et al. 2016). Although its anti-cancer capabilities are well documented, limited data exist regarding its potential as a therapeutic agent for TNBC or as an inhibitor of the PI3K/AKT/mTOR signaling in breast cancer. Therefore, the present study explored a structure-based virtual screening to identify natural compounds in Dysphania ambrosioides that could interfere with AKT-mediated signaling in TNBC.
Materials and methods
Protein preparation and active site detection
We obtained the three-dimensional (3D) crystal structure of two AKT isoforms (AKT1 and AKT2) proteins with protein data bank (PDB) IDs 4EKL and 2JDR from the RCSB (Research Collaboratory for Structural Bioinformatics) database. Prior to computational analysis, validation was performed to check for problems such as missing atoms or incorrect bond formations. The missing residues were added using the protein repair and analysis server (https://www.protein-science.com/) (Nnyigide et al. 2022), and the proteins were prepared as receptors. The technique here begins with the generation of a binding sphere (SBD) and removal of the co-crystalized Ipatasertib (GDC-0068) and A-443654 from AKT1 and AKT2, respectively. Autodock Vina in PyRx (https://pyrx.sourceforge.io) was utilized to insert the validated protein structures and generate PDBQT files for each protein (Oyedele et al. 2022; Younus et al. 2022).
The region in the AKT1 and AKT2 proteins that interacts with the ligands, known as the active site, was detected during the active site determination process. This procedure entailed analyzing the two-dimensional (2D) protein–ligand interaction for both proteins and their co-crystallized ligands in order to identify the particular residues required for ligand binding. Throughout the docking procedures with PyRx and BIOVIA Discovery Studio (https://discover.3ds.com/discovery-studio-visualizer-download), the active site residues served as a reference and guide in comparative analysis.
Ligands selection and preparation for docking
From the PubChem database (http://pubchem.ncbi.nlm.nih.gov), we downloaded the 3D conformer of an investigational drug (Capiversatib [AZD5363; PubChem ID 25227436]) and 25 phytochemicals found in Dysphania ambrosioides and utilized them as putative ligands for AKT1 and AKT2 proteins. PyRx’s OpenBabel was used to import SDF (structure data file) molecules, minimize energy, and generate PDB formats for each ligand’s 3D conformer. After optimization, we entered the chemicals into PyRx Autodock Vina to obtain their appropriate PDBQT forms (Trott and Olson 2010).
Molecular docking
The molecular docking experiment was carried out using AutoDock Vina in the PyRx virtual screening software (Dallakyan and Olson 2015). We fine-tuned the grid box sizes to encompass the defined binding pocket residues of AKT 1 and 2. Accordingly, the grid box dimension values are X = 30.4497, Y = 29.5329, Z = 27.2303 and X = 25.5233, Y = 26.6809, Z = 25.0, for AKT 1 and 2, respectively. Also, the default exhaustiveness value of 8 was used for both dockings.
We performed a molecular docking analysis to probe Dysphania ambrosioides phytochemicals and capivasertib (reference drug) binding interactions with AKT isoforms 1 and 2. The x-ray crystal structure of AKT 1 [PDB ID: 4EKL (Lin et al. 2012)] and AKT 2 [PDB ID: 2JDR (Davies et al. 2007)] in complex with GDC0068 and A-443654, respectively, were used. The interaction energy of the test ligands and reference drug (capivasertib) (AZD5363) with AKT were analyzed, and the top ligands having the lowest binding energy compared to the reference drug were selected. The Discovery Studio v21.1.0 (BIOVIA, Dassault Systѐmes, San Diego, CA, USA) was used to visualize the protein–ligand interactions. Further, the top two hits with considerable ADMET profiles were subjected to molecular dynamics simulation experiments.
Docking validation
The molecular docking protocol validation was done by redocking AKT 1 inhibitor GDC0068 (Ipatasertib) in the active pocket of AKT 1 and subsequent alignment of the X-ray crystallographic conformation of GDC0068 with the best-predicted docked pose. A perfect match between the X-ray crystallographic conformation of GDC0068 and the best-fitted docked pose was obtained, as indicated by an RMSD value of 0.674 (Fig. 3).
ADMET and drug-likeness analysis
The top five hit compounds having the highest binding energy with AKT 1 and AKT 2 proteins (Tables 1 and 2) were independently extrapolated and subjected to computational drug-likeness testing to quantify their drug-likeness. The respective compound’s absorption, distribution, metabolism, excretion, and toxicity (ADMET) characteristics were investigated using SwissADME (Daina et al. 2017) and ADMElab 2.0 (Xiong et al. 2021). Diverse parameters in line with ADMET and druggability were examined in reference to capivasertib (Pires et al., 2015; Yang et al. 2019).
Molecular dynamics
The GROMACS 2019.2 and GROMOS96 43a1 forcefield on the WebGRO (Abraham et al. 2015; Oostenbrink et al. 2004) were employed to perform a comprehensive atomistic 50 ns Molecular Dynamic (MD) simulation of the unliganded AKT1 (PDB ID: 4EKL) and AKT2 (PDB ID: 2JDR) alongside the top protein–ligand complexes as demonstrated earlier (Ogunyemi et al. 2022a, b; Olawale et al. 2023). According to Schüttelkopf and van Aalten (2004), the PRODRG webserver (http://davapc1.bioch.dundee.ac.uk/cgi-bin/prodrg) was used to create the necessary topology files for the ligand molecules. Using a four-point (TIP4P) water model and periodic boundary conditions, the apoAKT proteins and AKT-ligand complexes were solvated within a cubic box of the transferable intermolecular potential at a physiological concentration of 0.154 M established by neutralized NaCl ions. The biomolecular systems were reduced for 10,000 steps using the steepest descent algorithm in the constant atom number, constant pressure, and constant temperature ensemble (NVT) ensemble lasting 0.3 ns (NPT). Using velocity rescale, the constant temperature was kept at 310 K, and the Parrinello-Rahman barostat was used to maintain the constant pressure at 1 atm. A time step of 2 femtoseconds was used with the Leap-frog integrator. A snapshot was taken every 0.1 s with a total of 1000 frames from each system during the 50 ns production run for each system. Using VMD TK console scripts, the thermodynamic parameters RMSD, RMSF, SASA, RoG, and the number of H-bond were calculated from the trajectory files (Humphrey et al. 1996).
Results and discussion
Molecular docking analysis
In this study, we performed a structure-based virtual screening to investigate the antitumor activity of Dysphania ambrosioides. Our study explores AKT isoforms 1 and 2 as potential therapeutic targets for recalcitrant triple-negative breast cancer due to their role in cell proliferation, survival, and apoptosis inhibition. Briefly, about twenty-five (25) biologically active compounds found in Dysphania ambrosioides (Kandsi et al. 2021) and a reference drug (capivasertib) were docked in the binding pocket of AKT 1 (4EKL) and 2 (2JDR) isoforms. The binding energy of the ATP-competitive inhibitor (capivasertib) for AKT 1 and 2 were −9.5 and −8.4 kcal/mol, respectively. And these values were set as the cut-off point to identify lead-like compounds with potential AKT inhibitory activity. Since lower binding energy indicates a higher binding affinity for the target proteins (Simon et al. 2017), the top ligands having a binding strength that exceeds that of the reference drug for both AKT isoforms were identified accordingly (Fig. 2; Tables 1, 2). The top five ligands (Tables 1, 2) that bind with high affinity to AKT 1 and 2 isoforms were further subjected to computational drug-likeness testing using SwissADME and ADMETlab 2.0. Cynaroside and epicatechin gallate emerged as the top two hit compounds with fascinating pharmacokinetics and drug-likeness profiles (Fig. 3).
Structurally, AKT isoforms have three major domains; the N-terminal pleckstrin homology (PH) domain, a central catalytic kinase domain, and a C-terminal regulatory domain which houses the hydrophobic motif (HM) (Lazaro et al. 2020). The phosphorylation of the Thr 308/309 and Ser 473/474 in the catalytic domain and hydrophobic motif of AKT 1 and 2, respectively, is central to the optimal activation of their kinase activity (Hinz and Jücker 2019). Once activated, AKT 1 and 2 translocate intracellularly to phosphorylate their downstream substrates, most of which are involved in tumor initiation and propagation (Lazaro et al. 2020). However, the phosphorylation of substrates by AKT could be downregulated either by an allosteric or ATP-competitive AKT inhibitor (Lazaro et al. 2020). ATP-competitive AKT inhibitors specifically bind to the catalytic kinase domain of an activated AKT protein, thereby preventing ATP binding and phosphate transfer to target substrates (Shariati and Meric-Bernstam 2019). On the other hand, allosteric AKT inhibitors bind to the PH domain, disrupting AKT phosphorylation and activation (Shariati and Meric-Bernstam 2019). Due to limited efficacy and toxicity, the investigational drugs belonging to both classes of AKT inhibitors have yet to be approved for use in a clinical setting. Intensifying the need to increase the search for potent, discriminative, and safe AKT inhibitors.
Considering their appealing ADMET profile and superior binding strength (− 9.9 and − 10.2 kcal/mol) with AKT 1 and 2, we assessed the binding mode of cynaroside and epicatechin gallate with AKT 1 and 2 isoforms ATP-binding pockets. The analysis of the amino acid residues involved in the protein (AKT 1)-ligand interaction is summarized in Table 3. Cynaroside and epicatechin gallate interacted with more than five amino acid residues in the active pocket of AKT 1 through conventional hydrogen bonds, carbon-hydrogen bonds, and pi-interactions (Fig. 4). Similarly, the reference drug interacted with AKT 1 through conventional hydrogen bonds and pi-interactions. However, cynaroside and epicatechin gallate displayed four (Gly159, Gly162, Glu191, Glu234) and three (Lys158, Thr291, Glu228) conventional hydrogen bonds with residues in the active pocket of AKT 1, respectively. On the other hand, capivasertib has two conventional hydrogen bonds with only Asp292 residue of AKT 1 binding pocket. According to Lu et al. (2015), ATP usually binds to a cleft under the conserved G-loop (glycine-rich nucleotide motif) between the N-terminal and C-terminal lobes. Further, the G-loop (Thr160/Phe161) and αC-helix (Glu191/His194) residues have been said to function in the transduction of signal from ATP to the phosphorylated Thr308 of AKT 1 (Lu et al. 2015). Capivasertib was observed to exhibit pi-interactions with Phe161 and His194. However, cynaroside interacts with Glu159, Glu162, and Phe161 of the G-loop and Glu191 of the αC-helix via stable conventional hydrogen bonds and pi-interaction. Emphasizing its strong potential to interfere with ATP binding to AKT 1 and disrupts signal propagation from ATP to the pThr308.
The interaction pattern of cynaroside and epicatechin gallate with the active pocket of AKT 2 was also compared with that of the reference drug (Fig. 5). Cynaroside and epicatechin gallate also showed interactions with more than five residues in the binding pocket of AKT 2 through conventional hydrogen bonds, carbon-hydrogen bonds, and pi-interactions (Table 4). Conversely, the reference drug has a halogen bond in addition to the conventional hydrogen bond and pi-interactions. However, cynaroside displayed two conventional hydrogen bonds (Glu236 and Asn280) compared to the single conventional hydrogen bond (Leu158) observed in the capivasertib interaction with AKT 2. Furthermore, epicatechin gallate established two pi-alkyl (Leu158 and Val166) and a pi–sulfur interaction (Met282) with three conserved residues in AKT 2 activation loop (Huang et al. 2003). Compared to the reference drug, the relatively high binding affinity with AKT 1 and 2 active site residues suggests that these compounds could emerge as potent ATP-competitive AKT inhibitors.
ADMET and drug-likeness evaluation
Following our computational drug-likeness analysis, cynaroside and epicatechin gallate emerge as the top two hits with considerable ADMET profiles in reference to the investigational drug capivasertib. The human ether-a-go-go-related gene (hERG) potassium channel, otherwise known as KCNH2, plays a vital role in cardiac action and resting potential regulation. Thus, the inhibition of the hERG channels could prolong QT intervals and heighten the risk of ventricular arrhythmias (Lamothe et al. 2016; Vandenberg et al. 2012). A typical off-target activity that is often encountered during preclinical safety studies is the blockade of hERG channels which could result in cardiac arrest, palpitations, fainting, or sudden death. Interestingly, our ADMETlab 2.0 computational screening indicates that cynaroside (0.027) and epicatechin gallate (0.041) doesn’t interfere with hERG channels. However, the investigational drug capivasertib was predicted to have an hERG value of 0.538, suggesting a potential hERG inhibition at a relatively high concentration. The Food and Drug Administration Maximum recommended Daily Dose (FDAMDD) values of the hit compounds deduced from ADMETlab 2.0 further reiterates the better toxicity profile of cynaroside and epicatechin gallate compared to the reference drug capivasertib. Accordingly, cynaroside and epicatechin gallate have excellent FDAMDD scores of 0.031 and 0.055 respectively. Conversely, capiversatib has a poor FDAMDD value (0.922), indicating a highly toxic dose threshold in the human body (Table 5).
Further, we assessed the conformance of the top two hit compounds to Lipinski’s rule of five, Pfizer’s rule, GSK’s rule, and the Golden Triangle rule using ADMETlab 2.0. Similar to the reference drug capivasertib, epicatechin gallate was in agreement with the Lipinski, Pfizer, and Golden Triangle rules. However, cynaroside only conforms to the Pfizer and Golden Triangle rules. Together, our hit compounds were predicted to be less toxic and exhibit a favorable ADMET profile in reference to the investigational drug (capivasertib). In contrast to the reference drug (capivasertib) which has the potential of crossing the blood–brain barrier (BBB) with a relatively high BBB penetration value of 0.687 as computed by ADMETlab 2.0. Cynaroside and epicatechin gallate having respective BBB penetration values of 0.061 and 0.008 do not penetrate the BBB. Indicating that the two hit compounds have no effects on the central nervous system. Besides, cynaroside and epicatechin gallate have an impressive SAscore (synthetic accessibility score) of 3.924 and 3.608, respectively. On the other hand, the reference drug (capivasertib) has a SAscore of 3.256. Therefore, the computed SAscore values using ADMETlab 2.0 suggest an ease of synthesizing the hit compounds into a drug-like molecule.
In addition, the ADMETlab 2.0 screening revealed that epicatechin gallate has a superior clearance level (17.814) when compared to cynaroside (4.318) and the reference drug capivasertib (4.668). P-glycoprotein (Pgp), otherwise referred to as MDR1 (Multidrug resistance protein 1), is a member of the ATP-binding cassette (ABC) transporter superfamily. It is a promiscuous efflux transporter that functions mainly in the elimination of xenobiotics and toxins from the body (Koehn 2021). The inhibition of Pgp could result in delayed clearance of toxins from the body (Callaghan et al. 2014; Mealey and Fidel 2015). The hit compounds (cynaroside and epicatechin gallate) follow a similar trend with the reference drug capivasertib. As such, the hit compounds are unlikely to compromise xenobiotic or toxin clearance from the body. Further, cynaroside and epicatechin gallate, as well as the reference drug, were predicted to be non-carcinogenic and non-nephrotoxic using ADMETlab 2.0. Also, epicatechin gallate has a similar bioavailability score (0.55) to the reference drug (capivasertib), while cynaroside has a bioavailability value of 0.17.
Molecular dynamics analysis
The Root Mean Square Deviation computed from the trajectories obtained from the 50 ns atomistic MD simulation is shown in Fig. 6.
The RMSD values of the apo AKT1, cynaroside, and epicatechin gallate displayed a similar pattern and appeared more stable than the complex formed by the reference compound (Fig. 6a). The initial 8, 10, 9, and 15 ns of the apo AKT1 protein and AKT1-cynaroside complex, AKT1-epicatechin gallate complex, and AKT1- capivasertib complex showed an increasing trend before equilibration and stabilizing at average values of within 2 Å. In the case of AKT2 biosystems, the AKT2-epicatechin gallate complex showed less stability than those of Apo AKT2, AKT2-cynaroside complex, and AKT2-capivasertib (Fig. 6b). The apo-proteins and ligand-receptor complexes were stable upon equilibration since all RMSD values were under 2 Å. The tendency of the apo-protein and protein–ligand complexes to unfold away from their original structures was estimated using the Radius of Gyration (RoG), a thermodynamics parameter that measures the compactness of biomolecular systems. The RoG computed from the trajectories obtained from the MD simulations performed in this study is shown in Fig. 6.
Based on the RoG calculations, the apo AKT1 and AKT1 complexes with the capivasertib, cynaroside, and epicatechin gallate showed a stable RoG trend kept within 2 Å (Fig. 7a). Similar results were obtained for AKT2 systems (Fig. 7b).
The Surface Accessible Surface Area of the apo-proteins and the ligand–protein complexes were also computed from the trajectories obtained from the MD simulations, as depicted in Fig. 8.
The SASA plots show that the accessible area of all the complexes is greater than that of the apo-proteins.
The cynaroside and epicatechin gallate in complexes with the AKT1 and AKT2 displayed a slightly higher number of hydrogen bonds than that of apo-proteins and the complexes formed by the reference compounds.
The inherent flexibility of proteins is closely related to their biochemical functions. Thus, the detailed, atomistic protein–ligand binding in a simulated dynamic environment was modeled in this study using Root Mean Square Deviation (RMSF), as shown in Fig. 9.
The RMSF values of the apo-proteins and those of the complexes revealed that some binding site residues indicated fluctuations greater than 2 Å. In addition, other residues in proximity to the binding site also exhibit several peaks with significantly increased RMSF values (Fig. 10). Analyzing the binding interactions of a protein system with specific ligands created by molecular docking simulation is a typical application of MD Simulation. In ligand-receptor complexes, MDS can simulate both bound and non-bonded interactions. Consequently, an integrated approach utilizing molecular docking and dynamic simulation can provide detailed information on ligand-receptor interactions. Thus, the computational studies suggest that the hit compounds reported in this study exhibit dual targeting and stable binding interactions with AKT1 and AKT2. A recent study by Halder and Cordeiro (2021) reported similar in silico results and provided important guidelines for discovering novel AKT inhibitors. Therefore, these in silico hits can be exploited further for developing potent AKT inhibitors.
Conclusion
The development of a targeted therapeutic regimen for Triple-negative breast cancer patients remains an indisputable problem of clinical medicine. The hyperactivation of the AKT signaling pathway has been found to contribute enormously to the propagation of the aggressive phenotype of TNBCs, making AKT a vulnerable target for novel therapeutic intervention. Therefore, inhibiting this downstream signal transducer using highly selective and efficacious small molecule inhibitors could be a promising approach in TNBC management. In the present study, we identified two drug candidates (epicatechin gallate and cynaroside) that significantly inhibit AKT 1 and AKT 2 isoforms with high affinity than the reference drug. Furthermore, our computational drug-likeness evaluation showed that the lead-like molecules, particularly epicatechin gallate, demonstrate superior ADMET properties over the investigational drug capivasertib. Importantly, our in silico hits displayed stable binding interactions within the active pocket of AKT 1 and AKT 2 proteins through the 50 ns molecular dynamics trajectories. Our data indicate that the hit molecules could emerge as potent, safe, and discriminative AKT inhibitors. However, further experimental in vivo studies are required to establish their empirical application in a clinical setting.
Abbreviations
- ADMET:
-
Absorption, distribution, metabolism, excretion, and toxicity
- AKT:
-
Serine/threonine kinase
- AR:
-
Androgen receptor
- BAD:
-
Proapoptotic BCL-2 family protein
- BC:
-
Breast cancer
- BCL2:
-
B-cell lymphoma 2
- BL1:
-
Basal-like 1
- BL2:
-
Basal-like 2,
- BRCA1/2:
-
Breast cancer gene 1/2
- C:
-
Catalytic domain
- EGFR:
-
Epidermal growth factor receptor
- EMT:
-
Epithelial–mesenchymal transition
- ER:
-
Estrogen receptor
- ERBB2:
-
Erythroblastic oncogene B-2
- FDA:
-
Food and Drug Administration
- FGFR1:
-
Fibroblast growth factor receptor 1
- FKHR:
-
Forkhead protein
- Foxo:
-
Forkhead box protein O1
- HER2:
-
Human epidermal growth factor receptor 2
- IM:
-
Immunomodulatory
- LAR:
-
Luminal androgen receptor
- M:
-
Mesenchymal
- MAGI3:
-
Membrane-associated guanylate kinase, WW, and PDZ domain containing 3
- MD:
-
Molecular dynamics
- MDM-2:
-
Mouse double minute 2
- MSL:
-
Mesenchymal stem-like
- mTOR:
-
Mechanistic target of rapamycin
- mTORC2:
-
Mechanistic target of rapamycin complex 2
- N:
-
N-terminal domain
- NF-κB:
-
Nuclear factor κB;
- P21:
-
Cyclin-dependent kinases inhibitor (CDKI)
- p27:
-
Cyclin-dependent kinase inhibitor (KIP1)
- PDB:
-
Protein Data Bank
- PDK1:
-
3-Phosphoinositide-dependent kinase 1
- PDK2:
-
Pyruvate dehydrogenase kinase 2
- PH:
-
N-terminal pleckstrin homology domain
- PI3K:
-
Phosphoinositide 3-kinase
- PIK3CA:
-
Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit α
- PIP2:
-
Phosphatidylinositol 4,5-bisphosphate
- PIP3:
-
Phosphatidylinositol-3,4,5-trisphosphate
- PR:
-
Progesterone receptor
- PTEN:
-
Phosphatase and tensin homolog
- RCSB:
-
Research collaboratory for structural bioinformatics
- SDF:
-
Structure data file
- TN:
-
Triple negative
- TNBC:
-
Triple negative breast cancer
- TP53:
-
Tumor protein P53
- VEGFRA:
-
Vascular endothelial growth factor receptor A
References
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Aine M, Boyaci C, Hartman J, Häkkinen J, Mitra S, Campos AB, Nimeus E, Ehinger A, Vallon-Christersson J, Borg Å, Staaf J (2021) Molecular analyses of triple-negative breast cancer in the young and elderly. Breast Cancer Res 23(1):1–19. https://doi.org/10.1186/S13058-021-01392-0
Anders CK, Abramson V, Tan T, Dent R (2016) The evolution of triple-negative breast cancer: from biology to novel therapeutics. Am Soc Clin Oncol Educ Book 36:34–42. https://doi.org/10.1200/edbk_159135
Banerji S, Cibulskis K, Rangel-Escareno C, Brown KK, Carter SL, Frederick AM, Lawrence MS, Sivachenko AY, Sougnez C, Zou L, Cortes ML, Fernandez-Lopez JC, Peng S, Ardlie KG, Auclair D, Bautista-Piña V, Duke F, Francis J, Jung J et al (2012) Sequence analysis of mutations and translocations across breast cancer subtypes. Nature 486(7403):405–409. https://doi.org/10.1038/NATURE11154
Basu A, Lambring CB (2021) AKT isoforms: a family affair in breast cancer. Cancers 2021(13):3445. https://doi.org/10.3390/CANCERS13143445
Bellacosa A, de Feo D, Godwin AK, Bell DW, Cheng JQ, Altomare DA, Wan M, Dubeau L, Scambia G, Masciullo V, Ferrandina G, Panici PB, Mancuso S, Neri G, Testa JR (1995) Molecular alterations of the AKT2 oncogene in ovarian and breast carcinomas. Int J Cancer 64(4):280–285. https://doi.org/10.1002/IJC.2910640412
Callaghan R, Luk F, Bebawy M (2014) Inhibition of the multidrug resistance P-glycoprotein: time for a change of strategy? Drug Metab Disposit Biol Fate Chem 42(4):623–631. https://doi.org/10.1124/dmd.113.056176
Chandarlapaty S, Sawai A, Scaltriti M, Rodrik-Outmezguine V, Grbovic-Huezo O, Serra V, Majumder PK, Baselga J, Rosen N (2011) AKT inhibition relieves feedback suppression of receptor tyrosine kinase expression and activity. Cancer Cell 19(1):58–71. https://doi.org/10.1016/J.CCR.2010.10.031
Chen X, Ariss MM, Ramakrishnan G, Nogueira V, Blaha C, Putzbach W, Islam ABMMK, Frolov MV, Hay N (2020) Cell-autonomous versus systemic AKT isoform deletions uncovered new roles for AKT1 and AKT2 in breast cancer. Mol Cell 80(1):87-101.e5. https://doi.org/10.1016/J.MOLCEL.2020.08.017
Cheng JQ, Godwin AK, Bellacosa A, Taguchi T, Franke TF, Hamilton TC, Tsichlis PN, Testa JR (1992) AKT2, a putative oncogene encoding a member of a subfamily of protein-serine/threonine kinases, is amplified in human ovarian carcinomas. Proc Natl Acad Sci USA 89(19):9267–9271. https://doi.org/10.1073/PNAS.89.19.9267
Ciriello G, Magnani L (2021) The many faces of cancer evolution. IScience 24(5):102403. https://doi.org/10.1016/J.ISCI.2021.102403
da Silva SB, Barbosa JR, da Silva Martins LH, Rai M, Lopes AS (2021) Traditional uses, phytochemicals, and pharmacological properties of Chenopodium ambrosioides L. (Dysphania ambrosioides) L. Mosyakin & Clemants. Ethnopharmacol Wild Plants. https://doi.org/10.1201/9781003052814-14
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx. Methods Mol Biol (Clifton, NJ) 1263:243–250. https://doi.org/10.1007/978-1-4939-2269-7_19
Davies TG, Verdonk ML, Graham B, Saalau-Bethell S, Hamlett CCF, McHardy T, Collins I, Garrett MD, Workman P, Woodhead SJ, Jhoti H, Barford D (2007) A structural comparison of inhibitor binding to PKB, PKA and PKA-PKB chimera. J Mol Biol 367(3):882–894. https://doi.org/10.1016/J.JMB.2007.01.004
Davies BR, Greenwood H, Dudley P, Crafter C, Yu DH, Zhang J, Li J, Gao B, Ji Q, Maynard J, Ricketts SA, Cross D, Cosulich S, Chresta CC, Page K, Yates J, Lane C, Watson R, Luke R, Pass M (2012) Preclinical pharmacology of AZD5363, an inhibitor of AKT: pharmacodynamics, antitumor activity, and correlation of monotherapy activity with genetic background. Mol Cancer Ther 11(4):873–887. https://doi.org/10.1158/1535-7163.MCT-11-0824-T
Degenhardt RT, Farias IV, Grassi LT, Franchi GC, Nowill AE, da Bittencourt CMS, Wagner TM, de Souza MM, Cruz AB, Malheiros A (2016) Characterization and evaluation of the cytotoxic potential of the essential oil of Chenopodium ambrosioides. Rev Bras Farmacogn 26(1):56–61
Faje AT, Lawrence D, Flaherty K, Freedman C, Fadden R, Rubin K, Cohen J, Sullivan RJ (2018) High-dose glucocorticoids for the treatment of ipilimumab-induced hypophysitis is associated with reduced survival in patients with melanoma. Cancer 124(18):3706–3714. https://doi.org/10.1002/CNCR.31629
Gener P, Rafael D, Seras-Franzoso J, Perez A, Pindado LA, Casas G, Arango D, Fernández Y, Díaz-Riascos ZV, Abasolo I, Schwartz S (2019) The pivotal role of AKT2 during dynamic phenotypic change of breast cancer stem cells. Cancers 11(8):1058. https://doi.org/10.3390/CANCERS11081058
Gottifredi V (2020) Targeting DNA damage response kinases in cancer therapy. Mutat Res 821:111725. https://doi.org/10.1016/J.MRFMMM.2020.111725
Greer YE, Lipkowitz S (2015) TIC10/ONC201: a bend in the road to clinical development. Oncoscience 2(2):75–76. https://doi.org/10.18632/ONCOSCIENCE.133
Halder AK, Cordeiro M (2021) AKT inhibitors: the road ahead to computational modeling-guided discovery. Int J Mol Sci 22(8):3944. https://doi.org/10.3390/ijms22083944
Hinz N, Jücker M (2019) Distinct functions of AKT isoforms in breast cancer: a comprehensive review. Cell Commun Signal 17(1):1–29. https://doi.org/10.1186/S12964-019-0450-3
Hoxhaj G, Manning BD (2020) The PI3K–AKT network at the interface of oncogenic signaling and cancer metabolism. Nat Rev Cancer 20(2):74–88. https://doi.org/10.1038/S41568-019-0216-7
Huang X, Begley M, Morgenstern KA, Gu Y, Rose P, Zhao H, Zhu X (2003) Crystal structure of an inactive AKT2 kinase domain. Structure 11(1):21–30. https://doi.org/10.1016/S0969-2126(02)00937-1
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Kandsi F, Conte R, Marghich M, Lafdil FZ, Alajmi MF, Bouhrim M, Mechchate H, Hano C, Aziz M, Gseyra N (2021) Phytochemical analysis, antispasmodic, myorelaxant, and antioxidant effect of Dysphania ambrosioides (L.) mosyakin and clemants flower hydroethanolic extracts and its chloroform and ethyl acetate fractions. Molecules 26(23):7300. https://doi.org/10.3390/MOLECULES26237300
Khan MA, Jain VK, Rizwanullah M, Ahmad J, Jain K (2019) PI3K/AKT/mTOR pathway inhibitors in triple-negative breast cancer: a review on drug discovery and future challenges. Drug Discov Today 24(11):2181–2191. https://doi.org/10.1016/J.DRUDIS.2019.09.001
Koehn LM (2021) ABC transporters: an overview BT. The ADME encyclopedia: a comprehensive guide on biopharmacy and pharmacokinetics. Springer International Publishing, Cham, pp 1–10
Koo MM, von Wagner C, Abel GA, McPhail S, Rubin GP, Lyratzopoulos G (2017) Typical and atypical presenting symptoms of breast cancer and their associations with diagnostic intervals: evidence from a national audit of a cancer diagnosis. Cancer Epidemiol 48:140. https://doi.org/10.1016/J.CANEP.2017.04.010
Kumar Prusty R, Begum S, Patil A, Naik DD, Pimple S, Mishra G (2020) Knowledge of symptoms and risk factors of breast cancer among women: a community-based study in a low socio-economic area of Mumbai, India. BMC Women’s Health. https://doi.org/10.1186/s12905-020-00967-x
Lamothe SM, Guo J, Li W, Yang T, Zhang S (2016) The human ether-a-go-go-related Gene (hERG) potassium channel represents an unusual target for protease-mediated damage. J Biol Chem 291(39):20387–20401. https://doi.org/10.1074/jbc.M116.743138
Lazaro G, Kostaras E, Vivanco I (2020) Inhibitors in AKTion: ATP-competitive vs allosteric. Biochem Soc Trans 48(3):933–943. https://doi.org/10.1042/BST20190777
Lin K, Lin J, Wu WI, Ballard J, Lee BB, Gloor SL, Vigers GPA, Morales TH, Friedman LS, Skelton N, Brandhuber BJ (2012) An ATP-site on-off switch that restricts phosphatase accessibility of AKT. Sci Signal 5(223). https://doi.org/10.1126/SCISIGNAL.2002618
Lin J, Sampath D, Nannini MA, Lee BB, Degtyarev M, Oeh J, Savage H, Guan Z, Hong R, Kassees R, Lee LB, Risom T, Gross S, Liederer BM, Koeppen H, Skelton NJ, Wallin JJ, Belvin M, Punnoose E et al (2013) Targeting activated AKT with GDC-0068, a novel selective AKT inhibitor that is efficacious in multiple tumor models. Clin Cancer Res 19(7):1760–1772. https://doi.org/10.1158/1078-0432.CCR-12-3072
LoRusso PM (2016) Inhibition of the PI3K/AKT/mTOR pathway in solid tumors. J Clin Oncol Off J Am Soc Clin Oncol 34(31):3803–3815. https://doi.org/10.1200/JCO.2014.59.0018
Lu S, Deng R, Jiang H, Song H, Li S, Shen Q, Huang W, Nussinov R, Yu J, Zhang J (2015) The mechanism of ATP-dependent allosteric protection of AKT kinase phosphorylation. Structure (London, England: 1993) 23(9):1725–1734. https://doi.org/10.1016/J.STR.2015.06.027
Łukasiewicz S, Czeczelewski M, Forma A, Baj J, Sitarz R, Stanisławek A (2021) Breast cancer—epidemiology, risk factors, classification, prognostic markers, and current treatment strategies—an updated review. Cancers 13(17):4287. https://doi.org/10.3390/CANCERS13174287
Mahmoud R, Ordóñez-Morán P, Allegrucci C (2022) Challenges for triple negative breast cancer treatment: defeating heterogeneity and cancer stemness. Cancers 14(17):4280. https://doi.org/10.3390/CANCERS14174280
Manning BD, Toker A (2017) AKT/PKB signaling: navigating the network. Cell 169(3):381–405. https://doi.org/10.1016/J.CELL.2017.04.001
Marra A, Trapani D, Viale G, Criscitiello C, Curigliano G (2020) A practical classification of triple-negative breast cancer: intratumoral heterogeneity, mechanisms of drug resistance, and novel therapies. NPJ Breast Cancer 6(1):1–16. https://doi.org/10.1038/s41523-020-00197-2
Mealey KL, Fidel J (2015) P-glycoprotein mediated drug interactions in animals and humans with cancer. J Vet Intern Med 29(1):1–6. https://doi.org/10.1111/JVIM.12525
Mephors VC, Ogbole OO, Ajaiyeoba EO (2018) Plants used in treatment of five cancers in two local government areas in southwest Nigerian ethnomedicine. Niger J Nat Prod Med 21(1):54–60. https://doi.org/10.4314/njnpm.v21i1
Merckaert T, Zwaenepoel O, Gevaert K, Gettemans J (2021) An AKT2-specific nanobody that targets the hydrophobic motif induces cell cycle arrest, autophagy, and loss of focal adhesions in MDA-MB-231 cells. Biomed Pharmacother. https://doi.org/10.1016/J.BIOPHA.2020.111055
Millis SZ, Gatalica Z, Winkler J, Vranic S, Kimbrough J, Reddy S, O’Shaughnessy JA (2015) Predictive biomarker profiling of > 6000 breast cancer patients shows heterogeneity in TNBC with treatment implications. Clin Breast Cancer 15(6):473-481.e3. https://doi.org/10.1016/J.CLBC.2015.04.008
Nazmy MH, Hamdy Abu-Baih D, Abdul-Aziz El-Rehany M, Fathy M (2021) Pathways of triple-negative breast cancer. MJMR 32(4):1–3
Nnyigide OS, Nnyigide TO, Lee SG, Hyun K (2022) Protein repair and analysis server: a web server to repair PDB structures, add missing heavy atoms and hydrogen atoms, and assign secondary structures by amide interactions. J Chem Inf Model 62(17):4232–4246. https://doi.org/10.1021/ACS.JCIM.2C00571/SUPPL_FILE/CI2C00571_SI_001.PDF
O’Reilly KE, Rojo F, She QB, Solit D, Mills GB, Smith D, Lane H, Hofmann F, Hicklin DJ, Ludwig DL, Baselga J, Rosen N (2006) mTOR inhibition induces upstream receptor tyrosine kinase signaling and activates AKT. Can Res 66(3):1500–1508. https://doi.org/10.1158/0008-5472.CAN-05-2925
Ogunyemi OM, Gyebi GA, Ibrahim IM, Esan AM, Olaiya CO, Soliman MM, Batiha GE-S (2022a) Identification of promising multi-targeting inhibitors of obesity from Vernonia amygdalina through computational analysis. Mol Divers. https://doi.org/10.1007/s11030-022-10397-6
Ogunyemi OM, Gyebi GA, Saheed A, Paul J, Nwaneri-Chidozie V, Olorundare O, Olaiya CO (2022b) Inhibition mechanism of alpha-amylase, a diabetes target, by a steroidal pregnane and pregnane glycosides derived from Gongronema latifolium Benth [Original Research]. Front Mol Biosci. https://doi.org/10.3389/fmolb.2022.866719
Olawale F, Olofinsan K, Ogunyemi OM, Karigidi KO, Gyebi GA, Ibrahim IM, Iwaloye O (2023) Deciphering the therapeutic role of Kigelia africana fruit in erectile dysfunction through metabolite profiling and molecular modelling. Inform Med Unlocked 37:101190. https://doi.org/10.1016/j.imu.2023.101190
Oostenbrink C, Villa A, Mark AE, van Gunsteren WF (2004) A biomolecular force field based on the free enthalpy of hydration and solvation: the GROMOS force-field parameter sets 53A5 and 53A6. J Comput Chem 25(13):1656–1676. https://doi.org/10.1002/jcc.20090
Oyedele AQK, Adelusi TI, Ogunlana AT, Ayoola MA, Adeyemi RO, Babalola MO, Ayorinde JB, Isong JA, Ajasa TO, Boyenle ID (2022) Promising disruptors of p53-MDM2 dimerization from some medicinal plant phytochemicals: a molecular modeling study. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2022.2097313
Pires DE, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem. 58(9):4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Rasul HO, Aziz BK, Ghafour DD, Kivrak A (2022) In silico molecular docking and dynamic simulation of eugenol compounds against breast cancer. J Mol Model 28(1):1–18. https://doi.org/10.1007/S00894-021-05010-W/FIGURES/17
Risso G, Blaustein M, Pozzi B, Mammi P, Srebrow A (2015) AKT/PKB: One kinase, many modifications. Biochem J 468(2):203–214. https://doi.org/10.1042/BJ20150041
Rodrik-Outmezguine VS, Chandarlapaty S, Pagano NC, Poulikakos PI, Scaltriti M, Moskatel E, Baselga J, Guichard S, Rosen N (2011) mTOR kinase inhibition causes feedback-dependent biphasic regulation of AKT signaling. Cancer Discov 1(3):248–259. https://doi.org/10.1158/2159-8290.CD-11-0085
Schüttelkopf AW, van Aalten DM (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr D Biol Crystallogr 60(Pt 8):1355–1363. https://doi.org/10.1107/s0907444904011679
Shariati M, Meric-Bernstam F (2019) Targeting AKT for cancer therapy. Expert Opin Investig Drugs 28(11):977–988. https://doi.org/10.1080/13543784.2019.1676726
Sherman M, Gaebe K, Li AY, Habbous S, Sahgal A, Raphael MJ, Erickson AW, Das S (2022) Erythroblastic oncogene B-2 status and intracranial metastatic disease in patients with gastrointestinal cancer: a systematic review. J Neurooncol 160(3):735–742. https://doi.org/10.1007/S11060-022-04195-1
Siegel RL, Miller KD, Fuchs HE, Jemal A (2022) Cancer statistics, 2022. CA Cancer J Clin 72(1):7–33. https://doi.org/10.3322/CAAC.21708
Siegel RL, Miller KD, Wagle NS, Jemal A (2023) Cancer statistics, 2023. CA Cancer J Clin 73(1):17–48. https://doi.org/10.3322/CAAC.21763
Simon L, Imane A, Srinivasan KK, Pathak L, Daoud I (2017) In silico drug-designing studies on flavanoids as anticolon cancer agents: pharmacophore mapping, molecular docking, and Monte Carlo method-based QSAR modeling. Interdiscip Sci Comput Life Sci 9(3):445–458. https://doi.org/10.1007/S12539-016-0169-4
Skladanowski A, Bozko P, Sabisz M, Larsen AK (2007) Dual inhibition of PI3K/AKT signaling and the DNA damage checkpoint in p53-deficient cells with strong survival signaling: implications for cancer therapy. Cell Cycle (Georgetown, TX) 6(18):2268–2275. https://doi.org/10.4161/CC.6.18.4705
Sporikova Z, Koudelakova V, Trojanec R, Hajduch M (2018) Genetic markers in triple-negative breast cancer. Clin Breast Cancer 18(5):e841–e850. https://doi.org/10.1016/J.CLBC.2018.07.023
Staal SP, Huebner K, Croce CM, Parsa NZ, Testa JR (1988) The AKT1 proto-oncogene maps to human chromosome 14, band q32. Genomics 2(1):96–98. https://doi.org/10.1016/0888-7543(88)90114-0
Stein MN, Bertino JR, Kaufman HL, Mayer T, Moss R, Silk A, Chan N, Malhotra J, Rodriguez L, Aisner J, Aiken RD, Haffty BG, DiPaola RS, Saunders T, Zloza A, Damare S, Beckett Y, Yu B, Najmi S et al (2017) A first-in-human clinical trial of oral ONC201 in patients with refractory solid tumors. Clin Cancer Res 23(15):4163–4169. https://doi.org/10.1158/1078-0432.CCR-16-2658
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/CAAC.21660
Tauchen J, Huml L, Bortl L, Doskocil I, Jarosova V, Marsik P, Frankova A, Clavo Peralta ZM, Chuspe Zans ME, Havlik J, Lapcik O, Kokoska L (2019) Screening of medicinal plants traditionally used in the Peruvian Amazon for in vitro antioxidant and anticancer potential. Nat Prod Res 33(18):2718–2721. https://doi.org/10.1080/14786419.2018.1462180
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. https://doi.org/10.1002/JCC.21334
Turner KM, Sun Y, Ji P, Granberg KJ, Bernard B, Hu L, Cogdell DE, Zhou X, Yli-Harja O, Nykter M, Shmulevich I, Yung WKA, Fuller GN, Zhang W (2015) Genomically amplified AKT3 activates the DNA repair pathway and promotes glioma progression. Proc Natl Acad Sci USA 112(11):3421–3426. https://doi.org/10.1073/PNAS.1414573112
Unger H, van Golen KL (2009) AKT signaling in oncology. Belgian J Med Oncol 3(4):139–144
Vandenberg JI, Perry MD, Perrin MJ, Mann SA, Ke Y, Hill AP, Herg KHA (2012) hERG K channels: structure, function, and clinical significance. Physiol Rev 92:1393–1478. https://doi.org/10.1152/physrev.00036.2011
Waks AG, Winer EP (2019) Breast cancer treatment: a review. JAMA 321(3):288–300. https://doi.org/10.1001/JAMA.2018.19323
Wang DY, Jiang Z, Ben-David Y, Woodgett JR, Zacksenhaus E (2019) Molecular stratification within triple-negative breast cancer subtypes. Sci Rep. https://doi.org/10.1038/S41598-019-55710-W
Wang XY, Hao JM, Ren QR, Li HY, Wu JS, Zhu XH, Chen JY, Wang YN, Zhang LS (2021) Cytotoxicity and apoptosis induced by Chenopodium ambrosioides L. essential oil in human normal liver cell line L02 via the endogenous mitochondrial pathway rather than the endoplasmic reticulum stress. Int J Environ Res Public Health 18(14):7469. https://doi.org/10.3390/IJERPH18147469
Xiong G, Wu Z, Yi J, Fu L, Yang Z, Hsieh C, Yin M, Zeng X, Wu C, Lu A, Chen X, Hou T, Cao D (2021) ADMETlab 2.0: an integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucl Acids Res 49(W1):W5–W14. https://doi.org/10.1093/NAR/GKAB255
Yang H, Lou C, Sun L, Li J, Cai Y, Wang Z, Li W, Liu G, Tang Y (2019) AdmetSAR 2.0: web-service for prediction and optimization of chemical ADMET properties. Bioinformatics 35(6):1067–1069. https://doi.org/10.1093/bioinformatics/bty707
Yin L, Duan JJ, Bian XW, Yu SC (2020) Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res 22(1):1–13. https://doi.org/10.1186/S13058-020-01296-5/TABLES/3
Younus S, Vinod Chandra SS, Ibrahim J, Nair ASS (2022) A new approach used in docking study for predicting the combination drug efficacy in the EML4-ALK target of NSCLC. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2022.2091658
Zakaria A-S, Naglaa E-K, Loay IH, Nassar R, Darwish T, Mervat E-D, Allam M, Nabil H (2018). Triple negative breast cancer, clinicopathologic study of Egyptian patients, NCI experience. In: Cairo Univ, vol 86(5). www.medicaljournalofcairouniversity.net
Zohra T, Ovais M, Khalil AT, Qasim M, Ayaz M, Shinwari ZK (2018) Extraction optimization, total phenolic, flavonoid contents, HPLC-DAD analysis and diverse pharmacological evaluations of Dysphania ambrosioides (L.) Mosyakin & Clemants. Nat Product Res 33(1):136–142. https://doi.org/10.1080/14786419.2018.1437428
Author information
Authors and Affiliations
Contributions
LOA—conceptualization, methodology, data curation, writing—original draft, writing—review and editing. OKP—conceptualization, methodology, data curation, writing—original draft. FNA—methodology, data curation, writing—original draft, writing—review and editing. OMO—methodology, data curation, writing—original draft. ROA—data curation, writing—original draft. SFH—writing—review and editing. MAG—writing—review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no financial or non-financial competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Anifowose, L.O., Paimo, O.K., Adegboyega, F.N. et al. Molecular docking appraisal of Dysphania ambrosioides phytochemicals as potential inhibitor of a key triple-negative breast cancer driver gene. In Silico Pharmacol. 11, 15 (2023). https://doi.org/10.1007/s40203-023-00152-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40203-023-00152-6