Abstract
JAK/STAT pathway has been widely acknowledged in the development of human cancers. However, the role of different phosphorylated STAT proteins translocating into nucleus in transcription activation of target genes is not fully understood. In present research, ChIP-seq was carried on to investigate the genome-wide distribution of the activated STAT1, STAT2, STAT3, STAT5 and STAT6 in colorectal cancer HCT-116 cells. Our observations indicated that the homodimers rather than heterodimers of STAT protein predominantly occupied on genomic DNA. STAT3 accounted for the largest proportion among all STAT proteins HCT-116 cells. Furthermore, the biased binding motif targeted by different STAT homodimers suggested the distinct biological functions. Here, we noticed that NR5A2 was a specific co-activator of STAT3 by DNA motif analysis. Co-IP assay determined that NR5A2 indeed interacted with STAT3 homodimer rather than other homodimers or heterodimers. NR5A2 knockdown resulted in a reduced binding affinity of STAT3 homodimer in the original regions. Taken together, we characterize the genome-wide landscape of activated STAT proteins, and reveal the differences of binding patterns as well as the target genes and associated functions between homodimer and heterodimer of STAT proteins in HCT-116 cells. We also present some new findings and possible mechanisms regarding the role of NR5A2 on STAT3 in CRC. Our findings may provide new insights into the design of STAT inhibitors to treat CRC and other diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Janus kinase/signal transducer and activator of transcription (JAK/STAT) pathway responding to a large number of growth factors and cytokine receptors is critical in immune function, blood formation and cell growth via transmitting the signals of anti-apoptotic, proliferation and differentiation in normal circumstances [1]. However, once JAK/STAT signaling pathway is excessively and continuously activated, the disordered activities of oncogenes and tumor suppressor genes can promote the proliferation and migration of malignant cells, and enhance the occurrence and progression of a large proportion of cancers [2].
As well-acknowledged JAK/STAT signaling pathway [3], there are five key steps to complete the signal transduction process including (i) stimulation from cytokines to the ligands on cell surface; (ii) activation of JAK proteins by receptors via mutual phosphorylation at tyrosines; (iii) activation of STAT proteins by JAK via phosphorylation of tyrosine at Src homology 2 (SH2) domains; (iv) entry of the activated STAT hetero- or homodimers in nucleus for transcription regulation coordinated with other transcription factors; and (v) inhibition of JAKs and STATs by the up-regulated suppressors of cytokine singling (SOCS) as a feedback. Although the selection of JAK proteins for signal transmission depends on different cytokine receptors, the activated JAK protein kinases do not seem to have specificity for particular STAT substrates [4]. However, the types and distributions of activated STAT family bound on genome are inconsistent in different diseases. STAT1 contributes to governing cell cycle progression via affecting the expression of regulators for apoptosis [5], whereas STAT3 possesses oncogenic potentials, thus promoting cell proliferation and survival [6]. STAT4 and STAT6 expressed in immune cells majorly targets to a series of genes encoding cytokines, receptors and signaling factors for lymphocytes development and inflammatory response [7, 8]. STAT5 proteins are associated with cell differentiation lipid mobilization and hormone synthesis [9]. The above contents suggest that the defined gene population specifically activated by different STAT proteins are related to the genomic accessibility and recognition by the STAT proteins with other cofactors [10].
In colorectal cancer (CRC) cells, STAT3 has been determined to be constitutively activated in vivo [11], and widely explored to develop novel therapies for CRC [12]. Nevertheless, other STAT proteins except STAT4 are also reported to participate in CRC carcinogenesis and progression [13, 14]. To data, the components of human activated STAT dimers in the nucleus are not clear, which includes the synergic, supplementary or competitive relationship among different STATs in CRC cells. In current study, chromatin immunoprecipitation sequencing (ChIP-seq) was carried on to decipher the genome-wide accessibility and landscape of STAT1, STAT2, STAT3, STAT5A/B and STAT6 in human HCT-116 CRC cells. The bioinformatic analysis attempted to figure out the different landscapes of STAT dimers on genomic DNA, which is benefit to searching the precise therapeutic targets of JAK/STAT pathway and developing novel drugs for CRC and spreading to other diseases.
Materials and methods
Clinical study
Thirty CRC patients from 2015 to 2019 were enrolled in this study. All participants signed the informed consent. This study (2022-KY-0073-001) was approved by the ethics committee of Henan cancer hospital. The study was carried out under the standards of the declaration of Helsinki.
Immunohistochemistry (IHC) assay
CRC tissues were fixed at 4 °C overnight by 4% formaldehyde solution, then embedded by paraffin, and prepared 8 μm sections by microtome. Primary antibodies included STATs and p-STATs (STAT1, sc-464; STAT2, sc-514193; STAT3, sc-293151; STAT5A/B, sc-74442; STAT6, sc-374021; p-STAT1, sc-8394; p-STAT3, sc-8059; p-STAT6, sc-136019, Santa Cruz Biotechnology, Santa Cruz, USA; p-STAT2, AF3342; p-STAT5, AF3305, Affinity Biosciences, Cincinnati, USA) at working solution (1:200). The slides were dipped in hematoxylin for 10 min, 0.1% HCl-ethanol for 3 min, 0.5% NH3·H2O for 30 s, and then 0.5% eosin-ethanol for 30 s. The slides were rinsed tap by water for 5 s in experimental gap. Finally, the sections were dehydrated in graded ethanol, vitrified by xylene and mounted in neutral balsam. The positive staining was statistically analyzed using Image J.
Cell culture
CRC HCT-116, RKO and CACO-2 cells as well as normal colon epithelial HcoEpiC cells and intestinal fibroblasts CCD-18Co cells were cultured in cultured within Dulbecco’s Minimum Essential Medium (DMEM) containing 10% fetal bovine serum (FBS) (Thermo Fisher Scientific, Waltham, MA, USA). NR5A2 siRNA (5’-UCAUUGAGCAAAAGAAAAGUG-3’) synthesized from GenScript Biotech (Nanjing, Jiangsu, China) was transfected into HCT-116 cells using PolyFast Transfection Reagent (MedChemExpress LLC, Shanghai, China) following the instruction manual. Cells were further cultured 36 h after transfection, following collection at − 80 °C immediately for the subsequent experiments.
Immunofluorescence (IF) assay
HCT-116 cells were subcultured on µ-Slide 8 well (IBIDI, Martinsried, Bavaria, Germany) and grew to the density of 1 × 105/well. After gently taken away the medium, cells were crosslinked by 1% paraformaldehyde/PBS solution, then permeabilized by 0.1% Triton-X-100 and blocked with 1% BSA/PBS solution. Primary antibodies against p-STATs (1: 200, Refer to “IHC assay” for antibodies information) were sued to incubated at 4 °C overnight. After washing by PBS, secondary antibodies against rabbit or mouse (1:1000; Beyotime Biotechnology, Shanghai, China) were incubated at room temperature for 30 min. After washing by PBS and air drying in dark room, slides were dropped by Antifade mounting medium with DAPI and coated with cover glasses.
Chromatin immunoprecipitation sequencing (ChIP-seq) assay
ChIP-seq was conducted following previous study [15]. Briefly, genomic DNA of 1 × 107 HCT-116 cells were randomly broken into fragments around 200 bp by sonication. 10% of cell lysates were stored as input, and the rest lysates mixed with 800 ng IP-grade antibodies against STAT proteins were slowly rotated at 4 °C overnight. Next day, 20 μL Protein A beads (Thermo Fisher Scientific) were added for another 2 h rotation. Beads were processed by washing buffer of LiCl [0.25 M LiCl, 1% Triton-X 100, 2 mM EDTA, 1% sodium deoxycholate, 20 mM Tris–HCl (pH 8.0)], high [500 mM NaCl, 0.1% SDS, 2 mM EDTA, 1% Triton-X 100, 20 mM Tris–HCl (pH 8.0)] and low salt [150 mM NaCl, 0.1% SDS, 2 mM EDTA, 1% Triton-X 100, 20 mM Tris–HCl (pH 8.0)] successively. The bound DNA fragments were extracted by phenol chloroform, and eluted by sterile water.
For high throughput sequencing, 3’-dA overhangs were added to the different STAT proteins enriched or input DNA and then established a DNA library. The libraries were quantified using Qubit 4.0 (Thermo Fisher Scientific), and the fragment distribution was determined by Agilent Bioanalyzer 2100 (Agilent, USA). The libraries were sequenced with paired-end 2 × 150 on Illumina. High-quality clean reads were obtained by removing adapters and low-quality reads using Cutadapt v1.18 and Trimmomatic v0.35. FastQC was used to obtain high quality of the clean reads. Then, the processed reads were mapped into human genome (assembly GRCh38) with HISAT2 v2.1.0. Peak calling was carried out using MACS 2 (v2.1.1). Annotation files of human differential expressed genes (DEGs) were retrieved from Ensembl genome browser 96 database (http://www.ensembl.org/index.html). R package ClusterProfiler was used to annotate genes with GO terms and KEGG pathways. We also conducted GO and KEGG-based functional enrichment analysis using ClusterProfiler. The raw data were submitted to the ArrayExpress database with accession number E-MTAB-11113.
Immunoprecipitation (IP) assay
Antibodies of STAT proteins and NR5A2 (sc-393369, Santa Cruz Biotechnology) or IgG were used for target protein pulling down. Other experimental steps were identical with ChIP until the beads purification. Beads were washed by high and low salt washing buffer, and directly added loading buffer [0.28 M Tris–HCl, 30% glycerol, 1% SDS, 0.5 M DTT, 0.0012% bromophenol blue (pH 6.8)] at 100 °C water bath for 10 min, and detected STAT proteins and NR5A2 by WB assay.
WB assay
For regular WB assay, at least 5 × 106 CRC cells were added RIPA buffer [50 mM Tris–HCl, 150 mM NaCl, 1% Triton-X 100, 0.5% sodium deoxycholate, 0.1% SDS (pH 7.4)]. 30 μg total proteins were separated by 10% SDS gel electrophoresis, and transferred onto PVDF membranes (Millipore, Billerica, MA, USA). PVDF membranes were incubated with primary antibodies against STAT proteins and NR5A2 (1:2000) at 4 °C overnight, then washed by PBST for three times and incubated with the secondary antibody against rabbit for 1 h at room temperature. Protein signals were developed by ECL Plus reagents (Pierce, Rockford, IL, USA) and exposed under the Tanon 4600SF system (Tanon, Shanghai, China).
Results
The activity of JAK/STAT pathway in CRC
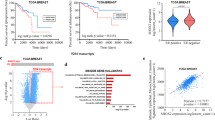
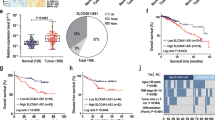
Initially, 30 CRC tissues and the adjacent tissues collected from Henan Cancer Hospital were detected the activity of JAK/STAT signaling pathway. We observed that p-STATs were all highly expressed in tumor tissues compared to para-carcinomatous tissues (Fig. 1A). For in vitro study, we detected the activities of JAK/STAT pathway in different CRC HCT-116, RKO and CACO-2 cells as well as normal colon epithelial HcoEpiC cells and intestinal fibroblasts CCD-18Co cells. The phosphorylation of STAT1, STAT2, STAT3, STAT5 and STAT6 investigated by WB assay were all highly expressed in CRC cells compared to normal colon cells (Fig. 1B), indicating that JAK/STAT pathway was sustainedly over-activated in CRC cells. Furthermore, the phosphorylated STAT3 seemed to be more robust in HCT-116 compared to other two CRC cell lines, which was further validated by IF assay (Fig. 1C). Taken together, we concluded that STAT proteins except STAT4 were all generally continuously activated in CRC in vivo and in vitro.
The JAK/STAT activity in CRC in vivo and in vitro. A IHC assay showing the p-STAT1, p-STAT2, p-STAT3, p-STAT5 and p-STAT6 in CRC and para-carcinomatous (PC) tissues. Images are captured with 200 × magnification. B WB assay showing the expression of STAT and p-STAT in CRC HCT-116, RKO and CACO-2 cells, normal colon epithelial HcoEpiC cells and intestinal fibroblasts CCD-18Co cells. C IF assay showing the location of p-STAT3 in CRC HCT-116, RKO and CACO-2 cells, normal colon epithelial HcoEpiC cells and intestinal fibroblasts CCD-18Co cells. Images are captured with 200 × magnification
The overview of genome-wide distribution of STAT proteins in HCT-116 cells
Next, HCT-116 cells were employed to study the genomic binding signatures of STAT proteins by ChIP-seq. The overall calling peaks of different STAT proteins indicated that STAT3 was the most robustly enriched on genome in HCT-116 cells (Fig. 2A). The genome-wide distribution of STAT proteins showed two groups of similar landscape (Group I: STAT1, STAT2 and STAT3; Group II: STAT5 and STAT6) (Fig. 2B), but appeared the obvious differences in some local areas such as Chromosome 13 (Fig. 2C). We noticed that STAT proteins were profoundly enriched on gene promoters and introns from the perspective of coding genes as well as distal intergenic regions (Fig. 2D, Table S1-5). We obtained the information of typical enhancer and super-enhancer of HCT-116 cells from SEdbV1.05 Database (http://www.licpathway.net/sedb/), and further analyzed in distal intergenic regions. Venn diagram showed that the occupancies of different STAT proteins had a low degree of overlap in promoter regions, enhancers and super-enhancers (Fig. 2E, G, H), but a much higher degree of overlap in intron (Fig. 2F). Taken together, we described the global distribution of STAT proteins on the genome in HCT-116 cells.
The overview of STAT proteins on genomic DNA. A Total calling peaks of STAT1, STAT2, STAT3, STAT5 and STAT6 in HCT-116 cells. B Circos diagram depicting whole-genome ChIP-seq data of STAT proteins. Outside-in track from 1 to 6: Cytoband, chromosomes are depicted qter to pter; genome-wide reads per kilobase per million mapped reads (RPKM) values of STAT1, STAT2, STAT3, STAT5 and STAT6. C A snapshot of the IGV genome browser showing sequencing reads of STAT1, STAT2, STAT3, STAT5 and STAT6 normalized by input on Chromosome 10 in HCT-116 cells. D The distribution of STAT1, STAT2, STAT3, STAT5 and STAT6 in genomic contexts of coding genes
Occupancies of distinct combinations of STAT proteins on STAT-binding areas
Due to STAT proteins functioning on transcription activation in the form of dimer in the nucleus, the interplay and combination of these five STAT proteins was investigated in pairs. STAT-binding areas could be classified into five clusters based on how many types of STAT on them (Fig. 3A). Take Cluster II as an example, this cluster containing with any two STAT proteins indicated three scenarios, namely two homodimers and one heterodimer. Cluster III, IV and V presented the superposition state of Cluster II essentially, and were supposed to display the similar results with Cluster II. We could conclude that 93.33% (336,930/361,008) of STAT-binding areas were recognized by homodimer of STAT proteins (Fig. 3B). Whereas only 0.84% areas in Cluster II (22,061/22,247) and all areas in Cluster III-V showed significant difference of binding strength compared between two STAT proteins in (|log2FC|> 0.585, p < 0.05) (Table S6), implying that heterodimers or multiple homodimers of STAT proteins filled in these areas. By comparison to the peak counts of STAT proteins in each cluster, we determined that the occupancy of STAT3 was the most abundant no matter in homo- or heterodimer, but not absolutely dominant in genomic DNA, while the other four STAT proteins also accounted for a certain proportion (Fig. 3C).
The combinations of STAT dimer in HCT-116 cells. A Upset plot showing the calling peaks with different combinations of STAT protein. The intersection is defined by the non-significant difference of binding strength compared between two STAT proteins in each area (|log2FC|< 0.585, p < 0.05). B Schematic diagram representing the possible combinations of different STAT dimer. C Pie chart displaying the peak accounts of single and combined STAT signal
Next, we investigated the preference of potential biological significance between homo- and heterodimers of STAT protein in CRC. Candidate genes with single STAT protein binding were picked up (Table S6) to carry out GO analysis. The top functions associated with these target genes indicated that STAT1, STAT2, STAT3, STAT5 and STAT6 governed different sets of genes. STAT1 mainly took responsible for the expression of genes such as HSPA1A, IMPDH2 and RAN for cell mitosis and DNA replication including nucleotide biosynthetic process, and spindle as well as microtubule organization (Fig. 4A, B). STAT2 affected genes such as ADIPOR1, BAX and HTRA2 that controlled translational initiation process, responded to external adverse stress, and regulated negative feedback of JAK-STAT pathway (Fig. 4C, D). STAT3 displayed a tissue-specific management on cell proliferation, migration and differentiation of intestinal epithelial cells (Fig. 4E, F). STAT5 majorly modulated glycosphingolipid biosynthetic and catabolic metabolism, and mediated the expression of multiple miRNAs (miRNA-197 and miRNA-494) (Fig. 4G, H). STAT6 governed the expression of enzymes for tricarboxylic acid cycle and glycolysis (Fig. 4I, J). Taken together, we characterized the combinations of STAT proteins on genome, and determined that different STAT homodimer managed a specific series of target genes in HCT-116 cells.
The involved genes and associated functions of each STAT homodimer. A Snapshots of IGV genome browser showing STAT1 homodimer on HSPA1A, IMPDH2 and RAN. B GO analysis showing the involved biological process of binding genes of STAT1 homodimer. C Snapshots of IGV genome browser showing STAT2 homodimer on ADIPOR1, BAX and HTR2A. D GO analysis showing the involved biological process of binding genes of STAT2 homodimer. E Snapshots of IGV genome browser showing STAT3 homodimer on FANCE, FANCF and MYD88. F GO analysis showing the involved biological process of binding genes of STAT3 homodimer. G Snapshots of IGV genome browser showing STAT5 homodimer on miR-197, miR-494 and PDGFB. H GO analysis showing the involved biological process of binding genes of STAT5 homodimer. I Snapshots of IGV genome browser showing STAT6 homodimer on ALDOA, ENO1 and PKLR. J GO analysis showing the involved biological process of binding genes of STAT6 homodimer
Genomic binding preference of STAT3 homodimer controlled by NR5A2 in HCT-116 cell
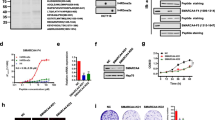
Next, the different binding areas bound by single STAT protein were collected, and extended additional 50 nucleotides up- and downstream to further analyze the cooperative DNA-binding proteins by MEME tool. Given the top five credible DNA motifs, we found that Kruppel-like factor 2 (KLF2) and zinc figure proteins seemingly co-localized with STAT homodimers. Considering about the importance of STAT3 on CRC, we unexpectedly noticed that nuclear receptor subfamily 5 group A member 2 (NR5A2) uniquely appeared near and affected the interaction of STAT3 on genomic DNA compared to other STAT proteins (Fig. 5). To verify the assumption of interaction between NR5A2 and STAT3 homodimer, co-IP assay showed that the interactions of STAT1, STAT2, STAT5 and STAT6 with NR5A2 were extremely weakened compared to STAT3 in HCT-116 cell, indicating that NR5A2 was likely to particularly link with STAT3 homodimer but not heterodimer with other STAT proteins (Fig. 6A, B). Finally, NR5A2 was silenced in HCT-116 cells, followed by STAT3 ChIP-seq again. Although the genome-wide peaks of STAT3 showed no difference (Fig. 6C), the areas bound with STAT3 homodimer showed the reduced STAT3 peaks compared to normal HCT-116 cells (Fig. 6D), implying that NR5A2 might facilitate the recruitment of STAT3 homodimer on target sites. The protein structures of NR5A2 and STAT3 were computed by I-TASSER server [16] via the full length amino acids, and protein rigid docking was conducted to imitate the interaction between STAT3 and NR5A2 by Z-DOCK tool [17] showed that coiled-coil domain of STAT3 (amino acids 139–318) was likely to approach NR5A2 (Fig. 6E). Taken together, we determined that NR5A2 could facilitate to formation of STAT3 homodimer in CRC.
The interaction between NR5A2 and STAT3 in HCT-116 cells. A WB assay showing the interaction of STAT1, STAT2, STAT3, STAT5 and STAT6 with NR5A2 by NR5A2 pull down in HCT-116 cells. B WB assay showing the interaction of NR5A2 with STAT1, STAT2, STAT3, STAT5 and STAT6 by STAT proteins respective pull down in HCT-116 cells. C Total calling peaks of STAT3 in HCT-116 cells with or without NR5A2 knockdown. “KD” means knockdown. D Total calling peaks of STAT3 homodimer in HCT-116 cells with or without NR5A2 knockdown. “KD” means knockdown. E Pymol showing the protein docking between NR5A2 and STAT3. Pink one is NR5A2 while yellow one is STAT3. The coiled-coil domain containing four α-helices is entangled with NR5A2
Discussion
JAK/ STAT signaling pathway is considered as an essential signal transduction pathway for cell development [18]. Under the stimulation from cytokines, tyrosine phosphorylation of STAT proteins are transiently activated and persistently over-expressed in multiple neoplasms including CRC [19,20,21]. Our results confirm that JAK/STAT pathway is continuously activated in CRC tissues and cell lines compared to normal colon cells (Fig. 1), and we determine that STAT3 accounts for the vast majority of the activated STAT proteins in CRC. From the similar pattern of STAT protein expressions in different CRC and normal colon cells, the genetic regulation of STAT proteins by cytokines is supposed to be identical, and the presence of predominant protein expression of p-STAT3 among all p-STAT proteins in CRC indicates that the activation and degradation processes of JAK/STAT pathway in cytoplasm determine the primary importance of STAT3 in CRC. Nevertheless, we still lack sufficient evidence to support this hypothesis by current data.
Phosphorylated STAT homodimers and heterodimers possess a DNA-binding capability and mediated transcriptional regulation via combination with co-activator proteins. Herein, our findings suggest that the vast majority of STAT proteins bound to genomic DNA appear as homologous dimers, while the heterodimers composed by different combination of STAT proteins only occupy a small proportion on genomic DNA (Fig. 3A). In general, STAT proteins are able to form either homo- or heterodimers by interacting via their SH2 domains when they are activated in cytoplasm. However, the amino acid sequences of SH2 domain in different human STAT proteins are actually not consistent, and the affinity between different SH2 sequences is certainly not as strong as the identical ones. Moreover, the initial specificity for the dimerization via this reciprocal SH2 interaction may be also affected by the specific interactions between STATs and their receptors [3] as well as the different splicing isoforms or mutations of the same STAT protein [22]. Therefore, the known structural mechanism of STAT proteins is still not sufficient to exactly figure out the properties that are more prone to homologous dimer.
Since we declare that the activated STAT proteins translocating into nucleus are mainly homologous dimers, the involved functions of different STAT proteins are questioned. GO analysis on target genes indicates a selective and biased pattern of target genes by distinct STAT proteins on genome in CRC cells (Fig. 4). We speculate that a series of other epigenetic factors or transcription factors coordinated with certain STAT may facilitate the recruitment of STAT towards the target binding sites such as cyclic AMP response element-binding protein (CBP)/EA1 binding protein p300 (p300), SMAD family members (SMADs), minichromosome maintenance complex component 5 (MCM5), glucocorticoid receptor (GR) and N-myc and STAT interactor (NMI), and these factors are tightly connected with tissue or disease specificity [23]. Mechanistically, these interactions appear to be divided into two distinct categories, one for DNA-binding stabilization and the other for enhancement of transcription factor activity without affecting the association with DNA.
In our study, the extent DNA motif assay and IP study both suggest that NR5A2 is a potential specific co-activator of STAT3 in CRC. NR5A2 also named as Liver receptor homolog-1 (LRH1) plays a constitutively active role in driving the transcription of its target genes for intestinal function, coordinating with cell regeneration and immunologic function with implications to regular intestinal diseases [24]. Previous study has also showed that NR5A2 can assist in DNMT1 recruitment on CpG islands in neurons [25], indicating that NR5A2, beyond the transcription factor, also acts as an epigenetic regulator to guide and stabilize the DNA–protein complex formation. In our system, we may infer that reinforce of STAT3 homodimer is likely to be attributed to NR5A2. The predicted protein docking indicates that NR5A2 seemingly approaches to the coiled-coil domain of STAT3, in which these α-helices is crucial for interplay between STAT3 and its receptor. Then, NR5A2 can further facilitate the effects of tyrosine phosphorylation on stimulation by epidermal growth factor (EGF) or interleukin-6 (IL-6) for dimer formation, nuclear translocation and genomic binding [26, 27]. Although we have discussed about the structural insights of NR5A2-STAT3 based on protein docking, this new crystal complex needs to be further deciphered and illustrated in vitro in future study.
Finally, more questions are raised about our observations. First, the function of STAT proteins in non-coding regions are not studied although 30% of them are observed to occupy at distal intergenic regions (Fig. 2D). Second, there are still some regions where homodimer and heterodimer of STAT proteins are overlapping in CRC (Fig. 4), but which combination of dimer drives the transcription activity of target gene more powerfully is unknown. More extensively, disclosure of the proteomics even non-coding RNA profiles physically linked with activated STATs within nucleus is a valuable research direction from health to disease.
Conclusion
In summary, we characterize the genome-wide landscape of activated STAT proteins, and reveal the differences of binding patterns as well as the target genes and associated functions between homodimer and heterodimer of STAT proteins in HCT-116 cells. We also present some new findings and possible mechanisms regarding the role of NR5A2 on STAT3 in CRC. Our findings may provide new insights into the design of STAT inhibitors to treat CRC and other diseases.
References
Vainchenker W, Constantinescu SN. JAK/STAT signaling in hematological malignancies. Oncogene. 2013;32:2601–13.
Groner B, von Manstein V. Jak Stat signaling and cancer: Opportunities, benefits and side effects of targeted inhibition. Mol Cell Endocrinol. 2017;451:1–14.
Darnell JE Jr. STATs and gene regulation. Science. 1997;277:1630–5.
Otero-Muras I, Yordanov P, Stelling J. Chemical reaction network theory elucidates sources of multi-stability in interferon signaling. PLoS Comput Biol. 2017;13: e1005454.
Zhang Y, Zhang Y, Yun H, Lai R, Su M. Correlation of STAT1 with apoptosis and cell-cycle markers in esophageal squamous cell carcinoma. PLoS ONE. 2014;9: e113928.
Sellier H, Rebillard A, Guette C, Barre B, Coqueret O. How should we define STAT3 as an oncogene and as a potential target for therapy? Jak-Stat. 2013;2: e24716.
Good SR, Thieu VT, Mathur AN, et al. Temporal induction pattern of STAT4 target genes defines potential for Th1 lineage-specific programming. J Immunol. 2009;183:3839–47.
Walford HH, Doherty TA. STAT6 and lung inflammation. Jak-Stat. 2013;2: e25301.
Heltemes-Harris LM, Farrar MA. The role of STAT5 in lymphocyte development and transformation. Curr Opin Immunol. 2012;24:146–52.
Murray PJ. The JAK-STAT signaling pathway: input and output integration. J Immunol. 2007;178:2623–9.
Tang S, Yuan X, Song J, Chen Y, Tan X, Li Q. Association analyses of the JAK/STAT signaling pathway with the progression and prognosis of colon cancer. Oncol Lett. 2019;17:159–64.
Wang SW, Sun YM. The IL-6/JAK/STAT3 pathway: potential therapeutic strategies in treating colorectal cancer (Review). Int J Oncol. 2014;44:1032–40.
Slattery ML, Lundgreen A, Kadlubar SA, Bondurant KL, Wolff RK. JAK/STAT/SOCS-signaling pathway and colon and rectal cancer. Mol Carcinog. 2013;52:155–66.
Dariya B, Muppala S, Srivani G, Momin S, Alam A, Saddala MS. Targeting STAT proteins via computational analysis in colorectal cancer. Mol Cell Biochem. 2021;476:165–74.
Zhou S, Shen Y, Zang S, Yin X, Li P. The epigenetic role of HTR1A antagonist in facilitaing GnRH expression for pubertal initiation control. Mol Ther Nucleic Acids. 2021;25:198–206.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER Suite: protein structure and function prediction. Nat Methods. 2015;12:7–8.
Pierce BG, Wiehe K, Hwang H, Kim BH, Vreven T, Weng Z. ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics. 2014;30:1771–3.
Jiang L, Zhao XH, Mao YL, Wang JF, Zheng HJ, You QS. Long non-coding RNA RP11–468E2.5 curtails colorectal cancer cell proliferation and stimulates apoptosis via the JAK/STAT signaling pathway by targeting STAT5 and STAT6. J Exp Clin Cancer Res. 2019;38:465.
Murata T, Noguchi PD, Puri RK. IL-13 induces phosphorylation and activation of JAK2 Janus kinase in human colon carcinoma cell lines: similarities between IL-4 and IL-13 signaling. J Immunol. 1996;156:2972–8.
Luan J, Fu J, Wang D, et al. miR-150-Based RNA interference attenuates Tubulointerstitial fibrosis through the SOCS1/JAK/STAT pathway in vivo and in vitro. Mol Ther Nucleic Acids. 2020;22:871–84.
Pu Z, Xu M, Yuan X, Xie H, Zhao J. Circular RNA circCUL3 accelerates the Warburg effect progression of gastric cancer through regulating the STAT3/HK2 Axis. Mol Ther Nucleic Acids. 2020;22:310–8.
Lim CP, Cao X. Structure, function, and regulation of STAT proteins. Mol BioSyst. 2006;2:536–50.
Decker T, Kovarik P. Transcription factor activity of STAT proteins: structural requirements and regulation by phosphorylation and interacting proteins. Cell Mol Life Sci. 1999;55:1535–46.
Fernandez-Marcos PJ, Auwerx J, Schoonjans K. Emerging actions of the nuclear receptor LRH-1 in the gut. Biochem Biophys Acta. 2011;1812:947–55.
Shen Y, Zhao H, Zhang L, et al. The roles of DNA methylation and hydroxymethylation at short interspersed nuclear elements in the hypothalamic arcuate nucleus during puberty. Mol Ther Nucleic Acids. 2021;26:242–52.
La Sala G, Michiels C, Kukenshoner T, et al. Selective inhibition of STAT3 signaling using monobodies targeting the coiled-coil and N-terminal domains. Nat Commun. 2020;11:4115.
Zhang T, Kee WH, Seow KT, Fung W, Cao X. The coiled-coil domain of Stat3 is essential for its SH2 domain-mediated receptor binding and subsequent activation induced by epidermal growth factor and interleukin-6. Mol Cell Biol. 2000;20:7132–9.
Acknowledgements
This work was supported by Science and Technology Development Plan of Suzhou (Medical and Health Science and Technology Innovation) (SKYD2022023), Start-up Funding of Suzhou Ninth People’s Hospital, Youth Program of Developing Public Health through Science and Education of Suzhou (KJXW2019069), Program of Developing Public Health through Science and Education of Wujiang District (wwk201811), The People’s Livelihood Science and Technology of Suzhou (SYSD2020043), Suzhou science and technology planning project (SKJY2021016), Joint Co-construction Project of Henan Medical Science and Technology Research Plan (LHGJ20210199), Key R & D and promotion projects in Henan Province of 2020 (scientific and technical program) (222102310266). We are grateful for the assistance from Shanghai Genefund Biotech Co. Ltd. in the generation and analysis of high-throughput sequencing data. We are also grateful for Coweldgen Scientific Co. Ltd. for STR authentication of cell lines.
Author information
Authors and Affiliations
Contributions
HY and SYF performed the cellular and molecular experiments and analyzed data. HY and ZY provided the clinical samples. TY and GY helped cell culture and molecular experiments. LC helped bioinformatic analysis. YT provided the major financial support for the project. SYH designed the research route, and was responsible for quality control of all raw data, and drafted and revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared that they have no conflicts of interest to this work. The datasets and supporting materials generated during and/or analysis during the current study are available from the corresponding author on reasonable request.
Ethical approval
The research protocol was approved by Ethics Committee of Henan Cancer Hospital (2022-KY-0073-001). The experiments were conducted in accordance with the Declaration of Helsinki (2000 version).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2022_815_MOESM1_ESM.xlsx
Supplementary file1 Table S1-5. The genomic annotation of STAT1 (S1), STAT2 (S2), STAT3 (S3), STAT5 (S4) and STAT6 (S5) in HCT116 cells. Column E lists the nearby gene element if exists. Column I-K list the information of probable typical enhancer (TE) and super-enhancer (SE) of HCT116 cells based from SEdbV1.05 Database. (XLSX 5445 KB)
13577_2022_815_MOESM6_ESM.xlsx
Supplementary file6 Table S6. The genomic annotation of binding areas with probable dimers with different STAT combinations. In Column D-H, the positive (“1”) and negative (“0”) abundance of the given protein are listed. In Column I, number means how many kind of STAT proteins bind on this area. In Column J to AD, difference of binding abundance between two STAT proteins are compared and shown by fold change and FDR. (XLSX 5909 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, Y., Shen, Y., Zhao, Y. et al. Genomic distribution of signal transducer and activator of transcription (STAT) family in colorectal cancer. Human Cell 36, 286–295 (2023). https://doi.org/10.1007/s13577-022-00815-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-022-00815-0