Abstract
Early blight (EB), caused by the fungus, Alternaria solani, is one of the most destructive diseases of tomatoes and other solanaceous crops; particularly in warm and humid climate. This study was targeted to explore the genetic and pathogenic diversity of A. solani from major tomato producing states of India. Thirty-three isolates were chosen for this study. These isolates exhibited considerable intra as well as inter-state variation. The phylogenetic tree generated with the ISSR sequences confirmed this result. Aggressiveness of the isolates towards susceptible tomato genotype was assessed in vitro, using detached leaf method. Considerable amount of variability in virulence was observed among the isolates. Specific activity of polygalacturonase and pectin methyl esterase was also estimated, to observe the relation among these, with the virulence of isolates. The information generated in the present study provides initial data on the population variability of the EB pathogen. It could be a valuable aid for tomato breeding strategies, aimed at obtaining cultivars with resilient resistance. This will provide a basis for planning disease protection strategies for sustainable agriculture which is required for producing crop plants which harmonize with the environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In developing countries like India, a major concern is the fact that agricultural land is being continuously limited. Decrease in area under crop with different biotic and abiotic stresses had affected productivity which has resulted in threat to food security. These stress conditions in agricultural ecosystems can occur at variable intensities. Continuous exposure to these stress factors; either biotic or abiotic, affects the growth, development and production of the plant. EB, caused by Alternaria solani, is one of the most destructive diseases in solanaceous vegetable crops (tomato, potato and eggplant etc.). It affects the aerial parts i.e. fruits, stem and leaves of tomatoes and stem, foliage and tubers of potatoes. Warm and humid (25–29 °C, 90–100%) environmental conditions are conducive to infection. Germination of Alternaria conidia requires free moisture and an optimum temperature of 28–30 °C. The average time for its germination has been notice is approximately 40 min (apsnet.org). EB is distributed worldwide and predominantly occurs wherever tomato and potato are grown. The losses due to early blight disease in solanaceous crops may be up to 80% [1]. It has already been established that variation in populations of plant pathogens directly affects disease management, especially the strategies related to the deployment of resistant cultivars and fungicide usage [2]. The present research is focused on the diversity present in identified A. solani isolates from different states of India. The authors hope to generate information which will be useful for integrated disease management (IDM) programs against EB and for developing new control strategies of the disease.
The interaction between the pathogen and its host possesses diversity that is mainly dependent on the geographical and environmental conditions prevailing in the area where the crop is grown. When a plant is attacked by a pathogen, the plant and the pathogen try to overcome each other by producing some biochemicals. The first line of barrier to colonize the pathogen in plant is cell wall of host plant. Secretion of cell wall degrading enzymes by the pathogen is the first step towards its establishment in the plant. Pectin is one of the major and most complex components of plant cell wall. Plant pathogenic fungi are known to produce a range of cell wall degrading enzymes that macerate plant cell walls; including pectolytic enzymes such as pectin methylesterase (PME), polygalacturonase (PG) and pectate lyase (PNL and PL, respectively) that may have important roles in the infectivity in the progress of disease symptoms [3, 4]. The plant cell contains cell wall and pectin in it. Hence, the importance of such enzymes in pathogenicity is supported by the ability of purified enzymes to reproduce disease symptoms [5] and by the correlation of the pectolytic enzyme level with the degree of the symptoms shown by the plant [6]. This has been taken in account here for assessing the pathogenicity of different isolates of Alternaria, as they are the necrotroph. Along with the above mentioned cell wall degrading enzymes, many DNA based markers have been utilized to investigate the genetic diversity among plant pathogens. Studies of genetic variations in the population of plant pathogens in the last decade has increased because of availability of molecular markers. These techniques consist of random amplified polymorphic DNA [7, 8], microsatellites, restriction fragment length polymorphism (RFLP) [9], amplified fragment length polymorphism (AFLP) [10], and inter simple sequence repeats (ISSR) [11]. Other markers, which are being used for this purpose, are internal transcribed regions [12], rRNA coding regions of nuclear DNA [13] and mitochondrial DNA [14].
Determination of genetic diversity using inter simple sequence repeats (ISSR) is widely used in genotyping the Alternaria species as well as other fungal genera [15, 16]. This marker analysis is based on PCR-technique. ISSRs are amplified by PCR, using microsatellite core sequences as primers, with a few selective nucleotides as anchors, into the non-repeat adjacent regions (16–18 bp). Approximately 10–60 fragments from multiple loci are generated simultaneously, depending on the size of genome, separated by gel electrophoresis and scored as the presence or absence of fragments of a particular size. ISSR primers produce dominant molecular markers. ISSR fingerprinting shows higher levels of polymorphisms, when compared to some of the other PCR-based techniques. Its strength is that it does not require the sequence information for use. Due to the fact that this marker analysis is reliable and reproducible, it is preferred over other dominant markers.
Material and Methods
Fungal isolates
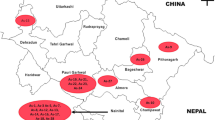
Totally, 33 isolates of A. solani (Table 1) were taken for the present study. Out of 33 isolates of A. solani, 28 were procured from National Bureau of Agriculturally Important Microorganisms (NBAIM), Mau Nath Bhanjan (Uttar Pradesh), 02 from Indian Agricultural Research Institute, New Delhi and and one each was collected from Kanpur, Varanasi, and Kalyani-West Bengal. Maintenance of the isolates and production of mycelium were performed as per instruction of the institutes, NBAIM and IARI.
Cultural Variation
The cultures were grown on the potato dextrose agar (PDA) plates at 25 ± 2 °C (12 h light/12 h dark) in a culture room. The isolates were characterized on the basis of colony color and radial growth after 5 and 10 days of inoculation. The experiment was performed in three replicates.
Detached Leaf Assay for Pathogenicity
The CO-3 variety of tomato, susceptible to A. solani [17] was grown in pots containing soil-compost mixture (2:1) in a greenhouse (25–28 °C). For detached leaf assay, healthy tomato leaves (5 weeks old plants) were surface sterilized using 2% of NaOCl for 10–15 s. Sterilized leaves were then rinsed thrice using sterile distilled water. Two (2) mm agar discs were placed on the adaxial side of leaves with the mycelium side facing down. Three leaves were inoculated per isolate and incubated at 25 ± 2 °C (12 h light/12 h dark) for 7 days. Disease severity was determined using the rating scale: 0 = no symptoms, 1 = necrosis around the lesion, 2 = necrosis covering ½ to ¾ of leaf, 3 = whole leaf necrosis, 4 = petiole necrosis (petiole detached from stem) and 5 = necrosis on plant stem [18]. The experiment was performed in triplicates.
Hydrolytic Enzyme Activities
Activities of cell wall degrading enzymes such as pectin methyl esterase (PME) and polygalacturonase (PG) were determined spectrometrically. Fungal strains grown in induction medium containing pectin as a substrate were incubated for 10 days at 25 ± 2 °C. After incubation, the contents were filtered and the filtrate was used as an enzyme extract.
Polygalacturonase (PG) Assay
To 0.8 ml of enzyme extract, 2 ml of assay buffer (2 mg/ml pectin in 0.1 M citrate buffer, pH 5.0) was added and the contents were incubated at 37 °C for 1 h. Five and half ml of phenol sulphuric acid reagent (PSA; 0.5 ml 80% phenol in 5 ml sulphuric acid) was added to 1 ml reaction mixture and absorbance of the resulting solution was measured at 480 nm. The activity of the enzyme was calculated by using a standard calibration curve obtained using galacturonic acid as reducing sugar. One unit corresponded to 1 µM of reducing sugar liberated from substrate in 1 h at 37 °C [19].
Pectin Methyl Esterase (PME) Assay
Enzyme extract (0.5 ml) was mixed with 7 ml assay buffer (10 mg/ml pectin in 0.02 M Tris–HCl buffer, pH 8.0). Initial pH of the solution was maintained at 8.0 by using 0.05 M NaOH. The reaction mixture was incubated for 1 h at 37 °C in a water bath and the initial pH was noted. The pH was again adjusted to 8.0 by titrating against 0.02 N NaOH containing 5 mM sodium azide. The volume that was required to bring back the pH was noted. The amount of enzyme that was utilized was calculated as 1 µM/h NaOH [20]. Calculations for the activity were done according to the formula given by Balaban et al. [20]. Both the biochemical assays were carried out in triplicates and repeated twice. Induction medium with no fungal inoculation served as the control.
Analysis of Genetic Variation Through ISSR Markers
Fungal Strains and Genomic DNA Isolation
For genomic DNA isolation, isolates were subcultured in 20 ml Potato dextrose broth (PDB) medium and incubated at 25 ± 2 °C for 10 days. The fungal material was removed from the broth and modified Doyle and Doyle [21] method was used for DNA isolation. Genomic DNA quality was checked with electrophoresis in 0.8% agarose gel and the quantity was measured with a nanodrop spectrophotometer at a wavelength of 260 nm (Nanovue, GE).
Development of ISSR Fingerprinting Method
The ISSR primers were custom synthesized from IDT, India. Initially, 25 primers were screened using five A. solani DNA samples. Finally, primers which produced reproducible and consistent profiles (11 ISSR) were selected for profiling all DNA samples (Table 2).
ISSR-PCR amplification was carried out in 25 μl reaction mixture in 200 μl PCR plates. Each reaction mixture contained 50 ng genomic DNA, 200 μM of each dNTPs, 0.5 unit of Taq polymerase, 1 X Taq polymerase buffer solution and 0.2 μM of primer. Amplifications were performed in a thermal cycler (Bio Rad, USA) programmed for an initial denaturation of 4 min at 94 °C, 45 cycles of denaturation at 94 °C for 1 min, annealing temperature specific for each primer for 1 min and extension at 72 °C for 2 min, followed by a final extension at 72 °C for 5 min. The PCR products were separated on 2% agarose gel in 1X TAE buffer at 65–70 V for 3–4 h. DNA fragments were visualized under UV light and documented using a Gel Documentation System (BioDoc-It 220 Imaging System, UK). All PCR amplifications were performed at least twice for each isolate to assure reproducibility.
The ISSR data for genotyping was assembled into a binary matrix by scoring unambiguous polymorphic bands manually. Presence of a band was denoted as “1”, and absence was marked as “0”. The dendrogram was constructed using the neighbour-joining (NJ) method through the software program Darwin 6.0 (Perrier & Jacquemoud-Collet 2006, https://darwin.cirad.fr/darwin).
Results and Discussion
Radial Growth and Color
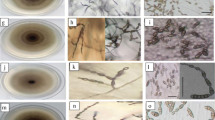
Radial growth observed for isolates presented in Table 3 was significantly not different for most of the isolates, with the lowest radial growth for 00113 (at 5th day after inoculation) and the highest radial growth for 00111 (at 10th day after inoculation). However, considerable variation was observed among the isolates in terms of colour (Table 3, Fig. 1).
Pathogenicity
Inoculation of detached tomato (susceptible line CO-3) leaves with A. solani isolates resulted in clearly defined necrosis for most of the isolates. Observations were noted on the 7th day after inoculation (Fig. 2). Eight isolates could not cause any necrotic lesion on the leaves and were categorized as nonpathogenic isolates. Sixteen isolates caused necrosis in the leaves around the lesion and they were rated as 1. Nine isolates were given the rating 2, as the necrosis caused by these covered ½ or ¾ area on the leaves surrounding the lesion. The results of the pathogenicity tests have been summarized in Table 3 and Fig. 2.
Scoring for virulence as 0 and 2 [18]
Biochemical Studies
Severe damage to plant tissue is a measure of fungal action for pathogenicity which results in disease production in the host [22]. The first line of defense in plants is the cell wall and it is composed of pectin (constituent of middle lamella and primary cell wall) and other compounds. Fungus has the ability to produce polygalacturonase (PG) and pectin methyl esterase (PME) enzymes that hydrolyze pectin [23]. It has been observed that pectic enzymes are sufficient to cause cell death and create huge losses to the plant [24]. Figure 3 shows the value of specific activity of polygalacturonase, which ranges from 0.012 (for isolate 2655) to 0.111 uM/ml (for isolate 2666). All the isolates with their respective PG activity values are shown in Table 4 and Fig. 3.
Another enzyme, pectin degrading pectin methylesterase (PME) was estimated for its specific activity in these isolates and it ranged from 0.63 (for 2665) to 3.54 Units/ml (for 0139). It is secreted in combination with pectin-degrading enzyme, polygalacturonase, and together, decays the plant cell wall to establish infection and absorb nutrients from the host [25]. The specific activity of PME is represented in Table 4 and the bar graph (Fig. 4).
Genetic Diversity Study Using ISSR Primers
Totally, twenty-five primers were selected for the present study and among these, only 11 primers gave amplification in all the isolates. Of these, eleven tested primers with total 56 alleles were amplified by PCR. Each of these primers generated 1–7 scorable bands, ranging from 200 to 1500 bp.
The dendrogram (Fig. 5) generated by DARWIN clearly showed two distinct groups of the isolates. The largest group, “Group 1”, comprised of nineteen isolates and “Group II” consisted of 13 isolates. However, a single isolate, 2665, could not be clustered into the above mentioned main clusters. Both the groups have isolates from diverse geographical distribution and with different virulence ability. Group 1 is divided into two subgroups that have 11 and 8 isolates. Five isolates in the subgroup showed no pathogenicity, while four showed high virulence. Two isolates were moderately virulent. This subgroup has four members from the hill collection i.e. 2664, 2663, 2669 and 2659. Two isolates in this SG are from Rajasthan, two from Karnataka and three belong to Uttar Pradesh, Andhra Pradesh and West Bengal, respectively.
The second subgroup comprised of eight isolates with moderate virulence, except 0112, that showed more virulence than other isolates in this group. Four isolates in this SG II have been isolated from Karnataka, two from Uttar Pradesh, one from New Delhi and one from Jammu.
The second group consisted of thirteen isolates among which four isolates showed more virulence and six revealed comparatively low virulence. Three isolates did not show any virulence in the present study. This group consisted of more diverse isolates; four from Himachal Pradesh, three from Uttar Pradesh, two from Karnataka, one from New Delhi, one from Chhattisgarh and one from Haryana. The variance in the pathogenicity levels was observed in the dendrogram that consisted of six isolates with lesser virulence and four isolates with more virulence. Three isolates in this group did not produce any symptoms during the detached leaf study. Both the analysis (dendrogram and PCO) revealed that the different isolates possessed diversity among themselves. The grouping among these isolates shows that ISSR primers can reveal diversity in same geographical regions too.
Discussion
Morphological, biochemical and genetic diversity existing in the pathogen population facilitates our understanding for potential appearance of new variants or pathotypes. The study has shown that there is substantial genetic diversity among the isolates. Alternaria solani, an asexually propagating fungus, is highly variable at a genetic level [26]. Cultural practices, environmental conditions prevailing in a particular area and plant materials also have contributed to the genetic diversity observed within geographic regions [27]. Several studies have been performed for evaluation of variability in Alternaria spp. and its correlation with disease incidence in the laboratory and field condition.
In the present study, simple pigmentation, growth characteristics, detached leaf inoculation with an accordant external condition; especially temperature and moisture, specific activity assay of pectin degrading enzymes along with ISSR DNA markers were used to understand the genetic diversity among the Alternaria solani isolates belonging to different states of India. The virulence of isolates was scored on the area covered on the leaf after 72 h of inoculation. The lesions caused by the strains varied in size indicate the level of virulence the selected isolates possessed, suggesting that there was significant differentiation in pathogenicity among the isolates of A. solani on the used susceptible cultivar of tomato i.e. CO-3. This provides experimental evidence and useful reference data for the reasonable use of disease-resistant varieties of tomato for disease control. Necrotrophic fungus creates a complex physiological system in the plant for their establishment that includes conidial attachment, germination, host penetration, lesion formation and expansion, and tissue maceration followed by sporulation [28]. The plant cell wall is composed of cellulose and hemicellulose in a strengthening network of cohesive pectin matrix cross-linked to lignin and proteins by ionic and covalent bonds. This is the first line of defense for pathogens, bacteria and insects.
In the present study, the authors also tried to corroborate the virulence of isolates with the production of cell wall degrading enzymes i.e. PGs and PMEs. It was found that these enzymes might have a role in the penetration of the fungal isolates in the leaf but it has been observed that many of the isolates which have high activity of these two have lesser virulence or no virulence as compared to the others. The correlation values were calculated for pathogenicity, PG and PME values, and the calculated results have shown a positive correlation between enzyme activity and pathogenicity. However, it may be conferred from the present study that not much correlation was found between the ability to produce the enzymes, in vitro, and pathogenicity. The absence/lesser ability to cause a lesion on the tomato leaf suggests that these enzymes are not concerned with the production of moderate disease symptoms in the tomato plant. The variation in pathogenic potential of sampled isolates ranged from no virulence (0) to moderately virulent (2), suggesting the existence of mild pathogenic variability and also that these have no relation with either geography or their in vitro production of polygalacturonase or pectin methyl esterase activity. The isolates might have expressed pathogenicity and symptoms in field conditions, but this study was performed under in vitro conditions to observe their virulence on the detached leaf which is quite different from its natural state. In this study, it has been observed that the isolates with high activity of pectin degrading enzymes i.e. PG and PME did not produce disease/symptoms during detached leaf assay e.g. 0139, 2408 have highest PME activity (3.54 and 3.13 Unit/ml) with 0.4 and 0.5 μmol/ml specific activity of PG, respectively. Also, the isolates that showed more virulence did not possess high activity of these two enzymes. This could be understood by a recent study performed in alfalfa [29] that showed the association of disease resistance in the plant with the expression of polygalacturonase-inhibiting proteins. A similar observation can be made in this study, where the isolates showed higher pectin degrading enzyme activity but did not produce any disease symptoms during detached leaf assay. An et al. [30] reported that an inhibitor protein CaPMEI1 is required for antifungal activity, basal disease resistance and abiotic stress tolerance. A possible reason for this could be that the machinery for producing disease in live and natural conditions is accompanied with associated biochemical and physiological changes inside the plant. Dendrogram constructed on the basis of ISSR markers did not show grouping of selected isolates. Grouping of the isolates are not in geographical congruency. The isolates that grouped together differed in their morphology, pathogenicity and at biochemical levels. However, subgroup 3 of group III has members with pathogenicity 1. Since A. solani is an asexually propagated fungus, genetic mechanisms that could explain such diversity mainly include simple mutations due to external effects. The significant amount of diversity among Indian isolates of A. solani can be explained mainly by evolution; resulting from natural and stress induced transposition. These genetic diversity studies can be useful for plant breeding programs for disease resistance as the genetic structure of pathogens reflects the history and the evolutionary potential of a pathogen.
References
Gwary DM, Nahunnaro H (1998) Epiphytotics of early blight of tomatoes in northeastern Nigeria. Crop Prot 17:619–624
Milgroom MG, Peever TL (2003) Population biology of plant pathogens: the synthesis of plant disease epidemiology and population genetics. Plant Dis 87:608–617
Bateman DF, Basham HG (1976) Degradation of plant cell walls and membranes by microbial enzymes. In: Heitefuss R, Williams PH (eds) Physiological plant pathology. Springer, Berlin, pp 316–355
Walton JD (1994) Deconstructing the cell wall. Plant Physiol 104:1113–1118
Holtz G, Knox-Davies PS (1985) Production of pectic enzymes by Fusarium oxysporum f. sp. cepae and its involvement in onion bulb rot. J Phytopathol 112:69–80
Baayen RP, van Dreven F, Krijger MC, Wallwijk C (1997) Genetic diversity in Fusarium oxysporum f. sp. dianthi and Fusarium redolens f. sp. dianthi. Eur J Plant Pathol 103:395–408
Lathar PK, Sharma A, Thakur I (2010) Isolation and random amplified polymorphic DNA (RAPD) analysis of wild yeast species from 17 different fruits. J Yeast Fungal Res 18:146–151
Fegan M, Manners JM, Maclean DJ, Irwin JA, Samuels KD, Holdom DG, Li DP (1993) Random amplified polymorphic DNA markers reveal a high degree of genetic diversity in the entomopathogenic fungus Metarhizium anisopliae var. anisopliae. J Gen Microbiol 139:2075–2081
Kim Y, Choi SJ, Choi C (2017) An Efficient PCR-RFLP Method for the rapid identification of Korean Pyropia Species. Molecules. https://doi.org/10.3390/molecules22122182
Nath VS, Senthil M, Hegde VM, Jeeva ML, Misra RS, Veena SS, Raj M (2013) Genetic diversity of Phytophthora colocasiae isolates in India based on AFLP analysis. 3. Biotechnology 3(4):297–305. https://doi.org/10.1007/s13205-012-0101-55
Velez P, Quintero CA, Merino G, Gasca-Pineda J, González MC (2016) An ISSR-based approach to assess genetic diversity in the marine arenicolous fungus Corollospora maritima sensu lato. Mycoscience 57(3):187–195
Del-Prado R, Cubas P, Lumbsch HT, Divakar PK, Blanco O, de Paz GA, Molina MC, Crespo A (2010) Genetic distances within and among species in monophyletic lineages of Parmeliaceae (Ascomycota) as a tool for taxon delimitation. Mol Phylogenet Evol 56(1):125–133
Fell JW, Boekhout T, Fonseca A, Scorzetti G, Statzell-Tallman A (2000) Biodiversity and systematics of basidiomycetous yeasts as determined by large-subunit rDNA D1/D2 domain sequence analysis. Int J Syst Evol Microbiol 50(3):1351–1371
Vialle A, Feau N, Allaire M, Didukh M, Martin F, Moncalvo JM, Hamelin RC (2009) Evaluation of mitochondrial genes as DNA barcode for Basidiomycota. Mol Ecol Resour 9:99–113
Bagherabadi S, Zafari D, Soleimani MJ (2015) Genetic diversity of Alternaria alternata isolates causing potato brown leaf spot using ISSR markers in Iran. J Plant Pathol Microb 6:286. https://doi.org/10.4172/2157-7471.1000286
Troncoso-Rojas R, Báez-Flores María Elena, Pryor Barry, Garcíaand Hugo S, Tiznado-Hernández Martín-Ernesto (2013) Inter simple sequence repeat polymorphism in Alternaria genomic DNA exposed to lethal concentrations of Isothiocyanates. Afr J Microbil Res 7(10):838–852
Upadhyay P, Singh PC, Sinha B, Singh M, Kumar R (2009) Sources of resistance against early blight (Alternaria solani) in tomato (Solanum lycopersicum). Ind J Agric Sci 79:752–753
Weber B, Halterman DA (2012) Analysis of genetic and pathogenic variation of Alternaria solani from a potato production region. Eur J Plant Pathol 134:847–858
Kumari S, Tayal P, Sharma E, Kapoor R (2014) Analyses of genetic and pathogenic variability among Botrytis cinerea isolates Microbio Res 169(11):862–872
Balaban MO, Arreola AG, Marshall M, Peplow A, Wei CI, Cornel J (1991) Inactivation of pectinesterase in orange juice by supercritical carbon dioxide. J Food Sci 56:743–746
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Casadevall A, Pirofski L-A (1999) Host–pathogen interactions: redefining the basic concepts of virulence and pathogenicity. Infect Immun 67:3703–3713
Cervone F, De Lorenzo G, Salvi G, Camardella L (1986). Molecular evolution of fungal polygalacturonase. In: Bailey J (ed) Biology and molecular biology of plant–pathogen interactions. NATOASI series, vol H1. Springer, Berlin, pp 385–392
Brett C, Waldron K (1990) Physiology and biochemistry of plant cell walls. In: Black M, Chapman J (eds) Topics in plant physiology. Unwin Hyman, London, pp 6–57
de Vries RP, Visser J (2001) Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol Mol Biol Rev 65:497–522
Martinez SP, Snowdon R, Pons-Kuhnemann J (2004) Variability of Cuban and international populations of Alternaria solani from different hosts and localities: AFLP genetic analysis. Eur J Plant Pathol 110(4):399–409
O’Donnell K, Kistler HC, Cigelnik E, Ploetz RC (1998) Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies. Proc Natl Acad Sci USA 95:2044–2449
Prins TW, Tudzynski P, Von Tiedemann A, Tudzynski B, ten Have A, Hansen ME (2000) Infection strategies of Botrytis cinerea and related necrotrophic pathogens. In: Kronstad J (ed) Fungal pathology. Kluwer, Dordrecht, pp 33–64. https://doi.org/10.1007/978-94-015-9546-9_2
Gui Z, Gao J, Xin N, Wang Y, Yongshuo P, Liu H, Yuan Q, Li X (2016) Association of polygalacturonase-inhibiting protein gene 2 (MsPGIP2) to common leaf spot resistance in alfalfa. Eur J Plant Pathol 144:245–256
An SH, Sohn KH, Choi HW, Hwang IS, Lee SC, Hwang BK (2008) Pepper pectin methylesterase inhibitor protein CaPMEI1 is required for antifungal activity, basal disease resistance and abiotic stress tolerance. Planta 228(1):61–78
Acknowledgements
The authors are thankful to University Grant Commission, New Delhi, India for providing Post-Doctoral [Fellowship F.15-1/2013-14/PDFWM-2013-14-GE-UTT-20193 (SA-II)] and Department of Botany, University of Delhi, Delhi for providing laboratory facilities.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest in this publication.
Additional information
Significance statement
Diversity assessment among A. solani isolates from different states of India will provide valuable aid for resistance breeding strategies and basis for planning disease protection for sustainable agriculture.
Rights and permissions
About this article
Cite this article
Upadhyay, P., Ganaie, S.H. & Singh, N. Diversity Assessment Among Alternaria solani Isolates Causing Early Blight of Tomato in India. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 89, 987–997 (2019). https://doi.org/10.1007/s40011-018-1017-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-018-1017-6