Abstract
Alzheimer’s disease is a neurological disorder that affects an individual’s memory, motor functions, behaviour, and thought process. It has been observed that the hippocampus is the first region that gets affected by Alzheimer’s. Hence, a study of the hippocampus region can identify genes responsible for the occurrence of the early stage of the disease. Most often, t-test and correlation are used to identify significant genes at the initial level. As the genes are differentially expressed, their classification power is generally high. These genes might appear significant, but their degree of specificity towards the disease might be low, leading to misleading interpretations. Similarly, there may be many false correlations between the genes that can affect the identification of relevant genes. This paper introduces a new framework to reduce the false correlations and find the potential biomarkers for the disease. The framework concerned uses the t-test, correlation, Gene Ontology (GO) categories, and machine learning techniques to find potential genes. The proposed framework detects Alzheimer-related genes and achieves more than 95% classification accuracy in every dataset considered. Some of the identified genes which are directly involved in Alzheimer are APP, GRIN2B, and APLP2. The proposed framework also identifies genes like ZNF621, RTF1, DCH1, and ERBB4, which may play an important role in Alzheimer’s. Gene set enrichment analysis (GSEA) is also carried out to determine the major GO categories: down-regulated and up-regulated.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Alzheimer’s disease is a prevalent form of dementia. It is an irreversible disease with a progressive loss of memory and worsening cognitive function. The leading cause of AD is said to be the abnormal deposits of protein forms amyloid plaques and tau tangles throughout the brain (Alzheimer’s 2015). Hippocampus is the brain region associated with all stages of semantic memory and is said to be affected first in AD (Duff and Covington 2020; Anand and Dhikav 2012). APOE is said to be the most common gene associated with AD (Alzheimer’s 2015). Apart from APOE, APP, PSEN1, and PSEN2 are also observed as the cause of AD (Lanoiselée 2017). Various studies have been carried out to identify the genes which are differentially expressed in the AD affected brains (Lanoiselée 2017; Ray 2017). T-test and gene correlation networks are the most common statistical techniques used to identify the significant genes. The t-test is used to test the significant difference in gene expression levels (Zhou 2008). For example, Zhu and Yang (2016) used the rejection region of the t-test to identify the candidate gene for AD. However, the t-test only gives the significant difference in the mean expression values of genes between control and disease sets, which is not enough to determine the significant influence of genes on the disease. There could be many other reasons apart from the disease, which can result in a change in the expression value of a particular gene. Ray (2017) analyzed the preservation patterns of gene co-expression networks during Alzheimer’s disease progression. Like the t-test, the correlation between two genes is not enough to tell that two correlated genes interact with each other. Hu and Yu (2020) constructed a co-expression network using WGNCA and analyzed their clinical features. As a result, they identified four genes(ENO2, ELAVL4, SNAP91, and NEFM) said to be associated with AD. Xia et al. (2014) constructed the co-expression network using the method proposed by Ruan and Zhang (2006). Then, they ranked the genes based on a new topological overlap formula, a modified version of the formula described in Ray and Zhang (2010), Ray et al. (2012). The main concern with constructing a co-expression network using this method is that it depends on the user-defined value \(\alpha \). Different values of \(\alpha \) result in a different number of edges. This means that every gene in the co-expression network is connected to its top \(\alpha \) co-expressed genes. It may impact the removal of positive edges.
As the gene expression datasets are vast, various machine learning techniques are used along with the other statistical methods. Takahiro et al. (2016); Nishiwaki et al. (2016) used the random forest to identify the AD-related genes. In AL-Dlaeen and Alashqur (2014), AL-Dlaeen et al. used a decision tree classifier to predict the AD. There are many other algorithms, such as the K-means clustering algorithm, Principal component analysis(PCA), ant colony algorithm (ACO), independent component analysis algorithm (ICA), the angle cosine distance algorithm, and Chebyshev inequality algorithm (ACD), which produce less efficient and unstable results (Zhu and Yang 2016). Sharma and Dey (2021) combined two feature selection techniques, LASSO and Random forest, for gene selection and achieved a high classification accuracy. In Ramaswamy (2021), Ramya et al. used the t-test, signal-to-noise ratio, and f-test for the initial selection of genes and then selected genes were used in a modified particle swarm optimization algorithm to obtain further refined genes. Cheng and Liu (2021) observed that the machine learning model’s average classification accuracy is higher than that of conventional methods. Apart from this, the authors also observed that machine learning approaches could also recognize oxidative phosphorylation genes in the Alzheimer’s pathway. Saputra (2020) compared different decision trees with particle swarm optimization as feature selection methods and observed that the random forest gives the best accuracy. Kuang et al. (2021) compared the performance of three machine learning algorithms, artificial neural network (ANN), and decision tree and logistic regression models, to predict the AD. They found that ANN worked better than the other two models, and observed that the age, daily routine, urine neuronal thread protein associated with AD, smoking, alcohol intake, and sex are the crucial factors.
Almost every feature selection technique is applied on differentially expressed genes, i.e., genes obtained after the t-test. As the genes are differentially expressed, their classification power is generally high. These genes might appear significant, but their degree of specificity towards the disease might be low, leading to misleading interpretations. Some genes are expressed in basic cellular pathways and possess a higher probability of being differentially expressed across several biological conditions (Crow and Lim 2019). Nevertheless, as AD’s causes probably include genetic, environmental, and lifestyle factors, different genes are identified as important in different AD datasets. Due to these various factors involved in AD, statistical methods and machine learning techniques alone are inadequate.
2 Dataset
The gene expression datasets GSE48350, GSE5281, and GSE28146, are downloaded from Gene Expression Omnibus (GEO), NCBI. The datasets GSE48350Footnote 1 (dataset 1) and GSE5281Footnote 2(dataset 2) contain gene expression data of control and Alzheimer’s disease patients. The dataset GSE28146Footnote 3(dataset 3) contains microarray data of the hippocampal gray matter. The GSE48350 and GSE5281 datasets contain samples from different brain regions. We took only Hippocampus data for analysis as it is said to be affected first in Alzheimer’s disease Anand and Dhikav (2012). Table 1 describes the data.
3 Proposed framework
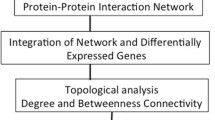
This paper introduces a new framework, including t-test, correlation network, GO similarity matrix, and feature selection for filtering genes of less interest. Figure 1 shows the proposed framework.
Initially, differentially expressed genes are identified using the t-test. Then, the identified genes are used to create two separate correlation networks for AD and control sets using Pearson’s correlation. There may be many false correlations, so a GO similarity matrix is introduced to reduce the false correlations. GO matrix consists of the number of similar GO terms between every pair of genes. Then, the GO similarity matrix is used to eliminate edges in the correlation networks that do not fall under the pre-defined criteria. The resultant correlation networks are then used for further analysis. Genes present in the control correlation network but not in the AD correlation network and vice versa are selected as the genes of interest. A separate Gene Set Enrichment Analysis (GSEA) has been carried out for selected genes to identify the affected GO categories. The feature selection algorithm is now applied to the selected genes to determine the most important genes from the important ones. In the final stage, the classification accuracy of the final set of genes is checked using a classification algorithm. All the components of the proposed framework are explained in detail in the following sections.
3.1 T-test
A t-test was performed on all the datasets, i.e., GSE48350, GSE5281, and GSE28146, to find the significant difference in the expression values of genes in control and AD patients using GEO2R [NCBI]. p value \(\le {0.05}\) and fold count, \(|logFC|\ge {0.8}\) are used as the threshold values. These are standard values used in the literature. As many genes have different probe ids, we took the average expression and fold count values. 696, 7222, and 1893 Differentially expressed genes (DEGs) are obtained from dataset 1, dataset 2, and dataset 3, respectively.
3.2 Gene co-expression network
Pearson’s correlation is used to calculate the correlation between each pair of genes after performing the t-test. \(\pm 0.8\) is taken as the threshold value as it is interpreted as strong/high correlation (Akoglu 2018; Mukaka 2012). They have pointed out that a correlation value of 0.7–0.9 indicates a high positive correlation and 0.9 as a very high positive correlation. Hence, a value of 0.8 is chosen as the threshold. All the correlation values which are greater than or equal to |0.8| are considered as 1, and the rest of the values are considered as 0. The resultant adjacency matrix is used to create the gene co-expression matrix. Two separate networks for control and AD are constructed using the binarised Pearson correlation values as edges.
3.3 GO similarity matrix
Gene ontology (GO) (Ashburner 2000) has become an accepted norm to evaluate the practical connections among gene products. GO is a scientific classification of biological terms identified with the properties of genes or their products. There are three GO categories: biological process, cellular component, and molecular function. Two proteins engaged with the same biological process are bound to interact than proteins engaged with various biological processes (Zhao and Wang 2018). Besides, two proteins need to come into close contact (essentially momentarily) to communicate; subsequently, co-localization can likewise be utilized to anticipate protein–protein interactions. Hence, the proposed framework uses GO categories for measuring the strength of the connection between genes in the correlation network.
GO similarity matrix consists of the GO similarity score between a pair of genes. Go similarity score is calculated as the number of common GO terms between two genes. For example, if Gene1 has 5 GO terms GO1, GO2, GO3, GO4, and GO5, and Gene2 has 4 GO terms GO1, GO3, GO5, and GO6. There are three common GO terms between the genes Gene1 and Gene2, which are GO1, GO2, and GO5. Hence, the GO similarity score (\(GO_{(Gene1, Gene2)} \)) between Gene1 and Gene2 is 3. GO categories of the differentially expressed genes (DEGs) identified by the t-test are used to construct the GO similarity matrix. The GO categories of all the DEGs are downloaded from DAVID (The Database for Annotation, Visualization, and Integrated Discovery) (Huang 2007). In the first dataset (GSE48350), out of 696 DEGs, 646 DEGs have known GO terms, and in the second dataset (GSE5281), out of 7222 DEGs, 6377 DEGs have known GO terms. In dataset 3 (GSE28146), out of 1893 DEGs, 1210 DEGs have known GO terms. All the three GO categories, i.e., Biological Process (BP), Molecular Function (MF), and Cellular Component (CC), are considered for the construction of the GO similarity matrix. Gene similarity matrix consists of the GO similarity score between all pairs of genes, as shown in Fig. 2.
This GO similarity matrix is used to create the GO network. To determine the cut-off score for the GO similarity score, 4000 genes (except the genes considered in the experiment) having nearly 11000 edges that are experimentally proven are taken [DAVID]. The GO similarities between the genes having experimentally proven interactions are analyzed. The average number of similar GO terms between two genes [having experimentally proven edges (interactions)] is 3.14. Hence, the ceiling value 4 is taken as the threshold value. All the edges whose weight (GO similarity score) is less than four are deleted. An edge between two genes is to be considered if they have at least four common GO terms.
3.4 Common genes and edges between GO and correlation networks
A combined network is constructed to take care of the false correlations by mapping gene correlation networks (Control and AD) to the GO network. As genes sharing more GO terms will tend to have a high biological association, combining the correlation and GO network helps to eliminate the edges with less biological significance (Martin et al. 2004; Zhao and Wang 2018). A combined AD network is constructed using the common edges between the AD correlation network and the GO network. A similar combined network is constructed for the control network using the control correlation network and GO network. Table 2 shows the count of edges in the correlation network and GO network.
The common control network consists of 240 genes and 673 edges in dataset 1, 2487 genes and 20486 edges in dataset 2, and 595 genes and 989 edges in dataset 3. The common AD network consists of 219 genes and 774 edges in dataset 1, 3138 genes and 15499 edges in dataset 2, and 12 genes and 7 edges in dataset 3.
3.5 Analysis of networks
Generally, a gene of interest behaves differently in normal and affected persons. Hence, both AD and control common networks are analyzed and culled the genes present in the AD network but not in the control network (AD-CTRL). As a result, 79 such genes are identified in dataset 1, 1107 genes in dataset 2, and 1 gene in dataset 3. Similarly, 100, 456, and 584 genes in dataset 1, dataset 2 and dataset 3 are identified, which are present in the control network but not in the AD network. Figure 3 shows the Venn diagram of genes.
3.6 Gene set enrichment analysis
The gene set enrichment analysis of AD and control networks is performed using GSEA 4.0 application, which can be downloaded from http://software.broadinstitute.org/gsea (Subramanian and Tamayo 2005). The all_GENE_ONTOLOGY database is used for this analysis. In dataset 1, we found that 44 and 13 GO terms are down-regulated and up-regulated, respectively, in the AD network. In the control network of dataset 1, 99, and 20 GO terms are down-regulated and up-regulated, respectively. Similarly, in dataset 2, 298 and 148 GO terms are down-regulated and up-regulated, respectively, in the AD network. In contrast, in the control network, 307 and 321 GO terms are down-regulated and up-regulated. We found a total of 11 and 21 GO terms, which got down-regulated in both the AD networks of dataset 1 and dataset 2 and control networks of dataset 1 and dataset 2, respectively. Similarly, 8 and 16 common GO terms got up-regulated in AD and control networks of dataset 1 and dataset 2. Tables 3 and 4 list the GO terms which got up-regulated/down-regulated in AD network but not in control network and vice versa. Tables 5 and 6 list the GO terms which got down-regulated and up-regulated in the control and AD networks, respectively. All GO terms related to dataset 3 are provided in supplementary data.
3.7 Classification
As all the genes in the combined network may be important concerning Alzheimer’s disease, only the top genes are picked up using feature selection are chosen for the discussion. Although any classification method can be used, the main purpose of this stage is to see the proposed approach’s effectiveness for selecting the potential candidate genes in discriminating between control and genes. To analyze whether the identified genes are able to classify the disease or not, the decision tree and random forest are used for the classification. As an input to the decision tree and random forest, the expression value of genes present in the AD network but not in the control network and genes present in the control network but not in the AD network are used. J48 decision tree and random forest are used with tenfold cross-validation. A total of 2 decision trees are constructed, one for dataset 1 and another for dataset 2. Feature selection is also performed using correlation-based feature subset selection for machine learning algorithms (Hall 2000). After performing feature selection, we got 13 genes out of 179 genes (79 + 100, Fig. 3), in dataset 1 (GSE48350), 101 genes out of 1563 (1107 + 456, Fig. 3) genes, and 54 genes out of 585 (1 + 584, Fig. 3) genes, in dataset 2 (GSE5281). Table 7 shows the accuracy obtained for each dataset.
4 Comparison
For the comparison purpose, we have considered two recently published frameworks: the first is based on Lasso and random forest (LASSO & RF) (Sharma and Dey 2021), and the second is based on t-test, genetic algorithm, and a modified particle swarm optimization algorithm (MPSO) (Ramaswamy 2021). For a fair comparison, if the number of genes obtained by the frameworks is more than 20, we chose only the top 20 genes for the comparison. Table 8 shows the top genes obtained from the different frameworks for different datasets. Tables 9 and 10 list all the identified genes in dataset 1 and dataset 2, respectively. Genes selected for dataset 3 are provided in supplementary data.
As observed from Table 8, though the significant genes obtained after feature selection are almost different for all the datasets, yet the accuracy of the genes acquired is nearly the same in all datasets (Table 7). Hence, this does not provide us with any inference. Therefore, we compared the degree of specificity of genes obtained by the proposed framework, LAASO & RF and MPSO, towards Alzheimer’s disease. We checked the direct interactions of the genes obtained with the AD pathway genes using the STRING database. We did not find any common pattern in the number of interactions, making it difficult to draw any conclusion. We further used DAVIDFootnote 4 to obtain the diseases of the genes obtained after feature selection which did not yield significant results as the number of genes is less, and some are not characterized. It is well known that interacting proteins regulate the function of a protein (Swamy 2021). Therefore, retrieving the interacting partners and the associated diseases can give us a deeper insight into the genes obtained from our framework. HIPPIEFootnote 5 is used to fetch the high confidence primary interacting proteins of the genes obtained from our analysis. The primary interacting genes are then subjected to DAVID analysis to obtain the corresponding diseases.
It is observed that in dataset 1 and dataset 2, primary interactions of the genes obtained by the proposed framework are directly associated with Alzheimer’s disease with high significance. In contrast, the interacting partners of genes obtained from other algorithms are not at all related to any neurological disorders. Although in dataset 3, genes obtained from the proposed framework, LASSO & RF, and MPSO framework have interacting partners implicated in Alzheimer’s disease. However, it is interesting to note that the significance and count of genes associated with AD in the proposed framework are quite high compared to the LASSO & RF and MPSO framework. The supplementary data provide the table of all the diseases related to the genes, the gene count, and their corresponding p values.
5 Results
Using the introduced framework, we are able to identify genes in all datasets that are directly or indirectly related to AD with a high classification power. More than 95% accuracy is achieved for classifying the disease and control using the identified genes. Tables 9 and 10 list the genes identified. The link between the identified genes and the AD pathway genes is analyzed to find out the importance of the identified genes in this work. As a result, it is found that most genes have either direct or one-hop interaction with the AD pathway genes. Table 11 shows some direct interactions between top genes of dataset 1 and AD pathway genes. As the top genes in both datasets are different, we tried to determine the relationship between both datasets’ top genes. STRING databaseFootnote 6 is used to find interactions between the genes, only interactions that are experimentally proven or are from the curated database with at least medium confidence value 0.4 (as mentioned in STRING database) are considered. All the top genes of dataset 1 have either direct or one-hop connections with at least one top gene of dataset 2 (a few of the interactions are shown in Table 12). We also checked the GO similarity between the top genes of both datasets and the GO similarity of top genes with the AD pathway genes to find the similarity between them. Also checked the primary interactions of the identified genes and found them related to AD with high significance compared to the genes identified by other considered frameworks. All the interactions, GO similarity, disease-associated, and primary interactions files can be downloaded from “Supplementary Data”.
In dataset 1, out of 13 identified genes, 5 (BTK, CD44, ERBB4, NSG1, and TAC1) are found to be related to AD in the recent literature. Similarly, many genes (ADAM22, AGFG1, GRIN2B, MPRIP, ZNF532, etc.), identified in dataset 2 are listed in the literature. Gene ATP2B3Footnote 7 has human phenotype ontology of ataxia, cerebellar atrophy, cerebellar hypoplasia, and clumsiness. Gene FGF12 has a human phenotype ontology of abnormal myelination, abnormality of vision, and absence of speech. In Keaney (2019) observed that the activation of phospholipase gamma 2, a genetic risk factor in AD, is decreased due to the blockade of BTK. Pinner (2017) investigated the expression values of CD44 splice variants in the hippocampus region of AD patients and compared it with the control patients and observed that the expression values of splice variants of CD44 are significantly higher in AD patients when compared to the normal person. The research suggested that some splice variants of CD44 contribute to AD pathology. Woo (2011) found that up-regulation of the immunoreactivity of ERBB4 may involve in Alzheimer’s disease progression. Abhik Ray and Gerecke (2003) observed that Neuregulin-1 and ERBB4 immunoreactivity is associated with plaques formation in the AD brain. Norstrom (2010) Norstrom et al. reported that NEEP21 protein (gene name: NSG1) affects the processing of APP and A\(\beta \) production. Magistri (2015) analyzed that in the hippocampus region of the brain in AD patients, TAC1 is down-regulated compared to controls hippocampus.
The GSEA analysis shows that out of 22, 12 GO terms that got down-regulated in the control network are not being regulated in the AD network and vice versa, which indicates that there may be a disturbance in the regulation of those 12 GO terms. Similarly, out of 21, 18 GO terms that got up-regulated in the control network is not being regulated in the AD network vice versa. Tables 3 and 4 list the GO categories which may got disturbed. In the identified GO terms, we find that some are found to be disturbed in the Alzheimer’s disease, like, GO SYNAPTIC VESICLE MEMBRANE, GO AXON, GO TRANSPORT VESICLE MEMBRANE, GO VESICLE MEDIATED TRANSPORT IN SYNAPSE, GO NEGATIVE REGULATION OF RNA BIOSYNTHETIC PROCESS, GO REGULATION OF CELL POPULATION PROLIFERATION, GO NEGATIVE REGULATION OF TRANSCRIPTION BY RNA POLYMERASE II, GO RESPONSE TO WOUNDING, GO SKELETAL SYSTEM DEVELOPMENT, GO POSITIVE REGULATION OF RNA BIOSYNTHETIC PROCESS, and GO ZINC ION BINDING. Blennow and Bogdanovic (1996) found that the level of synaptic vesicle membrane protein rab3a was reduced in Alzheimer’s disease in the hippocampus. In the studies, it is found that in Alzheimer’s disease, the amyloid-beta disturbed the vesicle transport in synapse in the hippocampus (Seifert and Eckenstaler 2016; Kelly and Ferreira 2007). Wu and Zhang (2016) observed that the cell proliferation gets slowdown when the APP is overexpressed. Watt (2010) discussed the role of Zinc in Alzheimer’s disease. Zinc binds to amyloid-beta, advancing its conglomeration into neurotoxic species, and disturbance of zinc homeostasis in the brain results in synaptic and memory deficiencies. Kiecolt-Glaser and Marucha (1995) observed that wound healing took a long time significantly in AD patients than in controls. Chen and Lo (2017) conclude that AD increase the risk of osteoporosis (Skelton disorder). The overexpression of amyloid-beta might happen in both cerebrum and bone, meddling with the RANKL signalling cascade, improving osteoclast activities, and prompting osteoporosis.
6 Conclusions
In summary, in this paper, a framework that includes t-test, correlation, GO categories, and machine learning techniques is developed to identify the potential biomarkers for Alzheimer’s disease. The GO categories are analyzed and used to create a more biologically significant network, which helps in eliminating false correlations. Feature selection is used to list out the top genes. Then, using the J48 decision tree and random forest, their classification power is estimated and obtained more than 95% accuracy for all the datasets. Biological interactions between the top genes of all datasets are studied in which the top genes either have direct or one-hop experimentally proven interactions with one another. Biological interactions between top genes and AD pathway genes are also studied. As a result, many of the genes were found to have direct experimentally proven interactions with the AD pathway genes. Primary interactions of selected genes show that the genes selected by the proposed framework are associated with Alzheimer’s disease. Gene set enrichment analysis of AD and control networks is also carried out and found that GO terms which got up-regulated/down-regulated in AD network but not in control network and vice versa, may get disturbed in Alzheimer’s disease. The literature shows that the genes identified by the decision tree classifier whose logFC values indicate that these genes that need to be up-regulated are down-regulated and vice versa. The results consist of the genes and GO terms that are related to Alzheimer’s disease in the literature, which adds more credibility to the results. The results show that even though the classification power of genes identified by other frameworks are high or the same, the genes identified by the proposed framework have a high degree of association with AD in comparison to the genes identified by the other frameworks considered. In future, the proposed framework can be applied to other diseases too, and an automated tool based on the proposed framework can be developed.
Notes
References
Akoglu H (2018) User’s guide to correlation coefficients. Turk J Emerg Med 18(3):91–93
AL-Dlaeen D, Alashqur A(2014) Using decision tree classification to assist in the prediction of Alzheimer’s disease. 6th International conference on computer science and information technology (CSIT), Amman, pp. 122-126
Alzheimer’s Disease Fact Sheet, National Institute of Aging, U.S. Department of Health and Human Services. 2015. https://www.nia.nih.gov/health/alzheimers-disease-fact-sheet
Anand KS, Dhikav V (2012) Hippocampus in health and disease: an overview. Ann Indian Acad Neurol 15(4):239–46
Ashburner M et al (2000) Gene ontology: tool for the unification of biology. Nat Genet 25(1):25–9
Blennow K, Bogdanovic N et al (1996) Synaptic pathology in Alzheimer’s disease: relation to severity of dementia, but not to senile plaques, neurofibrillary tangles, or the ApoE4 allele. J Neural Transm 103(5):603–618
Chaudhury AR, Gerecke KM et al (2003) Neuregulin-1 and ErbB4 immunoreactivity is associated with neuritic plaques in Alzheimer disease brain and in a transgenic model of Alzheimer disease. J Neuropathol Exp Neurol 62(1):42–54
Chen Y-H, Lo RY (2017) Alzheimer’s disease and osteoporosis. Ci Ji Yi Xue za Zhi (Tzu-chi Med J) 29(3):138–142
Cheng J, Liu HP et al (2021) Machine learning compensates fold-change method and highlights oxidative phosphorylation in the brain transcriptome of Alzheimer’s disease. Sci Rep 11:13704
Crow M, Lim N et al (2019) Predictability of human differential gene expression. Proc Natl Acad Sci 116(13):6491–6500
Duff MC, Covington NV et al (2020) Semantic memory and the hippocampus: revisiting, reaffirming, and extending the reach of their critical relationship. Front Hum Neurosci 13:471
Hall M (2000) Correlation-based feature selection for machine learning. Dep Comput Sci 19
Hu R-T, Yu Q et al (2020) Co-expression network analysis reveals novel genes underlying Alzheimer’s disease pathogenesis. Front Aging Neurosci 12:432
Huang DW et al (2007) The DAVID gene functional classification tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol 8(9):R183
Keaney J et al (2019) Inhibition of Bruton’s tyrosine kinase modulates microglial phagocytosis: therapeutic implications for Alzheimer’s disease. J Neuroimmune Pharmacol Off J Soc NeuroImmune Pharmacol 14(3):448–461
Kelly BL, Ferreira A (2007) $A\beta $ disrupted synaptic vesicle endocytosis in cultured hippocampal neurons. Neuroscience 147(1):60–70
Kiecolt-Glaser JK, Marucha PT et al (1995) Slowing of wound healing by psychological stress. Lancet 346(8984):1194–1196
Kuang J, Zhang P, Cai T et al (2021) Prediction of transition from mild cognitive impairment to Alzheimer’s disease based on a logistic regression-artificial neural network-decision tree model. Geriatr Gerontol Int 21(1):43–47
Lanoiselée H-M et al (2017) APP, PSEN1, and PSEN2 mutations in early-onset Alzheimer disease: a genetic screening study of familial and sporadic cases. PLoS Med 14(3):e1002270
Magistri M et al (2015) Transcriptomics profiling of Alzheimer’s disease reveal neurovascular defects, altered amyloid-$\beta $ homeostasis, and deregulated expression of long noncoding RNAs. J Alzheimer’s Dis JAD 48(3):647–665
Martin D, Brun C, Remy E et al (2004) GOToolBox: functional analysis of gene datasets based on Gene Ontology. Genome Biol 5:R101
Mukaka MM (2012) Statistics corner: a guide to appropriate use of correlation coefficient in medical research. Malawi Med J J Med Assoc Malawi 24(3):69–71
Nishiwaki K, Kanamori K, Ohwada H (2016) Finding a disease-related gene from microarray data using random forest. IEEE 15th international conference on cognitive informatics and cognitive computing (ICCICC), Palo Alto, pp. 542-546
Norstrom EM et al (2010) Identification of NEEP21 as a $\beta $-amyloid precursor protein-interacting protein in vivo that modulates amyloidogenic processing in vitro. J Neurosci Off J Soc Neurosci 30(46):15677–15685
Pinner E et al (2017) CD44 splice variants as potential players in Alzheimer’s disease pathology. J Alzheimer’s Dis JAD 58(4):1137–1149
Ramaswamy R et al (2021) Feature selection for Alzheimer’s gene expression data using modified binary particle swarm optimization. IETE J Res 2021:1–12
Ray M, Zhang W (2010) Analysis of Alzheimer’s disease severity across brain regions by topological analysis of gene co-expression networks. BMC Syst Biol 4:136
Ray M, Yunis R, Chen X, Rocke DM (2012) Comparison of low and high dose ionizing radiation using topological analysis of gene co-expression networks. BMC Genomics 13(1):190
Ray S et al (2017) A comprehensive analysis on preservation patterns of gene co-expression networks during Alzheimer’s disease progression. BMC Bioinform 18(1):579
Ruan J, Zhang W (2006) Identification and evaluation of functional modules in gene co-expression networks. Syst Biol Comput Proteomics Lecture Notes Comput Sci 4532(1):57–76
Saputra RA et al (2020) Detecting Alzheimer’s disease by the decision tree methods based on particle swarm optimization. J Phys Conf Ser 1641:012025
Seifert B, Eckenstaler R et al (2016) Amyloid-beta induced changes in vesicular transport of BDNF in hippocampal neurons. Neural Plast 2016:4145708
Sharma A, Dey P (2021) A machine learning approach to unmask novel gene signatures and prediction of Alzheimer’s disease within different brain regions. Genomics 113(4):1778–1789
Subramanian A, Tamayo P et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci 102(43):15545–15550
Swamy K et al (2021) Protein complexes form a basis for complex hybrid incompatibility. Front Genet 12:144
Takahiro K, Kazutaka N, Hayato O (2016) Finding unknown disease-related genes by comparing random forest results to secondary data in medical science study. Proceedings of the 7th international conference on computational systems-biology and bioinformatics (CSBio ’16), pp. 24-27
Watt NT et al (2010) The role of Zinc in Alzheimer’s disease. Int J Alzheimer’s Dis 2011:971021
Woo R-S et al (2011) Expression of ErbB4 in the neurons of Alzheimer’s disease brain and APP/PS1 mice, a model of Alzheimer’s disease. Anat Cell Biol 44(2):116–27
Wu Y, Zhang S et al (2016) Regulation of global gene expression and cell proliferation by APP. Sci Rep 6:22460
Xia J, Rocke DM, Perry G, Ray M (2014) Differential network analyses of Alzheimer’s disease identify early events in Alzheimer’s disease pathology. Int J Alzheimer’s Dis. https://doi.org/10.1155/2014/721453
Zhao C, Wang Z (2018) GOGO: an improved algorithm to measure the semantic similarity between gene ontology terms. Sci Rep 8:15107
Zhou S (2008) Probability theory and mathematical statistics, 4th edn. Higher Education Press, Beijing
Zhu G, Yang P (2016) Identifying the candidate genes for Alzheimer’s disease based on the rejection region of T-test. International conference on machine learning and cybernetics (ICMLC), Jeju, vol. 2, pp. 732–736
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sahu, S., Dholaniya, P.S. & Rani, T.S. Identifying the candidate genes using co-expression, GO, and machine learning techniques for Alzheimer’s disease. Netw Model Anal Health Inform Bioinforma 11, 10 (2022). https://doi.org/10.1007/s13721-021-00349-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13721-021-00349-9