Abstract
MicroRNAs serve a crucial role in the regulation of malignant biological behavior of Ewing’s sarcoma (ES). Abnormal expression of miR-107 has been reported in a cohort of cancers, while its exact function in ES remains unclear. Hence, we explored the expression of miR-107 in ES cells and detected its effects on the malignant phenotype of ES cells. Firstly, we perceived the under-expression of miR-107 in human ES cells contrast with the human mesenchymal stem cells. Over-expression of miR-107 restrained cell proliferation and tube formation, arrested cell cycle progression, and facilitated cell apoptosis in SK-ES-1 and RD-ES cell lines. Furthermore, hypoxia inducible factor-1β (HIF-1β) was assumed as a target gene of miR-107. We confirmed the target role of HIF-1β in ES cells. Finally, restoring the expression of HIF-1β could partly abolish miR-107-mediated tumor suppression in ES cells. In conclusion, our results advised that miR-107 suppressed the malignant biological ability of ES cells through targeting HIF-1β.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ewing’s sarcoma (ES) is a primary malignant bone tumor in children and adolescents, accounting for 6–8% of the primary bone tumors, with rapid progression and high mortality [1]. Since majority part of death cases accompanied with tumor metastasis, the high recurrence and metastasis have become a big obstacle in the treatment of ES.
MiRNAs (microRNAs) are kinds of single-stranded non-coding and small molecule RNAs, which could regulate the expression of genes transcriptionally [2]. Abnormal expression of miRNAs has been widely reported in the tumorigenesis and development of variety tumors, including ES [3, 4]. Among the miRNAs being reported, miR-107 is a newly reported miRNA, whose expression was abnormal in many kinds of cancers. In osteosarcoma, miR-107 was down-regulated in OS tissues, and overexpression of miR-107 inhibited cell proliferation and promoted cell apoptosis of OS in vitro [5]. However, its putative role in ES has not been identified yet. In this paper, we will explore the expression and its putative biological effects in ES and unravel potential molecular mechanisms might be involved in ES tumorigenesis.
Materials and methods
Cell lines and cell culture
Human Ewing’s sarcoma cell lines A673, SK-ES-1 and RD-ES were procured from the American Type Culture Collection (ATCC). MSCs were grown in IMDM, 10% Fetal Calf Serum (FCS) and PDGF-BB (10 ng/ml). A673, RD-ES and SK-ES-1 cells were grown in 1640 medium augment with 10% Fetal Bovine Serum (FBS), streptomycin (100 µg/ml), and penicillin (100 U/ml). Human mesenchymal stem cells (MSCs) used in our experiments were obtained from normal adult human bone marrow withdrawn from bilateral punctures of the posterior iliac crests of three normal volunteers. MSCs were cultured at low confluence in IMDM, 10% FBS, and 10 ng/ml PDGF-BB (PeProtechEC). All cells were incubated at 37 °C in 5% CO2 cell culture incubator. All volunteers have written informed consent forms and use of these tissue samples has been approved by Ethics Committee of Chongqing Medical University.
Oligonucleotide transfection
MiR-107 mimic and scramble mimic oligonucleotides were purchased from Dharmacon (Dharmacon, Austin, USA) and transfected into SK-ES-1 and RD-ES cells with 50 nM using Dharmfect 1 (Dharmacon, Austin, USA) according to the manufacturer’s protocols. Medium was changed 6 h after transfection. Cells were grown for about 48 h and collected for further experiments.
RNA extraction and RT-quantitative PCR
To quantitate miR-107 expression in Ewing’s sarcoma cells, total RNA was extracted from the cells with the Trizol reagent (Invitrogen, USA) according to the protocol. RT-quantitative PCR analysis was used to explore the content of RNA transcripts. In a nutshell, cDNA was interosculated through M-MLV reverse transcriptase (Invitrogen, USA) and RT primer was applied to the reverse transcription of mRNA. RT-qPCR was performed on the CFX96TM Real-Time PCR Detection System (Bio-Rad, USA) using the Quanti-Tect SYBR Green PCR mixture. The cycling contexts were used by TaqMan probes (Invitrogen, USA): 95 °C for 30 s (initial denature); then 40 cycles of 95 °C for 10 s, 60 °C for 30 s and 72 °C for 30 s. GAPDH were used for mRNA normalization.
Vector construction and dual-luciferase reporter analysis
To demonstrate whether HIF-1β was regulated by miR-107 through precisely targeting its 3′-UTR, we performed Dual-luciferase reporter assays. The full-length of 3′-URT of the HIF-1β mRNA was amplified from genomic DNA utilizing primers of HIF-1β-UTR-F/R and next cloned into the XbaI and NotI sites of the pGL-3 vector (Promega, USA). The mutant construct of 3′-UTR of HIF-1β was introduced through the QuickChange site-directed mutagenesis kit (Stratagene, USA). The luciferase reporter structure containing the HIF-1β target sequence acted as the positive control, and the pRL-TK vector acted as the internal control. Approximately, 1 × 105 cells were seeded into each well of 24-well plates for 24 h before transfection. Cells were transfected with the pGL-3 firefly luciferase reporter (50 ng/well), pRL-TK Renilla luciferase reporter (10 ng/well), and the miR-107/scramble mimic (40 nM). All of the transfections were accomplished using Lipofectamine 2000 (Invitrogen, USA). 48 h later after transfection, Passive Lysis Buffer (Promega, USA) was used to prepare the cell lysates, and the activity of luciferase was estimated by the Dual-Luciferase Reporter analysis (Promega, USA). Results were normalized to the Renilla luciferase.
Cell cycle and cell proliferation assays
For ES cell proliferation assays, about 8–10 × 103 cells were resuspended into each well of 24-well plates. Cells were incubated into plate compared with CCK-8 (10 µl, Dojindo) at 37 °C for 2 h. Proliferation rates were determined at 0, 24, 48 and 72 h after transfection. The absorbance of per well was measured with a microplate reader set at 450 nM. All proliferation experiments were performed in triplicity. Cell cycle assays were enforced in SK-ES-1 and RD-ES cells 48 h after transfection. After cells were harvested, cells were washed with cold PBS, fixed in cold ethanol (70%), and further hatched with the propidium and RNase A, and finally analyzed through FACS.
Cell apoptosis analysis
In the cause of explore the apoptosis of RD-ES and SK-ES-1 cells, flow cytometric analysis was equipped with PE-Annexin V/7AAD (BD Pharmingen, USA) by the instructions.
Tube formation assays
24 h after transfection with miR-107/scramble mimic, about 1 × 104 cells were resuspended into each well of 48-well plates with Matrigel (60 µl, BD Bioscience, USA). Microscope was used to observe the structure of tubes after 12-h incubation. The branch point of tube structure was quantified by the protocols.
Protein extraction and western blotting
Immunoblot analysis was accomplished by standard methods. Proteins were analyzed by 10% SDS-PAGE, and then electroblotted onto PVDF membranes (Millipore, USA). Membranes were incubated overnight at 4 °C with primary antibody after blockage for 2 h with 5% non-fat dried milk. Detection was accomplished through peroxidase-conjugated secondary antibodies by the enhanced chemiluminescence system (Millipore, USA). Primary antibodies were HIF-1β (Cell Signaling Technology, USA) and GAPDH (ZSGB-BIO, Beijing, China). After rinsing with TBST (10 mM Tris, pH 8.0, 150 mM NaCl, and 0.1% Tween20), the membranes were incubated for 2 h at room temperature with goat anti-rabbit antibody (ZSGB-BIO, Beijing, China) at 1:20,000.
Plasmid construction
To over-express HIF-1β, the human HIF-1β gene open-reading frame (ORF) was inserted into pcDNA-3.1 construct to generate the pcDNA-3.1-HIF-1β construct. The control was empty pcDNA-3.1. Cells were transfected with miR-107 mimic or scramble mimic (60 nM) into six-well plates by the manufacturer’s protocols.
Statistical analysis
Each experience should be repeated at least three times. Student’s t test was enforced and two-group data were analyzed through one-way analysis of variance. All of statistical analyses were enforced by SPSS 15.0 software. P values <0.05 were considered statistically significant.
Results
Aberrant expression of miR-107 in human ES cell lines
To explore the putative effects of miR-107 in ES, we firstly conducted its expression in a cohort of ES cell lines. Compared with the human mesenchymal cells (hMSCs), which were reported to be the original cells of ES, miR-107 was consistently under-expressed in human ES cell lines (Fig. 1a). These results made us speculate that miR-107 might act as a tumor suppressor in human ES cells. To further identify the effects of miR-107 in ES, the following biological function assays were carried out in SK-ES-1 and RD-ES cell lines. The expression of miR-107 was restored in ES cells upon transfection with miR-107 mimic (Fig. 1b).
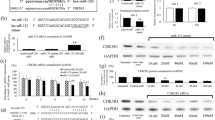
The expression of miR-107 in Ewing’s sarcoma (ES) cell lines. a MiR-107 was under-expressed in ES cells contrasted with human mesenchymal stem cells (MSCs) by the real-time quantitative PCR analysis. b The miR-107 expression in SK-ES-1 and RD-ES cells transfected with miR-107 mimic was statistically upregulated contrasted with scramble group; **P < 0.01
Overexpression of miR-107 inhibits cell proliferation in human ES cell lines
To detect the effects of miR-107 on the proliferation of ES cells, CCK-8 assays were performed. There was no significant deviation in the growth rates between the SK-ES-1 and RD-ES cells transfected with miR-107 or scramble mimic within 24 h of transfection. However, upon 48 h of transfection, significant suppressed growth rates were found in miR-107 group comparing with the scramble group (Fig. 2a), demonstrating that transfection with miR-107 inhibited cells proliferation of ES cells.
Over-expression of miR-107 inhibited cell proliferation, arrested G0/G1 cell cycle, promoted cell apoptosis and restrained angiogenesis. a Cell viability was detected by cell counting kit-8 (CCK-8) analysis upon 72 h of transfection. MiR-107 significantly inhibited cell proliferation of ES cells. b Cell cycle analysis according to fluorescence-activated cell sorting (FACS) (left side) at 48 h after transfection, and histogram (right side) suggested miR-107 arrested cell cycle in the G0/G1 phase. c Cell apoptosis analysis by FACS (left side) at 48 h after transfection and histogram (right side) suggested miR-107 promoted early apoptosis in ES cells. d Cell angiogenesis analysis by microscope (left side) at 12 h after transfection and histogram (right side) showed miR-107 restrained angiogenesis in cells; *p < 0.05, **p < 0.01
Overexpression of miR-107 arrests cell cycle and induces cell apoptosis in human ES cell lines
Next, the effects of miR-107 on cell cycle progression of ES cells were analyzed using FCM assays. Compared with cells transfected with scramble mimic, cells treated with miR-107 mimic showed increased ratio of cells in G1 phase and decreased ratio in S phase in both cell lines (Fig. 2b). The influence on the cell count of G2 phase was not that significant. Furthermore, we also detected the effects of miR-107 on cell apoptosis of ES cells. The results suggested that the early apoptotic rate of cells transfected with miR-107 mimic was significantly higher than the cells transfected with scramble mimic (Fig. 2c). Taken together, our results suggested that miR-107 inhibited the cell proliferation and promoted cell apoptosis of ES cells.
Transfection with miR-107 inhibit tube formation of ES cell lines
Tube formation of tumor cells performs an important role during the progression of tumor invasion and metastasis [6]. MiR-107 has been reported to perform pivotal role in the angiogenesis of endothelial progenitor cells [7]. Thus, we speculated that miR-107 might influence tube formation of human ES cells. We detected the influence of miR-107 on the tube formation of RD-ES and SK-ES-1 cells using tube formation assays. Tubes formation of SK-ES-1 and RD-ES cells was quantitated according to branchpoint counting. As shown in Fig. 2d, the number of branches in miR-107 mimic group was less than that in the scramble group, demonstrating that miR-107 could suppress the tube formation of ES cells in vitro.
MiR-107 regulated the HIF-1β gene in human ES cell lines directly
To investigate the mechanisms involved in miR-107-mediated suppressive effects on human ES cells, the target genes of miR-107 were searched by the target prediction softwares, miRanda and PicTar. Among the results predicted, hypoxia inducible factor-1β (HIF-1β) caught our attention most, as it served a vital role in regulation of malignant phenotype of many tumors (Fig. 3a). Relative luciferase activity was conducted to prove the target role of HIF-1β in human ES cells, on account of miR-107 over-expression suppressed the activity of the luciferase reporter which contained the 3′-UTR of HIF-1β, but did not influence the luciferase activity of the mutated 3′-UTR structure (Fig. 3b). Moreover, miR-107 over-expression also affected the protein and mRNA levels of HIF-1β upon transfection. The protein and mRNA levels of HIF-1β in miR-107 mimic groups were substantially decreased than scramble mimic group after transfection (Fig. 3c, d). Thus, HIF-1β was hypothesized to serve as a functional target gene of miR-107 in human ES cells.
Hypoxia-inducible factor-1β (HIF-1β) is a direct target gene of miR-107 in ES cells. a Graphical presentation of 3′-UTR of HIF-1β involving the suggested miR-107 target site. b Relative luciferase activity of the bespoken HIF-1β reporter structure in SK-ES-1 and RD-ES cells. MiR-107 significantly inhibited the luciferase activity of the report gene involving 3′-UTR of HIF-1β WT instead of 3′-UTR of HIF-1β MUT in either cell lines. c MiR-107 over-expression significantly decreased the mRNA level of HIF-1β in RD-ES and SK-ES-1 according to real-time quantitative PCR. d MiR-107 Over-expression significantly decreased the protein level of HIF-1β in RD-ES and SK-ES-1 according to western blot assays; *P < 0.05, **P < 0.01
HIF-1β was involved in miR-107-mediated tumor suppression
The previous observations showed that HIF-1β was a target gene of miR-107 in human ES cells, but the outcome of miR-107/HIF-1β reciprocity in human ES was unknown. Hence, we decided to take the “rescue” methodology to detect whether HIF-1β was involved in miR-107-mediated tumor suppression. To recover the HIF-1β expression in ES cells which had been transfected by miR-107 mimic before. Western blot assays were used to score the HIF-1β expression in SK-ES-1 cells. The level of HIF-1β expression was restored upon transfection with HIF-1β constructs in SK-ES-1 cells which have been treated with miR-107 mimic before (Fig. 4a). MiR-107-mediated inhibition on cell proliferation and induction on cell apoptosis were mitigated by transfection of HIF-1β constructs (Fig. 4b, c). Furthermore, this phenomenon was also observed in tube formation assays. Although transfection with miR-107 decreased the number of branches, transfection with HIF-1β constructs re-increased the number of new branches comparing with the cells transfected with control constructs (Fig. 4d). These results demonstrated that HIF-1β might be involved in miR-107-mediated tumor suppression in human ES cells.
HIF-1β participates in miR-107-mediated tumor suppression. a The restored protein expression of HIF-1β was rescued through western blot analysis. b To perceive and explore the effect of HIF-1β on cell proliferation of SK-ES-1 cells, CCK-8 assays were used. Restored HIF-1β expression rescinded miR-107-mediated inhibition on cell proliferation. c Cell apoptosis analysis showed restoration HIF-1β expression suppression early apoptosis in SK-ES-1 cells. d Cell angiogenesis analysis by microscope (left side) and histogram (right side) showed restoration HIF-1β expression promoted angiogenesis in SK-ES-1 cells; *P < 0.05
Discussion
Ewing’s sarcoma (ES) is a kind of the malignant tumor originated from bone and soft tissues with low differentiation, which is characterized with small round cells, along with high malignant degree, short duration and fast metastasis [8]. In recent years, with the rapid development of operative treatment combined with radiotherapy and chemotherapy, the survival rate of ES patients has been improved, while is still unsatisfactory [9]. Thus, understanding the mechanisms of carcinogenesis of ES would provide new therapeutic strategies in the future.
MiRNAs are kinds of single-stranded non-coding small molecule RNAs [10]. miRNAs might regulate the expression of thousands of downstream genes, and different miRNAs also regulate the same proteins through diverse of cross accesses [11]. These complex network structures adjust the cellular metabolic in vivo. Once the network was disordered, tumorigenesis might be happened [12]. Except for miR-124, let-7a we reported previously, many miRNAs were also reported to perform as tumor suppressor in ES, such as miR-34 family [13]. Moreover, Lida et al. found that miR-125b might be related with chemoresistance of ES, as miR-125b was found to be up-regulated in Dox-resistant ES cells and knock-down of miR-125b showed enhanced sensitivity to doxorubicin [14]. Understanding the miRNA expression profile might provide potential for prediction of disease progression and survival of ES, and provide new strategy for tumor therapy, especially for the one with chemoresistance.
The effects of miRNA in different tumor development and progression were duplicity. MiR-107 is a newly reported miRNA, whose effects in the development and progression of tumor were contradictory. In lung cancer, miR-107 was reported to inhibit cell cycle progression and induce apoptosis [15, 16], while in gastric and breast cancer cells, miR-107 could promote tumor invasion and metastasis [17, 18]. Herein, we found that the expression of miR-107 was suppressed in human ES cell lines. Moreover, recovered the expression of miR-107 significantly suppressed the malignant biological behaviors of SK-ES-1 and RD-ES cells; as cell proliferation was decreased, cell apoptosis was induced and the cell cycle was redistributed in the G0/G1 phase. All these results demonstrated that miR-107 might perform as a tumor suppressor in ES cells.
Angiogenesis is vital for tumor growth and its metastases, as tumors cannot grow in dimensions greater than 2 mm without angiogenesis [19]. Tube formation of tumor cells was the first step for supplement of oxygen and nutrition. Importantly, the presence of blood lakes lines, as present as the tube formation of ES cells, is a striking feature of ES, which might be correlated with poor clinical outcomes of tumor [20]. MiR-107 was reported to perform suppressive effects on angiogenesis of many cancers, such as through targeting Dicer-1-mediated suppression on VEGF, miR-107 participated in the regulation of post-stroke angiogenesis [21]. Herein, we found that miR-107 over-expression could significantly inhibit the tube formation of ES cells in vitro, proposing that miR-107 might participate in the angiogenesis of ES.
HIF-1β was a subunit and constitutive expression of HIF-1, which participated in tumorigenesis and progression in tumors [22]. Tumor cell proliferation under the condition of the hypoxia might be monitored by the HIF-1β gene [23], and the expression of HIF-1β was positively correlated with the malignant biological of the tumor in vivo [24]. Moreover, miR-107 was reported to regulate cell migration and promote HIF-1-regulated angiogenesis under hypoxia [25]. Among the target gene predicted by biological informatics, we also found that HIF-1β was one of the putative target genes. To investigate whether HIF-1β was involved in miR-107-mediated tumor suppression in human ES cells. Firstly, we accomplished studies to give evidence of the target role of HIF-1β in human ES cells, as the luciferase activity containing the 3′-UTR of HIF-1β was decreased, and the protein and mRNA levels were suppressed upon transfection with miR-107. Videlicet, miR-107 under-expression may increase the expression of HIF-1β and thus contribute to tumorigenesis and progression in ES. As expected, the suppression of miR-107 could be rescued through restoring the expression of HIF-1β in cells. Our results are partially consistent with Yamakuchi et al. who suggested that miR-107 was reported to promote HIF-1-regulated angiogenesis under hypoxia. We also found that miR-107 could inhibit the tube formation of ES cells and suppression could be abolished by re-expression of HIF-1β.
In conclusion, our results showed that miR-107 was suppressed in ES cells, and overexpression of miR-107 could restrain the malignant biological ability of ES cells through inhibiting the HIF-1β expression. Here, we show that miR-107 is crucial for ES progression, and this miRNAs may provide invaluable information for ES treatment.
References
Thorn D, Mamot C, Krasniqi F, Metternich F, Prestin S. Multimodality treatment in Ewing’s sarcoma family tumors of the maxilla and maxillary sinus: review of the literature. Sarcoma. 2016;2016:3872768. doi:10.1155/2016/3872768.
Rupaimoole R, Calin GA, Lopez-Berestein G, Sood AK. miRNA deregulation in cancer cells and the tumor microenvironment. Cancer Discov. 2016;6(3):235–46. doi:10.1158/2159-8290.CD-15-0893.
Slattery ML, Herrick JS, Mullany LE, Wolff E, Hoffman MD, Pellatt DF, et al. Colorectal tumor molecular phenotype and miRNA: expression profiles and prognosis. Mod Pathol. 2016;29(8):915–27. doi:10.1038/modpathol.2016.73.
Zhang Z, Li Y, Huang L, Xiao Q, Chen X, Zhong J, et al. Let-7a suppresses macrophage infiltrations and malignant phenotype of Ewing sarcoma via STAT3/NF-kappaB positive regulatory circuit. Cancer Lett. 2016;374(2):192–201. doi:10.1016/j.canlet.2016.02.027.
Zhang ZC, Liu JX, Shao ZW, Pu FF, Wang BC, Wu Q, et al. In vitro effect of microRNA-107 targeting Dkk-1 by regulation of Wnt/beta-catenin signaling pathway in osteosarcoma. Medicine. 2017;96(27):e7245. doi:10.1097/md.0000000000007245.
Xu H, Zhang Y, Pena MM, Pirisi L, Creek KE. Six1 promotes colorectal cancer growth and metastasis by stimulating angiogenesis and recruiting tumor-associated macrophages. Carcinogenesis. 2017;. doi:10.1093/carcin/bgw121.
Meng S, Cao J, Wang L, Zhou Q, Li Y, Shen C, et al. MicroRNA 107 partly inhibits endothelial progenitor cells differentiation via HIF-1beta. PLoS ONE. 2012;7(7):e40323. doi:10.1371/journal.pone.0040323.
Tarazona N, Navarro L, Cejalvo JM, Gambardella V, Perez-Fidalgo JA, Sempere A, et al. Primary paraesophageal Ewing’s sarcoma: an uncommon case report and literature review. Onco Targets Ther. 2015;8:1053–9. doi:10.2147/OTT.S80879.
Grassetti L, Torresetti M, Brancorsini D, Rubini C, Lazzeri D, Di Benedetto G. A peculiar case of large primary cutaneous Ewing’s sarcoma of the foot: case report and review of the literature. Int J Surg Case Rep. 2015;15:89–92. doi:10.1016/j.ijscr.2015.08.024.
Ren G, Yu B. Post-transcriptional control of miRNA abundance in Arabidopsis. Plant signal Behav. 2012;7(11):1443–6. doi:10.4161/psb.21956.
Sen CK, Ghatak S. miRNA control of tissue repair and regeneration. Am J Pathol. 2015;185(10):2629–40. doi:10.1016/j.ajpath.2015.04.001.
Osaki M, Okada F, Ochiya T. miRNA therapy targeting cancer stem cells: a new paradigm for cancer treatment and prevention of tumor recurrence. Therap Deliv. 2015;6(3):323–37. doi:10.4155/tde.14.122.
Nugent M. microRNA and bone cancer. Adv Exp Med Biol. 2015;889:201–30. doi:10.1007/978-3-319-23730-5_11.
Iida K, Fukushi J, Matsumoto Y, Oda Y, Takahashi Y, Fujiwara T, et al. miR-125b develops chemoresistance in Ewing sarcoma/primitive neuroectodermal tumor. Cancer Cell Int. 2013;13(1):21. doi:10.1186/1475-2867-13-21.
Cui J, Mo J, Luo M, Yu Q, Zhou S, Li T, et al. c-Myc-activated long non-coding RNA H19 downregulates miR-107 and promotes cell cycle progression of non-small cell lung cancer. Int J Clin Exp Pathol. 2015;8(10):12400–9.
Zhang Z, Zhang L, Yin ZY, Fan XL, Hu B, Wang LQ, et al. miR-107 regulates cisplatin chemosensitivity of A549 non small cell lung cancer cell line by targeting cyclin dependent kinase 8. Int J Clin Exp Pathol. 2014;7(10):7236–41.
Ayremlou N, Mozdarani H, Mowla SJ, Delavari A. Increased levels of serum and tissue miR-107 in human gastric cancer: correlation with tumor hypoxia. Cancer Biomark. 2015;15(6):851–60. doi:10.3233/CBM-150529.
Wang S, Ma G, Zhu H, Lv C, Chu H, Tong N, et al. miR-107 regulates tumor progression by targeting NF1 in gastric cancer. Sci Rep. 2016;6:36531. doi:10.1038/srep36531.
Villanueva MT. Angiogenesis: a sudden rush of blood to the tumour. Nat Rev Cancer. 2015;15(3):135. doi:10.1038/nrc3914.
van der Schaft DW, Hillen F, Pauwels P, Kirschmann DA, Castermans K, Egbrink MG, et al. Tumor cell plasticity in Ewing sarcoma, an alternative circulatory system stimulated by hypoxia. Can Res. 2005;65(24):11520–8. doi:10.1158/0008-5472.can-05-2468.
Li Y, Mao L, Gao Y, Baral S, Zhou Y, Hu B. MicroRNA-107 contributes to post-stroke angiogenesis by targeting Dicer-1. Sci Rep. 2015;5:13316. doi:10.1038/srep13316.
Deng B, Du J, Hu R, Wang AP, Wu WH, Hu CP, et al. MicroRNA-103/107 is involved in hypoxia-induced proliferation of pulmonary arterial smooth muscle cells by targeting HIF-1beta. Life Sci. 2016;147:117–24. doi:10.1016/j.lfs.2016.01.043.
Lauzier MC, Michaud MD, Dery MA, Richard DE. HIF-1 activation during tumor progression: implications and consequences. Bull Cancer. 2006;93(4):349–56.
Meijer TW, Kaanders JH, Span PN, Bussink J. Targeting hypoxia, HIF-1, and tumor glucose metabolism to improve radiotherapy efficacy. Clin Cancer Res. 2012;18(20):5585–94. doi:10.1158/1078-0432.CCR-12-0858.
Yamakuchi M, Lotterman CD, Bao C, Hruban RH, Karim B, Mendell JT, et al. P53-induced microRNA-107 inhibits HIF-1 and tumor angiogenesis. Proc Natl Acad Sci USA. 2010;107(14):6334–9. doi:10.1073/pnas.0911082107.
Acknowledgements
This project is Funded by the National Nature Science Foundation of China (Grant No. 81502329); Program of Science and Technology of ChongQing Commission (Grant No. KJ1600228); Programs of Yongchuan Hospital of ChongQing Medical University (Grant Nos. YJZQN 201514; YCZQN201511).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no potential conflicts of interest.
Ethical statement
All procedures performed in studies involving human participants were in accordance with the ethical standards of the Ethics Committee of Chongqing Medical University and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Chen, J., Zhou, X., Xiao, Q. et al. MiR-107 suppresses cell proliferation and tube formation of Ewing sarcoma cells partly by targeting HIF-1β. Human Cell 31, 42–49 (2018). https://doi.org/10.1007/s13577-017-0183-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-017-0183-9