Abstract
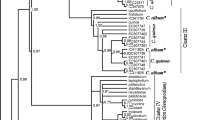
Korean Chrysanthemum species naturally distributed throughout Korea is regarded as one of the most popular ornamentals and medicinal herbs. Phylogenetic studies were conducted to evaluate interspecific relationships in 19 populations of Korean Chrysanthemum species including 16 taxa and two outgroups using internal transcribed spacer (ITS1, ITS 2) region sequences of nuclear ribosomal DNA (nrDNA) and chromosome number. Phylogenetic analysis was conducted using strict consensus methods. The taxa have well-developed series of polyploids (4x, 6x, 8x) from diploids (2x) with basic chromosome number (n = 9). The length of ITS sequences aligned was 663–664 bp, and the length of ITS 1 and ITS 2 regions were 268–269 bp and 231 bp, respectively. The divergence rate was very low within Chrysanthemum zawadskii subspecies from 0.0 to 1.4%. In the phylogenetic tree obtained by strict consensus analyses, the taxa were divided into three major groups. The C. lineare appeared as the most basal clade within the Korean Chrysanthemum. This result suggest that Chrysanthemum morphologically independent group such as C. zawadskii and C. indicum subspecies were unresolved with ITS region sequences. These findings contribute to our understanding of the phylogeny, origin, and classification of Chrysanthemum species. Chrysanthemum species could have been derived from the same ancestor to advance polypoidy for the phylogenetic analysis based on ITS sequence and chromosome data.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Literature Cited
Abd El-Twab, M.H., A.M. Mekawy, and M.S. El-Katatny. 2008. Karyomorphological studies of some species of Chrysanthemum sensu lato in Egypt. Chromosome Bot. 3:41–47.
Abd El-Twab, M.H. and K. Kondo. 2003. Physical mapping of 45S rDNA loci by fluorescent in situ hybridization and evolution among polyploidy Dendranthema species. Chromosome Sci. 7:71–76.
Abd El-Twab, M.H. and K. Kondo. 2006. FISH physical mapping of 5S, 45S and Arabidopsis-type telomere sequence repeats in Chrysanthemum zawadskii showing intra-chromosomal variation and complexity in nature. Chromosome Bot. 1:1–5.
Ahn, S.Y., K.S. Cho, K.O. Yoo, and J.T. Suh. 2010. Phylogenetic relationship of Ligularia species based on RAPD and ITS sequences analyses. Kor. J. Hort. Sci. Technol. 28:638–647.
Baldwin, B.G., M.J. Sanderson, J.M. Porter, M.F. Wojciechowski, C.S. Campbell, and M.J. Donoghue. 1995. The ITS region of nuclear ribosomal DNA: A valuable source of evidence on angiosperm phylogeny. Ann. Missouri Bot. Garden 82:247–277.
Dowrick, G.J. 1952. The Chromosome of Chrysanthemum 1. The species. Heredity 6:365–375.
Felsenstain, J. 1985. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39:783–791.
Hong, S.Y., K.S. Cho, and K.O. Yoo. 2012. Phylogenetic analysis of Korean native Aster plants based on internal transcribed spacer (ITS) sequences. Kor. J. Hort. Sci. Technol. 30:178–184.
Hwang, Y.J., A. Younis, K.B. Ryu, K.B. Lim, C.H. Eun, J.H. Lee, S.H. Sohn, and S.J. Kwon. 2013. Karyomorphological analysis of wild Chrysanthemum boreale in Korea. Flower Res. J. 21:182–189.
Iwatsuki, K., Y. Takasi, E.B. David, and O. Hideaki. 1997. Flora of Japan. III. Angiospermae Dicotyledoneae Sympetalae. Kodansha Ltd., Tokyo, Japan.
Inceer, H. and S. Hayirlioglu-Ayaz. 2010. Chromosome numbers in Tripleurospermum Sch. Bip. (Asteraceae) and closely related genera: Relationships between ploidy level and stomatal length. Plant Syst. Evol. 285:149–157.
Kim, J.S. 1999. Phytogenetic relationship among Korean Dendranthema using the analysis of chromosome characters and cpDNA intergenic region sequences. MS Diss., Kyungpook Natl. Univ., Daegu, Korea p. 6–78.
Kim, J.S. 2003. A study of diversity and speciation in Dendranthema zawadskii and its related species (Asteraceae). PhD. Diss., Kyoto Univ., Japan p. 1–74.
Kim, S.J., C.H. Lee, J. Kim, and K.S. Kim. 2014. A Phylogenetic analysis of Korean native Chrysanthemum species based on morphological characteristics. Sci. Hort. 175:278–289.
Kimura, M. 1980. A simple method for estimating evolutionary rate base substitution comparative studies nucleotide sequences. J. Mol. Evol. 16:111–120.
Kondo, K., M.H. Abd El-Twab, R. Idesawa, S. Kimura, and R. Tanaka. 2003. Genome phyogenetics in Chrysanthemum sensulato, p. 117–200. In: A.K. Sharma and A. Sharma (eds). PlantGenome: Biodiversity and evolution, Vol. 1A, Phanerogams. Plymouth, Enfield.
Korea National Arboretum (KNA). 2013. Korea arboretum. http://www.kna.go.kr.
Korea Seed and Variety Service (KSVS). 2012. Searching plant variety protection database. http://www.seed.go.kr.
Lau, D.T., P.C. Shaw, J. Wang, and P.P. But. 2001. Authentication of medicinal Dendrobium species by the internal transcribed spacer of ribosomal DNA. Planta Med. 67:456–460.
Lee, Y.N. 1975. Taxonomic study on white flowered wild chrysanthemum in Asia. Kor. J. Plant Res. 14:63–79.
Lee, Y.N. 2006. New flora of Korea II. Kyohaksa, Seoul, Korea.
Lee, C.H. and K.S. Kim. 2000. Genetic diversity of Chrysanthemum zawadskii Herb. and the related groups in Korea using RAPD. J. Kor. Soc. Hort. Sci. 41:230–236.
Lee, J.H., C.B. Park, C.G. Park, and S.G. Moon. 2010. A phylogenetic analysis of Korean Artemisia L. based on ITS sequences. Kor. J. Plant Res. 23:293–302.
Masuda, Y., T. Yukawa, and K. Kando. 2009. Molecular phylogenetic analysis of members of Chrysanthemum and its related genera in the tribe Anthemideae, the Asteaceae in East Asia on the basis of the internal transcribed spacer (ITS) region and the external transcribed spacer (ETS) region of nrDNA. Chromosome Bot. 4:25–36.
Ministry for Food, Agriculture, Forestry, and Fisheries (MIFAFF). 2012. The present condition of floriculture cultivation. MIFAFF, Seoul, Korea.
Nakata, M., R. Tanaka, K. Taniguchi, and N. Shimotomai. 1987. Species of wild Chrysanthemum in Japan: Cytological and cytogenetically views on its entity. Acta Phytotax. Geobot. 38:241–259.
Nicolson, D.H. 1999. Report of the general committee: 8. Taxon 48:373–378.
Oberprieler, C., R. Vogt, and L.E. Watson. 2007. XVI. Tribe Anthemideae cass, p. 342–374. In: J.W. Kadereit and C. Jeffrey (eds.). The families and genera of vascular plants VIII. Flowering plants Eudicots, Asterales (ed.). Heidelberg and Berlin: Springer-Verlag.
Palibin, J.W. 1898. Conspectus florae Koreae (I). Act. Hot. Petrop. 114–123.
Park, K.R. and J.H. Lee. 2004. The genetic evidence for polyploid origin of Chrysanthemum zawadskii complex. J. Basic Sci. 9:99–112.
Saitou, N. and M. Nei. 1987. The Neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4:406–425.
Soltis, D.E., P.S. Soltis, J.C. Pires, A. Kovarik, and J.A. Tate, and E. Mavrodiev. 2004. Recent and recurrent polyploidy in Tragopogon (Asteraceae): Cytogenetic, genomic and genetic comparisons. Biol. J. Linn. Soc. 82:485–501.
Suh, Y., L.B. Thien, H.E. Reeve, and E.A. Zimmer. 1993. Molecular evolution and phylogenetic implications of internal sequences of nuclear ribosomal DNA in Winteraceae. Amer. J. Bot. 80:1042–1055.
Swofford, D.L. 1998. PAUP: Phylogenetic analysis using parsimony and other methods. Ver. 4.02b Sinauer Asso. Inc., Sunderland, Massachusetts, USA.
Trehane, P. 1995. Proposal to conserve Chrysanthemum L. with a conserved type (Compositae). Taxon 44:439–441.
Tsai, C.C., C.I. Peng, S.C. Huang, P.L. Huang, and C.H. Chou. 2004. Determination of the genetic relationship of Dendrobium species (Orchidaceae) in Taiwan based on the sequence of the internal transcribed spacer of ribosomal DNA. Sci. Hort. 101:315–325.
Yapwattanaphun, C., S. Subhadrabandhu, C. Honsho, and K. Yonemori. 2004. Phylogenetic relationship of Mangosteen (Garcinia mangostana) and several wild relatives (Garcinia spp.) revealed by ITS sequence data. J. Amer. Soc. Hort. Sci. 129:368–373.
Zhao, H.E., Z.H. Liu, X. Hu, J.L. Yin, W. Li, G.Y. Rao, X.H. Zhang, C.L. Huang, N. Anderson, Q.X. Zhang, and J.Y. Chen. 2009. Chrysanthemum genetic resources and related genera of Chrysanthemum collected in China. Genet. Res. Crop. Evol. 56:937–946.
Zhao, H.B., F.D. Chen, S.M. Chen, G.S. Wu, and W.M. Guo. 2010. Molecular phylogeny of Chrysanthemum, Ajania and its allies (Anthemideae, Asteraceae) as inferred from nuclear ribosomal ITS and chloroplast trnL-F IGS sequences. Plant Syst. Evol. 284:153–169.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, S.J., Cho, K.S., Yoo, K.O. et al. Sequence analysis of the internal transcribed spacer (ITS) region of the nuclear ribosomal DNA (nrDNA) Chrysanthemum species in Korea. Hortic. Environ. Biotechnol. 56, 44–53 (2015). https://doi.org/10.1007/s13580-015-0085-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-015-0085-2