Abstract
Objective
Due to the importance of Chronic obstructive pulmonary disease (COPD) as the fourth cause of mortality worldwide and the lack of studies evaluating the prevalence of bacterial infections in disease exacerbation, this systematic review and meta-analysis was performed to determine the prevalence rate of bacterial infections in COPD patients.
Methods
PubMed, ISI Web of Science, and Scopus databases were systematically searched for population-based prevalence studies (1980–2018). MeSH terms for “Bacterial infections” and “AECOPD” were used as search keywords. The selected studies were filtered according to the inclusion and exclusion criteria. Fixed and random-effects models were used for estimation of summary effect sizes. Between-study heterogeneity, as well as publication bias, were calculated.
Results
Finally, 118 out of 31,440 studies were selected. The overall estimation of the prevalence of bacterial infection was 49.59% [95% confidence interval (CI) 0.4418–0.55]. The heterogeneity in estimating the pooled prevalence of bacterial infections was shown in the studies (Cochran Q test: 6615, P < 0.0001, I2 = 98.23%). In addition, S. pneumoniae, H. influenzae, M. catarrhalis, A. baumannii, P. aeruginosa, and S. aureus were the most prevalent reported bacteria.
Conclusions
Our results as the first meta-analysis for the issue demonstrated that bacterial infections are an important risk factor for AECOPD. Further studies must be performed for understanding the exact role of bacterial agents in AECOPD and help physicians for more applicable preventive and therapeutic measurements.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction and objectives
Chronic Obstructive Pulmonary Disease (COPD) is a prevalent type of obstructive lung disease involved ~ 174 million people worldwide [132]. The main clinical symptoms of COPD include cough, shortness of breath, and sputum production [19], with periods of acute exacerbations (AECOPD) that characterizes by increased cough, more shortness of breath, sputum color changing, and increase in sputum production [11, 19]. Among several risk factors suggested to play role in COPD/AECOPD pathogenesis, respiratory bacterial infections are correlated with approximately 50% of cases [11], as studies have demonstrated a significant correlation between the lower airway bacterial load and COPD consequences [86, 118]. Accordingly, many experimental and review studies have reported a wide range of bacterial species involved in COPD/AECOPD patients, among which respiratory microbiome is a key player [28, 104]. Due to demographic and geographical variations, different clinical specimens, and various detection methods (including molecular methods such as PCR and RT-PCR and serology), different bacterial species have been isolated from the patients. Since there are few studies tried to pool and meta-analyze these heterogeneous studies altogether, the prevalence and geographical distribution of respiratory bacterial infection in AECOPD has not yet been clearly defined. In the previous study, a systematic review and meta-analysis was performed by the present authors on 28 studies selected out of 26,078 articles to determine the prevalence of viral infections in AECOPD, resulting in the about 43% overall viral prevalence rate [52]. To the best of our knowledge, there is no systematic review and meta-analysis on the prevalence of bacterial infections in AECOPD. Accordingly, in the present study, we aimed to determine the frequency of bacterial infections in COPD/AECOPD patients through a systematic review and meta-analysis study.
Data source and study selection
Search strategy
A systematic search was performed in the main databases, including PubMed, Scopus, and ISI Web of Science to identify available articles to May 2018. According to MeSH terms, searches were performed using the following keywords: “chronic obstructive pulmonary disease”, “COPD”, “Exacerbation”, “infection”, “microbe”, “bacteria”, and “colonization”, alone or combined together with the Boolean operators “OR”, “AND”, and “NOT” in the Keywords/Title/Abstract fields. In addition, the reference list of selected full-text papers was precisely searched manually to find additional citations not retrieved in the first step of the systematic search. Gray literature, dissertations, and relevant proceedings of international congresses were not explored. Finally, we restricted our search to the original articles or abstracts published which reported the prevalence of bacterial infections in COPD patients. The literature search was conducted by two independent researchers in two stages. Disagreements among researchers were resolved by discussion or, if necessary, by a third researcher. Journals and authors were not blinded during study selection.
Inclusion and exclusion criteria
A protocol for inclusion and exclusion criteria was defined for eligible peer-reviewed publications according to the following inclusion criteria: (A) Articles published up to May 2018, (B) The articles in English language reporting the prevalence of bacterial infections in COPD patients, (C) All studies included samples from sputum, nasopharyngeal swab, bronchoalveolar lavage, brushing, and nasal lavage, and (D) reported data related to a group of individuals taken from the general population. The main exclusion criteria were: (A) Studies with unknown sample origins, (B) Studies that failed to present data clearly, (C) Studies conducted on animal models, (D) Studies with overlapping subjects, time, and place of sample collection, and (F) Congress abstracts, review articles, case report articles, meta-analysis or systematic reviews, and duplicate publication of the same study.
Quality assessment and data extraction
The preferred reporting items for systematic reviews and meta-analysis (PRISMA) guidelines were used to assess the quality of the included studies by two researchers, independently. The PRISMA statement consists of a 27-item checklist and four-phase flow diagram [75]. A complete information list was extracted from the articles into a Microsoft Excel worksheet. These data were included the first author’s name, publication date, sample size, the prevalence of bacterial infection, Age mean, detection method, biological sample, smoking status, risk factors, and reference. Furthermore, unclear data were consulted and achieved consensus before recording an entry in the dataset. Cohen’s κ was acceptable between the researchers as the agreement coefficient and was considered equal to 0.80.
Statistical methods
Pooled relative frequency (RF) and its corresponding 95% CI were used to evaluate the prevalence of bacterial infections in COPD. The heterogeneity and the variation in the pooled estimations were assessed using Cochran’s Q test and I2 index, respectively, and was considered significant at P < 0.05 level [71]. The pooled RF was calculated by a random effect model while significantly heterogeneity existed between the individual studies, and otherwise, this pooled effect sizes were derived from a fixed-effect model. A funnel plot was established for checking the existence of publication bias. The funnel plot asymmetry was measured by Egger’s linear regression test and Begg’s test (P < 0.05 levels were considered statistically significant for publication bias) [51]. Finally, the subgroup analysis was used in the year of publication, age average, biological sample, detection method, smoking status, and risk factors. All statistical analyses were conducted by data analysis and statistical software (STATA) (version 11.0; Stata Corporation, College Station, TX) and MedCalc software.
Results
Literature review
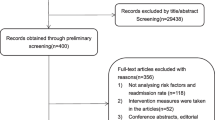
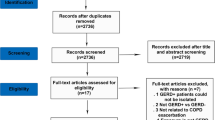
The study selection process and the flowchart of the literature search are shown in Fig. 1. Through our search in PubMed, Scopus, and ISI web of science database and references, 31,440 articles were identified for “Exacerbation” in the title or abstract. In primary screening, 29,679 publications were excluded according to COPD Mesh terms in the title or abstract (COPD, chronic obstructive pulmonary disease). The retained publications were screened according to “Infection” Mesh terms, including “Infect*” OR “Microb*” OR “Bacteria” OR “Bacterium” OR “Colonization” in the title and abstract that resulted in 868 publications. Then, after manually assessing for bacterial infections in the title and abstract and eligibility evaluation, finally, 118 papers were retained for full-text evaluation. Two independent researchers did the literature search in two stages. Disagreements among researchers were resolved by discussion or, if necessary, by a third researcher. Journals and authors were not blinded during study selection.
Study characteristics
Out of all studies entered into the meta-analysis, 32 studies (27%) were published before 2005 and others (73%) were after 2005. Most studies were conducted in Spain (14.4%) and only less than 1% were published in England and Japan (Table 1).
Overall prevalence
According to included studies, the pooled estimation for the prevalence of bacterial infection in COPD patients was 0.4959 (95% CI 0.4418–0.55). Total patients analyzed for the pooled prevalence of bacterial infection were 19,409 among which, 8447 cases were positive for bacterial infections. The most common isolated pathogens were including H. influenzae, S. pneumoniae, Klebsiella pneumoniae, S. aureus, M. catarrhalis, A. baumannii, and P. aeruginosa. The heterogeneity for estimating the pooled prevalence among the studies was shown; Cochran Q test: 6615, P < 0.0001, I2 = 98.23% (Fig. 2 and Table 2).
Subgroup analysis
As we analyzed for the time trend of bacterial infections in AECOPD, the LOWESS Smoother plot showed a significant shift in bacterial infections prevalence in 2005. In fact, subsequent to a decreasing rate of bacterial prevalence in AECOPD studies, an increasing shift is seen after 2005 and continues almost steadily. Therefore, it seems that bacterial infection was noted to be more prevalent in studies published after 2005 (0.4939; 95% CI 0.4281–0.5598). As shown in Fig. 3, culture is constantly used in most studies but a dramatic shift was seen in applying PCR-based methods for detecting bacterial infections in AECOPD. More than 60 years old was the most common group of patients (0.5114; 95% CI 0.446–0.5767, P < 0.001). Sputum was the most used clinical specimen (82.2%) (0.4943; 95% CI 0.4328–0.5558, P < 0.001) and culture was the most common detection method (83.8%) (0.4956; 95% CI 0.4375–0.5538, P < 0.001). More information on Subgroup analysis is presented in Table 2. In addition, the prevalence of bacterial infection in COPD patients increased over age and decreased in men, approximately (Fig. 4).
Publication bias
Although publication bias was statistically significant in some cases based on Egger’s regression test, no publication bias was detected according to the Begg’s adjusted rank correlation test (Table 2 and Fig. 5).
Discussion
To the best of our knowledge, this is the first systematic review and meta-analysis about the prevalence of bacterial infections in AECOPD patients, showing that the prevalence of bacterial infections is 49.5% (95% CI 0.4418–0.55). Although about 20% of the AECOPD cases may be a co-infection of simultaneous bacterial and viral infections, according to the principal objective of our study, we determined only the role of bacterial infections in AECOPD patients. The majority of the most common microorganisms, including S. pneumoniae, H. influenza, and M. catarrhalis, were a part of the respiratory microbiome. It seems reasonable, since microbiome dysbiosis is a major cause of chronic respiratory complications that can disturb homeostasis in the lung resulting in lung inflammation and infection [104]. Sethi et al. isolated bacterial agents from sputum samples of 40–60% of COPD patients [107]. A study in 2003 identified bacterial infections in 43.3% of AECOPD patients as the major risk factor with the dominance of H. influenzae (13.3%) [76]. Several other studies have isolated H. influenzae, Streptococcus pneumoniae, Moraxella catarrhalis, and Pseudomonas as the most common bacteria from both stable and exacerbated COPD [11, 37, 72]. In most studies, the average age of patients was above 60 years. Previous studies have shown that young age groups of COPD patients may be more likely to show worse clinical outcomes [48]. Another influencing factor is the bacterial detection method. As shown in the subgroup analysis for the year of publication, after a decreasing rate of bacterial prevalence in AECOPD studies, an increasing shift is seen after 2005. This shift may be due to three possible reasons: first, molecular techniques (with higher sensitivity and being independent of previous antibiotic administration) are mainly introduced almost after 2005; second, bacterial culture systems have been progressed more frequently; and third, it is simply due to the slight changes in the prevalence of bacterial caused AECOPD before and after 2005. In this meta-analysis, the most common detection method was the culture (83.8%) (0.4956; 95% CI 0.4375–0.5538, P < 0.001). For bacterial detection, culture is the gold standard method, but has some shortcomings such as time-consuming and low sensitivity [38, 61]. Despite being expensive and more complicated, molecular techniques have several advantages including higher sensitivity, being able to detect noncultivable bacteria, being independent of previous antibiotic administration, and can quantitatively determine the load of bacteria [38]. In addition, since it is proved that changing in the total pattern of microbiome content is the cause of asthma and COPD rather than changing in an individual bacterial species, next-generation sequencing techniques would be needed to give a comparison between healthy and patients respiratory microbiome. We found ELISA as the third most common method in 21 studies (0.5099; 95% CI 0.3767–0.6423, P < 0.001), as a fast but low specific method. In fact, sandwich and indirect ELISA methods used to evaluate microbial antigens and/or antibodies are indirectly detected the pathogens using nonspecific polyclonal antibodies [110]. As an important note regarding the serology assay studies, although serology could indicate passive infections and not necessarily active infection, these studies using serology assays (mostly for detecting M. pneumoniae, M. catarrhalis, and Chlamydophila pneumonia) evaluated current active bacterial infections in AECOPD patients. Regarding the advantages and disadvantages of both culture and molecular methods, it seems that the combination of these two methods may give more reliable results [61]. Of note, three studies used microarray as the high-throughput technique for bacterial detection. Due to the high capacity of the microarray to specifically identify a wide range of microbial species as well as their different characteristics, using microarray developed for simultaneous detection of many bacterial species and strains would be of great attention in the future. A mechanism related to bacterial infection is the bacterial load. In COPD exacerbations, the bacterial load increases with a significant decrease in microbiome complexity [11, 37]. In one study, Sethi et al. demonstrated that the bacterial load of H. haemolyticus, M. catarrhalis, and H. influenzae in the sputum of AECOPD patients was significantly different from stable patients. In addition, they indicated a significant negative correlation between the sputum bacterial load for H. parainfluenzae and S. pneumoniae and the exacerbation occurrence, whereas H. influenzae was detected in high concentrations in AECOPD patients [112]. In terms of clinical specimens, sputum was the most prevalent sample in this meta-analysis. Although sputum is not an ideal representative for lower respiratory tract microflora (due to the possible contamination by oral flora), the sampling simplicity and feasibility have introduced sputum as the most prevalent lower respiratory sample. In addition, bronchoalveolar lavage and protected specimen brush sampling may be applicable only for the milder severity states of AECOPD but not for a more severe disease state. Although the application of PCR or real-time PCR methods in sputa could correctly detect oral flora, it would be unable to ascertain if these are commensals or have converted into a pathogenic state. One of the main factors in COPD pathogenesis is airway inflammation leading to progressive airway injuries, though producing inflammatory mediators such as cytokines by T cells, macrophages, and neutrophils [11, 57]. Studies indicated that bacterial infection can trigger airway immune system that resulted in inflammation [45]. A study showed that the neutrophilic inflammation (including neutrophil elastase, TNF-α, and IL-8) in COPD-mediated bacterial exacerbation was higher than nonbacterial exacerbation [111]. Neutrophil migration into airways may occur due to bacterial products. The elastase produced by neutrophils can act synergistically with bacterial products to inhibit tracheobronchial ciliary function [11, 110]. After inflammation, another risk factor considered here was diabetes. This disease in individuals with COPD more occurs than in healthy persons. The association between COPD and diabetes is varied and different studies known that diabetes affects 2–37% of COPD patients [102].
Conclusion
The current study provides the first overall bacterial infection prevalence in AECOPD patients worldwide and information about bacterial species in different geographical areas. It seems that besides other risk factors such as viral infections and environmental conditions, bacterial infections caused mainly by the dysbiotic respiratory microbiome may consider as the major risk factor of AECOPD for which more accurate and applicable detection and therapeutic methods. Further studies with more developed methods must be performed for understanding the exact role of bacterial agents in AECOPD.
References
Adlowitz DG, et al. Human serum and mucosal antibody responses to outer membrane protein G1b of Moraxella catarrhalis in chronic obstructive pulmonary disease. FEMS Immunol Med Microbiol. 2005;46:139–46.
Agmy G, et al. Bacterial profile, antibiotic sensitivity and resistance of lower respiratory tract infections in upper Egypt. Mediterr J Hematol Infect Dis. 2013;5:1.
Alamoudi OS. Bacterial infection and risk factors in outpatients with acute exacerbation of chronic obstructive pulmonary disease: a 2-year prospective study. Respirology. 2007;12:283–7.
Aleemullah M, et al. Bacteriological profile of patients with AECOPD-hospital based study. Int J Curr Microbiol App Sci. 2016;5:84–90.
Aydemir Y, et al. Relationship between the GOLD combined COPD assessment staging system and bacterial isolation. Int J Chronic Obstruct Pulm Dis. 2014;27:1045–1051.
Bafadhel M, et al. Airway bacteria measured by quantitative polymerase chain reaction and culture in patients with stable COPD: relationship with neutrophilic airway inflammation, exacerbation frequency, and lung function. Int J Chronic Obstruct Pulm Dis. 2015;10:1075.
Bafadhel M, et al. Acute exacerbations of chronic obstructive pulmonary disease: identification of biologic clusters and their biomarkers. Am J Respir Crit Care Med. 2011;184:662–71.
Bari M, et al. Microbes responsible for acute exacerbation of COPD. Mymensingh Med J MMJ. 2010;19:576–85.
Barker BL, et al. Association between pathogens detected using quantitative polymerase chain reaction with airway inflammation in COPD at stable state and exacerbations. Chest. 2015;147:46–55.
Bathoorn E, et al. Change in inflammation in out-patient COPD patients from stable phase to a subsequent exacerbation. Int J Chronic Obstruct Pulm Dis. 2009;4:101.
Beasley V, et al. Lung microbiology and exacerbations in COPD. Int J Chronic Obstruct Pulm Dis. 2012;7:555.
Biagini M, Rossi M. Bacterial pathogens in sputum and degree of bronchial obstruction in patients with acute exacerbation of chronic obstructive pulmonary disease. Recenti Prog Med. 2002;93:470–3.
Blasi F, et al. Chlamydia pneumoniae infection in acute exacerbations of COPD. Eur Respir J. 1993;6:19–22.
Boixeda R, et al. Bacterial flora in the sputum and comorbidity in patients with acute exacerbations of COPD. Int J Chronic Obstruct Pulm Dis. 2015;10:2581.
Boixeda R, et al. Pneumonia as comorbidity in chronic obstructive pulmonary disease (COPD). Differences between acute exacerbation of COPD and pneumonia in patients with COPD. Archivos de Bronconeumología (English Edition). 2014;50:514–20.
Braeken D, et al. Bacterial aetiology and mortality in COPD patients with CAP: results from the German Competence Network, CAPNETZ. Int J Tuberc Lung Dis. 2017;21:236–43.
Cabello H, et al. Bacterial colonization of distal airways in healthy subjects and chronic lung disease: a bronchoscopic study. Eur Respir J. 1997;10:1137–44.
Cazzola M, et al. Bronchial hyperresponsiveness and bacterial respiratory infections. Clin Ther. 1991;13:157–71.
Celli B, Barnes P. Exacerbations of chronic obstructive pulmonary disease. Eur Respir J. 2007;29:1224–38.
Chang C, et al. The changes and clinical implications of serum procalcitonin in acute exacerbations of chronic obstructive pulmonary disease. Chin J Tuberc Respir Dis. 2006;29:444–7.
Chang C, et al. Value of serum procalcitonin in diagnosing bacterial lower respiratory tract infections in people with exacerbation of chronic obstructive pulmonary disease. J Peking Univ Health Sci. 2006;38:389–92.
Chang C, et al. Bacterial infection, airway and systemic inflammation and clinical outcomes before and after treatment of AECOPD, a longitudinal and cross-sectional study. COPD J Chronic Obstruct Pulm Dis. 2015;12:19–30.
Char A, et al. Evidence of mycobacterial disease in COPD patients with lung volume reduction surgery; the importance of histological assessment of specimens: a cohort study. BMC Pulm Med. 2014;14:124.
Chen L, et al. Composition of pathogenic bacteria of chronic obstructive pulmonary disease patients and drug resistance of gram-negative bacilli for various antibiotics. Biomed Res. 2018;29:827–9.
Chin C, et al. Haemophilus influenzae from COPD patients with exacerbations induce more inflammation than colonizers. Am J Respir Crit Care Med. 2005;172:85–91.
Christol TJ, et al. Antibiogram pattern of Moraxella catarrhalis isolates in acute exacerbation chronic obstructive pulmonary disease. Chemotherapy. 2011;57:94–6.
Cukic V. The most common detected bacteria in sputum of patients with the acute exacerbation of COPD. Materia socio-medica. 2013;25:226.
D’Anna SE, et al. Bacterial–viral load and the immune response in stable and exacerbated COPD: significance and therapeutic prospects. Int J Chronic Obstruct Pulm Dis. 2016;11:445.
Dai M-Y, et al. Respiratory infectious phenotypes in acute exacerbation of COPD: an aid to length of stay and COPD Assessment Test. Int J Chronic Obstruct Pulm Dis. 2015;10:2257.
Desai H, et al. Bacterial colonization increases daily symptoms in patients with chronic obstructive pulmonary disease. Ann Am Thorac Soc. 2014;11:303–9.
Diederen BM, et al. The role of atypical respiratory pathogens in exacerbations of chronic obstructive pulmonary disease. Eur Respir J. 2007;30:240–4.
Domenech A, et al. Some pneumococcal serotypes are more frequently associated with relapses of acute exacerbations in COPD patients. PLoS one. 2013;8:e59027.
Domenech A, et al. Infectious etiology of acute exacerbations in severe COPD patients. J Infect. 2013;67:516–23.
Duan Z, et al. Chlamydia pneumoniae infection in patients with chronic obstructive pulmonary disease. Chin J Tuberc Respir Dis. 2001;24:208–11.
ElFeky DS, et al. Sputum bacteriology in patients with acute exacerbation of chronic obstructive pulmonary disease. Int J Curr Microbiol App Sci. 2016;5:289–305.
Eller J, et al. Infective exacerbations of chronic bronchitis: relation between bacteriologic etiology and lung function. Chest. 1998;113:1542–8.
Erkan L, et al. Role of bacteria in acute exacerbations of chronic obstructive pulmonary disease. Int J Chronic Obstruct Pulm Dis. 2008;3:463.
Fournier P-E, et al. Clinical detection and characterization of bacterial pathogens in the genomics era. Genome Med. 2014;6:114.
Fruchter O, et al. Airway bacterial colonization and serum C-reactive protein are associated with chronic obstructive pulmonary disease exacerbation following bronchoscopic lung volume reduction. Clin Respir J. 2016;10:239–45.
Furqan S, Paracha SAU. Frequency of Streptococcus pneumonia and Haemophilus influenza in acute exacerbation of chronic obstructive airway disease and their sensitivity to levofloxacin. Age (years). 2014;50:28.
Gallego M, et al. Pseudomonas aeruginosa isolates in severe chronic obstructive pulmonary disease: characterization and risk factors. BMC Pulm Med. 2014;14:103.
Garcha DS, et al. Changes in prevalence and load of airway bacteria using quantitative PCR in stable and exacerbated COPD. Thorax. 2012;67:1075–80.
Garcia-Nuñez M, et al. Bronchial microbiome, PA biofilm-forming capacity and exacerbation in severe COPD patients colonized by P. aeruginosa. Future Microbiol. 2017;12:379–92.
Geelen TH, et al. The host immune response contributes to Haemophilus influenzae virulence. Respir Med. 2014;108:144–52.
Groenewegen KH, Wouters EF. Bacterial infections in patients requiring admission for an acute exacerbation of COPD; a 1-year prospective study. Respir Med. 2003;97:770–7.
Guo Z, et al. Bacteriology in acute exacerbation in patients hospitalized frequently for acute exacerbation of chronic obstructive pulmonary disease. Zhonghua yi xue za zhi. 2014;94:729–32.
Hashemi SH, et al. High seroprevalence of Bordetella pertussis in patients with chronic obstructive pulmonary disease: a case-control study. Tanaffos. 2015;14:172.
Holm KE, et al. The impact of age on outcomes in chronic obstructive pulmonary disease differs by relationship status. J Behav Med. 2014;37:654–63.
Huang YJ, et al. A persistent and diverse airway microbiota present during chronic obstructive pulmonary disease exacerbations. OMICS. 2010;14:9–59.
Huang YJ, et al. Airway microbiome dynamics in exacerbations of chronic obstructive pulmonary disease. J Clin Microbiol. 2014;52:2813–23.
Huedo-Medina TB, et al. Assessing heterogeneity in meta-analysis: Q statistic or I2 index? Psychol Methods. 2006;11:193.
Jafarinejad H et al. Worldwide prevalence of viral infection in AECOPD patients: a meta-analysis. Microb Pathogen. 2017.
Jassem E, et al. Occurrence of non-spore forming anaerobic bacteria in the upper airways of patients with chronic obstructive pulmonary disease. Med Dosw Mikrobiol. 1996;48:49–54.
Karnak D, et al. Chlamydia pneumoniae infection and acute exacerbation of chronic obstructive pulmonary disease (COPD). Respir Med. 2001;95:811–6.
Keith ER, et al. Characteristics of Streptococcus pseudopneumoniae isolated from purulent sputum samples. J Clin Microbiol. 2006;44:923–7.
Kieszko R et al. Bacterial exacerbations of chronic obstructive pulmonary disease. The role of lung function in aetiology of exacerbation. Annales Universitatis Mariae Curie-Sklodowska. Sectio D: Medicina. 2003.
King PT, et al. Bacteria in COPD; their potential role and treatment. Transl Respir Med. 2013;1:13.
Kuwal A, et al. A Prospective study of bacteriological etiology in hospitalized acute exacerbation of COPD patients: relationship with lung function and respiratory failure. Turk Thorac J. 2018;19:19.
Lacoma A, et al. Value of procalcitonin, C-reactive protein, and neopterin in exacerbations of chronic obstructive pulmonary disease. Int J Chronic Obstruct Pulm Dis. 2011;6:157.
Larsen MV, et al. Bacteriology in acute exacerbation of chronic obstructive pulmonary disease in patients admitted to hospital. Scand J Infect Dis. 2009;41:26–32.
Laupland KB, Valiquette L. The changing culture of the microbiology laboratory. Can J Infect Dis Med Microbiol. 2013;24:125–8.
Lee HY, et al. Association between Helicobacter pylori seropositivity and mild to moderate COPD: clinical implications in an Asian country with a high prevalence of H. pylori. Int J Chronic Obstruct Pulm Dis. 2016;11:2055.
Leitao Filho F et al. Comparisons between sputum molecular pathogen detection methods and microbiome sequencing in COPD Exacerbations. In: C13. The MICROBIOME IN CHRONIC AIRWAYS DISEASE, American Thoracic Society. 2018; pp. A4430-A4430.
Lieberman D, et al. Infectious etiologies in acute exacerbation of COPD. Diagn Microbiol Infect Dis. 2001;40:95–102.
Lieberman D, et al. Serological evidence of Mycoplasma pneumoniae infection in acute exacerbation of COPD. Diagn Microbiol Infect Dis. 2002;44:1–6.
Lieberman D, et al. Serological evidence of Legionella species infection in acute exacerbation of COPD. Eur Respir J. 2002;19:392–7.
Lode H, et al. A prediction model for bacterial etiology in acute exacerbations of COPD. Infection. 2007;35:143.
Mantero M, et al. Role of Streptococcus pneumoniae infection in chronic obstructive pulmonary disease patients in Italy. Ther Adv Respir Dis. 2017;11:403–7.
Martínez-Solano L, et al. Chronic Pseudomonas aeruginosa infection in chronic obstructive pulmonary disease. Clin Infect Dis. 2008;47:1526–33.
Messous S, et al. Prevalence of Chlamydophila pneumoniae and Mycoplasma pneumoniae IgM and IgG antibodies in Tunisian patients presenting with exacerbation of chronic obstructive pulmonary disease. Medecine et maladies infectieuses. 2017;47:158–63.
Miller JJ. The inverse of the Freeman-Tukey double arcsine transformation. Am Stat. 1978;32:138.
Miravitlles M, Anzueto A. Chronic respiratory infection in patients with chronic obstructive pulmonary disease: what is the role of antibiotics? Int J Mol Sci. 2017;18:1344.
Miravitlles M, et al. Relationship between bacterial flora in sputum and functional impairment in patients with acute exacerbations of COPD. Chest. 1999;116:40–6.
Mogulkoc N, et al. Acute purulent exacerbation of chronic obstructive pulmonary disease and Chlamydia pneumoniae infection. Am J Respir Crit Care Med. 1999;160(1):349–53.
Moher D, et al. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst Rev. 2015;4:1.
Monso E, et al. Bacterial infection in exacerbated COPD with changes in sputum characteristics. Epidemiol Infect. 2003;131:799–804.
Montero M, et al. Mortality of COPD patients infected with multi-resistant Pseudomonas aeruginosa: a case and control study. Infection. 2009;37:16–9.
Murphy TF, et al. Pseudomonas aeruginosa in chronic obstructive pulmonary disease. Am J Respir Crit Care Med. 2008;177:853–60.
Nagai K, et al. Antimicrobial susceptibilities and serotypes of Streptococcus pneumoniae in southwestern Japan and correlation of penicillin-binding protein 2b and 2x mutations in susceptibilities of penicillin G and cefotaxime. Diagn Microbiol Infect Dis. 2000;37:107–13.
Nakou A, et al. A prospective study on bacterial and atypical etiology of acute exacerbation in chronic obstructive pulmonary disease. Future Microbiol. 2014;9(11):1251–60.
Naveed A et al. Sputum culture and antibiogram in infective acute exacerbation of chronic obstructive pulmonary disease in a tertiary care hospital in India. Indian J Chest Dis Allied Sci. 2018;60:13–18.
Navratilova L, et al. The Streptococcus milleri group in chronic obstructive pulmonary disease. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2016;160:378–84.
Noweta K, et al. Exacerbations of chronic obstructive pulmonary disease and the role of sputum bacteriological examination. Adv Respir Med. 2006;74:396–402.
Nseir S, et al. Factors predicting bacterial involvement in severe acute exacerbations of chronic obstructive pulmonary disease. Respiration. 2008;76:253–60.
Nseir S, et al. Multiple-drug–resistant bacteria in patients with severe acute exacerbation of chronic obstructive pulmonary disease: prevalence, risk factors, and outcome. Crit Care Med. 2006;34:2959–66.
O’donnell R, et al. Inflammatory cells in the airways in COPD. Thorax. 2006;61:448–54.
Papaetis G, et al. Serological evidence of Mycoplasma pneumoniae infection in patients with acute exacerbation of COPD: analysis of 100 hospitalizations. Adv Med Sci. 2010;55:235–41.
Papaetis GS, et al. Serological diagnosis of Chlamydophila pneumoniae infection in Greek COPD patients by microimmunofluorescence and ELISA. Med Sci Monit. 2008;14:MT27–35.
Papi A, et al. Infections and airway inflammation in chronic obstructive pulmonary disease severe exacerbations. Am J Respir Crit Care Med. 2006;173:1114–21.
Parameswaran GI, et al. Effects of bacterial infection on airway antimicrobial peptides and proteins in COPD. Chest. 2011;140:611–7.
Parameswaran GI, et al. Moraxella catarrhalis acquisition, airway inflammation and protease–antiprotease balance in chronic obstructive pulmonary disease. BMC Infect Dis. 2009;9:178.
Paran H, et al. Serological, clinical and radiological findings in adults with bronchopulmonary infections caused by Chlamydia trachomatis. Isr J Med Sci. 1986;22:823–7.
Patel I, et al. Relationship between bacterial colonisation and the frequency, character, and severity of COPD exacerbations. Thorax. 2002;57:759–64.
Peng C, et al. C-reactive protein levels predict bacterial exacerbation in patients with chronic obstructive pulmonary disease. Am J Med Sci. 2013;345:190–4.
Perotin JM, et al. Detection of multiple viral and bacterial infections in acute exacerbation of chronic obstructive pulmonary disease: a pilot prospective study. J Med Virol. 2013;85:866–73.
Philit F, et al. Infectious agents associated with exacerbations of chronic obstructive bronchopneumopathies and asthma attacks. Rev Mal Respir. 1992;9:191–6.
Przybyłowska D, et al. Potential respiratory pathogens colonisation of the denture plaque of patients with chronic obstructive pulmonary disease. Gerodontology. 2016;33:322–7.
Ra SW, et al. Sputum bacteriology and clinical response to antibiotics in moderate exacerbation of chronic obstructive pulmonary disease. Clin Respir J. 2018;12:1424–32.
Rashid MHU, Ahmed I. Pattern of sputum bacteriology in acute exacerbations of chronic obstructive pulmonary disease. J Enam Med Coll. 2018;8:80–4.
Reissig A, et al. Microbiological diagnosis and antibiotic therapy in patients with community-acquired pneumonia and acute COPD exacerbation in daily clinical practice: comparison to current guidelines. Lung. 2013;191:239–46.
Roche N, et al. Yield of sputum microbiological examination in patients hospitalized for exacerbations of chronic obstructive pulmonary disease with purulent sputum. Respiration. 2007;74:19–25.
Rogliani P, et al. Chronic obstructive pulmonary disease and diabetes. COPD Res Pract. 2015;1:3.
Rosell A, et al. Microbiologic determinants of exacerbation in chronic obstructive pulmonary disease. Arch Intern Med. 2005;165:891–7.
Saeedi P, et al. The transient but not resident (TBNR) microbiome: a Yin Yang model for lung immune system. Inhal Toxicol. 2015;27:451–61.
Saldias P, et al. Etiology and biomarkers of systemic inflammation in mild to moderate COPD exacerbations. Rev Med Chil. 2012;140:10–8.
Saxena S et al. Bacteriological profile in acute exacerbation of chronic obstructive lung disease (AECOPD). AIMDR. 2016;2:1–6.
Sethi S. Bacteria in exacerbations of chronic obstructive pulmonary disease: phenomenon or epiphenomenon? Proc Am Thorac Soc. 2004;1:109–14.
Sethi S, et al. Determinants of bacteriological outcomes in exacerbations of chronic obstructive pulmonary disease. Infection. 2016;44:65–76.
Sethi S, et al. New strains of bacteria and exacerbations of chronic obstructive pulmonary disease. N Engl J Med. 2002;347:465–71.
Sethi S, Murphy TF. Bacterial infection in chronic obstructive pulmonary disease in 2000: a state-of-the-art review. Clin Microbiol Rev. 2001;14:336–63.
Sethi S, et al. Airway inflammation and etiology of acute exacerbations of chronic bronchitis. Chest. 2000;118:1557–65.
Sethi S, et al. Airway bacterial concentrations and exacerbations of chronic obstructive pulmonary disease. Am J Respir Crit Care Med. 2007;176:356–61.
Sethi S, et al. Inflammatory profile of new bacterial strain exacerbations of chronic obstructive pulmonary disease. Am J Respir Crit Care Med. 2008;177:491–7.
Sharma P, et al. Sputum bacteriology and antibiotic sensitivity pattern in COPD exacerbation in India. Egypt J Chest Dis Tuberc. 2017;66:593–7.
Shashibhushan B, et al. Bacteriological profile and antibiotic sensitivity pattern in sputum culture of chronic obstructive pulmonary disease patients. Int J Adv Med. 2016;3:671–4.
Shimizu K, et al. Pathogens in COPD exacerbations identified by comprehensive real-time PCR plus older methods. Int J Chronic Obstruct Pulm Dis. 2015;10:2009.
Simpson JL, et al. COPD is characterized by increased detection of Haemophilus influenzae, Streptococcus pneumoniae and a deficiency of Bacillus species. Respirology. 2016;21:697–704.
Singh R, et al. Inflammatory thresholds and the species-specific effects of colonising bacteria in stable chronic obstructive pulmonary disease. Respir Res. 2014;15:114.
Soler N, et al. Bronchoscopic validation of the significance of sputum purulence in severe exacerbations of chronic obstructive pulmonary disease. Thorax. 2007;62:29–35.
Soler N, et al. Bronchial microbial patterns in severe exacerbations of chronic obstructive pulmonary disease (COPD) requiring mechanical ventilation. Am J Respir Crit Care Med. 1998;157:1498–505.
Sriram KB, et al. Non-typeable Haemophilus influenzae detection in the lower airways of patients with lung cancer and chronic obstructive pulmonary disease. Multidiscip Respir Med. 2018;13:11.
Su J, et al. Sputum bacterial and fungal dynamics during exacerbations of severe COPD. PLoS One. 2015;10:e0130736.
Suseela KV, et al. Bacterial profile and antibiotic susceptibility in chronic obstructive pulmonary disease patients with acute exacerbation: a cross sectional study in a tertiary care hospital. Indian J Microbiol Res. 2016;3:317–21.
Tandon MK, et al. Oral immunotherapy with inactivated nontypeable Haemophilus influenzae reduces severity of acute exacerbations in severe COPD. Chest. 2010;137(4):805–11.
Tufvesson E, et al. Patients with chronic obstructive pulmonary disease and chronically colonized with Haemophilus influenzae during stable disease phase have increased airway inflammation. Int J Chronic Obstruct Pulm Dis. 2015;10:881.
Tumkaya M, et al. Relationship between airway colonization, inflammation and exacerbation frequency in COPD. Respir Med. 2007;101:729–37.
Umut S, et al. Determination of the etiological organism during acute exacerbations of COPD and efficacy of azithromycin, ampicillin-sulbactam, ciprofloxacin and cefaclor. J Chemother. 1999;11:211–4.
van der Valk P, et al. Clinical predictors of bacterial involvement in exacerbations of chronic obstructive pulmonary disease. Clin Infect Dis. 2004;39:980–6.
Varma-Basil M, et al. Role of Mycoplasma pneumoniae infection in acute exacerbations of chronic obstructive pulmonary disease. J Med Microbiol. 2009;58:322–6.
Von Hertzen L, et al. Chlamydia pneumoniae antibodies in chronic obstructive pulmonary disease. Int J Epidemiol. 1996;25:658–64.
Wani UA, et al. Seroprevalence of Helicobacter pylori in COPD patients in Kashmir, India. COPD. 2019;1:57–8.
Wedzicha JA. Role of viruses in exacerbations of chronic obstructive pulmonary disease. Proc Am Thorac Soc. 2004;1:115–20.
Ye F, et al. Spectrum and antimicrobial resistance of common pathogenic bacteria isolated from patients with acute exacerbation of chronic obstructive pulmonary disease in mainland of China. Chin Med J. 2013;126:2207–14.
Zhao M, et al. A clinical study of bacterial infection in patients with chronic obstructive pulmonary disease. Chin J Tuberc Respir Dis. 1999;22:88–91.
Zhou Y, et al. Study on the relationship between airway bacterial infections and acute exacerbations in patients with chronic obstructive pulmonary disease. Zhonghua liu xing bing xue za zhi. 2007;28:503–6.
Author information
Authors and Affiliations
Contributions
Study design: SAJ and AA. Acquisition of data: MM, MM, and JS. Analysis and interpretation of data: MM. Drafting of the manuscript: MM and AA. Critical revision of the manuscript for important intellectual content: SAJ, JS, and MM.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Ethical approval
This study does not need ethical approval and patient consent. All analyses were according to previously published studies.
Rights and permissions
About this article
Cite this article
Moghoofei, M., Azimzadeh Jamalkandi, S., Moein, M. et al. Bacterial infections in acute exacerbation of chronic obstructive pulmonary disease: a systematic review and meta-analysis. Infection 48, 19–35 (2020). https://doi.org/10.1007/s15010-019-01350-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s15010-019-01350-1