Abstract
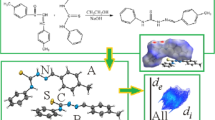
Thiosemicarbazides are sulfur-based nitrogen-rich donor ligands, characterized by π-delocalized electronic charge and flexible molecular chain. The flexible molecular chain and polydentate nature is responsible for their wide biological and synthetic profile to synthesize several classes of heterocycles. In this context, the molecular and crystal structures coupled with Hirshfeld surface (HS) analysis of benzylidene thiosemicarbazides: (E)-2-(4-hydroxybenzylidene) hydrazine-1-carbothioamide, (3a) and (E)-2-(3-nitrobenzylidene) hydrazine-1-carbothioamide, (3b) are reported. Consequently, single crystal x-ray revealed interesting results, and the analyzed compounds were found to be polymorphs of the previously reported structures each having widely different unit cell. Briefly, compound (3a) crystallized with a single molecule in the asymmetric unit whereas its polymorph has six. In contrast, compound (3b) crystallized with Z = 2 having single molecule in asymmetric unit of the polymorph. Subsequently, HS analysis successfully visualized, explored, and quantified weak interactions present in the crystal lattices of the polymorphs. HS analysis revealed oxygen atoms of the nitro group significantly influence HS of molecule (3b).
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thiosemicarbazide (TSC) the simplest hydrazine of thiocarbamic acid is sulfur-based nitrogen-rich donor ligand [1, 2]. TSC possess delocalized π-electronic system in the flexible molecular configuration. The thiocarbonyl carbon show different orientations, creating different conformers as: S=C and C=S. The S=C conformer (2) has thiocarbonyl carbon directed towards intramolecular bonding. Conversely in C=S conformer (1), thiocarbonyl carbon is turned away from intramolecular OH…N bonding (Fig. 1) [3]. TSC ligands display antitumor, antimicrobial, and anti-HIV potential [1]. TSCs are employed in the efficient synthesis of oxadiazoles, pyrazoles, tetrazoles, thiadiazoles, triazepines, triazoles, thiosemicarbazones and metal complexes [4,5,6]. Thiosemicarbazones (TSCZ) possess [C=N–NH–C=S] moiety, interact biomolecules [7, 8] and display anticancer [9] antimicrobial [10], antinociceptive, as well as anti-tubercular potential [11]. Polydentate ligands synthesize metal complexes with tunable properties [12,13,14]. α-N-heterocyclic TSCZ and their metal complexes possess significant antiproliferative potency dependent upon the nature of the heterocyclic ring [15]. TSCZ complexes are highly active against cell lines like MCF-7 [16], SK-BR-3 [17], HCT-116 [18], Hela, and A549 [19]. TSCZ based molybdenum complexes inhibit tyrosinase enzyme by interacting copper ions in the active site [20]. Therefore, some TSCZ of pharmacological interests are presented below (Fig. 2).
Polymorphs are crystalline forms of same chemical composition with different molecular packing. They hold significance in drug molecules, which may form multiple crystalline phases of similar compositions and different internal structures. The most stable polymorph is used as drug to evade the possibility of conversion into another form [21]. Furthermore, hydrogen bonding with receptors holds significance in drug design [22]. Consequently, research has been focused to explore crystal packing of nitrogen and sulfur-based organic compounds. In this context, the centrosymmetric N–H…S=C hydrogen bond dimer in TSCZ is reported to be the result of higher polarizability of lone pair electrons on sulfur atom [23]. Similarly, short centrosymmetric R22(8) N–H···S synthon is involved in supramolecular architecture of thiones [24]. Recently, intermolecular hydrogen bonds via H–N…S is revealed to be responsible for one-dimensional chain of TSCZ [25]. In the current research, H–N…S hydrogen bonds form inversion dimers enclosing R22(6) rings, which in turn are linked to extended R22(22) dimers, creating sheets of molecules of (3a). Likewise, the presence of N–H…S, C–H…S and C–H…O hydrogen bonds link the molecules into chains in (3b).
In view of the pharmacological significance of TSCZ and polymorphs in drug molecules, the single crystal analysis of TSCZ (3a–3b) is presented. A search of the cambridge structural database [26] for compounds incorporating the structure of (3a) produced 3 hits, including the structure of a second triclinic polymorph of (3a) VAGCIR [27] as well as Zn (II), GETZAH [28], and Cu(I), SUTLUO [29], complexes in which (3a) acts as a monodentate ligand coordinating through sulfur atom. A corresponding search for the structure of (3b) produced a single hit, amazingly also of a second polymorph of (3b) but in this instance with both structures crystallizing in the monoclinic space group P21/c but with vastly different unit cell dimensions KOMVUE [30]. No structures with (3b) acting as a ligand were found in this case.
Experimental
Materials and methods
Ethanol (C2H5OH, 99.50%), sulfuric acid (H2SO4, 98%), thiosemicarbazide (NH2C(S)NHNH2, 99%), 4-hydroxybenzaldehyde (4-OH-ArC(O)H, 99%), and 3-nitroaldehyde (3-NO2-ArC(O)H, 99%) were obtained from Sigma-Aldrich. Consequently, the substituted benzaldehydes (1a–1b) (1 mmol) and thiosemicarbazide (2) (1.2 mmol) were dissolved in dry distilled ethanol (15 mL) at room temperature. After 15 min, concentrated sulfuric acid (1 drop) was added to the reaction mixture, and the mixture was constantly refluxed for next 3 h to yield the respective thiosemicarbazones (3a–3b) in higher yield. After the required time, the reaction mixture was cooled to room temperature, and ethanol was rotary evaporated to obtain precipitates. The precipitates were later collected and purified by recrystallization from ethanol at room temperature. Structural variations of the synthesized TSCZ (3a–3b) are described in Table 1. The physico-chemical data of the synthesized TSCZ (3a–3b) have been reported previously [31].
X-ray structure determination
The details of crystal data collected and refined is provided in Table 2. All x-ray measurements were carried out on an Agilent Duo diffractometer using Cu-Kα radiation (λ = 1.54184 Å) with data collection, reduction and absorption corrections controlled using CrysAlisPro at 100(2) K [32]. Data were reduced and multi-scan absorption corrections were applied using CrysAlisPro [32]. The crystal structure was solved by direct methods using SHELXT [33] and refined using full-matrix least-squares procedures via SHELXL-2018/3 [34] and TITAN2000 [35]. All non-hydrogen atoms were refined anisotropically, and all hydrogen atoms bound to carbon were placed in the calculated positions, and their thermal parameters were refined isotropically with Ueq = 1.2–1.5 Ueq(C). The N–H and O–H hydrogen atoms were located in a difference Fourier map and their coordinates were refined with Ueq = 1.2 Ueq(N) and 1.5 Ueq(O) respectively. Molecular plots and the packing diagrams were drawn using Mercury [36]. Additional metrical data were calculated using PLATON [37] and tabular material was produced using WINGX [38].
Results and discussion
Synthesis and chemistry
The synthesis of 1-benzylidene thiosemicarbazides (3a–3b) was performed as given below (Scheme 1). Briefly, substituted benzaldehydes (1a–1b) were dissolved in dry distilled ethanol by continuous stirring at room temperature using catalytic amount of concentrated sulfuric acid. Once the benzaldehydes were completely dissolved, thiosemicarbazide (2) dissolved in dry ethanol was gently poured and the reaction mixture was allowed to reflux for next 3 h to get TSCZ (3a–3b) in higher yield. Upon completion of the reaction, ethanol was rotary evaporated to get precipitates of the target compounds (3a–3b) that were purified by recrystallization from ethanol at room temperature. Structural assignments of the synthesized TSCZ (3a–3b) are based on their consistent spectral (FT-IR, 1H/13C NMR) and % elemental analysis (CHNS) data reported previously [31].
Molecular and crystal structures of (3a–3b)
Details of the crystal and data collection, as well as structure refinements of 1-benzylidene thiosemicarbazides (3a–3b) are given in Table 2.
Crystal structure of (3a)
The molecule of (3a) is a second discrete triclinic polymorph of the crystallized compound (E)-2-(4-hydroxybenzylidene)hydrazine-1-carbothioamide. The compound is crystallized with a single molecule in the asymmetric unit (Fig. 3). In stark contrast, the earlier polymorph, VAGCIR [27] reported in 2009, crystalizes with six unique, reasonably planar molecules in the asymmetric unit. Conversely, the crystallized molecule reported in this work is not planar, the benzene ring being inclined to the reasonably planar hydrazine carbothioamide part of the molecule by 28.82(6)° The planarity of this non-ring segment (max deviation 0.0665 Å) is supported by the formation of intramolecular N1-H1…N3 hydrogen bond.
Crystal structure of (3b)
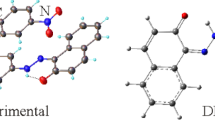
The molecule of (3b) is a second monoclinic polymorph of the synthesized compound (E)-2-(3-nitrobenzylidene)hydrazine-1-carbothioamide. The alternative polymorph was reported in 2009 [30]. Contrary to the original polymorph, the compound reported here crystallizes with two unique molecules, M1 and M2, in the asymmetric unit, linked by N21-HN22…O12 hydrogen bond (Fig. 4). In the numbering scheme, the two unique molecules are differentiated by respective leading 1 and 2 characters. Intramolecular N11-HN11…N13 and corresponding N21-HN21…N23 hydrogen bonds also effect the conformations of the molecules. Both molecules are almost planar with rms deviations of 0.1395 Å for M1 and 0.0626 Å for M2 from the least squares planes through all non-hydrogen atoms of the molecules. Bond distances and angles in the two molecules are reasonably comparable and are also like those reported for the polymorph.
Crystal packing of (3a)
In the crystal structure of (3a), O4-H4O…S1 and C5-H5…S1 hydrogen bonds (Table 3) combine to form rows of molecules (3a) along c, (Fig. 5). The N2-H1N4…S1 hydrogen bonds form inversion dimers that enclose R22(6) rings and these link to more extensive R22(22) dimers, formed by N10-H1N1…O2 hydrogen bonds, resulting in sheets of (3a) molecules formed along the bc diagonal (Fig. 6). A peculiar feature of the packing in this structure is the presence of close π…π contacts between adjacent aromatic rings, with a centroid-to-centroid distances of 3.9465(13) Å. These combine with previously mentioned inversion related to N1-H1N1…O4 hydrogen bonds to stack the (3a) molecules along the a axis (Fig. 7). This extensive series of contacts combine to stack the molecules along the a axis direction (Fig. 8). Hydrogen bonds for (3a) are detailed in Table 3.
Crystal packing of (3b)
In the crystal of (3b), N–H…S, C-H…S and C-H…O hydrogen bonds (Table 4) link the molecules M1 and M2 into chains along c (Fig. 9). Chains of both the molecules also form along the bc diagonal through N–H…S, C-H…S and N–H…O hydrogen bonds (Fig. 10). However, unlike (3a) no close π…π contacts were observed here. The presence of various hydrogen bonds combines to stack the molecules along the a axis direction (Fig. 11).
Hirshfeld surface analysis of (3a) and (3b)
Details of the intermolecular interactions present in the molecules (3a) and (3b) were obtained from Hirshfeld surface (HS) analysis [39]. The surfaces and two dimension fingerprint plots were generated using Crystal Explorer [40].
Hirshfeld surface analysis of (3a)
Figure 12 shows HS of the asymmetric unit of compound (3a). The corresponding bold red circles represent classical O–H…S and N–H…S hydrogen bonds. Fingerprint plots depict H…H contacts are the most prevalent (Fig. 13). Moreover, significant contributions come from H…S/S…H, H…C/C…H, H…O/O…H, C…S, S…C and N…N contacts. The weak contacts, < 1% of the total are confined to Table 5.
Full 2D fingerprint plots for the asymmetric unit of (3a), (a), together with (b-j) separate contact types and included HS for the individual contacts. Minor contacts contributing less than 1% to the total HS are not shown here but, for completeness, are shown in Table 5
Hirshfeld surface analysis of (3b)
Figure 14 shows HS of the two molecules comprising the asymmetric unit of (3b). Consequently, the bold red circles represent classical N–H…S and N–H…O hydrogen bonds. The weaker C-H…O hydrogen bond appears as faint red circle. The major impact of introducing two additional oxygen atoms of the nitro group in (3b) on the HS contributions is displayed in fingerprint plots in Fig. 15. H…O/O…H contacts contribute most to the HS at 21.7% as compared to 9.6% in (3a). Moreover, other substantial contributions come from H…H, H…S/S…H, H…C/C…H and H…N/N…H contacts. Furthermore, indication of nitro group on the HS is the presence of C…O/O…C, N…O/O…N and O…S/S…O contacts at moderate levels that are insignificant among the contacts for (3a).
Full 2D fingerprint plots for the asymmetric unit of (3b), (a), together with (b-l) separate contact types and included surface areas for the individual contacts. Minor contacts contributing less than 1% to the total surface area are not shown but, for completeness, are shown in Table 5
Conclusions
Given our interests in the synthesis, characterization and structure determination of organic molecules with potential pharmacological applications, herein the molecular, crystal structures and Hirshfeld surface analyses of (E)-2-(4-hydroxybenzylidene) hydrazine-1-carbothioamide, (3a) and (E)-2-(3-nitrobenzylidene) hydrazine-1-carbothioamide, (3b) is reported. Immediate interest in these structures lies in the fact that they are both polymorphs of previously reported structures but with significantly different cell occupancies and dimensions. This implores the question of how these structurally different materials would differ in their pharmacological activity. Another important feature of these structures is the singular differences in the Hirshfeld surface contacts for the two structures. Whereas (3a) has a more conventional surface contact distribution with more than 50% being from H…H and H…C/C…H contacts with a substantial contribution also from H…S/S…H interactions, the situation for (3b) changes dramatically with the presence of the nitro group. While the H…S/S…H contacts remain similar, H…O/O…H becomes the dominant contact, with the H…H contact more than halved. It would be interesting to see if this trend extended to similar systems.
References
P.T. Acharya, Z.A. Bhavsar, D.J. Jethava, D.B. Patel, H.D. Patel, J. Mol. Struct. 1226 129268 (2021).https://doi.org/10.1016/j.molstruc.2020.129268
G.A. Gazieva, A.N. Kravchenko, Russ. Chem. Rev. 81, 494–523 (2012). https://doi.org/10.1070/RC2012v081n06ABEH004235
A. Sykula, A.K. Baron, A. Dzeikala, A. Bodzioch, E.L. Chruscinska, Chem. Phy. 517, 91–103 (2019). https://doi.org/10.1016/j.chemphys.2018.09.033
A.A. Hassan, A.M. Shawky, J. Hetero. Chem. 48, 495–516 (2011). https://doi.org/10.1002/jhet.553
J.S. Casas, M.S.G. Tasende, J. Sordo, J. Coord. Chem. Rev. 209, 197–261 (2000). https://doi.org/10.1016/S0010-8545(00)00363-5
M. Khosravi, S. Saeednia, P. Iranmanesh, M.H. Ardakani, J. Clust. Sci. 34, 311–322 (2023). https://doi.org/10.1007/s10876-021-02210-5
P. Kalaivani, C. Umadevi, R. Prabhakaran, F. Dallemer, P.S. Mohan, Inorg. Chim. Acta 438, 264–276 (2015). https://doi.org/10.1016/j.ica.2015.08.031
G. Kalaiarasi, C. Umadevi, A. Shanmugapriya, P. Kalaivani, F. Dallemer, R. Prabhakaran, Inorg. Chim. Acta 453, 547–558 (2016). https://doi.org/10.1016/j.ica.2016.09.006
I. Dilović, M. Rubčić, V. Vrdoljak, S.K. Pavelić, M. Kralj, I. Piantanida, M. Cindric, Bioorg. Med. Chem. 16, 5189–5198 (2008). https://doi.org/10.1016/j.bmc.2008.03.006
M. Jagadeesh, H.K. Rashmi, Y.S. Rao, R.A. Sreenath, B. Prathima, P.U.M. Devi, A.V. Reddy, Spectrochim. Acta. A Mol. Biomol Spectrosc. 115, 583–587 (2013). https://doi.org/10.1016/j.saa.2013.06.071
M. Wujec, E. Kedzierska, E. Kusmierz, T. Plech, A. Wrobel, A. Paneth, J. Orzelska, S. Fidecka, P. Paneth, Molecules 19(4), 4745–4759 (2014). https://doi.org/10.3390/molecules19044745
H.E. Hosseini, H. Aghabozorg, M. Shamsipur, M. Mirzaei, M. Ghanbari, J. Iran. Chem. Soc. 8(3), 762–774 (2011)
M. Mirzaei, H.E. Hosseini, M. Shamsipur, M. Saeedi, M. Ardalani, A. Bauzá, J.T. Mague, A. Frontera, M. Habibi, RSC Adv. 5, 72923–72936 (2015)
M. Kheirkhahi, B. Shaabani, H.S. Kafil, J. Iranian, Chem. Soc. 18, 3429–3441 (2021). https://doi.org/10.1007/s13738-021-02281-1
A.I. Matesanz, C. Hernández, P. Souza, J. Inorg. Biochem. 138, 16–23 (2014). https://doi.org/10.1016/j.jinorgbio.2014.04.017
G. Prakash, R. Manikandan, P. Viswanathamurthi, K. Velmurugan, R. Nandhakumar, J. Photochem. Photobiol. B 138, 63–74 (2014). https://doi.org/10.1016/j.jphotobiol.2014.04.019
B.M. Zeglis, V. Divilov, J.S. Lewis, J. Med. Chem. 54, 2391–2398 (2011). https://doi.org/10.1021/jm101532u
M.A. Hussein, T.S. Guan, R.A. Haque, M.B.K. Ahamed, A.M.S.A. Majid, Polyhedron 85, 93–103 (2015). https://doi.org/10.1016/j.poly.2014.02.048
N.K. Singh, A.A. Kumbhar, Y.R. Pokharel, P.N. Yadav, J. Inorg. Biochem. 210, 111134 (2020). https://doi.org/10.1016/j.jinorgbio.2020.111134
S.E. Bakır, O. Sacan, M. Şahin, R. Yanardag, B. Ülküseven, J. Mol. Struct. 1194, 35–41 (2019). https://doi.org/10.1016/j.molstruc.2019.05.077
J.R. Cox, L.A. Ferris, V.R. Thalladi, Angew. Chem. 119(23), 4411–4414 (2007). https://doi.org/10.1002/ange.200605257
J. Sun, S. Cai, H. Mei, J. Li, N. Yan, Q. Wang, Z. Lin, D. Huo, Chem. Biol. Drug Des. 76, 245–254 (2010). https://doi.org/10.1111/j.1747-0285.2010.01006.x
S.B. Novaković, B. Fraisse, G.A. Bogdanović, A.S. Biré, Cryst. Growth Des. 7(2), 191–195 (2007). https://doi.org/10.1021/cg060497+
D. Dey, T.P. Mohan, B. Vishalakshi, D. Chopra, Cryst. Growth Des. 14, 5881–5896 (2014). https://doi.org/10.1021/cg501103c
U.M. Osman, A.S.N. Farizal, S. Arshad, M.A. Kadir, X-ray Struct. Anal Online 33, 3–4 (2017). https://doi.org/10.2116/xraystruct.33.3
C.R. Groom, I.J. Bruno, M.P. Lightfoot, S.C. Ward, Acta Cryst B. Struct. Sci. Cryst. Eng. Mater. 72, 171–179 (2016). https://doi.org/10.1107/S2052520616003954
N.N. Ji, Z.Q. Shi, Chinese J. Spectrosc. Lab. 26, 1156 (2009)
X.M. Yang, Acta Crystallogr. E. 63, m284–m285 (2007). https://doi.org/10.1107/S1600536806054183
H.G. Zheng, D.X. Zeng, X.Q. Xina, W.T. Wong, Polyhedron 16, 3499–3503 (1997). https://doi.org/10.1016/S0277-5387(97)00129-0
D.H. Wu, Z.F. Li, Y.H. Zhang, Acta Crystallogr. E. 65, o163-0163 (2009). https://doi.org/10.1107/S1600536808042645
H. Aziz, A. Mahmood, S. Zaib, A. Saeed, Z. Shafiq, J. Pelletier, J. Sevigny, J. Iqbal, Bioorg. Chem. 102, 104088 (2020).https://doi.org/10.1016/j.bioorg.2020.104088
CrysAlisPro, Agilent Technologies Ltd, Yarnton, (2018).
G.M. Sheldrick, SHELXT Acta Crystallogr. A 71, 3–8 (2015). https://doi.org/10.1107/S2053273314026370
G.M. Sheldrick, SHELXL Acta Crystallogr. C 71, 3–8 (2015). https://doi.org/10.1107/S2053229614024218
K.A. Hunter, J. Simpson, TITAN2000: Inoversity of Otago, Dunedin New Zealand (1999).
C.F. Macrae, I.J. Bruno, J.A. Chisholm, P.R. Edgington, P. McCabe, E. Pidcock, L.R. Monge, R. Taylor, J. van de Streek, J.P.A. Wood, Mercury J. Appl. Crystallogr. 41, 466–470 (2008). https://doi.org/10.1107/S0021889807067908
A.L. Spek, PLATON Acta Crystallogr. D65, 148–155 (2009). https://doi.org/10.1107/S090744490804362X/
L.J. Farrugia, WinGX J. Appl. Crystallogr. 45, 849–854 (2012). https://doi.org/10.1107/S0021889812029111
M.A. Spackman, D. Jayatilaka, Cryst. Eng. Comm. 11, 19–32 (2009). https://doi.org/10.1039/B818330A
M.J. Turner, J.J. McKinnon, S.K. Wolff, D.J. Grimwood, P.R. Spackman, D. Jayatilaka, M.A. Spackman, CrystalExplorer17. University of Western Australia Nedlands, Western Australia (2017). http://hirshfeldsurface.net
Acknowledgements
Dr. Hamid Aziz is highly grateful to the Higher Education Commission (HEC), Pakistan for providing indigenous scholarship as the financial support for the research work performed. We also thank the University of Otago for purchase of the diffractometer and the Chemistry Department, University of Otago for support of the work of Jim Simpson.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aziz, H., Saeed, A., Simpson, J. et al. New polymorphs of 1-benzylidene thiosemicarbazides: crystal structure and hirshfeld surface analysis. J IRAN CHEM SOC 20, 1831–1842 (2023). https://doi.org/10.1007/s13738-023-02800-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13738-023-02800-2