Abstract
Adverse environmental conditions greatly influence crop production every year and threaten food security. Plants have a range of signaling networks to combat these stresses, in which several stress-responsive genes and regulatory proteins function together. One such important family of proteins, the Stress Associated Protein (SAP) family, has been identified as a novel regulator of multiple stresses. The SAPs possess a characteristic N-terminal A20 zinc-finger domain combined with either AN1 or C2H2 at the C-terminus. SAPs provide tolerance against various abiotic stresses, including cold, salt, drought, heavy metal, and wounding. The majority of SAPs are stress-inducible and have a function in conferring stress tolerance in transgenics. The role of SAPs in regulating biotic stress responses is a newly emerging field among researchers. SAPs interact with many other proteins to execute their functions; however, the detailed mechanism of these interactions needs to be elucidated. In this context, the present review provides a detailed view of the evolution and functions of SAPs in plants. The involvement in crosstalk between abiotic and biotic stress signaling pathways makes SAPs ideal targets to develop crops with tolerance against multiple stresses without any yield penalty. Altogether, we provide current knowledge on SAPs for investigating their role in stress response, which can further be exploited to develop climate-resilient crops through transgene-based, breeding-mediated, or genome-editing approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
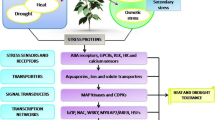
Globally, plant growth, development, and yield are challenged by environmental stresses. However, the plants counteract efficiently by activating a cascade of signals that regulates the expression of various stress-responsive genes like AP2/ERF, MAPKs, MYB, NAC, Stress associated proteins (SAPs), ZIP, etc. Currently, SAPs belonging to the zinc finger protein family have gained recognition for their significant role against various abiotic stresses and plant development (Dixit et al. 2018). They are Zinc finger proteins with characteristic AN1/A20 domains at C and N terminals, respectively (Mukhopadhyay et al. 2004) and act as regulatory proteins with diverse functions in various organisms (Takatsuji 1999). The basic domain organization of SAPs in plants is composed of the A20 domain at N terminal and AN1 domain at C terminal separated by a variable stretch of amino acids (Vij and Tyagi 2008). The A20 domain of SAPs was first identified as a tumor necrosis factor (TNF) induced gene from the human umbilical cord consisting of the Cx2-4Cx11Cx2C consensus sequence, where x can be any amino acid (Opipari et al. 1990). Additionally, the AN1 domain was first discovered in Xenopus laevis, a protein encoded by animal hemisphere 1 (AN1) maternal RNA and comprises of Cx4Cx9-12Cx1-2Cx4Cx2Hx5Hx6 consensus sequence at C terminal (Jin et al. 2007). OsSAP1 was the first SAP that was reported in rice, and its expression was induced in response to several abiotic stresses, like ABA, cold, drought, salt, heavy metals, and wounding. Also, the overexpression of OsSAP1 in transgenic tobacco enhanced the tolerance to salt, dehydration, and cold stress (Mukhopadhyay et al. 2004). Since then, several homologs of SAPs have been identified and characterized in diverse plant species like Arabidopsis thaliana (Vij and Tyagi 2006), Solanum lycopersicum (Solanke et al. 2009), Gossypium hirsutum (Gao et al. 2016), Malus domestica (Dong et al. 2018), Glycine max (Zhang et al. 2019), Brassica napus (He et al. 2019), Populus trichocarpa (Li et al. 2019), and Cucumis sativa (Lai et al. 2020) (Fig. 1).
Molecular phylogenetic analysis of AtSAP5 homologs in different plants along with their domain architecture. The phylogenetic tree was constructed using MEGA 7.0 by Maximum Likelihood method based on the JTT matrix-based model with a bootstrap value of 1000 replicates. The tree with the highest log likelihood (-1551.81) is shown. The percentage of trees in which the associated taxa clustered together is shown next to the branches. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The domains on the right were predicted using NCBI CDD and visualized using Domain Graph v 1.0 (http://dog.biocuckoo.org/). Zm-Zea mays; Ph-Panicum hallii; Si-Setaria italica; Os-Oryza sativa; Pv-Panicum virgatum; Gm-Glycine max; Mt-Medicago truncatula; Eg-Eucalyptus grandis; At-Arabidopsis thaliana; Br-Brassica rapa; Cs-Cucumis sativus; Pp-Prunus persica; Md-Malus domestica; Bd-Brachypodium distachyon; Ma-Musa acuminata; St-Solanum tuberosum
Most SAPs have been characterized to confer stress tolerance in transgenic plants (Giri et al. 2013). For instance, the overexpression of SAP (AtSAP5) from Arabidopsis in cotton conferred tolerance to drought and salt stress (Hozain et al. 2012). Similarly, overexpression of SAP (AlSAP) from Aeluropus littoralis improves the tolerance against cold, drought, and salt stress in transgenic rice (Ben et al. 2012). Hitherto, various homologs of AtSAP5 were identified in different plant species. Apart from abiotic stress tolerance, SAPs are also associated with pathogen defense. A study has revealed that GhSAP17A/D negatively regulates the defense response of cotton to Vertillium dahlia (Gao et al. 2016). Besides, AtSAP5 and its homolog Pha13 from orchid could mediate antiviral immunity in plants (Chang et al. 2018). SAPs also intervene in physiological processes, like growth and development in plants. For example, elevated expression of GhSAPs in the stamen and pistil indicates their possible role in regulating flower development in cotton (Gao et al. 2016). In summary, SAPs are now recognized as potential regulators of plant development and stress responses. Despite the recent progress, understanding the mode of action and precise functions in stress response remains elusive. In this context, the present review enumerates the progress made in understanding the evolution and interplay of SAPs during the stress response and the mechanism underlying plant defense.
Structural features and evolution of SAPs
The identification of SAPs in plants was influenced by their pre-existing occurrence in humans. In humans, the Zinc finger protein A20 serves as an important regulator of innate and adaptive immunity (Mc Dermott and Aksentijevich 2002; Ngo et al. 2014; Peckham et al. 2017). It plays a major role in innate immunity by inhibiting NF-kB signaling, required for immune cell activation in humans (Lu et al. 2013; Ma and Malynn 2012; Wertz et al. 2004). In addition, characterization of other A20 like domain-containing proteins, such as ZNF216, AWP1, and ZNF216 from mice unraveled their involvement in NF-kB signaling. Further, a study revealed that the human zinc finger protein, ZNF216, can regulate immune responses due to the presence of an additional domain, AN1, at the C-terminal (Heyninck and Beyaert 2005). Additionally, many structural studies revealed that four cysteine residues at C-terminal were found to be conserved between OsSAP1, ZNF216, and AWP1 (Mukhopadhyay et al. 2004). However, domain analysis of A20/AN1 protein revealed diversity in domain architecture and variation in domain organization across taxa. Besides, domain comparison of SAPs from plants and animals reported maximum diversity in domain architecture of animal origin SAPs (Vij and Tyagi 2008).
The A20/AN1 zinc finger domains containing SAPs were also identified in some primitive organisms like protists, algae, and fungi. In some protists and fungi, like Plasmodium falciparum, Entamoeba histolytica, Saccharomyces cerevisiae, the AN1 domain exists alone, indicating its primitive nature (Jin et al. 2007; Vij and Tyagi 2008). SAPs from animals showed the presence of additional domains, like OTU, AAA, R3H, UBQ, UIM, and VPS9, associated with specific functions (Hurley et al. 2006). On the contrary, plants show much simpler domain organization (Fig. 2; Table 1) and consist of only one additional domain, i.e., C2H2 linked with AN1. In addition, only a few domains are shared between plants and animals, such as AN1 + A20, AN1, A20, and 2AN1. Among all reported domain architectures in plant SAPs, A20/AN1 is most prevalent, indicating its distinct lineage-specific expansion (LSE) and dynamic evolution in land plants (Table 1). The distinct LSE of a particular domain organization in plants might be due to whole-genome duplication (Lespinet et al. 2002). The presence of distinct domain organization enables SAPs to regulate diverse physiological functions in plants (Vij and Tyagi 2008). Further, the identification and characterization of SAPs in various plant species showed the occurrence of intron-less genes. For example, among 30 SAP genes of Malus domestica, 25 were intron-less (Dong et al. 2018), and in Glycine max, 18 from 27 SAP genes were reported to be intron-less (Zhang et al. 2019). Hence, the occurrence of intron-less SAP encoding genes supports their primitive origin and resemblance with prokaryotic genes. Altogether, these facts indicated that due to reduced post-transcriptional processing of SAPs, they have a primary role in early responses to stress conditions (Grzybowska 2012).
In addition, Kang et al. (2013) had reported that AtSAP5 shared close homology with animal proteins like Rabex-5 and A20 protein. These animal orthologs of AtSAP5 could perform E3 ubiquitin ligase activity and similar action (Kang et al. 2013). Further insights in this study were gained by the prediction of three-dimensional structures of AtSAP5 and its homologs in different plants, Oryza sativa (OsSAP7), Orchid (Pha13), Brassica rapa, Glycine max, wheat, and Cucumis sativus (Fig. 3). The three-dimensional and secondary structure distribution used for the modelling were conserved in all plants. However, due to the lack of X-ray crystallographic structural information of SAPs, formulating a conclusion on the mechanism and alteration of their conformation during ubiquitination will be a daunting task. Nevertheless, the predominance of random coil and alpha-helix in AtSAP5 and its homologs, and their resemblance with animal ortholog Rabex-5, suggested the function as E3 ubiquitin ligase in ubiquitination (Penengo et al. 2006).
The three-dimensional structures of candidate SAPs reported in different plant species predicted using SOPMA tool (https://npsa-prabi.ibcp.fr/cgi-bin/secpred_sopma.pl)
Multitude function of stress associated proteins in plants
SAPs are widely characterized in plants and have been recognized to play a crucial role in enhancing tolerance against abiotic stresses in plants. Comprehensive studies on SAPs revealed multiple modes of action in plants, ranging from ubiquitin ligase (Kang et al. 2011, 2017; Liu et al. 2019; Lloret et al. 2017) to redox sensors (Stroher et al. 2009; Giri et al. 2011), and in the regulation of stress-responsive genes (Li et al. 2019; Lloret et al. 2017; Giri et al. 2011; Hozain et al. 2012; Ben et al. 2012). Moreover, SAPs are also involved in other physiological processes of plants, such as regulation of phytohormone signaling (Chang et al. 2018; Liu et al. 2019; Ben et al. 2020), defense responses against pathogens (Chang et al. 2018; Liu et al. 2019; Sreedharan et al. 2012), cell elongation (Liu et al. 2011), glandular trichome development (Wang et al. 2020) and cell expansion (Lloret et al. 2017), which altogether indicate the diverse functions of SAPs in plants.
SAPs and ubiquitination
Ubiquitination is reversible and often controlled by the alternating action of ubiquitin ligase and deubiquitinases, which are key regulators of various stress responses in plants and animals. The cytoplasm localized mammalian A20 protein mediates the dual function of ubiquitination and de-ubiquitination in NF-kB signaling in humans. The OTU domain of A20 protein at N –terminal is responsible for the K-63 linked de-ubiquitination of receptors like RIP1 (Receptor Interacting serine/threonine Kinase Protein 1) and NEMO (Nf- Kappa B essential modulator), which further disrupts the downstream signaling of NF-kB activation (Mauro et al. 2006; Wertz et al. 2004). Also, the Zinc finger 4 and Zinc finger 7 domain at the C-terminal of the A20 protein is crucial in K48 linked ubiquitination of RIP receptors. Thus, A20 protein disrupts the receptor of NF-kB signaling and hinders the shuttling of NF-kB into the nucleus, thus inhibiting the signaling (Verhelst et al. 2012; Tokunaga et al. 2012). Similar functions of SAPs are reported in plants due to the presence of A20 domain at the N-terminus. Some SAPs have E3 ligase activity and often facilitate abiotic and biotic stress tolerance in plants. In contrast, some play an essential role in ubiquitination by interacting with other proteins of ubiquitination pathways. In 2011, Kang et al. had reported the E3 ligase activity of the AtSAP5 in Arabidopsis. The domain mapping with in-vitro ubiquitin assay revealed ligase activity in the AN1 domain of AtSAP5 due to the replacement of cys6/7 by histidine and his4/5 by cysteine (Kang et al. 2011). This conformational alteration in the AN1 domain resembled the structure of RING Finger proteins, which function as ligase enzyme during ubiquitination (Kang et al. 2011). Further, it was revealed that a highly conserved diaromatic patch in animal orthologs of AtSAP5, i.e., A20 protein and Rabex-5, was important for binding of polyubiquitin. However, the dialipathic patch in AtSAP5 mediates linkage-specific polyubiquitin binding and recognizes K-63 linked polyubiquitination to regulate the downstream signaling of target protein (Choi et al. 2012). A study by Kang et al. (2013) revealed the direct interaction of AtSAP5 with AtMBP-1 to regulate developmental processes and stress response. The expression of AtMBP-1 makes plants hypersensitive to ABA and abiotic stress. However, yeast two-hybrid screening showed direct interaction of AtMBP-1 with AtSAP5. The co-expression analysis of these proteins resulted in the ubiquitination of AtMBP-1 in-vivo, restoring the developmental abnormalities caused by the expression of AtMBP-1 (Kang et al. 2013). Similarly, the ligase activity of SAPs was also reported in other plants, like wheat, where in-vitro ubiquitination assay performed with TaSAP5 resulted in the formation of polyubiquitin chain (Zhang et al. 2017). It was observed that the TaSAP5 mediated the ubiquitination of DRIP (DREB2A–Interacting protein), which is responsible for destabilizing DREB2A, a drought tolerance protein. TaSAP5 enhances DRIP degradation by ubiquitination process, resulting in increased accumulation of DREB2A (Dehydration Responsive Element Binding protein 2A) proteins, thereby indicating the function of TaSAP5 in mediating drought tolerance in wheat (Zhang et al. 2017).
Many studies have reported the interaction of SAPs with other ubiquitination pathway proteins. For instance, Kang et al. (2017) have reported the interaction of AtSAP9 with AtRAD23d (Radiation sensitive 23) protein, which is involved in the transfer of ubiquitinated protein to ubiquitin proteasomes (Kang et al. 2017). Similarly in tomato, Liu et al. (2019) observed the interaction of SlSAP4 with SlRAD23d, mediated by the A20 domain of SlSAP4 and the UBA domain of SlRAD23d. The study also revealed the interaction of SISAP4 with ubiquitin and ubiquitin ribosomal fusion protein (Liu et al. 2019). Interestingly, PpSAP1 regulates leaf morphological alteration by interacting with polyubiquitin proteins (Lloret et al. 2017). Chang et al. (2018) reported E3 ligase and ubiquitin-binding activity in the A20 domain of Pha13, a homolog of AtSAP5. The detailed analysis confirmed the interaction of Pha13 with multiple ubiquitinated proteins (Chang et al. 2018). The deletion and mutant studies of AN1 and A20 domains of Pha13 demonstrated the importance of A20 domain for its function (Chang et al. 2018). However, further structural analysis of A20/AN1 domains is needed to understand their coordination in ubiquitination. Functional analysis of domains across different plant species will assist in understanding the functional deviation in domains of SAPs in different plant species.
SAP-mediated redox regulation
The redox-dependent function of SAPs was reported in AtSAP12 (Ströher et al. 2009). The structural analysis of AtSAP12 showed the presence of 16 cysteine residues in two AN1 domains, of which 12 residues are involved in coordinating zinc ions, indicating their role in redox homeostasis (Ströher et al. 2009). Further, the 2D SDS-PAGE analysis showed the formation of monomers, dimers, and oligomers under oxidizing and reducing conditions, indicating redox-dependent conformational changes by the formation of intra- or inter-molecular disulfide bonds in AtSAP12. Similarly, OsSAP1/11 from rice showed redox-dependent conformational changes to promote its interaction with OsRLCK253, leading to activation of stress response in rice (Giri et al. 2011). However, this hypothesis needs further experimental proof. Although in animals, it has been reported that the NF-kB is dependent on the A20 domain activity for redox control (Bubici et al. 2006). Besides, the function of cysteine residues in mediating the formation of disulphide linkages in SAPs for conformational changes during stress-induced redox changes remains elusive.
Regulation of growth and development by SAPs
Interests have been developed in examining the role of SAPs in various other physiological processes, including growth and development. It has been reported that expression of MtSAP1 in Medicago truncatula was enhanced by six to eight folds during seed maturation, thereby increasing tolerance to the desiccation phase; however, the expression was reduced after few hours of seed imbibition (Gilles et al. 2011). MtSAP1 expression is necessary to accumulate storage proteins, such as globulin, vecilin, and legumin, for efficient germination of seeds (Gilles et al. 2011). Further, overexpression of MtSAP1 in tobacco confers tolerance to abiotic stress by enhancing chlorophyll content and proline accumulation (Charrier et al. 2013). Nevertheless, in rice, OsDOG, dwarf rice with overexpression of gibberellin-induced gene, an A20/AN1 ZFP family protein, acts as a negative regulator of gibberellic acid mediated cell-elongation (Liu et al. 2011). The study reported a reduction in cell elongation of leaf sheath and internodes in the OsDOG overexpressing lines due to a reduced level of GA1. The study reported the novel function of OsDOG in regulating GA homeostasis by modulating GA metabolism genes. Although, the detailed molecular mechanism underlying this signaling needs further work (Liu et al. 2011).
The transcriptional analysis provides more profound knowledge on the molecular functions of SAPs in various biological processes. For instance, the transcriptome data revealed the significant expression of cotton SAPs in pistil and stamen, suggesting their possible role in flower development (Gao et al. 2016). Likewise, overexpression of AtSAP9 delayed the flowering in transgenic lines by reducing the expression of genes involved in flowering time, such as FT (Flowering locus T), SOC1 (Suppressor of overexpression of CO1), and CO1 (Constants). Hence, AtSAP9 is directly or indirectly involved in determining flowering time by CO-dependent pathway (Kang et al. 2017). The orchid SAP, Pha13, a homolog of AtSAP5, is associated with the expression of NPR and NPR-1 independent genes, which are important in salicylic acid-mediated immune pathways (Chang et al. 2018). Besides, the expression of the JA/ET signaling responsive genes were downregulated in SISAP4 silenced Solanum lycopersicum plants after infection with Botrytis cinerea, indicating the role of SlSAP4 in JA/ET signaling (Liu et al. 2018). The SlSAP10 silenced plants witnessed an accelerated yellowing and low chlorophyll content. These results suggest that SlSAP10 plays a crucial role in maintaining the integrity of chlorophyll in plants (Liu et al. 2018).
Apart from growth and development, SAPs also regulate the phytohormone responses in plants. Recently, the A20 domain of LmSAP in Lobularia maritima was identified in regulating GA homeostasis in plants. Overexpression of LmSAP and LmSAPΔAN1 led to the upregulation of GA biosynthetic genes, thus increasing the endogenous GA content, which further affects the GA responsive genes (Ben et al. 2020). Consequently, it is evident that SAP controls diverse functions in plants, and more comprehensive studies are required to unravel the molecular mechanisms underlying these novel functions of SAPs in plants.
Stress associated proteins and molecular stress response
SAPs regulate defense responses in plants and confer stress tolerance. Therefore, SAPs could be important targets for developing stress-tolerant plants, leading to increased crop productivity under prevailing environmental disturbances. Till now, many abiotic stress tolerant transgenic plants harboring different SAPs have been developed (Table 2). SAPs are not only confined to controlling abiotic stress tolerance but are also associated with pathogen defense.
SAPs and multiple abiotic stress tolerance
SAPs mediates stress response by regulating the expression of the stress-responsive genes in plants. Numerous studies showed that the overexpression of SAPs resulted in differential expression of various stress-responsive genes, thereby affecting stress tolerance. The change in expression of stress-related genes was observed in Oryza sativa, Arabidopsis, and Aeluropus littoralis when their respective SAPs, namely OsSAP11, AtSAP5, and AlSAP, were overexpressed in individual host plants (Ben et al. 2012; Giri et al. 2011; Hozain et al. 2012). Further, the transcriptome data also revealed the function of SAPs in regulating the expression of stress-responsive genes (Fig. S1). However, the actual mechanism of the regulation remains elusive and needs further investigation. Similarly, overexpression of PpSAP1 in plum leads to modification of leaf morphology to circumvent drought stress (Lloret et al. 2017). The change in morphology is caused by the downregulation of genes involved in cell growth, like GRF-1 (Growth Regulating Factor), TIP (Tonoplast Intrinsic Protein), and TOR (Target of Rapamycin), in response to overexpressed PpSAP1. Overexpression of PtSAP13 gene in Populus trichocarpa resulted in upregulation of stress-related genes, like BZIP860 (basic leucine zipper transcription factor), DI19-4 (drought-induced gene family), GPX-8 (Glutathione peroxidase 8), NADP-ME2 (NADP malic enzyme 2), RAS1 (Response to ABA and Salt 1), WRKYs (transcription factor associated with drought stress), GSTUs (Glutathione S Transferase), MYBs (MYB domain transcription factor). These genes play a vital role in drought response in plants, and their upregulation conferred tolerance to drought in transgenic Populus plants (Li et al. 2019). Similar results were observed in Glycine max, where overexpression of GmSAP16 affected the expression of stress-responsive genes, such as GmDREB1;1 (Transcription factor involved in drought stress), GmNCED3, and GmRD22, thereby regulating drought stress (Zhang et al. 2019). The elucidation of the function of the first reported SAP, i.e., OsSAP1, in response to various stresses, such as cold, salt, desiccation, and heavy metals, brought new insights for further research in understanding the mechanism of these proteins in stress response (Mukhopadhyay et al. 2004). Consequently, it was identified that SAPs of Sorghum bicolor (SbSAP14), Lobularia maritima (LmSAP), Leymus chinesis (LcSAP), Brassica napus (BnSAP), Sacharum officinarum (ShSAP1), Malus domestica (MdSAP15), Gossypium hirsutum (GhSAP17 A/D), Prunus (PpSAP1), Aeluropus littoralis (AlSAP), Medicago tranculata, and Arabidopsis thaliana (AtSAP5) were induced by multiple abiotic stresses like salt, drought, cold, heat and phytohormone ABA (Ben et al. 2012, 2020; Dong et al. 2018; Gilles et al. 2011; Gao et al. 2016; He et al. 2019; Kang et al. 2011; Liu et al. 2016; Lloret et al. 2017; Wang et al. 2013). Additionally, expression analysis of wheat SAP, TaSAP17D, indicated its involvement in various stresses, such as salt, PEG (Polyethylene glycol), cold, and ABA treatment. Interestingly, transcript levels of TaSAP17D were initially enhanced by NaCl treatment but reduced during later time points of stress (Xu et al. 2018). Similar results were reported in Glycine max, where GmSAP16 showed increased transcript level under drought, salt, and ABA treatment (Zhang et al. 2019), and PtSAP13 was upregulated in Populus trichocarpa, under salt stress (Li et al. 2019). Recently, the transcriptome analysis of cucumber SAPs revealed an upregulation of all CsSAPs under drought treatments (Lai et al. 2020). The result showed that CsSAP5, CsSAP6, CsSAP9, and CsSAP10 were responsive to salt and cold treatment (Lai et al. 2020). Similarly, five SAP genes of the barley, i.e., HvSAP5, HvSAP6, HvSAP11, HvSAP12, and HvSAP15, were found to be upregulated in response to salt stress (Baidyussen et al. 2020).
Extensive studies on SAPs suggested their involvement in early stress response in plants. For example, OsSAP1 transcript abundance was enhanced after 15 min of salt stress and ABA treatment (Mukhopadhayay et al. 2004). Nearly all SAPs in cotton were upregulated within one hour of salt treatment (Gao et al. 2016). However, the time period varies across different plant species. For instance, LcSAP was upregulated to a maximum level within six hours of exposure to NaCl, and the abundance of transcript remains stable for 24 h (Liu et al. 2016). TaSAP17D also showed an early response, and its expression increased within a half-hour of salt stress (Xu et al. 2018). However, LmSAP expression increased after 12 h of treatment with heavy metals (Ben et al. 2018). Seven PtSAP genes were induced in Populus trichocarpa within an hour of salt treatment and reached its maximum expression level in 6 h (Li et al. 2019). Altogether, these studies support the fact that SAPs have a crucial function during early stress response but can vary in different plant species.
The function of SAPs in regulating environmental stress has been successfully deployed in some plants to develop tolerance. Overexpression of SAPs in different plants enhances the growth under stress conditions. For example, the overexpression of OsSAP8 in rice conferred tolerance to salt and drought stress at the anthesis stage without any growth penalty (Kannegati and Gupta 2008). Further, overexpression of AtSAP10 elicited enhanced tolerance to heavy metal and heat stress in transgenic Arabidopsis (Dixit and Dhankher 2011). Similarly, overexpression of MusaSAP1 imparted stress tolerance in transgenic plants due to reduced membrane damage, inhibition of malondialdehyde formation, and strong up-regulation of polyphenol oxidase in response to salt and drought stress (Sreedharan et al. 2012). Likewise, overexpression of SbSAP14 in rice seedlings showed a high germination and survival rate with better tolerance to salt stress than wild type (Wang et al. 2013).
SAPs also impart better adaptive abilities to transgenic plants by regulating distinct morpho-physiological, biochemical, and molecular responses during stress. For example, the overexpression of PtSAP13 in Arabidopsis enhanced flavonoid biosynthesis, leading to improved tolerance to salt stress (Li et al. 2019). Similarly, overexpression of LmSAP in tobacco seedlings enhances their tolerance to heavy metals, like copper, cadmium, and manganese, by increasing the activities of enzymes, such as superoxide dismutase (SOD), catalase (CAT), peroxidase (POD). In addition, transcript levels of specific genes like metallothioneins (Met1, Met2, Met3, Met4, and Met5), CCH (Copper transport proteins), cysteine, and histidine domain-containing protein RAR1 (Rar1), and Ubiquitin like protein 5 (PUB1) were also increased in tobacco seedlings. Further, the expression of AlSAP from Aeluropus littoralis in rice enhances the yield under drought conditions by increasing the tillering and panicle fertility (Thaura et al. 2017). Also, overexpression of PpSAP1 in transgenic plum plants led to leaf shape alterations, hence increasing water retention under drought stress (Lloret et al. 2017). MdSAP15 overexpression in Arabidopsis regulates plant physiological traits during osmotic stress by maintaining chlorophyll content and proline concentration (Dong et al. 2018). Recently, a study indicated that overexpression of GmSAP16 in Arabidopsis improved drought tolerance by regulating stomatal closure and reducing the rate of water loss (Zhang et al. 2019). Altogether, these studies elucidated the function of SAPs in enhancing stress tolerance as well as yield.
SAPs in pathogen defense
SAPs are well characterized in response to abiotic stress. However, SAPs are not confined to abiotic stress response but also regulate biotic stress response in plants. The plant innate immunity, like mammals, depends upon the intracellular nucleotide-binding/ leucine-rich repeat proteins or extracellular transmembrane anchored receptor-like kinases (RLK) or receptor-like proteins (RLP), which further activates downstream signaling (Feehan et al. 2020). The interaction of OsSAP11 with OsRLCK253 via A20 domain directs their function in plant innate immunity (Giri et al. 2011). Defense signaling hormones often induce certain SAPs. MusaSAP1 was induced by wounding and methyl jasmonate treatment, indicating its role in biotic stress response in plants (Sreedharan et al. 2012). In Rice, OsSAP1, is also responsive to various biotic stress and its overexpression in tobacco increases resistance against virulent pathogen Pseudomonas syringae pv. tabaci (Tyagi et al. 2014). Similarly, GhSAP17A/D was induced in response to salicylic acid and methyl jasmonates (Gao et al. 2016). Further, overexpression of AtSAP9 enhances sensitivity for P. syringae pv. Phaseolicola, indicating a negative function of AtSAP9 in immunity (Kang et al. 2017). In the case of Phalaenopsis aphrodite (orchid), Pha13 was induced by exogenous salicylic acid treatment. The Cymibidium mosaic virus-induced gene silencing system (CymMV-VIGS) showed that Pha13 silencing reduces the SA-related genes, thereby affecting the immune response. However, overexpression of Pha13 in orchid and Arabidopsis enhances the resistance to different viruses (Chang et al. 2018). Overexpression of Pha13 in Arabidopsis plants also enhanced their resistance to P. syringae pv. Tomato DC30000. Moreover, overexpression of A20 and AN1 domain double mutants of Pha13 elucidated the involvement of AN1 domain with PhaNPR1 expression, and both the domains are required for imparting resistance to different viruses (Chang et al. 2018). SlSAPs were induced on treatment with salicylic acid, methyl jasmonate, and 1-aminocyclopropane-1-carboxylic acid (ACC). SlSAPs expression was induced within 24 h of Botrytis cinerea infection, leading to enhanced immunity against the necrotrophic fungus through interaction with SIRAD23 and activation of JA/ET signaling. Virus-induced gene silencing (VIGS) and disease phenotyping assays identified that silencing of SlSAP4 and SlSAP10 decreases resistance to B. cinerea, depicting the crucial role of SISAP4 in providing resistance to B. cinerea (Liu et al. 2018). Hence, SAPs play an important role in regulating stress response against pathogen attacks; however, more work is required to understand the hidden molecular mechanisms of their functions.
Furthering the research on stress associated proteins
Currently, only limited knowledge is available about the function of SAPs in regulating various biological processes in plants, including stress responses. The current review summarizes the available knowledge on SAPs in different plant species. In particular, SAPs involved in regulating multiple stresses, should be of great interest to develop stress-tolerant cultivars. To our knowledge, the functional aspects of SAPs in plants against stress responses remain poorly understood. Hence, the advancement in omics approaches can provide in-depth knowledge for future research on the molecular functions of SAP proteins. For example, recently, a transcriptome study by Muthuramalingam et al. (2020) provides annotation of OsSAPs and unveils the role of these proteins in regulating hormonal pathways. Similarly, Lai et al. (2020) revealed transcriptional changes in CsSAP genes in cucumber in response to cold, drought, and salt stresses.
The studies on SAPs hypothesize that these proteins are functionally involved in biological processes such as metabolism, hormonal signaling, translation, developmental regulation, and stress responses. Genome-wide studies have unraveled the tissue level expression profiles of these genes in various plants species. However, it would be worth understanding the interaction of SAPs with various downstream genes involved in biological processes. The role of SAPs as E3 ubiquitin ligase to mediate signal transduction is well discussed in many plant species; however, the detailed mechanism remains elusive. Thus, discovering crystal structures of SAPs (NMR based or X-ray crystallography) having E3 ubiquitin ligase activity will delineate their mechanism of action and underlying pathways associated with ubiquitination. Most of the studies have reported the functions of A20 domain; however, very little information is available for the AN1 domain. Only in AtSAP5, Kang et al. (2011) observed the E3 ligase activity in the AN1 domain whereas most of the studies have suggested the involvement of A20 domain in E3 ligase activity. It would be interesting to unveil such mechanisms in detail to understand their role in plant immunity. Similarly, overexpression of SAPs enhances disease resistance in different plants, indicating their role in pathogen defense, and provides another crucial area of research to explore the detailed signaling underlying these disease resistance mechanisms. In summary, plant SAPs need to be studied comprehensively by employing combinatorial omics approaches and system biology to unravel the molecular mechanisms regulating stress-responsive metabolic and physiological processes in plants.
Conclusions
The zinc finger SAP family could either act as positive or negative regulators in providing plant immunity against biotic and abiotic stresses by the expression of several stress-responsive genes. SAPs can also be utilized to generate disease-resistant plants owing to their role in biotic stress response, hence fulfilling another major objective of the agronomic sector. However, the full potential of SAPs can only be utilized once their mode of action is understood efficiently. We proposed a presumed working model to understand the mechanism of SAPs in regulating the stress responses in plants (Fig. 4.). According to the model, biotic and abiotic stresses induce redox instability, thereby triggering ROS release, leading to the activation of receptors (RLCK), which interacts with SAPs via its A20 domain resulting in the formation of oligomers. SAPs also act as an E3 ubiquitin ligase and can participate in K63 and K48 linked polyubiquitination. In the case of K48 linked polyubiquitination, SAPs might interact with shuttling factors like Rad23b (in the nucleus), leading to degradation of proteins via 26S Proteasome. SAPs also play an important role in regulating stress-responsive genes, either by directly interacting with DNA or interacting with transcription factors, leading to the induction of signalling molecules, such as GA20ox, GA3ox, and GA2ox. Pattern-triggered immunity is generated when bacterial flagellin (Flg22) triggers SA production, followed by activation of NPR1. The activated NPR1 is transported to the nucleus and causes activation of salicylic acid-responsive genes. Salicylic acid might regulate SAPs at the transcriptional and post-transcriptional level, activating other SA-responsive genes leading to pathogen defense. Conclusively, the research exploring the precise involvement of SAPs in stress response requires further attention to gain functional insights that would enable genetic manipulation of these proteins for developing climate-resilient crop species.
A presumed model to depict the mechanism of SAPs in stress responses. The signal of any kind of stress (Biotic or abiotic stress) induces redox instability, triggering ROS release (Reactive Oxygen Species). This will trigger the activation of receptors (RLCK), which will interact with SAPs via its A20 domain and resulting in the formation of oligomers. SAPs also act as E3 ubiquitin ligase facilitated by its A20 domain and can participate in K63 linked polyubiquitination and K48 linked polyubiquitination. In case of K48 linked polyubiquitination, SAPs might interact with shuttling factors like Rad23b (in the nucleus), leading to degradation of proteins via 26S Proteasome. SAPs also play an important role in the regulation of stress-responsive genes, either by directly interacting with DNA or through binding with transcription factors which leads to induction of signalling molecules and causes their up-regulation (GA20ox, GA3ox and GA2ox). On the contrary, pattern-triggered immunity is generated when bacterial flagellin (Flg22) triggers the production of salicylic acid. This leads to monomeric forms of NPR1, which transports to the nucleus and causes salicylic acid-responsive genes. Salicylic acid might regulate SAPs at transcriptional regulation or post-transcriptional level where SAPs assist in the activation of other SA responsive genes, causing pathogen defense. Image created with BioRender.com
Abbreviations
- ABA:
-
Abscisic acid
- ACC:
-
1-Aminocyclopropane-carboxylic acid
- ATAF:
-
Arabidopsis Transcription Activation Factor
- bZIP:
-
Basic Leucine Zipper
- CAT:
-
Catalase
- CCH:
-
Copper Transport Proteins
- CK:
-
Cytokinin
- CO1:
-
Constant1
- DI19-4:
-
Drought-Induced gene family
- DREB:
-
Dehydration Responsive Element Binding protein
- DRIP:
-
DREB2A Interacting Protein
- ENO-1:
-
Enolase 1
- ET:
-
Ethylene
- FT:
-
Flowering locus T
- GA:
-
Gibberellic Acid
- GPX-8:
-
Glutathione Peroxidase 8
- GRF-1:
-
Growth Regulating Factor-1
- GSTUs:
-
Glutathione S Transferase
- JA:
-
Jasmonic Acid
- LEA:
-
Late Embryogenesis Abundant Protein
- LOS-2:
-
Low expression of Osmotically Responsive Gene-2
- LSE:
-
Lineage Specific Expansion
- MAPK:
-
Mitogen Activate Protein Kinase
- NADP:
-
Nicotinamide adenine dinucleotide phosphate
- NADP-ME:
-
NADP Malic Enzyme
- NAM:
-
No Apical Meristem
- NCED:
-
9-cis-Epoxycarotenoid dioxygenase gene
- NEMO:
-
NF-kappa B Essential Modulator
- NMR:
-
Nuclear Magnetic Resource Imaging
- NPR1:
-
Nonexpresser of PR1 gene
- OTU:
-
Ovarian Tumour
- PEG:
-
Polyethylene Glycol
- POD:
-
Peroxidase
- PUB1:
-
Plant U-box Protein 1
- RAD23:
-
Radiation sensitive 23
- RAS1:
-
Response to ABA and Salt 1
- RIP1:
-
Receptor Interacting serine/threonine Kinase Protein1
- RLCK253:
-
Receptor Like Cytoplasmic Kinase 253
- ROS:
-
Reactive Oxygen Species
- SA:
-
Salicylic Acid
- SAPs:
-
Stress Associated Proteins
- SOC1:
-
Suppressor of Overexpression of CO1
- SOD:
-
Superoxide Dismutase
- TIP:
-
Tonoplast Intrinsic Protein
- TOR:
-
Target of Rapamycin
- UBA:
-
Ubiquitin Associated Domain
- UIM:
-
Ubiquitin Interacting Motif
- VIGS:
-
Virus Induced Gene Silencing
- WRKYs:
-
Transcription factor associated with drought stress
- ZFPs:
-
Zinc Finger Proteins
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868
Agarwal PK, Agarwal P, Reddy MK, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25:1263–1274
Agarwal PK, Jha B (2010) Transcription factors in plants and ABA dependent and independent abiotic stress signalling. Biol Plant 54:201–212
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Atkinson NJ, Lilley CJ, Urwin PE (2013) Identification of genes involved in the response to simultaneous biotic and abiotic stress. Plant Physiol 162:2028–2041
Baidyussen A, Aldammas M, Kurishbayev A, Myrzabaeva M, Zhubatkanov A, Sereda G, Porkhun R, Sereda S, Jatayev S, Langridge P, Schramm C, Jenkins LD, Soole KL, Shavrukov Y (2020) Identification, gene expression and genetic polymorphism of zinc finger A20/AN1 stress-associated genes, HvSAP, in salt stressed barley from Kazakhstan. BMC Plant Biol 20:156
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 21:1–36
Ben SR, Fabre D, Mieulet D, Meynard D, Dingkuhn M, Al-Doss A, Guiderdoni E, Hassairi A (2012) Expression of the Aeluropus littoralis AlSAP gene in rice confers broad tolerance to abiotic stresses through maintenance of photosynthesis. Plant Cell Environ 35:626–643
Ben SR, Romdhane BW, Mihoubi W, Hsouna BA, Brini F (2020) A Lobularia maritima LmSAP protein modulates gibberellic acid homeostasis via its A20 domain under abiotic stress conditions. PLoS One 15:e0233420
Bubici C, Papa S, Dean K, Franzoso G (2006) Mutual cross-talk between reactive oxygen species and nuclear factor-kappa B: molecular basis and biological significance. Oncogene 25:6731–6748
Busk PK, Page M (1998) Regulation of abscisic acid-induced transcription. Plant Mol Biol 37:425–435
Chang L, Chang HH, Chang JC, Lu HC, Wang TT, Hsu DW, Tzean Y, Cheng AP, Chiu SY, Yeh HH (2018) Plant A20/AN1 protein serves as the important hub to mediate antiviral immunity. PloS Pathogens 14:e1007288
Charrier A, Lelièvre E, Limami AM, Planchet E (2013) Medicago truncatula stress associated protein 1 gene (MtSAP1) overexpression confers tolerance to abiotic stress and impacts proline accumulation in transgenic tobacco. J Plant Physiol 170:874–877
Choi H, Han S, Shin D, Lee S (2012) Polyubiquitin recognition by AtSAP5, an A20-type zinc finger containing protein from Arabidopsis thaliana. Biochem Biophys Res Commun 419:436–440
Christine GG, Marie LG, Elisabeth P, Pascale S, Anis ML, Eric L (2011) A stress-associated protein containing A20/AN1 zing-finger domains expressed in Medicago truncatula seeds. Plant Physiol Biochem 49:303–310
Cushman JC, Bohnert HJ (2000) Genome approaches to plant stress tolerance. Current Opinion in Plant Biology; 3: 117–124. Emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Dixit A, Tomar P, Vaine E, Abdullah H, Hazen S, Dhankher OP (2018) A stress-associated protein AtSAP13, from Arabidopsis thaliana provides tolerance to multiple abiotic stresses. Plant Cell Environ 41:1171–1185
Dixit AR, Dhankher OP (2011) A novel stress-associated protein “AtSAP10” from Arabidopsis thaliana confers tolerance to nickel, manganese, zinc, and high temperature stress. PLoS ONE 6:e20921
Dong Q, Duan D, Zhao S, Xu B, Luo J, Wang Q, Huang D, Liu C, Li C, Gong X, Mao K, Ma F (2018) Genome-wide analysis and cloning of the apple stress-associated protein gene family reveals MdSAP15, which confers tolerance to drought and osmotic stresses in transgenic arabidopsis. Int J Mol Sci 19:2478
Duan W, Sun B, Li TW, Tan BJ, Lee MK, Teo TS (2000) Cloning and characterization of AWP1, a novel protein that associates with serine/threonine kinase PRK1 in vivo. Gene 256:113–121
Feehan JM, Castel B, Bentham AR, Jones JD (2020) Plant NLRs get by with a little help from their friends. Curr Opin Plant Biol 56:99–108
Foyer CH, Noctor G (2016) Stress-triggered redox signaling: what’s in prospect. Plant Cell Environ 39:951–964
Gao W, Lu L, Xinquan T, Jinjing J, Huili L, Hui Z, Fuchun X, Chunpeng S (2016) Genome-wide identification and expression analysis of stress-associated proteins (SAPs) containing A20/AN1 zinc finger in cotton. Mol Genet Genom 291:2199–2213
Giri J, Dansana PK, Kothari KS, Sharma G, Vij S, Tyagi AK (2013) SAPs as novel regulators of abiotic stress response in plants. BioEssays 35:639–648
Giri J, Vij S, Dansana PK, Tyagi AK (2011) Rice A20/AN1 zinc-finger containing stress-associated proteins (SAP1/11) and a receptor-like cytoplasmic kinase (OsRLCK253) interact via A20 zinc-finger and confer abiotic stress tolerance in transgenic Arabidopsis plants. New Phytol 191:721–732
Gilles GC, Gervais ML, Planchet E, Satour P, Limami AM, Lelievre E (2011) A stress-associated protein containing A20/AN1 zing-finger domains expressed in Medicago truncatula seeds. Plant Physiol Biochem 49:303–310
Grzybowska EA (2012) Human intronless genes: functional groups, associated diseases, evolution, and mRNA processing in absence of splicing. Biochem Biophys Res Commun 424:1–6
He X, Xie S, Xie P, Yao M, Liu W, Qin L, Liu Z, Zheng M, Liu H, Guan M (2019) Genome-wide identification of stress-associated proteins (SAP) with A20/AN1 zinc finger domains associated with abiotic stresses responses in Brassica napus. Environ Exp Bot 165:108–119
Hershko A, Ciechanover A (1998) The ubiquitin system. Annu Rev Biochem 67:425–479
Heyninck K, Beyaert R (2005) A20 inhibits NF-kappaB activation by dual ubiquitin-editing functions. Trends Biochem Sci 30:1–4
Hishiya A, Iemura S, Natsume T, Takayama S, Ikeda K, Watanbe K (2006) A novel ubiquitin-binding protein ZNF216 functioning in muscle atrophy. EMBO J 25:554–564
Hozain MD, Abdelmageed H, Lee J, Kang M, Fokar M, Allen RD, Holaday AS (2012) Expression of AtSAP5 in cotton upregulates putative stress-responsive genes and improves the tolerance to rapidly developing water deficit and moderate heat stress. J Plant Physiol 169:1261–1270
Huang J, Wang MM, Jiang Y, Bao YM, Xi H, Sun H, Xu QD, Lan XH, Zhang SH (2008) Expression analysis of rice A20/AN1-type zinc finger genes and characterization of ZFP177 that contributes to temperature stress tolerance. Gene 420:135–144
Hurley JH, Lee S, Prag G (2006) Ubiquitin-binding domains. Biochem J 399:361–372
Irulappan V, Senthil-Kumar M (2018) Morpho-physiological traits and molecular intricacies associated with tolerance to combined drought and pathogen stress in plants. In: Gosal S, Wani S (eds) Biotechnologies of crop improvement, vol 3. Springer, Cham. https://doi.org/10.1007/978-3-319-94746-4_4
Jedmowski C, Ashoub A, Momtaz O, Brüggemann W (2015) Impact of drought, heat, and their combination on chlorophyll fluorescence and yield of wild barley (Hordeum spontaneum). J Bot 2015:120868
Jia H, Li J, Zhang J, Ren Y, Hu J, Lu M (2016) Genome-wide survey and expression analysis of the stress-associated protein gene family in desert poplar, Populus euphratica. Tree Genet Genom. https://doi.org/10.1007/s11295-016-1033-8
Jin Y, Meng W, Junjie F, Ning X, Lian Z, Jia Y, Zheng Z, Guoying W (2007) Phylogenetic and expression analysis of ZnF-AN1 genes in plants. Genomics 90:265–275
Kang M, Abdelmageed H, Lee S, Reichert A, Mysore KS, Allen RD (2013) AtMBP-1, an alternative translation product of LOS2, affects abscisic acid responses and is modulated by the E3 ubiquitin ligase AtSAP5. Plant J 76:481–493
Kang M, Fokar M, Abdelmageed H, Allen RD (2011) Arabidopsis SAP5 functions as a positive regulator of stress responses and exhibits E3 ubiquitin ligase activity. Plant Mol Biol 75:451–466
Kang M, Lee S, Abdelmageed H, Reichert A, Lee HK, Fokar M, Mysore KS, Allen RD (2017) Arabidopsis stress associated protein 9 mediates biotic and abiotic stress responsive ABA signaling via the proteasome pathway. Plant Cell Environ 40:702–716
Kanneganti V, Gupta AK (2008) Overexpression of OsiSAP8, a member of stress associated protein (SAP) gene family of rice confers tolerance to salt, drought and cold stress in transgenic tobacco and rice. Plant Mol Biol 66:445–462
Kothari KS, Dansana PK, Giri J, Tyagi AK (2016) Rice stress associated protein 1 (OsSAP1) interacts with aminotransferase (OsAMTR1) and pathogenesis-related 1a protein (OsSCP) and regulates abiotic stress responses. Front Plant Sci 7:1057
Kranner I, Minibayeva FV, Beckett RP, Seal CE (2010) What is stress? Concepts, definitions and applications in seed science. New Phytol 188:655–673
Lai W, Zhou Y, Pan R, Liao L, He J, Liu H, Yang Y, Liu S (2020) Identification and expression analysis of stress-associated proteins (SAPs) containing A20/AN1 zinc finger in cucumber. Plants 9:400
Lespinet O, Wolf YI, Koonin EV, Aravind L (2002) The role of lineage-specific gene family expansion in the evolution of eukaryotes. Genome Res 12:1048–1059
Li J, Sun P, Xia Y, Zheng G, Sun J, Jia H (2019) A stress-associated protein, PtSAP13, from populus trichocarpa provides tolerance to salt stress. Int J Mol Sci 20:5782
Li W, Yi Wang, Li R, Chang X, Yuan X, Jing R (2021) Cloning and characterization of TaSAP7-A, a member of the stress-associated protein family in common wheat. Front Plant Sci. https://doi.org/10.3389/fpls.2021.609351
Liu S, Wang J, Jiang S, Wang H, Gao Y, Zhang H, Li D, Song F (2019) Tomato SlSAP3, a member of the stress-associated protein family, is a positive regulator of immunity against Pseudomonas syringae pv. tomato DC3000. Mol Plant Pathol 20:815–830
Liu S, Yuan X, Wang Y, Wang H, Wang J, Shen Z, Gao Y, Li D, Song F (2018) Tomato stress-associated protein 4 contributes positively to immunity against necrotrophic fungus botrytis cinerea. Mol Plant Microbe Interact. https://doi.org/10.1094/mpmi-04-18-0097-r
Liu J, Yang X, Yang X, Xu M, Liu J, Xue M, Ma P (2016) Isolation and characterization of LcSAP, a Leymus chinensis gene which enhances the salinity tolerance of Saccharomyces cerevisiae. Mol Biol Rep 44(1):5–9. https://doi.org/10.1007/s11033-016-4091-y
Liu Y, Xu Y, Xiao J, Ma Q, Li D, Xue Z, Chong K (2011) OsDOG, a gibberellin-induced A20/AN1 zinc-finger protein, negatively regulates gibberellin-mediated cell elongation in rice. J Plant Physiol 168:1098–1105
Lloret A, Conejero A, Leida C, Petri C, Gil-Muñoz F, Burgos L, Badenes LM, Rios G (2017) Dual regulation of water retention and cell growth by a stress-associated protein (SAP) gene in Prunus. Sci Rep 7:332
Lu TT, Onizawa M, Hammer GE, Turer EE, Yin Q, Damko E, Agelidis A, Shifrin N, Advincula R, Barrera J, Malynn BA, Wu H, Ma A (2013) Dimerization and ubiquitin mediated recruitment of A20, a complex deubiquitinating enzyme. Immunity 38:896–905
Ma A, Malynn BA (2012) A20: linking a complex regulator of ubiquitylation to immunity and human disease. Nat Rev Immunol 12:774–785
Mauro C, Pacifico F, Livorgna A, Mellone S, Lannettu A, Acquaviva R, Formisano S, Vito P, Leonardi A (2006) ABIN-1 binds to NEMO/IKK gamma and co-operates with A20 in inhibiting NF-kappaB. J Biol Chem 281:18482–18488
McDermott MF, Aksentijevich I (2002) The autoinflammatory syndromes. Curr Opin Allergy Clin Immunol 2:511–516
Mittler R (2006) Abiotic stress, the field environment and stress combination. Trends Plant Sci 11:15–19
Mittler R (2017) ROS are good. Trends Plant Sci 22:11–19
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Moore M, Gossmann N, Dietz K-J (2016) Redox regulation of cytosolic translation in plants. Trends Plant Sci 21:388–397
Mukhopadhyay A, Vij S, Tyagi AK (2004) Overexpression of a zinc-finger protein gene from rice confers tolerance to cold, dehydration, and salt stress in transgenic tobacco. Proc Natl Acad Sci USA 101:6309–6314
Muthuramalingam P, Jeyasri R, Selvaraj A, Kalaiyarasi D, Aruni W, Pandian STK, Manikandan R (2020) Global transcriptome analysis of novel stress associated protein (SAP) genes expression dynamism of combined abiotic stresses in Oryza sativa (L.): an in silico approach. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1747548
Nakamura N (2018) Ubiquitin system. Int J Mol Sci 19:1080
Ngo ST, Steyn FJ, McCombe PA (2014) Gender differences in autoimmune disease. Front Neuroendocrinol 35:347–369
Opipari AW, Boguski MS, Dixit VM (1990) The A20 cDNA induced by tumor necrosis factor alpha encodes a novel type of zinc finger protein. J Biol Chem 265:14705–14708
Pandey P, Ramegowda V, Senthil-Kumar M (2015a) Shared and unique responses of plants to multiple individual stresses and stress combinations: physiological and molecular mechanisms. Front Plant Sci 6:723
Pandey P, Sinha R, Mysore KS, Senthil-Kumar M (2015) Impact of concurrent drought stress and pathogen infection on plants. In: Mahalingam R (ed) Combined stresses in plants. Springer, Cham. https://doi.org/10.1007/978-3-319-07899-1_10
Peckham D, Scambler T, Savic S, McDermott MF (2017) The burgeoning field of innate immune-mediated disease and autoinflammation. J Pathol 241:123–139
Petrov VD, Van Breusegem F (2012) Hydrogen peroxide—a central hub for information flow in plant cells. AoB Plants 2012:pls014
Penengo L, Mapelli M, Murachelli AG, Confalonieri S, Magri L, Musacchio A, Fiore PPD, Schneider TR (2006) Crystal structure of the ubiquitin binding domains of Rabex-5 reveals two modes of interaction with ubiquitin. Cell 124(6):1183–1195. https://doi.org/10.1016/j.cell.2006.02.020
Prasad PVV, Pisipati SR, Momcilovic I, Ristic Z (2011) Independent and combined effects of high temperature and drought stress during grain filling on plant yield and chloroplast EF-Tu expression in spring wheat. J Agron Crop Sci 197:430–441
Prasch CM, Sonnewald U (2013) Simultaneous application of heat, drought, and virus to Arabidopsis plants reveals significant shifts in signaling networks. Plant Physiol 162:1849–1866
Ramu VS, Paramanantham A, Ramegowda V, Mohan-Raju B, Udayakumar M, Senthil-Kumar M (2016) Transcriptome analysis of sunflower genotypes with contrasting oxidative stress tolerance reveals individual- and combined- biotic and abiotic stress tolerance mechanisms. PLoS ONE 11:e0157522
Rivero RM, Mestre TC, Mittler R, Rubio F, Garcia-Sanchez F, Martinez V (2014) The combined effect of salinity and heat reveals a specific physiological, biochemical and molecular response in tomato plants. Plant Cell Environ 37:1059–1073
Roos G, Messens J (2011) Protein sulfenic acid formation: from cellular damage to redox regulation. Free Radic Biol Med 51:314–326
Sharma G, Giri J, Tyagi A (2015) Rice OsiSAP7 negatively regulates ABA stress signaling and imparts sensitivity to water deficit stress in Arabidopsis. Plant Sci 237:80–92
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene network involved in drought stress tolerance and response. J Exp Bot 58:221–227
Sinha R, Gupta A, Senthil-Kumar M (2016) Understanding the impact of drought on foliar and xylem invading bacterial pathogen stress in chickpea. Front Plant Sci 7:902
Solanke AU, Sharma MK, Tyagi AK, Sharma AK (2009) Characterization and phylogenetic analysis of environmental stress-responsive SAP gene family encoding A20/AN1 zinc finger proteins in tomato. Mol Genet Genom 282:153–164
Sreedharan S, Shekhawat UK, Ganapathi TR (2012) MusaSAP1, a A20/AN1 zinc finger gene from banana functions as a positive regulator indifferent stress response. Plant Mol Biol 80:503–517
Ströher E, Wang XJ, Roloff N, Klein P, Husemann A, Dietz KJ (2009) Redox-dependent regulation of the stress-induced zinc-finger protein SAP12 in Arabidopsis thaliana. Mol Plant Pathol 2:357–367
Takatsuji H (1999) Zinc-finger proteins: the classical zinc finger emerges in contemporary plant science. Plant Mol Biol 39:1073–1078
Thaura GH, Michael GS, Donaldo M, Denis F, Alexandra P, Walid BR, Rania BS, Satoshi O, Maria CR, Manabu I, Joe T, Abdullah AD, Emmanuel G, Afif H (2017) Expression of the Aeluropus littoralis AlSAP gene enhances rice yield under field drought at the reproductive stage. Front Plant Sci 8:994
Tokunaga F, Nishimasu H, Ishitani R, Goto E, Noguchi T, Mio K, Kamei K, Ma A, Iwai K, Nureki O (2012) Specific recognition of linear polyubiquitin by A20 zinc finger 7 is involved in NF-kappaB regulation. EMBO J 31:3856–3870
Tyagi H, Jha S, Sharma M, Giri J, Tyagi AK (2014) Rice SAPs are responsive to multiple biotic stresses and overexpression of OsSAP1, an A20/AN1 zinc-finger protein, enhances the basal resistance against pathogen infection in tobacco. Plant Sci 225:68–76
Verhelst K, Carpentier I, Kreike M, Meloni L, Verstrepen L, Kensche T, Dikic I, Beyaert R (2012) A20 inhibits LUBAC-mediated NF-kappaB activation by binding linear polyubiquitin chains via its zinc finger 7. EMBO J 31:3845–3855
Vij S, Tyagi AK (2006) Genome-wide analysis of the stress associated protein (SAP) gene family containing A20/AN1 zinc-finger(s) in rice and their phylogenetic relationship with Arabidopsis. Molecular Genet Genom 276:565–575
Vij S, Tyagi AK (2008) A20/AN1 zinc-finger domain-containing proteins in plants and animals represent common elements in stress response. Funct Integr Genom 8:301–307
Wang Y, Zhang L, Zhang L, Xing T, Peng J, Sun S, Chen G, Wang X (2013) A novel stress-associated protein SbSAP14 from Sorghum bicolor confers tolerance to salt stress in transgenic rice. Mol Breed 32:437–449
Wang Z, Kuang J, Han B, Chen S, Liu A (2020) Genomic characterization and expression profiles of stress-associated proteins (SAPs) in castor bean (Ricinus communis). Plant Divers. https://doi.org/10.1016/j.pld.2020.07.010
Wertz IE, O’Rourke KM, Zhou H, Eby M, Aravind L, Seshagiri S, Wu P, Wiesmann C, Baker R, Boone DL, Ma A, Eugene VK, Dixit MV (2004) De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-kappaB signalling. Nature 430:694–699
Willems P, Mhamdi A, Stael S, Storme V, Kerchev P, Noctor G, Gevaert K, Van Breusegem F (2016) The ROS wheel: refining ROS transcriptional footprints. Plant Physiol 171:1720–1733
Xiang-Zhan Z, Wei-Jun Z, Xin-You C, Xi-Yan C, Shu-Ping Z, Tai-Fei Y, Jun C, Yong-Bin Z, Ming C, Shou-Cheng C, Zhao-Shi X, You-Zhi M (2019) Genomic analysis of stress associated proteins in soybean and the role of GmSAP16 in abiotic stress responses in arabidopsis and soybean. Front Plant Sci 10:1453
Xu Q, Mao X, Wang Y, Wang J, Xi Y, Jing R (2018) A wheat gene TaSAP17-D encoding an AN1/AN1 zinc finger protein improves salt stress tolerance in transgenic Arabidopsis. J Integr Agric 17:507–516
Xuan N, Jin Y, Zhang H, Xie Y, Liu Y, Wang G (2011) A putative maize zinc-finger protein gene, ZmAN13, participates in abiotic stress response. Plant Cell Tissue Organ Cult 107:101–112
Yoon SK, Bae EK, Lee H, Choi YI, Han M, Choi H, Kang KS, Park EJ (2018) Downregulation of stress-associated protein 1 (PagSAP1) increases salt stress tolerance in poplar (Populus alba × P. glandulosa). Trees 32:823–833
Yvona M, Vileb D, Braultc V, Blanca S, van Munstera M (2017) Drought reduces transmission of Turnip yellows virus, an insect-vectored circulative virus. Virus Res 241:131–136
Zagorchev L, Seal CE, Kranner I, Odjakova M (2013) A central role for thiols in plant tolerance to abiotic stress. Int J Mol Sci 14:7405–7432
Zhang N, Yin Y, Liu X, Tong S, Xing J, Zhang Y, Pudake NR, Izquierdo ME, Peng M, Xin M, Hu Z, Ni Z, Sun Q, Yao Y (2017) The E3 ligase TaSAP5 alters drought stress responses by promoting the degradation of DRIP proteins. Plant Physiol 175:1878–1892
Zhang XZ, Zheng WJ, Cao XY, Cui XY, Zhao SP, Yu TF, Chen J, Zhou BY, Chen M, Chai CS, Xu SZ, Ma YZ (2019) Genomic analysis of stress associated proteins in soybean and the role of GmSAP16 in abiotic stress responses in arabidopsis and soybean. Front Plant Sci. https://doi.org/10.3389/fpls.2019.01453
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgements
Authors' research in this area was supported by the DST INSPIRE Faculty Grant of Department of Science & Technology (DST), Ministry of Science & Technology, Government of India (File No. DST/INSPIRE/04/2016/002341).
Author information
Authors and Affiliations
Contributions
MM conceived and outlined the review; VS prepared the first draft, tables and figures; PC and SR critically revised the work and provided additional inputs. All the authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shukla, V., Choudhary, P., Rana, S. et al. Structural evolution and function of stress associated proteins in regulating biotic and abiotic stress responses in plants. J. Plant Biochem. Biotechnol. 30, 779–792 (2021). https://doi.org/10.1007/s13562-021-00704-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-021-00704-x