Abstract
LINC00941, also known as lncRNA-MUF, is an intergenic non-coding RNA located on chromosome 12p11.21. It actively participates in a complex competing endogenous RNA network, regulating the expression of microRNA and its downstream proteins. Through transcriptional and post-transcriptional regulation, LINC00941 plays a vital role in multiple signaling pathways, influencing cell behaviors such as tumor cell proliferation, epithelial–mesenchymal transition, migration, and invasion. Noteworthy is its consistently high expression in various tumor types, closely correlating with clinicopathological features and cancer prognoses. Elevated LINC00941 levels are associated with adverse clinical outcomes, including increased tumor size, extensive lymphatic metastasis, and distant metastasis, leading to poorer survival rates across different cancers. Additionally, LINC00941 and its associated genes are linked to various targeted drugs available in the market. In this comprehensive review, we systematically summarize existing studies, detailing LINC00941's differential expression, clinicopathological and prognostic implications, regulatory mechanisms, and associated therapeutic drugs. Our analysis includes relevant charts and incorporates bioinformatics analyses to verify LINC00941's differential expression in pan-cancer and explore potential transcriptional regulation patterns of downstream targets. This work not only establishes a robust data foundation but also guides future research directions. Given its potential as a significant cancer biomarker and therapeutic target, further investigation into LINC00941's differential expression and regulatory mechanisms is essential.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Long non-coding RNA (lncRNA) are RNA transcripts synthesized by RNA polymerase II that exceed 200 nucleotides in length and do not encode proteins [1, 2]. They have been demonstrated to play crucial roles in various biological processes, including inflammation, cell growth, and differentiation [2]. In tumor cells, lncRNAs exert significant influence on cellular processes such as differentiation, proliferation, metastasis, invasion, and apoptosis [3, 4]. Furthermore, current studies have revealed the potential of lncRNAs as biomarkers for cancer diagnosis and prognosis [5,6,7].
LINC00941, also known as lncRNA-MUF (MSC up-regulatory factor), is an intergenic non-coding RNA located on chromosome 12p11.21 in humans, with a length of 1895nt [8]. LINC00941 exhibits distinct distributions within cells: predominantly in the cytoplasm, it primarily influences post-transcriptional regulation, whereas its presence in the nucleus is associated with gene regulation [9]. It exhibits abnormally high expression levels in 12 different cancers and is closely associated with poor prognosis and clinical features in cancer patients, including degree of tumor–node–metastasis (TNM) stage and clinical stage.
Competing endogenous RNA (ceRNA) refers to a class of RNA molecules that competitively bind to microRNA (miRNA) and regulate the expression of downstream mRNA targets [10]. Increasing evidence suggests that lncRNAs can function as important ceRNAs, exerting regulatory effects [1]. LINC00941 serves as a ceRNA for five miRNAs, including miR-205-5p, miR-873-3p, miR-877-3p, miR-335-5p, and miR-34a-5p, promoting cancer cell proliferation, metastasis, invasion, and apoptosis. Additionally, LINC00941 is involved in five signaling pathways: PI3K/AKT [11, 12], TGF-β/SMAD [13, 14], Wnt/β-catenin [3, 8, 15], LIMK1/Cofilin-1 [16], and Hippo signaling pathways [17]. Moreover, LINC00941 can up-regulate the expression of downstream proteins, such as SOX2 [18], ANXA2 [8, 11], PKM [12], CAPRIN2 [3], hnRNPK [15], and YAP1 [17], by recruiting transcription factors.
This article comprehensively explores the aberrant expression, molecular mechanisms, and prognostic relevance of LINC00941 across various documented tumor types. It delves into the ceRNA regulatory network involving LINC00941 and its profound influence on the biological behavior of cancer cells. Additionally, we have broadened the scope of possible regulatory mechanisms and potential targeted drugs associated with LINC00941 by consulting online databases and conducting bioinformatics analysis. Our manuscript serves as a foundational reference, offering valuable insights for future research endeavors into the role of LINC00941 in tumor development.
Abnormal expression of LINC00941 in cancers
As depicted in Table 1, our analysis revealed the up-regulation of LINC00941 in various digestive system tumor tissues and cell lines. Specifically, elevated expression was observed in gastric cancer (GC) [19, 20], laryngocarcinoma (LC) [12, 21], and colorectal cancer (CRC) [14, 22]. Furthermore, LINC00941 exhibited high expression in cell lines of oral squamous cell carcinoma (OSCC) [3, 15] and esophageal squamous cell carcinoma (ESCC) [7]. In respiratory tumors, such as non-small cell lung cancer (NSCLC) [23] and pancreatic adenocarcinoma (PAAD) [24], LINC00941 displayed elevated expression in both corresponding tumor tissues and cell lines. Aberrantly high expression of LINC00941 was also identified in endocrine system cancer cell lines and tissues, including pancreatic ductal adenocarcinoma (PDAC) [17] and pancreatic cancer (PC) [16]. Additionally, current evidence indicates high expression of LINC00941 in a glioblastoma multiforme (GBM) cell line [13].
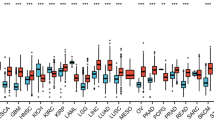
To explore the expression patterns of LINC00941 in cancer, we obtained the expression profile data of LINC00941 from the TCGA database (https://portal.gdc.cancer.gov/) (Fig. 1). We downloaded the TPM expression data of LINC00941 for 32 cancer types from the UCSC Xena database and performed a log2(TPM +1) transformation on them. We calculated the median expression of all lncRNAs and the rank percentage of LINC00941 among all lncRNAs with non-zero expression in these 32 cancer types. As shown in Fig. 1A, LINC00941 was highly expressed in 13 tumors (0.75–1.0 quantile, Q4) including cholangiocarcinoma (CHOL), head and neck squamous cell carcinoma (HNSC), PAAD, esophageal carcinoma (ESCA), stomach adenocarcinoma (STAD), adrenocortical carcinoma (ACC), lung squamous cell carcinoma (LUSC), bladder urothelial carcinoma (BLCA), sarcoma (SARC), colon adenocarcinoma (COAD), GBM, skin cutaneous melanoma (SKCM) and pheochromocytoma and paraganglioma cancer (PCPG); highly expressed in 14 types of tumors (0.5–0.75 quantile, Q3) including testicular germ cell tumors (TGCT), ovarian cancer (OV), rectum adenocarcinoma (READ), cervical esophageal squamous cell carcinoma (CESC), thyroid carcinoma (THCA), kidney renal clear cell carcinoma (KIRC), lung adenocarcinoma (LUAD), breast invasive carcinoma (BRCA), prostate Adenocarcinoma (PRAD), neuroblastoma (NBL), endometrioid cancer (UCEC), kidney papillary cell carcinoma (KIRP), uterine carcinosarcoma (UCS) and liver hepatocellular carcinoma (LIHC); and moderately expressed (0.25–0.5 quantile, Q2) in 5 types of tumors including lower grade glioma (LGG), kidney chromophobe (KICH), acute myeloid leukemia (AML), thymoma (THYM) and acute lymphoblastic leukemia (ALL). These suggest the value of LINC00941 in pan-cancer. We compared the expression difference of LINC00941 between normal and tumor samples in each cancer type. As shown in Fig. 1B, we found that LINC00941 was significantly upregulated in 18 tumor types including ALL, AML, BLCA, CHOL, COAD, ESCA, GBM, HNSC, KIRC, LIHC, LUSC, PAAD, READ, SARC, SKCM, STAD, TGCT, and THCA; significantly down-regulated in 9 tumors including ACC, BRCA, KICH, LGG, NBL, PCPG, PRAD, UCEC and UCS; and showed no significant difference in 5 tumors including CESC, KIRP, LUAD, OV and THYM. After comparing the reported tumor information, it was found that the results of 7 TCGA tumors (COAD, ESCA, GBM, LIHC, LUSC, PAAD, and THCA) were consistent with existing reports (Table S1). However, some TCGA results differ from the existing reports such as TGCA-HNSC and TGCA-LUAD. Previous studies have shown that LINC00941 is highly expressed in OSCC [7], LC [12] and NSCLC [23]. The reason for this discrepancy may be due to differences in cancer subtypes in the TCGA database, detection methods and small sample size.
A pan-cancer analysis of LINC00941 A quantile expression of LINC00941 in 32 cancer types; B LINC00941 is dysregulated in 32 cancer types. (*** means p< 0.001, ** means p < 0.01, * means p < 0.05, ns means no significant difference). Please check GDC (https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables/tcga-study-abbreviations) for the full name of the TCGA abbreviations and OCG (https://ocg.cancer.gov/programs/target/data-matrix) for the full name of the TARGET abbreviations
Furthermore, we performed a correlation analysis between the expression of LINC00941 and three types of RNA modification genes using the TCGA database. These included 10 m1A modification genes, 13 m5C modification genes, and 21 m6A modifying genes. The results revealed significant associations between LINC00941 and RNA-modifying genes in various cancers, although the specific correlations varied (Figure S2).
In conclusion, the dysregulation of LINC00941 in cancer has been broadly established. However, further investigation is required to explore its differential expression in specific cancer types by expanding the sample size and including different sample types. Additionally, it is worth noting that although a certain correlation exists, the relationship between LINC00941 and common epigenetic modifications has not been extensively studied, underscoring the need for further experimental verification.
Molecular mechanism of LINC00941 affecting cancer development
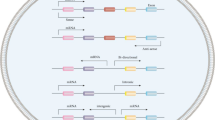
Currently, an increasing number of studies have provided evidence that lncRNAs participate in intricate regulatory networks and play a significant role in cancer pathogenesis [25]. Among them, LINC00941 has been shown to recruit transcription factors, thereby regulating the expression of six downstream factors (CTCF, YBX1, ILF2, ANXA2, PKM, and HNRNPK) at the transcriptional level. Moreover, LINC00941 functions as a ceRNA for six miRNAs, enabling the regulation of six downstream factors (including MYC, PMEPA, Snail1, VEGFA, ATXN2, and ROCK1) at the post-transcriptional level. Additionally, LINC00941 is involved in six key signaling pathways, namely the focal adhesion kinase (FAK)/AKT signaling pathway, PI3K/AKT signaling pathway, TGF-β/SMAD signaling pathway, Wnt/β-catenin signaling pathway, LIMK1/Cofilin-1 signaling pathway, and Hippo signaling pathway.
LINC00941 regulates gene expression at the transcriptional level
LncRNAs exert their influence on cancer progression by recruiting various molecular complexes, including transcription factors, RNA-binding proteins, DNA-binding proteins, or histone modification complexes, thereby modulating gene expression [26] In OSCC, the acetylation activity of EP300 is facilitated by its interaction with HAT, leading to the formation of the EP300-HAT domain. This, in turn, promotes H3K27ac modification on the LINC00941 promoter and enhances LINC00941 expression. Upregulation of LINC00941 occurs through the recruitment of the transcription factor CTCF, which induces DNA circularization via homodimerization, ultimately upregulating the expression of its downstream gene, CAPRIN2. The upregulated CAPRIN2 protein phosphorylates the WNT co-receptor LRP6, resulting in increased activity of the canonical Wnt/β-catenin signaling pathway and ultimately contributing to the development of OSCC [3, 27,28,29,30]. In ESCC, LINC00941 plays a role in promoting disease progression by directly recruiting YBX1 and the transcription factor ILF2 to the SOX2 promoter. This interaction activates the expression of SOX2 and enhances its mRNA stability, contributing to ESCC progression [18]. Similarly, in hepatocellular carcinoma (HCC), LINC00941 forms a complex with ANXA2 and disrupts the formation of the GSK-3β/β-catenin complex. This activation of the Wnt/β-catenin signaling pathway promotes HCC progression [8]. Additionally, in PC, LINC00941 competes with NEDD4L for binding to ANXA2, preventing the ubiquitination and degradation of ANXA2. This, in turn, activates the FAK/AKT signaling pathway and contributes to the progression of PC [11]. Importantly, a study demonstrated that METTL14 upregulated the m6A modification of LINC00941, leading to enhanced recognition of LINC00941 by IGF2BP2. This interaction subsequently stabilized LINC00941 and facilitated the progression of PC [31].
LINC00941 is primarily localized in the nucleus, although it can also be found in the cytoplasm [14, 18]. Within the nucleus, lncRNAs are known to recruit transcriptional regulators to target gene promoters, thereby modulating gene transcription [32]. Hence, it is plausible that LINC00941 regulates the transcription levels of ANXA2 [8], PKM [12], and HNRNPK [15] by recruiting transcription factors. To identify potential transcription factors recruited by LINC00941, we employed the Signaling Pathways Project (SPP) database [33] and the ChIP-Atlas database [34] (Fig. 2). These databases were utilized to predict regulatory transcription factors within the 2000 bp upstream regions of ANXA2, PKM, and HNRNPK. Through screening and applying certain criteria (Fold Enrichment > 50, p < 0.05 for ChIP-Atlas database, and p < 0.05 for SPP database), we identified JUN as a potential candidate for ANXA2 and PKM. To further validate the role of JUN in regulating these downstream factors, we retrieved the DNA sequences of the 2000 bp upstream of ANXA2 and PKM from the UCSC Genome Browser, with the transcription start site (TSS) as the reference point. These sequences were then analyzed using the JASPAR 2022 database [35] with a relative profile score threshold set at 80%, leading to the prediction of nine binding sites on ANXA2 and six binding sites on PKM. In summary, it is likely that LINC00941 regulates downstream gene transcription by recruiting JUN, although the specific mechanism remains to be elucidated. Further experimental verification is warranted to confirm the transcriptional regulatory role of LINC00941.
Regulation of gene expression by LINC00941 at the post-transcriptional level
CeRNA encompasses RNA transcripts containing microRNA response elements (MREs), such as mRNAs, transcribed pseudogenes, and lncRNAs [36]. CeRNAs possess the capability to modulate the expression of downstream targets by sequestering miRNAs that share the same MRE as the downstream targets [37]. When lncRNAs function as ceRNAs, they bind to miRNA-containing RNA-induced silencing complexes (miRNA-RISC), thereby diminishing the inhibitory effect of miRNAs on mRNA expression and leading to an elevation in mRNA expression levels [37].
As depicted in Fig. 3A, Table 1, and Table S2, LINC00941 exerts its regulatory role by engaging in six ceRNA axes involving five miRNAs, consequently promoting cancer progression. These axes encompass the LINC00941/miR-205-5p/MYC axis in CRC [22], the LINC00941/miR-877-3p/PMEPA axis in ESCC [22], the LINC00941/miR-34a-5p/Snail1 axis in GBM and HCC [8, 23], the LINC00941/miR-877-3p/VEGFA axis in NSCLC [23], the LINC00941/miR-873-3p/ATXN2 axis in PAAD [24], and the LINC00941/miR-335-5p/ROCK1 axis in PC [16].
Involvement of LINC00941 in multiple signaling pathways
LncRNAs play a crucial role in regulating various cellular signals in cancer, thereby influencing cancer progression [38]. As depicted in Fig. 3B, LINC00941 participates in five signaling pathways to facilitate the occurrence and advancement of cancer.
The PI3K/AKT signaling pathway is a vital pathway involved in normal cellular processes, regulating cellular glycolysis and influencing diverse cellular functions [39, 40]. Within the PI3K/AKT signaling, the PI3K/AKT/mTOR pathway plays a significant role by responding to upstream signals in cancer and regulating cell survival, metabolism, growth, and protein synthesis [41]. In LC, LINC00941 activates the PI3K/AKT/mTOR pathway, leading to upregulation of PKM expression. This activation enhances glucose uptake and conversion of glucose to pyruvate, thereby promoting glycolysis in LC cells and facilitating cancer progression [12]. FAK is a non-receptor tyrosine kinase and adapter protein that predominantly regulates adhesion signal transduction and cell migration [42]. FAK influences various physiological functions of cancer cells through its kinase and scaffold functions. It is often overexpressed in cancer and is considered a promising therapeutic target [43]. In PC, LINC00941 upregulates ANXA2 expression and activates the FAK/PI3K/AKT signaling, promoting the proliferation, invasion, and metastasis of PC cells [11].
The TGF-β/SMAD signaling pathway represents a classic SMAD-dependent pathway in TGF-β signaling. Transforming growth factor β (TGF-β) is a pleiotropic cytokine that modulates multiple cellular responses, influences the immune system and the tumor microenvironment, and promotes tumor cell invasion and metastasis [44, 45]. SMAD proteins serve as downstream signal transduction molecules of TGF-β family receptors and consist of two globular domains (MH1 and MH2 domains) connected by an unstructured linker [45, 46]. SMAD4, a shared component among all SMAD proteins, plays a crucial role in cancer metastasis and acts as an essential effector of TGF-β-induced epithelial–mesenchymal transition (EMT) [47, 48]. Phosphorylation of the two C-terminal Ser residues of SMADs by receptor-mediated targeting enables them to bind to the MH2 domain of SMAD4 or the cognate MH2 structure [48]. In CRC, nucleotides 1465 to 1895 of LINC00941 compete with β-TrCP for binding to the MH2 domain of both SMAD4 and full-length SMAD4. This interaction regulates the ubiquitination and degradation of SMAD4, stabilizes SMAD4, activates the TGF-β/SMAD signaling pathway, accelerates the EMT process in CRC cells, and promotes CRC cell metastasis [14].
Wnt/β-catenin signaling pathway plays a crucial role in the proliferation, metastasis, and invasion of cancer cells [49,50,51]. In HCC, nucleotides 800 to 1600 of LINC00941 specifically bind to ANXA2. Additionally, ANXA2 interacts with GSK-3β and disrupts the formation of the GSK-3β/β-catenin complex. Consequently, β-catenin accumulates continuously and translocates into the nucleus, activating the Wnt/β-catenin signaling pathway. This activation promotes the expression of downstream genes, facilitates the EMT process, and ultimately contributes to the progression of HCC in vivo [8]. In OSCC, hnRNPK directly enhances the expression of LINC00941 [15]. Furthermore, hnRNPK overexpression inhibits the phosphorylation of β-catenin, leading to its degradation through the ubiquitin–proteasome system. This results in the activation of the Wnt/β-catenin signaling pathway, promoting cell proliferation and metastasis [15].
The LIMK1/Cofilin-1 signaling pathway plays a significant role in the growth, invasion, and metastasis of cancer cells [52]. LIMK1, a serine/threonine protein kinase, phosphorylates and inactivates the downstream actin regulator, Cofilin-1. This affects the formation of actin stress fibers and adhesion plaques, thereby modulating the cytoskeleton system and influencing tumor cell metastasis [53, 54]. In PC, LINC00941 acts as a molecular sponge for miR-335-5p, leading to upregulation of ROCK1 and activation of the LIMK1/Cofilin-1 pathway. Consequently, the growth, EMT, and metastasis of PC cells are promoted [16].
The Hippo signaling pathway, a highly conserved pathway, is involved in various biological processes such as cell growth, apoptosis, and cancer development [55]. The core of the mammalian Hippo pathway consists of a kinase cascade involving mammalian sterile20-like kinase 1/2 (MST1/2) and large tumor suppressor 1/2 (LATS1/2), along with downstream effectors YAP and TAZ, which function as transcriptional coactivators. These core components govern cell proliferation, survival, metastasis, stemness, and differentiation [56]. In PDAC, LINC00941 interacts with mammalian MST1, promoting protein phosphatase 2A (PP2A)-mediated dephosphorylation of MST1 and activating the Hippo signaling pathway. This leads to the nuclear translocation of the transcriptional coactivator YAP, initiation of downstream target gene transcription, and promotion of the glycolysis process in PDAC cell lines, thereby enhancing cell proliferation [17].
LINC00941 drives tumor progression by modulating the diverse biological behaviors of cancer cells.
LncRNA plays a pivotal role in regulating the occurrence and development of cancer by influencing multiple cellular processes [57]. Figure 4 and Table 1 provide a comprehensive overview of the impact of LINC00941 on cancer progression, highlighting its modulation of downstream genes involved in pivotal cellular functions such as cell proliferation, apoptosis, EMT, invasion, and migration.
Upregulation of LINC00941 promotes cell proliferation while inhibiting apoptosis
Cell proliferation refers to the intricate process by which cells absorb nutrients, undergo growth, and divide, ultimately leading to an increase in cell population. Uncontrolled cell proliferation is a hallmark of cancer [58]. LINC00941 engages in various regulatory axes, exerting transcriptional and post-transcriptional control to facilitate cell proliferation. In CRC, LINC00941 is implicated in the LINC00941/miR-205-5p/MYC axis [22]. In ESCC, it participates in the LINC00941/miR-877-3p/PMEPA1 axis [7] as well as the LINC00941/ILF2/YBX1/SOX2 axis [18]. The LINC00941/miR-34a-5p/Snail1 axis has been identified in GBM [13], while the LINC00941/ANXA2/GSK-3β axis and LINC00941/miR-34a-5p/Snail1 axis are involved in HCC [8]. Additionally, LINC00941 contributes to cell proliferation in LC through the LINC00941/PKM axis [12], in NSCLC through the LINC00941/miR-877-3p/VEGFA axis [23], in OSCC through the LINC00941/hnRNPK axis [15] and the LINC00941/CTCF/CAPRIN2 axis [3], in PAAD through the LINC00941/miR-873-3p/ATXN2 axis [24], in PC through the LINC00941/miR-335-5p/ROCK1 axis and the LINC00941/ANXA2 axis [11, 16], and in PDAC through the LINC00941/MST1/YAP1 axis [17].
Apoptosis, a programmed cell death mechanism regulated by specific genes, represents a normal physiological response of cells [59]. However, in the context of cancer, apoptosis is frequently suppressed, leading to uncontrolled proliferation of cancer cells [60]. In GBM, LINC00941 activates the TGF-β/SMAD2/3 signaling pathway via the LINC00941/miR-34a-5p/Snail1 axis, thereby suppressing apoptosis [13].
Upregulation of LINC00941 promotes EMT, invasion, and migration
EMT is a pivotal process through which epithelial cells acquire mesenchymal characteristics, playing a significant role in cancer cell metastasis [61]. In HCC, LINC00941 upregulation triggers the EMT process by activating the Wnt/β-catenin signaling pathway via its interaction with ANXA2 [8]. Moreover, LINC00941 contributes to EMT promotion in various tumor cells through its involvement in the ceRNA axes. Notably, it participates in the LINC00941/miR-877-3p/PMEPA1 axis in ESCC [7], the LINC00941/miR-34a-5p/Snail1 axis in HCC [8], and the LINC00941/miR-335-5p/ROCK1 axis in PC [16].
Invasion and migration characterize the active motility and infiltration of malignant cells into surrounding tissues from the primary tumor site [62]. LINC00941 actively engages in multiple regulatory axes, promoting cell invasion and migration at both transcriptional and post-transcriptional levels. For example, in CRC, LINC00941 is implicated in the LINC00941/miR-205-5p/MYC axis [22]. In ESCC, it participates in the LINC00941/miR-877-3p/PMEPA1 axis [7] and the LINC00941/ILF2/YBX1/SOX2 axis [18]. In GBM, the LINC00941/miR-34a-5p/Snail1 axis comes into play [13], while in HCC, LINC00941 contributes to cell invasion and migration through the LINC00941/ANXA2/GSK-3β axis and the LINC00941/miR-34a-5p/Snail1 axis [8]. Additionally, LINC00941 is involved in the LINC00941/PKM axis in LC [12], the LINC00941/miR-877-3p/VEGFA axis in NSCLC [23], the LINC00941/hnRNPK axis and the LINC00941/CTCF/CAPRIN2 axis in OSCC [3, 15], the LINC00941/miR-873-3p/ATXN2 axis in PAAD [24], and the LINC00941/miR-335-5p/ROCK1 axis and the LINC00941/ANXA2 axis in PC [11, 16].
The potential implications of LINC00941 in clinical diagnosis and treatment
Given their substantial differential expression and extensive regulatory roles, lncRNAs hold tremendous promise as emerging biomarkers and therapeutic targets, thus playing a significant role in clinical diagnosis and treatment [63]. High expression of LINC00941 has been correlated with poorer prognosis, including overall survival (OS) and disease-free survival (DFS), as well as various clinicopathological characteristics such as tumor size, degree of metastasis, and clinical stage. Notably, several downstream targets of LINC00941 have demonstrated drug-targetable properties.
The association between LINC00941 and prognostic value and clinicopathological features of cancer patients
Table 2 highlights the close link between aberrant LINC00941 expression and the prognosis and clinicopathological features of different tumor types. In ten cancer types, elevated levels of LINC00941 were consistently associated with worse overall survival of patients. These cancer types include CRC [14], GC [19, 20], HCC [8, 21], LC [12], LUAD [64, 65], NSCLC [23], PAAD [24], PC [11, 16], PDAC [17], and papillary thyroid cancer (PTC) [66]. Furthermore, in PAAD [24] and PC [11, 16, 31], high LINC00941 expression was associated with reduced disease-free survival.
In LC, increased LINC00941 expression correlated with advanced tumor stage, deeper tumor invasion depth, and higher lymph node metastasis [12]. Similarly, in CRC, high LINC00941 expression was typically linked to a higher rate of lymph node metastasis, larger tumor size, more distant tumor metastasis [14, 22]. In GC, elevated LINC00941 expression was associated with advanced clinical stage and more advanced TNM stage [67, 68]. However, some studies have also reported that increased LINC00941 expression is related to more advanced tumor depth and a higher rate of distant metastasis [69]. In HCC, increased LINC00941 expression was significantly correlated with a higher rate of lymph node metastasis, and larger tumor size [8, 21]. Similarly, in PC, increased LINC00941 expression was significantly correlated with distant metastasis and higher lymph node metastasis [16]. In PDAC, elevated LINC00941 expression was significantly associated with larger tumor size and a higher tumor stage [17]. Lastly, in PTC, increased LINC00941 expression was significantly associated with a more advanced TNM stage [66].
Drugs targeting downstream effectors of LINC00941
LINC00941 and its downstream targets hold substantial potential as therapeutic targets for drug intervention. Figure 5 illustrates that a search in the CADDIE database (https://exbio.wzw.tum.de/caddie/) [70] has identified numerous approved drugs that target the downstream effectors of LINC00941. These include marketed drugs such as warfarin, acetylsalicylic acid, nadroparin, and resveratrol, which target MYC. Drugs targeting ATXN2 encompass raloxifene, nadroparin, nicardipine, amsacrine, and maprotiline. Marketed drugs targeting VEGFA include celecoxib, denibulin, enalapril, and carvedilol. Lastly, drugs targeting ROCK1 consist of midostaurin, lapatinib, bosutinib, imatinib, salicylamide, adenine, ruxolitinib, vandetanib, imatinib, and pazopanib.
Conclusions and future prospects
LINC00941, recently identified as an oncogenic factor, exhibits widespread upregulation in various types of cancers. Current research reveals the abnormal overexpression of LINC00941 in cancer cells or tissues across 12 different cancer types. However, the existing studies suffer from limitations such as small sample sizes and a narrow range of cancer types. Furthermore, there is a lack of differential research focusing on clinically relevant detection carriers suchas serum and cerebrospinal fluid. Moving forward, it is imperative to expand the sample size and include diverse sample types to gain further insights into the potential value of LINC00941 in non-invasive cancer diagnosis.
LINC00941 plays a pivotal role in extensive regulatory networks within cancer, exerting its influence through multiple mechanisms. It acts as a miRNA sponge, indirectly modulating the translation of five downstream mRNAs, and facilitates the transcription of downstream effector genes by recruiting transcription factors. Additionally, LINC00941 is involved in five important signaling pathways. However, the current understanding of the molecular mechanisms underlying LINC00941's actions remains limited and incomplete. Further research is necessary to unveil a more comprehensive and detailed regulatory network associated with LINC00941, providing a solid foundation for future studies focused on clinical treatment.
The clinical potential of LINC00941 has been observed across various cancer types. Elevated expression of LINC00941 shows a strong correlation with the prognosis and clinicopathological features of ten different cancers, making it a promising biomarker for prognostic evaluation in diverse malignancies. However, the current research faces challenges due to small sample sizes and a limited range of cancer types investigated. To validate the prognostic value of LINC00941, larger and more diverse sample sizes, including multi-center clinical studies, are needed in the future. It is worth noting that research on drugs targeting LINC00941 is currently scarce, and no drugs directly targeting LINC00941 have been reported. Although drugs targeting downstream effectors exist, their correlation with LINC00941 remains unexplored. Therefore, further exploration and experimental validation of LINC00941 and the drugs within its regulatory network are crucial to uncover the potential therapeutic value of LINC00941 in cancer treatment. Enhancing our understanding of the prognosis and drug interactions involving LINC00941 will contribute to a better grasp of its pivotal role in cancer treatment and facilitate its clinical translation.
This work presents a comprehensive review of the differential expression, molecular mechanisms, and clinical significance of LINC00941. LINC00941 emerges as a critical oncogene with the potential to serve as a biomarker and therapeutic target in various cancers. However, further research on LINC00941 is imperative to unravel its specific molecular mechanisms in different cancer types and uncover its diagnostic and therapeutic value in cancer management.
Data availability
Not applicable.
Abbreviations
- ACC:
-
Adrenocortical carcinoma
- ALL:
-
Acute lymphoblastic leukemia
- AML:
-
Acute myeloid leukemia
- BLCA:
-
Bladder urothelial carcinoma
- BRCA:
-
Breast invasive carcinoma
- ceRNA:
-
Competing endogenous RNA
- CESC:
-
Cervical esophageal squamous cell carcinoma
- CHOL:
-
Cholangiocarcinoma
- COAD:
-
Colon adenocarcinoma
- CRC:
-
Colorectal cancer
- DFS:
-
Disease-free survival
- EMT:
-
Epithelial–mesenchymal transition
- ESCA:
-
Esophageal carcinoma
- ESCC:
-
Esophageal squamous cell carcinoma
- FAK:
-
Focal adhesion kinase
- GBM:
-
Glioblastoma multiforme
- GC:
-
Gastric cancer
- HCC:
-
Hepatocellular carcinoma
- HNSC:
-
Head and neck squamous cell carcinoma
- KICH:
-
Kidney chromophobe
- KIRC:
-
Kidney renal clear cell carcinoma
- KIRP:
-
Kidney papillary cell carcinoma
- LATS1/2:
-
Large tumor suppressor 1/2
- LC:
-
Laryngocarcinoma
- LGG:
-
Lower grade glioma
- LIHC:
-
Liver hepatocellular carcinoma
- LncRNA:
-
Long non-coding RNA
- LUAD:
-
Lung adenocarcinoma
- LUSC:
-
Lung squamous cell carcinoma
- miRNA:
-
MicroRNA
- miRNA-RISC:
-
MiRNA-containing RNA-induced silencing complex
- MREs:
-
MicroRNA response elements
- MST1/2:
-
Mammalian sterile20-like kinase 1/2
- NBL:
-
Neuroblastoma
- NSCLC:
-
Non-small cell lung cancerr
- OV:
-
Ovarian cancer
- OS:
-
Overall survival
- OSCC:
-
Oral squamous cell carcinoma
- PAAD:
-
Pancreatic adenocarcinoma
- PC:
-
Pancreatic cancer
- PCPG:
-
Pheochromocytoma and paraganglioma cancer
- PDAC:
-
Pancreatic ductal adenocarcinoma
- PP2A:
-
Protein phosphatase 2A
- PRAD:
-
Prostate adenocarcinoma
- PTC:
-
Primes papillary thyroid cancer
- READ:
-
Rectum adenocarcinoma
- SARC:
-
Sarcoma
- SKCM:
-
Skin cutaneous melanoma
- SPP:
-
Signaling pathways project
- STAD:
-
Stomach adenocarcinoma
- TGCT:
-
Testicular germ cell tumors
- TGF- β:
-
Transforming growth factor-β
- THCA:
-
Thyroid carcinoma
- THYM:
-
Thymoma
- TNM:
-
Tumor node metastasis
- TSS:
-
Transcription start site
- UCEC:
-
Endometrioid cancer
- UCS:
-
Uterine carcinosarcoma
- UCSC Genome Browser:
- CADDIE database:
- ChIP-Atlas:
- JASPAR 2022 database:
- SPP database:
- TCGA database:
References
Yan H, Bu P. Non-coding RNA in cancer. Essays Biochem. 2021;65:625–39.
Li C, Ni YQ, Xu H, et al. Roles and mechanisms of exosomal non-coding RNAs in human health and diseases. Signal Transduct Target Ther. 2021;6:383.
Ai Y, Wu S, Zou C, Wei H. LINC00941 promotes oral squamous cell carcinoma progression via activating CAPRIN2 and canonical WNT/beta-catenin signaling pathway. J Cell Mol Med. 2020;24:10512–24.
Zhang Q, Shen J, Wu Y, Ruan W, Zhu F, Duan S. LINC00520: A Potential Diagnostic and Prognostic Biomarker in Cancer. Front Immunol. 2022;13:845418.
Bolha L, Ravnik-Glavac M, Glavac D. Long Noncoding RNAs as Biomarkers in Cancer. Dis Markers. 2017;2017:7243968.
Zhang Q, Zhong C, Duan S. The tumorigenic function of LINC00858 in cancer. Biomed Pharmacother. 2021;143:112235.
Zhang Y, Zhu H, Sun N, et al. Linc00941 regulates esophageal squamous cell carcinoma via functioning as a competing endogenous RNA for miR-877-3p to modulate PMEPA1 expression. Aging (Albany NY). 2021;13:17830–46.
Yan X, Zhang D, Wu W, et al. Mesenchymal Stem Cells Promote Hepatocarcinogenesis via lncRNA-MUF Interaction with ANXA2 and miR-34a. Cancer Res. 2017;77:6704–16.
Morgenstern E, Kretz M. The human long non-coding RNA LINC00941 and its modes of action in health and disease. Biol Chem. 2023. https://doi.org/10.1515/hsz-2023-0183.
Kunej T, Obsteter J, Pogacar Z, Horvat S, Calin GA. The decalog of long non-coding RNA involvement in cancer diagnosis and monitoring. Crit Rev Clin Lab Sci. 2014;51:344–57.
Wang J, He Z, Liu X, et al. LINC00941 promotes pancreatic cancer malignancy by interacting with ANXA2 and suppressing NEDD4L-mediated degradation of ANXA2. Cell Death Dis. 2022;13:718.
Li Z, Jin Q, Sun Y. LINC00941 promoted in vitro progression and glycolysis of laryngocarcinoma by upregulating PKM via activating the PI3K/AKT/mTOR signaling pathway. J Clin Lab Anal. 2022;36:e24406.
Shree B, Tripathi S, Sharma V. Transforming Growth Factor-Beta-Regulated LncRNA-MUF Promotes Invasion by Modulating the miR-34a Snail1 Axis in Glioblastoma Multiforme. Front Oncol. 2021;11:788755.
Wu N, Jiang M, Liu H, et al. LINC00941 promotes CRC metastasis through preventing SMAD4 protein degradation and activating the TGF-beta/SMAD2/3 signaling pathway. Cell Death Differ. 2021;28:219–32.
Liu J, Li Z, Zhang T, et al. Long Noncoding RNA LINC00941 Promotes Cell Proliferation and Invasion by Interacting with hnRNPK in Oral Squamous Cell Carcinoma. Nutr Cancer. 2022;74:2983–95.
Wang J, He Z, Xu J, Chen P, Jiang J. Long noncoding RNA LINC00941 promotes pancreatic cancer progression by competitively binding miR-335-5p to regulate ROCK1-mediated LIMK1/Cofilin-1 signaling. Cell Death Dis. 2021;12:36.
Xu M, Cui R, Ye L, et al. LINC00941 promotes glycolysis in pancreatic cancer by modulating the Hippo pathway. Mol Ther Nucleic Acids. 2021;26:280–94.
Lu JT, Yan ZY, Xu TX, et al. Reciprocal regulation of LINC00941 and SOX2 promotes progression of esophageal squamous cell carcinoma. Cell Death Dis. 2023;14:72.
Luo C, Tao Y, Zhang Y, et al. Regulatory network analysis of high expressed long non-coding RNA LINC00941 in gastric cancer. Gene. 2018;662:103–9.
Beeraka NM, Gu H, Xue N, et al. Testing lncRNAs signature as clinical stage-related prognostic markers in gastric cancer progression using TCGA database. Exp Biol Med (Maywood). 2022;247:658–71.
Fang Y, Yang Y, Zhang X, et al. A Co-Expression Network Reveals the Potential Regulatory Mechanism of lncRNAs in Relapsed Hepatocellular Carcinoma. Front Oncol. 2021;11:745166.
Chang L, Zhou D, Luo S. Novel lncRNA LINC00941 Promotes Proliferation and Invasion of Colon Cancer Through Activation of MYC. Onco Targets Ther. 2021;14:1173–86.
Ren MH, Chen S, Wang LG, Rui WX, Li P. LINC00941 Promotes Progression of Non-Small Cell Lung Cancer by Sponging miR-877-3p to Regulate VEGFA Expression. Front Oncol. 2021;11:650037.
Fang L, Wang SH, Cui YG, Huang L. LINC00941 promotes proliferation and metastasis of pancreatic adenocarcinoma by competitively binding miR-873-3p and thus upregulates ATXN2. Eur Rev Med Pharmacol Sci. 2021;25:1861–8.
Chan JJ, Tay Y. Noncoding RNA:RNA Regulatory Networks in Cancer. Int J Mol Sci. 2018;19:1310.
Tan YT, Lin JF, Li T, Li JJ, Xu RH, Ju HQ. LncRNA-mediated posttranslational modifications and reprogramming of energy metabolism in cancer. Cancer Commun (Lond). 2021;41:109–20.
Lupianez DG, Kraft K, Heinrich V, et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell. 2015;161:1012–25.
Debaugny RE, Skok JA. CTCF and CTCFL in cancer. Curr Opin Genet Dev. 2020;61:44–52.
Li Y, Haarhuis JHI, Sedeno Cacciatore A, et al. The structural basis for cohesin-CTCF-anchored loops. Nature. 2020;578:472–6.
Fang C, Wang Z, Han C, et al. Cancer-specific CTCF binding facilitates oncogenic transcriptional dysregulation. Genome Biol. 2020;21:247.
Lu J, Yu L, Xie N, Wu Y, Li B. METTL14 Facilitates the Metastasis of Pancreatic Carcinoma by Stabilizing LINC00941 in an m6A-IGF2BP2-Dependent Manner. J Cancer. 2023;14:1117–31.
Ulitsky I, Bartel DP. lincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154:26–46.
Becnel LB, Ochsner SA, Darlington YF, et al. Discovering relationships between nuclear receptor signaling pathways, genes, and tissues in Transcriptomine. Sci Signal. 2017. https://doi.org/10.1126/scisignal.aah6275.
Zou Z, Ohta T, Miura F, Oki S. ChIP-Atlas 2021 update: a data-mining suite for exploring epigenomic landscapes by fully integrating ChIP-seq, ATAC-seq and Bisulfite-seq data. Nucleic Acids Res. 2022;50:W175–82.
Castro-Mondragon JA, Riudavets-Puig R, Rauluseviciute I, et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 2022;50:D165–73.
Karreth FA, Pandolfi pp. ceRNA cross-talk in cancer: when ce-bling rivalries go awry. Cancer Discov. 2013;3:1113.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi pp. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146:353–8.
Peng WX, Koirala P, Mo YY. LncRNA-mediated regulation of cell signaling in cancer. Oncogene. 2017;36:5661–7.
Cheng SC, Quintin J, Cramer RA, et al. mTOR- and HIF-1alpha-mediated aerobic glycolysis as metabolic basis for trained immunity. Science. 2014;345:1250684.
Sun Q, Chen X, Ma J, et al. Mammalian target of rapamycin up-regulation of pyruvate kinase isoenzyme type M2 is critical for aerobic glycolysis and tumor growth. Proc Natl Acad Sci U S A. 2011;108:4129–34.
Tian T, Li X, Zhang J. mTOR Signaling in Cancer and mTOR Inhibitors in Solid Tumor Targeting Therapy. Int J Mol Sci. 2019;20:755.
Molli PR, Li DQ, Murray BW, Rayala SK, Kumar R. PAK signaling in oncogenesis. Oncogene. 2009;28:2545–55.
Dawson JC, Serrels A, Stupack DG, Schlaepfer DD, Frame MC. Targeting FAK in anticancer combination therapies. Nat Rev Cancer. 2021;21:313–24.
Katsuno M, Adachi H, Banno H, Suzuki K, Tanaka F, Sobue G. Transforming growth factor-beta signaling in motor neuron diseases. Curr Mol Med. 2011;11:48–56.
Xie F, Ling L, van Dam H, Zhou F, Zhang L. TGF-beta signaling in cancer metastasis. Acta Biochim Biophys Sin (Shanghai). 2018;50:121–32.
Akhurst RJ, Hata A. Targeting the TGFbeta signalling pathway in disease. Nat Rev Drug Discov. 2012;11:790–811.
Heldin CH, Moustakas A. Signaling Receptors for TGF-beta Family Members. Cold Spring Harb Perspect Biol. 2016;8:a022053.
Massague J. TGFbeta signalling in context. Nat Rev Mol Cell Biol. 2012;13:616–30.
Muralidhar S, Filia A, Nsengimana J, et al. Vitamin D-VDR Signaling Inhibits Wnt/beta-Catenin-Mediated Melanoma Progression and Promotes Antitumor Immunity. Cancer Res. 2019;79:5986–98.
Yu F, Yu C, Li F, et al. Wnt/beta-catenin signaling in cancers and targeted therapies. Signal Transduct Target Ther. 2021;6:307.
Reya T, Clevers H. Wnt signalling in stem cells and cancer. Nature. 2005;434:843–50.
Lee MH, Kundu JK, Chae JI, Shim JH. Targeting ROCK/LIMK/cofilin signaling pathway in cancer. Arch Pharm Res. 2019;42:481–91.
Lu G, Zhou Y, Zhang C, Zhang Y. Upregulation of LIMK1 Is Correlated With Poor Prognosis and Immune Infiltrates in Lung Adenocarcinoma. Front Genet. 2021;12:671585.
Ishaq M, Lin BR, Bosche M, et al. LIM kinase 1 - dependent cofilin 1 pathway and actin dynamics mediate nuclear retinoid receptor function in T lymphocytes. BMC Mol Biol. 2011;12:41.
Dey A, Varelas X, Guan KL. Targeting the Hippo pathway in cancer, fibrosis, wound healing and regenerative medicine. Nat Rev Drug Discov. 2020;19:480–94.
Ma S, Meng Z, Chen R, Guan KL. The Hippo Pathway: Biology and Pathophysiology. Annu Rev Biochem. 2019;88:577–604.
Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118.
Liu L, Michowski W, Kolodziejczyk A, Sicinski P. The cell cycle in stem cell proliferation, pluripotency and differentiation. Nat Cell Biol. 2019;21:1060–7.
Elmore S. Apoptosis: a review of programmed cell death. Toxicol Pathol. 2007;35:495–516.
Xu X, Lai Y, Hua ZC. Apoptosis and apoptotic body: disease message and therapeutic target potentials. Biosci Rep. 2019;39:495.
Babaei G, Aziz SG, Jaghi NZZ. EMT, cancer stem cells and autophagy; The three main axes of metastasis. Biomed Pharmacother. 2021;133:110909.
Lusby R, Dunne P, Tiwari VK. Tumour invasion and dissemination. Biochem Soc Trans. 2022;50:1245–57.
Winkle M, El-Daly SM, Fabbri M, Calin GA. Noncoding RNA therapeutics - challenges and potential solutions. Nat Rev Drug Discov. 2021;20:629–51.
Shao J, Zhang B, Kuai L, Li Q. Integrated analysis of hypoxia-associated lncRNA signature to predict prognosis and immune microenvironment of lung adenocarcinoma patients. Bioengineered. 2021;12:6186–200.
Chen H, Zhou C, Hu Z, et al. Construction of an algorithm based on oncosis-related LncRNAs comprising the molecular subtypes and a risk assessment model in lung adenocarcinoma. J Clin Lab Anal. 2022;36:e24461.
Gugnoni M, Manicardi V, Torricelli F, et al. Linc00941 Is a Novel Transforming Growth Factor beta Target That Primes Papillary Thyroid Cancer Metastatic Behavior by Regulating the Expression of Cadherin 6. Thyroid. 2021;31:247–63.
Zou Z, Ma T, He X, et al. Long intergenic non-coding RNA 00324 promotes gastric cancer cell proliferation via binding with HuR and stabilizing FAM83B expression. Cell Death Dis. 2018;9:717.
Wang S, Cheng Y, Yang P, Qin G. Silencing of Long Noncoding RNA LINC00324 Interacts with MicroRNA-3200-5p to Attenuate the Tumorigenesis of Gastric Cancer via Regulating BCAT1. Gastroenterol Res Pract. 2020;2020:4159298.
Liu H, Wu N, Zhang Z, et al. Long Non-coding RNA LINC00941 as a Potential Biomarker Promotes the Proliferation and Metastasis of Gastric Cancer. Front Genet. 2019;10:5.
Hartung M, Anastasi E, Mamdouh ZM, et al. Cancer driver drug interaction explorer. Nucleic Acids Res. 2022;50:W138–44.
Wang Y, Zhao D, Wang H, et al. Long non-coding RNA-LINC00941 promotes the proliferation and invasiveness of glioma cells. Neurosci Lett. 2023;795:136964.
Pan H, Wei W, Fu G, Pan J, Jin B. LINC00941 Promotes Cell Malignant Behavior and Is One of Five Costimulatory Molecule-Related lncRNAs That Predict Prognosis in Renal Clear Cell Carcinoma. Medicina (Kaunas). 2023;59:187.
Funding
This study was supported by the Qiantang Scholars Fund in Hangzhou City University (No. 210000–581835).
Author information
Authors and Affiliations
Contributions
QY, XS, YC, ZW, WH, QX, and YM collected and analyzed the literature, drafted the figures, and wrote the manuscript. SD and HL conceived the idea and gave the final approval of the submitted version. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yan, Q., Su, X., Chen, Y. et al. LINC00941: a novel player involved in the progression of human cancers. Human Cell 37, 167–180 (2024). https://doi.org/10.1007/s13577-023-01002-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-023-01002-5