Abstract
CLLU1, a disease-specific gene associated with chronic lymphoid leukemia (CLL), is located on chromosome 12q22. Previous studies considered CLLU1 to be a non-coding RNA; however, recent research has discovered that its coding sequence region possesses the potential to encode a short peptide similar to interleukin-4. Remarkably, abnormally elevated expression of CLLU1 has only been detected in chronic lymphoid leukemia among all hematological cancers. High CLLU1 expression often indicates more malignant pathological features and an unfavorable prognosis for patients. Importantly, the expression level of CLLU1 remains unaffected by the passage of time or therapeutic interventions, thus rendering it a novel prognostic marker. This article provides a comprehensive summary of relevant research findings on CLLU1 in the context of CLL prognosis and clinical applications, aiming to guide subsequent theoretical and clinical investigations in this field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chronic lymphoid leukemia (CLL) is the most prevalent type of leukemia in adults, characterized by an abnormal increase in dysfunctional mature B lymphocytes [1, 2]. It primarily affects the elderly population, with a median age at diagnosis of 70 years, and shows a higher incidence among men [3, 4]. Clinical presentation of CLL exhibits substantial heterogeneity [5], leading to a wide range of survival outcomes, varying from months to decades [6, 7].
The current clinical staging systems for CLL, namely Rai and Binet, face limitations in accurately predicting cancer progression and prognosis at early stages. As a result, the search for more precise prognostic markers is a crucial direction in current clinical research [5, 8, 9]. Nevertheless, representative CLL prognostic marker genes have yet to be identified. The advancement of cytogenetics [10, 11], molecular biology, and immunology has contributed to the discovery of numerous factors associated with CLL prognosis [12]. Notably, current research has highlighted various biological markers with significant prognostic value, including gene mutations, chromosomal abnormalities, immune markers, and serum biochemical indicators [3, 13,14,15].
The immunoglobulin heavy chain variable region (IgVH) gene undergoes mutations in at least 50% of CLL patients [16]. In 2006, Chronic Lymphocytic Leukemia Up-regulated 1 (CLLU1) was identified as the first disease-specific gene in CLL. Notably, significant differential expression of CLLU1 was observed between CLL cases with IgVH gene mutations and those without [17]. CLLU1 is located on chromosome 12q22 and comprises three exons, flanked by BTG1 and EEA1. It encodes six mRNA transcripts that do not exhibit sequence homology with any known genes. Among these transcripts, CLLU1-203 and the coding sequence (CDS) display the highest expression levels. The majority of these transcripts cluster on chromosome 12q22, with most being non-coding, while a few, such as cDNA 4 and 5, potentially encode a peptide similar to interleukin-4 (IL-4). The CDS likely encodes a short peptide chain consisting of 121 amino acids (Fig. 1) [13, 17,18,19,20,21].
CLLU1 transcript structure. A Genomic localization and transcript variants of CLLU1. CLLU1 is situated on chromosome 12q22 in close proximity to BTG1 and EEA1. Buhl AM et al. identified seven distinct cDNAs, namely cDNA 1, cDNA 1a, cDNA 2, cDNA 3, cDNA 4, cDNA 5, and cDNA 6, through the reverse transcription of CLLU1 mRNA. Notably, transcripts cDNA 4 and cDNA 5 are noteworthy as they encompass coding sequences (CDS) capable of encoding proteins. B Structure of the CDS-encoded IL-4-like short peptide. The structure of a short peptide has been accurately predicted using AlphaFold, a cutting-edge computational tool available at alphafold.ebi.ac.uk
In recent years, CLLU1 has emerged as a prognostic marker contributing to the prediction of CLL disease activity and prognosis. It has become a research focal point regarding CLL prognostic factors. However, there is currently no existing review summarizing the research on CLLU1 in CLL. Therefore, this article aims to provide an overview of the feasibility and future research directions concerning CLLU1 as a prognostic marker and therapeutic target in CLL.
CLLU1 is a specific gene associated with CLL
CLLU1 is the first disease-specific gene identified in CLL [5, 22]. High expression of CLLU1 is exclusive to CLL patients and not observed in other hematological malignancies [20, 22]. CLLU1 expression in B cells of CLL patients is significantly higher compared to normal B cells [18, 22]. The expression of CLLU1 in CLL patients showed a significant increase compared to normal B cells, with the transcripts CLLU1-203 and the CDS region transcripts being the predominant forms. CLLU1 content was notably higher in wild-type (WT)-IgVH CLL patients compared to mutant (mut)-IgVH patients, correlating with a poorer prognosis in WT-IgVH CLL patients [17]. CLLU1 expression remains stable over time and is unaffected by therapeutic interventions, making it an intrinsic marker of CLL clones [22].
Furthermore, abnormal CLLU1 expression has been observed during the early onset of CLL, serving as an independent predictor of prognosis for CLL patients of different ages [18]. The detection of CLLU1 content does not necessitate excessive purification of blood or bone marrow samples [23], as accurate results can be obtained through direct PCR detection [13, 22]. This method is also applicable for detecting residual CLL cells after treatment [24]. The simplicity of the detection method suggests potential widespread use in routine hospital settings [25].
To explore the potential aberrant expression profile of CLLU1 in other cancers, we downloaded the expression levels (TPM) of CLLU1 from the UCSC database (https://xenabrowser.net/) for 33 tumors in the TCGA dataset. Corresponding tissue expression levels from the GTEx database were included as control samples. After removing data with zero expression and applying log2 (TPM + 0.001) transformation, we grouped the sample data into tumor and normal groups. Using the unpaired Wilcoxon test (R version 4.3.1), we calculated the significance of CLLU1 expression differences between normal and tumor samples for each cancer type and marked them with asterisks based on their significance [26]. As shown in Fig. 2A, CLLU1 expression levels were significantly downregulated in four cancers (ESCA, KIRC, OV, and THCA) (p < 0.05) and significantly up-regulated in three cancers (COAD, DLBC, and LAML) (p < 0.05). However, no significant changes in CLLU1 expression were observed in 24 cancers (ALL, BLCA, BRCA, CESC, CHOL, GBM, HNSC, KICH, KIRP, LGG, LIHC, LUAD, LUSC, PAAD, PCPG, PRAD, READ, SKCM, STAD, TGCT, THYM, UCEC, UCS, and UVM). In MESO and SARC, no comparison was made due to the lack of reference data for CLLU1 expression in corresponding normal tissues in the database.
The diagnostic, prognostic, and therapeutic value of CLLU1. A *** indicates adjusted-p < 0.001. ** indicates adjusted-p < 0.01. * indicates adjusted-p < 0.05. "ns" indicates no significant difference. CLLU1 showed significant downregulation in four cancers (ESCA, KIRC, OV, THCA) (p < 0.05) and significant upregulation in three cancers (COAD, DLBC, LAML) (p < 0.05), while no significant changes were observed in 24 cancers (ALL, BLCA, BRCA, CESC, CHOL, GBM, HNSC, KICH, KIRP, LGG, LIHC, LUAD, LUSC, PAAD, PCPG, PRAD, READ, SKCM, STAD, TGCT, THYM, UCEC, UCS, and UVM). In THCA, high expression of CLLU1 was associated with a higher risk of overall survival (B). In LIHC, DLBC, and UCEC, high expression of CLLU1 was a risk factor for progression-free interval (C). In UCEC, high expression of CLLU1 was a risk factor for disease-free interval, while in LGG, it was a protective factor for disease-free interval (D). In DLBC, patients with high expression of CLLU1 were at a higher risk for disease-specific survival (E). F The x-axis indicates the correlation between the drug and the expression of CLLU1. Positive values in green indicate a positive correlation, while negative values in red indicate a negative correlation. The sensitivity of 13 drugs was positively correlated with high expression of CLLU1, including Megestrol acetate, BAY-876, Isotretinoin, Imiquimod, Fluphenazine, Zoledronate, S-63845, CB-839, Nandrolone phenpropionate, Acetylcysteine, JNJ-54302833, HPI-1, and AMG-176. There were four drugs whose sensitivity was negatively correlated with high expression of CLLU1, including VS-4718, 6-Thioguanine, Veliparib, and LOR-253. OS, overall survival; DSS, disease-specific survival; PFI, progression-free interval; DFI, disease-free interval; HR, hazard ratio. Please check the following link for the full name of TCGA abbreviations. Please check the following link for the full name of TCGA abbreviations. Link: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations
CLLU1 as a prognostic marker in CLL
Prognostic markers play a crucial role in identifying high-risk patients and facilitating timely intervention for better outcomes [27]. CLLU1 has emerged as a novel prognostic marker with potential value in CLL [14]. Patients with high CLLU1 expression exhibit significantly worse overall survival (OS) and progression-free survival (PFS) compared to those with low expression [14, 18]. Notably, in patients with IgVH gene mutations, the 6-year survival rate is significantly lower in those with high CLLU1 expression (50%) compared to those with low expression (76%) [7, 14]. Josefsson et al. reported a median survival time of 5.0 years for CLL patients with CLLU1 expression higher than 40 times, compared to 8.1 years for patients with lower expression [18]. Subsequent studies have consistently demonstrated a negative correlation between CLLU1 expression and CLL prognosis, with high expression levels associated with shorter treatment duration and lower overall survival rates [3, 13]. Assessment of blood CLLU1 levels serves as a reliable marker of tumor burden, offering independent predictive power [3, 9]. Furthermore, CLLU1 expression analysis complements current techniques for minimal residual disease (MRD) monitoring in CLL patients [28, 29]. High CLLU1 expression is also significantly associated with unfavorable pathological features, including WT-IgVH gene, ZAP-70 positivity, CD38 positivity, and 13q deletion [5, 13, 14].
To investigate whether CLLU1 has a similar prognostic value in other cancers, we obtained patient survival data [OS, disease-specific survival (DSS), progression-free interval (PFI), and disease-free interval (DFI)] from TCGA Pan-Cancer (PANCAN). Using the one-way COX test, we calculated the risk score of CLLU1 across different cancer types and depicted the results in Fig. 2B–E. In THCA, high CLLU1 expression was associated with a higher hazard ratio (HR) for OS (p < 0.05). Similarly, in DLBC, high CLLU1 expression correlated with a higher HR for DSS (p < 0.05). Moreover, in LIHC, DLBC, and UCEC, high CLLU1 expression was linked to a higher HR for PFI (p < 0.05). In UCEC, high CLLU1 expression was associated with a higher HR for DFI, while LGG exhibited a lower HR for DFI (p < 0.05).
CLLU1 and other biomarkers in CLL
The addition of additional prognostic markers in CLL can enhance risk prediction by reducing the impact of tumor heterogeneity [30]. We compared CLLU1 with other biomarkers in Table 1. The Rai and Binet staging systems are widely used for prognostic assessment in CLL, based on physical examination and categorization of patients according to the number of involved lymphatic areas [31]. However, these staging systems have limitations, as more than 70% of patients classified as low risk at diagnosis still progress. Therefore, while these systems serve as immediate prognostic indicators at the time of diagnosis, they cannot predict disease progression or survival time after progression. CLLU1 expression was found to be correlated with Rai and Binet staging system, with lower CLLU1 levels observed in patients with Binet A stage compared to Binet B and C patients [7, 13].
Table 2 and Fig. 3 display the current prognostic indicators of CLL, including four categories: gene mutation, chromosomal abnormality, immune markers, and serum biochemical indicators. To date, only two molecules have been identified with CLL-specific genetic mutation signatures: the IgVH structure and CLLU1 expression level [22]. CLL can be divided into two types based on the presence or absence of somatic mutations in the IgVH gene: mutated and unmutated. Patients with IgVH mutation have slower disease progression, longer median survival time, and treatment time, whereas CLL patients without IgVH gene mutation exhibit rapid disease progression, poor prognosis, and shorter survival time [5, 22]. However, the IgVH gene serves as a late prognostic indicator and cannot predict early outcomes. Studies have shown that patients without IgVH gene mutations have higher levels of CLLU1 [7, 13]. Regarding chromosomal abnormalities, del(17p) and del(11q) have been identified as markers of advanced disease [18], and high-risk cytogenetic features increase the risk of early death [18]. However, these abnormalities do not show significant correlation with CLLU1 and are considered independent prognostic factors.
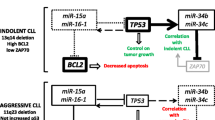
Commonly used molecular markers in CLL. ZAP-70 plays a crucial role in BCR signaling, promoting enhanced proliferation through pathways, such as NF-κB signaling and CD38, a transmembrane glycoprotein. Loss or mutation of TP53 and chromosomal abnormalities involving 11q- and 17p-disrupt cell repair mechanisms. Although the exact reasons for the elevated expression of LPL and CLLU1 in CLL remain unknown, these markers are associated with increased risk in patients
In terms of immune markers, CD38 expression in CLL was initially considered as a surrogate marker for IgVH gene analysis, but subsequent studies showed insufficient correlation with IgVH mutation status [32]. Elevated CD38 levels (over 30%) are associated with shorter survival [5, 33]. Multiple studies have reported higher CLLU1 levels in CD38-positive patients [7, 13, 14]. ZAP-70, initially recognized as one of the most differentially expressed genes between mutated and unmutated IgVH gene CLL cases, was proposed as a surrogate marker for IgVH mutations [34]. ZAP-70 aids in distinguishing patients requiring early treatment [5, 35, 36]. However, the correlation between ZAP-70 and CLLU1 is not limited to CLL [3, 13]. ZAP-70-positive patients have been shown to have higher CLLU1 levels [13, 14] although there are reports suggesting no association between CLLU1 and ZAP-70 [7].
There are currently limited studies on serum biochemical indicators in CLL, with only a mention of significant differences in lipoprotein lipase (LPL) expression between patients with wild-type IgVH and mutated CLL [37]. Multiple studies have confirmed high expression of LPL in wild-type IgVH CLL compared to mutated CLL.
CLLU1 and CLL treatment
Determining the optimal timing for CLL treatment is crucial due to the heterogeneity of patients, with some requiring early treatment and others not needing it in the early stages [4, 5]. Bubl et al. [13] conducted a Cox multiple regression analysis to assess the relationship between CLLU1 expression levels and the initiation of CLL therapy. The study revealed that doubling the expression of cDNA1 transcripts increased the risk of early treatment by 19%, while doubling the amount of CDS increased the risk by 47%. Patients with cDNA1 levels above the median had a median time to treatment initiation of 1.19 years, compared to 8.6 years for those with levels below the median. Patients with higher CDS levels started treatment at a median of 1.19 years, whereas those with lower CDS levels initiated treatment after 13 years [13]. Josefsson et al. found that patients with high CLLU1 expression had a median time to first treatment of 9.0 months, while patients with low CLLU1 expression had a median time to first treatment of 4.6 years [18]. These findings indicate that patients with high CLLU1 expression require early clinical intervention to achieve molecular remission, thereby improving survival rates and prolonging the overall survival period. Additionally, Josefsson et al. [18]observed that each doubling of CLLU1 expression was associated with a 7% increase in the risk of premature death. The expression level of CLLU1 can predict the timing of CLL treatment initiation, and patients with high CLLU1 expression should be closely monitored for clinical changes that may necessitate early intervention.
To further investigate the association between CLLU1 and clinical treatment, we performed a drug sensitivity analysis. Using the CellMiner database (http://discover.nci.nih.gov/cellminer/) [38], we obtained RNA expression data and relevant drug information. Missing values in the supplementary drug susceptibility data were imputed using the nearest neighbor mean. Pearson correlation coefficients were calculated to assess the relationship between gene expression and different drugs. The analysis results were screened based on a significance threshold of P value < 0.05. As depicted in Fig. 2F, high expression of CLLU1 was positively correlated with the sensitivity of 13 drugs, including Megestrol acetate, BAY-876, Isotretinoin, Imiquimod, Fluphenazine, Zoledronate, S-63845, CB-839, Nandrolone phenpropionate, Acetylcysteine, JNJ-54302833, HPI-1, and AMG-176. Conversely, four drugs showed a negative correlation between sensitivity and high expression of CLLU1, namely VS-4718, 6-Thioguanine, Veliparib, and LOR-253.
Concluding remarks and future perspectives
CLLU1 stands as a specific marker for CLL, expressed early in the disease and exhibiting relative stability. Evaluating the blood CLLU1 level can serve as a monitoring indicator for early treatment of CLL patients, with its expression level closely linked to prognosis, often indicating a poor prognosis for patients with high CLLU1 expression. Assessing the blood CLLU1 level provides a reliable marker of tumor burden and addresses the gap in MRD monitoring technology for CLL patients. Future studies can incorporate blood tests for CLLU1 to detect residual CLL cells post-treatment, monitor molecular responses to therapy, and guide decisions regarding consolidation or maintenance therapy. This approach will ultimately enhance our understanding of the interplay between treatment response kinetics, disease relapse, and long-term survival.
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
Abbreviations
- CDS:
-
Coding sequence
- CLL:
-
Chronic lymphoid leukemia
- CLLU1:
-
Chronic lymphocytic leukemia up-regulated 1
- DFI:
-
Disease-free interval
- DSS:
-
Disease-specific survival
- HR:
-
Hazard ratio
- IgVH:
-
Immunoglobulin heavy chain variable region
- IL-4:
-
Interleukin-4
- LPL:
-
Lipoprotein lipase
- MRD:
-
Minimal residual disease
- Mut:
-
Mutant
- OS:
-
Overall survival
- PFI:
-
Progression-free interval
- PFS:
-
Progression-free survival
- WT:
-
Wild-type
References
Hallek M, Al-Sawaf O. Chronic lymphocytic leukemia: 2022 update on diagnostic and therapeutic procedures. Am J Hematol. 2021;96:1679–705. https://doi.org/10.1002/ajh.26367.
Pekarsky Y, Zanesi N, Croce CM. Molecular basis of CLL. Semin Cancer Biol. 2010;20:370–6. https://doi.org/10.1016/j.semcancer.2010.09.003.
Sevov M, Rosenquist R, Mansouri L. RNA-based markers as prognostic factors in chronic lymphocytic leukemia. Expert Rev Hematol. 2012;5:69–79. https://doi.org/10.1586/ehm.11.80.
Zhang S, Kipps TJ. The pathogenesis of chronic lymphocytic leukemia. Annu Rev Pathol. 2014;9:103–18. https://doi.org/10.1146/annurev-pathol-020712-163955.
Sevinc M, Karabulut A, Eskazan AE, Catal Tatonyan S, Ozbek U, Soysal T. The impact and prognostic significance of chronic lymphocytic leukemia upregulated 1 (CLLU1) gene expression in patients with chronic lymphocytic leukemia: a single center experience. Lab Med. 2020;51:259–64. https://doi.org/10.1093/labmed/lmz058.
Caligaris-Cappio F, Ghia P. The nature and origin of the B-chronic lymphocytic leukemia cell: a tentative model. Hematol Oncol Clin North Am. 2004;18:849–62. https://doi.org/10.1016/j.hoc.2004.04.004.
Chen L, Li J, Zheng W, Zhang Y, Wu Y, Li L, et al. The prognostic evaluation of CLLU1 expression levels in 50 Chinese patients with chronic lymphocytic leukemia. Leuk Lymphoma. 2007;48:1785–92. https://doi.org/10.1080/10428190701534416.
Binet JL, Auquier A, Dighiero G, Chastang C, Piguet H, Goasguen J, et al. A new prognostic classification of chronic lymphocytic leukemia derived from a multivariate survival analysis. Cancer. 1981;48:198–206. https://doi.org/10.1002/1097-0142(19810701)48:1%3c198::aid-cncr2820480131%3e3.0.co;2-v.
Bosch F, Dalla-Favera R. Chronic lymphocytic leukaemia: from genetics to treatment. Nat Rev Clin Oncol. 2019;16:684–701. https://doi.org/10.1038/s41571-019-0239-8.
Tausch E, Schneider C, Robrecht S, Zhang C, Dolnik A, Bloehdorn J, et al. Prognostic and predictive impact of genetic markers in patients with CLL treated with obinutuzumab and venetoclax. Blood. 2020;135:2402–12. https://doi.org/10.1182/blood.2019004492.
Deaglio S, Vaisitti T, Aydin S, Ferrero E, Malavasi F. In-tandem insight from basic science combined with clinical research: CD38 as both marker and key component of the pathogenetic network underlying chronic lymphocytic leukemia. Blood. 2006;108:1135–44. https://doi.org/10.1182/blood-2006-01-013003.
Rawstron AC, Bennett FL, O’Connor SJ, Kwok M, Fenton JA, Plummer M, et al. Monoclonal B-cell lymphocytosis and chronic lymphocytic leukemia. N Engl J Med. 2008;359:575–83. https://doi.org/10.1056/NEJMoa075290.
Buhl AM, Jurlander J, Geisler CH, Pedersen LB, Andersen MK, Josefsson P, et al. CLLU1 expression levels predict time to initiation of therapy and overall survival in chronic lymphocytic leukemia. Eur J Haematol. 2006;76:455–64. https://doi.org/10.1111/j.0902-4441.2005.t01-1-EJH2530.x.
Gonzalez D, Else M, Wren D, Usai M, Buhl AM, Parker A, et al. CLLU1 expression has prognostic value in chronic lymphocytic leukemia after first-line therapy in younger patients and in those with mutated IGHV genes. Haematologica. 2013;98:274–8. https://doi.org/10.3324/haematol.2012.070201.
Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Dohner H, et al. iwCLL guidelines for diagnosis, indications for treatment, response assessment, and supportive management of CLL. Blood. 2018;131:2745–60. https://doi.org/10.1182/blood-2017-09-806398.
Dohner H, Stilgenbauer S, Benner A, Leupolt E, Krober A, Bullinger L, et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med. 2000;343:1910–6. https://doi.org/10.1056/NEJM200012283432602.
Buhl AM, Jurlander J, Jorgensen FS, Ottesen AM, Cowland JB, Gjerdrum LM, et al. Identification of a gene on chromosome 12q22 uniquely overexpressed in chronic lymphocytic leukemia. Blood. 2006;107:2904–11. https://doi.org/10.1182/blood-2005-07-2615.
Josefsson P, Geisler CH, Leffers H, Petersen JH, Andersen MK, Jurlander J, et al. CLLU1 expression analysis adds prognostic information to risk prediction in chronic lymphocytic leukemia. Blood. 2007;109:4973–9. https://doi.org/10.1182/blood-2006-11-054916.
Smolej L. The prognostic role of CLLU1 in chronic lymphocytic leukaemia. Eur J Haematol. 2006;77:177. https://doi.org/10.1111/j.1600-0609.2006.00694.x.
Oppliger Leibundgut E, Rogenmoser-Dissler D, de Beer D, Porret NA, Zibellini S, Durig J, et al. CLLU1 expression distinguishes chronic lymphocytic leukemia from other mature B-cell neoplasms. Leuk Res. 2012;36:1204–7. https://doi.org/10.1016/j.leukres.2012.05.015.
Grzybowska EA, Wakula M. Protein binding to cis-motifs in mRNAs coding sequence is common and regulates transcript stability and the rate of translation. Cells. 2021;10:2910. https://doi.org/10.3390/cells10112910.
Buhl AM, Novotny GW, Josefsson P, Nielsen JE, Pedersen LB, Geisler C, et al. The CLLU1 expression level is a stable and inherent feature of the chronic lymphocytic leukemia clone. Leukemia. 2009;23:1182–6. https://doi.org/10.1038/leu.2009.16.
Jebaraj BM, Kienle D, Lechel A, Mertens D, Heuberger M, Ott G, et al. Telomere length in mantle cell lymphoma. Blood. 2013;121:1184–7. https://doi.org/10.1182/blood-2012-08-452649.
Buhl AM, James DF, Neuberg D, Jain S, Rassenti LZ, Kipps TJ. Analysis of CLLU1 expression levels before and after therapy in patients with chronic lymphocytic leukemia. Eur J Haematol. 2011;86:405–11. https://doi.org/10.1111/j.1600-0609.2011.01588.x.
Matsuda K. PCR-based detection methods for single-nucleotide polymorphism or mutation: real-time PCR and its substantial contribution toward technological refinement. Adv Clin Chem. 2017;80:45–72. https://doi.org/10.1016/bs.acc.2016.11.002.
Shen J, Wang Q, Liang C, Su X, Ke Y, Mao Y, et al. Novel Insights into miR-944 in cancer. Novel Insights Cancers (Basel). 2022;14:4232. https://doi.org/10.3390/cancers14174232.
Wu L, Qu X. Cancer biomarker detection: recent achievements and challenges. Chem Soc Rev. 2015;44:2963–97. https://doi.org/10.1039/c4cs00370e.
Romero D. MRD to help assess response in CLL. Nat Rev Clin Oncol. 2019;16(2):68. https://doi.org/10.1038/s41571-018-0159-z.
Sampath D, Calin GA. miRs: fine-tuning prognosis in CLL. Blood. 2009;113:5035–6. https://doi.org/10.1182/blood-2009-02-205419.
Wos J, Szymanska A, Lehman N, Chocholska S, Zarobkiewicz M, Pozarowski P, et al. Can galectin-3 be a novel biomarker in chronic lymphocytic leukemia? Cells. 2023;3:30. https://doi.org/10.3390/cells13010030.
Hallek M. Chronic lymphocytic leukemia: 2020 update on diagnosis, risk stratification and treatment. Am J Hematol. 2019;94:1266–87. https://doi.org/10.1002/ajh.25595.
Deaglio S, Vaisitti T, Aydin S, Bergui L, D’Arena G, Bonello L, et al. CD38 and ZAP-70 are functionally linked and mark CLL cells with high migratory potential. Blood. 2007;110:4012–21. https://doi.org/10.1182/blood-2007-06-094029.
Hamblin TJ, Orchard JA, Ibbotson RE, Davis Z, Thomas PW, Stevenson FK, et al. CD38 expression and immunoglobulin variable region mutations are independent prognostic variables in chronic lymphocytic leukemia, but CD38 expression may vary during the course of the disease. Blood. 2002;99:1023–9. https://doi.org/10.1182/blood.v99.3.1023.
Crespo M, Bosch F, Villamor N, Bellosillo B, Colomer D, Rozman M, et al. ZAP-70 expression as a surrogate for immunoglobulin-variable-region mutations in chronic lymphocytic leukemia. N Engl J Med. 2003;348:1764–75. https://doi.org/10.1056/NEJMoa023143.
Wiestner A, Rosenwald A, Barry TS, Wright G, Davis RE, Henrickson SE, et al. ZAP-70 expression identifies a chronic lymphocytic leukemia subtype with unmutated immunoglobulin genes, inferior clinical outcome, and distinct gene expression profile. Blood. 2003;101:4944–51. https://doi.org/10.1182/blood-2002-10-3306.
Rassenti LZ, Huynh L, Toy TL, Chen L, Keating MJ, Gribben JG, et al. ZAP-70 compared with immunoglobulin heavy-chain gene mutation status as a predictor of disease progression in chronic lymphocytic leukemia. N Engl J Med. 2004;351:893–901. https://doi.org/10.1056/NEJMoa040857.
Rosenwald A, Alizadeh AA, Widhopf G, Simon R, Davis RE, Yu X, et al. Relation of gene expression phenotype to immunoglobulin mutation genotype in B cell chronic lymphocytic leukemia. J Exp Med. 2001;194(11):1639–47. https://doi.org/10.1084/jem.194.11.1639.
Luna A, Elloumi F, Varma S, Wang Y, Rajapakse VN, Aladjem MI, et al. CellMiner cross-database (CellMinerCDB) version 1.2: exploration of patient-derived cancer cell line pharmacogenomics. Nucleic Acids Res. 2021;49(D1083):D1093. https://doi.org/10.1093/nar/gkaa968.
Acknowledgements
We thank platforms such as PubMed for searching the literature
Funding
This study was supported by the Natural Science Foundation project of Ningbo Science and Technology Bureau (2022J040).
Author information
Authors and Affiliations
Contributions
CR, CL, JS, YZ, QW, FY, YC, YL, MG, PG, YX, and SD collected and analyzed the literature, drafted the figures and wrote the paper; YX and SD conceived and gave the final approval of the submitted version. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interests
The authors declare that they have no competing interest.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rong, C., Liang, C., Shen, J. et al. CLLU1 as an emerging biomarker in chronic lymphoid leukemia. Human Cell 37, 625–632 (2024). https://doi.org/10.1007/s13577-024-01051-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-024-01051-4