Abstract
Cold shock proteins (CSPs) are greatly conserved family of structurally related DNA binding proteins which are produced during temperature drift. A 213 bp long cspA gene was cloned and sequenced from Pseudomonas koreensis P2 in the present study. The expression analysis of the cspA showed > 2.5 folds increase in the mRNA level at 15 °C while the expression was almost on par at 30 °C and 5 °C indicating its role in moderately low temperature. In silico analyses of the gene showed that the gene codes for 7.69 kDa protein which was phylogenetically very similar to CspA present in Pseudomonads. Amino acid composition of the CspA from P. koreensis was different from that of mesophilic Pseudomonas and tiny/small amino varied significantly between CspA of cold adaptive and mesophilic species. The CspA from P. koreensis P2 contained RNP motifs involved in binding of DNA and RNA. Phylogenetic analyses revealed that the CspA protein of P. koreensis P2 was more close to CspA of distant subgroups of Pseudomonas like P. fluorescens and P. putida subgroup indicating a possible intra-specific gene transfer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Three-fourth of our biosphere’s temperature is below 5 °C which poses as major deterrent in sustenance of life in such harsh conditions (Casanueva et al. 2010; Rodrigues and Tiedje 2008). But many of the life forms have adapted well in such environments during the course of evolution. Microorganisms, the most ancient colonizers of this planet, are well known inhabitants of the extreme ecological niches due to their enormous metabolic diversity. Most of the known psychrophilic microorganisms fit into various species of archaea, bacteria, yeast, fungi and algae (Thieringer et al. 1998; D’Amico et al. 2006; Piette et al. 2011). To survive at lower temperatures, the expression of a considerable number of genes coding for a number of proteins involved in cold acclimatization and survival are up or down regulated by psychrophiles. Cold shock proteins (CSPs) constitute a highly conserved family of structurally related DNA-binding proteins which are released on temperature drift (Moon et al. 2009). CSPs are distributed in prokaryotic and eukaryotic kingdom to cope up with cold shock due and abrupt downshift in temperature. In prokaryotes, CSPs are found in a diverse group of bacteria like Bacillus sp., Streptococcus sp., Thermotoga sp., Listeria sp., Arthrobacter sp., Escherichia coli and Pseudomonas sp. (Goldstein et al. 1990; Graumann et al. 1997; Fang et al. 2012; Lee et al. 2013; Hoffmann et al. 2013; Lee et al. 2014; Bisht et al. 2014). Among prokaryotes, CSPs were first identified in E. coli and further discoveries showed that it codes for eight more similar proteins (CspB–CspI). In E. coli, CspA is the major CSP which accumulate up to 10% of total proteins upon exposure to lower temperature (Goldstein et al. 1990).Upon temperature downshift, cell growth is arrested transiently when synthesis of most proteins shuts excluding cold inducible proteins (Polissi et al. 2003). After this transient acclimatization the cells get adapted to lower temperature and recommence the growth but with a lower rate. The bulk protein synthesis restarts in the cell with decline in the expression of the cold-inducible proteins (Phadtare 2004).
In this study, we cloned and characterized cspA gene from Pseudomonas koreensis P2-a cold adaptive bacteria isolated from Himalayan region of Arunachal Pradesh, India. The real time expression of cspA gene under different temperature was also estimated using quantitative-real time PCR. Consequently, we implemented a composite approach of structure prediction for modeling of CspA protein with further studies to find its role during survival in adverse environmental conditions and its adaptation towards low temperature.
Materials and methods
Bacterial culture
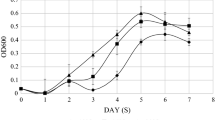
The bacterial strain, Pseudomonas koreensis P2 (NAIMCC-B-01747) was previously isolated by our group from soils collected from Sela Lake, Arunachal Pradesh, India. It was found to grow at temperatures ranging from 4 to 35 °C with optimum growth at 15 °C. Pure cultures were maintained in glycerol at − 20 °C and used for further studies.
Amplification of cspA gene
cspA genes of eight different Pseudomonas spp. available in the NCBI databases were downloaded and aligned (Supplementary figure 1) using ClustalW (Thompson et al. 2002). Primers cspA F (ATGTCTAATCGCCAAACC) and cspA R (TTACTCTGGGCGAACTTG) were then designed from the alignment for amplification of full length gene of CspA. Genomic DNA was extracted from P. koreensis P2 using standard protocol (Ausubel et al. 2003). PCR amplification of cspA was performed as described by Rai et al. (2015). Primer pair, cspA F and cspA R was used for amplification of cspA gene from chromosomal DNA of P. koreensis P2. The resultant amplicon was purified using PCR purification kit procured from Nucelo-pore (ThermoFisher Scientific, USA).
Cloning and sequencing
The purified amplicon thus obtained was successfully cloned in pJET1.2 vector, following manufacturer’s protocol with minor modifications and transformed into chemi-competent E. coli DH5α cells. The cloning procedure employed a positive selection in which clones grown on Luria–Bertani supplemented with Ampicillin (LB Amp+) plates were selected and the cloned gene was amplified using gene specific primers to ensure positive clones. Clones were then sequenced by an automatic ABI-3130 XL Sequencer (Applied Biosystems) using pJET1.2 forward and reverse primer. The curated sequence of cspA, after the removal of vector sequence by VecScreen (NCBI), was translated into its amino acids sequence using EXPASY server (Artimo 2012).
Analysis of expression of cspA gene at different temperatures
In order to measure the expression levels of cspA gene under different temperature regimes, the exponentially grown P. koreensis P2 was inoculated (@5%w/v) in nutrient broth in a shaking incubator at 5 °C, 15 °C and 30 °C for 12 h at 150 rpm. The low temperature conditions viz., 5 °C and 15 °C served as treatments which were named as T1 and T2 respectively; whereas culture incubated at 30 °C served as control. All the experimental runs were performed in triplicates. After 12 h of incubation, cultures under each temperature treatment were pelleted down for total RNA isolation. The isolation of total RNA was performed using GeneJETTM RNA purification kit (Fermentas). The quality of the total RNA was determined by gel electrophoresis using 1.2% formaldehyde agarose gel and visualizing the gels in BioradTM Chemidoc XRS gel documentation system and its quantity was determined using a Nanodrop. Prior to qPCR, the cDNA synthesis from the total RNA isolated from all the temperature treatments was performed using iScript™ cDNA synthesis kit (Biorad). The cDNA synthesized from all the treatments (T1 and T2) and control served as template for setting up reaction mix for qRT-PCR. qRT PCR was performed in triplicate in a reaction volume of 20 µl consisting of 10 µl 2 × SYBER green Master mix, 0.5 µl QN ROX Reference Dye, 1 µl each of primer F and primer R respectively (10 pmol), 1 µl cDNA and 6.5 µl RNase free water. The cspA specific primers described by Ivancic et al. (2013) were used for quantification of transcripts. The 16S rRNA gene served as an endogenous control. Following thermal cycling conditions were set:initial denaturation at 95 °C for 2 min, followed by 40 cycles of amplification by 3 step cycling, Denaturation at 95 °C for 5 s, annealing at 55 °C for 30 s, extension at 65 °C for 1 min. The gene quantification was performed as per the MIQE guidelines (Bustin et al. 2009).
Prediction of primary and secondary structure
The physico-chemical characteristics were studied by computing Extinction Coefficient, theoretical isoelectric point (pI), molecular weight, total number of positive and negative residues (Gill et al. 1989), Aliphatic Index (Atsushi 1980), Grand Average Hydropathy (GRAVY) (Kyte and Russell 1982) and Instability Index (Guruprasad et al. 1990) using Expasy’s ProtParam server (Gasteiger 2005). Amino acid compositions of CspA from cold adaptive P. syringae NCPPB 3739 and mesophilic P. aeruginosa DSM 22644 and P. stutzeri DSM 50227 were compared. For this amino acid sequences were retrieved from UniProt database. Principal Component Analysis (PCA) was carried out to identify important amino acids. Based on PCA results, a biplot was drawn to group different strains and to also identify the important amino acids for each group. Further, hierarchical clustering techniques were also performed and dendrogram was drawn to double check the grouping of strains. All the statistical analyses were carried out using R version 3.4.4 (2018-03-15). Secondary structure of CspA was also predicted using PDBSum Server (Laskowski 2001). PDBSum server provides 3D protein structure information regarding the motifs, domains, helices, beta sheets and strands, angles, etc. Subcellular localization of protein was determined using CELLO V.2.5 (Yu et al. 2006) and PSORTb (Nancy et al. 2010).
Phylogenetic analysis of CspA protein
A total of 35 amino acid sequence of CspA proteins of different bacteria were retrieved from NCBI. Phylogenetic analysis of all the retrieved protein sequences along with CspA protein of Pseudomonas koreensis P2 was performed based on the results of multiple alignments using ClustalW (Thompson et al. 2002). A phylogenetic tree was constructed by maximum likelihood method using MEGA 6.06 (Tamura 2013) with the bootstrap test replicated 1000 times. The evolutionary distances were computed using JTT matrix (Jones et al. 1992). All the positions containing missing data and gaps were removed.
Sequence analysis and molecular modeling
Similarity search of CspA sequence were performed by BLASTp (Altschul et al. 1990) at NCBI (http://www.ncbi.nlm.nih.gov) with PDB (Berman et al. 2000) as a reference database to identify the suitable templates for modeling of CspA. The search revealed that no single template with lower e-value and acceptable identity was able to satisfy 100% query coverage. Hence, the combination of multiple templates was opted to enhance the query coverage. We used I-TASSER server (http://zhang.bioin-formatics.ku.edu/I-TASSER) to develop high quality 3D model (Roy et al. 2010). Further, SWISSMODEL (Schwede 2003) and Phyre 2 (Kelley et al. 2015) were also used to check the structure reliability.
Assessment of predicted model
The stereo-chemical quality of predicted model was checked by analyzing the overall structure and residue-by-residue geometry of proteins. The quality assessment of the refined energy minimized CspA model was performed by PROCHECK (Laskowski et al. 2001) and QMEAN Z-score estimation using QMEAN server (Benkert et al. 2009).VERIFY 3-D (Eisenberg et al. 1997) and ERRAT server (Colovos and Yeates 1997) were used for structural validation.
Structural analysis of CspA model
Binding pockets of CspA proteins were predicted using CastP server (Binkowski et al. 2003).Detection of RNA and DNA binding specificities of CspA protein was performed using BindUP server (Paz et al. 2016). BindUP predicts nucleic acid binding proteins (NABPs) on the basis of electrostatic patches on protein surfaces using NAbind algorithm (Stawiski et al. 2003; Shazman and Mandel-Gutfreund 2008).
Results and discussion
Cloning and sequencing of cspA gene
PCR amplification resulted in approximately 200 bp amplicon (Fig. 1a, b). Cloning, sequencing and vector screening revealed that cspA of P. koreensis P2 is 213 bp long. When the sequence was searched against NCBI database, its identity as cspA gene was confirmed. The complete sequence of cspA gene has been submitted in DNA Data Bank of Japan (DDBJ) with accession no. LC214053.
Real time quantification of cspA gene expression at different temperature variables
The role of cspA gene for cold adaptation was better elucidated by the qRT-PCR based gene quantification studies. The size of the cDNA of cspA gene was found to be about 150 bp, this cDNA was further utilized for its copy number estimation using qRT-PCR. As compared to the control (30 °C), the relative quantity of cspA did not change significantly at 5 °C while at 15 °C, it increased to 2.57 ± 0.23 (Fig. 2). The results indicated two probable events, first, probably the cspA gene has minimal contribution in the cold adaptation at very low temperatures like 5 °C and have more profound role at moderately low temperatures; second, at very low temperatures, more cold inducible genes are responsible for the of P. koreensis P2. It is pertinent to mention that the optimum growth temperature for the bacterial strain is also 15 °C which indicates that probably at 5 °C, the machineries required for the transcription of cspA are inadequately formed leading to lower transcription. Similar induction of cspA mRNA synthesis has already been demonstrated in various cold adaptive isolates of P. fluorescens, E. coli, Arthrobacter protophormiae, Caulobacter spp. etc. (Ray 1994; Panicker et al. 2002; Mazzon et al. 2012; Bisht et al. 2014). It has been reported that, cspA mRNA levels are inversely related with temperature (Ivancic et al. 2013; Song et al. 2012). Goldenberg et al. (1996) suggested that the cspA mRNA (t1/2 = 10 s) is rapidly degraded at 37 °C while at low temperatures it becomes much more stable (t1/2 = 20 min).
Sequence analysis and secondary structure
Molecular weight and pI of the CspA protein was calculated 7.69 and 6.55 kDa respectively. The molecular formula of CspA protein was found as C342H524N94O105S2. CspA from P. koreensis P2 comprised of 70 amino acids residues. Bi-Plot analyses and UPGMA clustering based on Eucledian distance revealed that in terms of amino acid composition both the cold adaptive CspA (P. koreensis and P. syringae) clustered together while P. aeruginosa and P. stutzeri were placed in different cluster (Fig. 3). This indicates that the amino acid composition of CspA in cold adaptive and mesophilic species of Pseudomonas is different. It was observed that Lys (L), Pro (P) and Thr (T) were the predominant amino acids explaining the variations while Ser (S), Ile (I), Arg (R), His (H) and Tyr (Y) were important in CspA of P. aeruginosa and P. stutzeri.
Metpally and Reddy (2009) reported that neutral and small amino acid groups were greatly favored in psychrophiles. It was observed that tiny/small amino acids viz. Ala (A), Gly (G), Ser (S), Asn (N), Asp (D), Pro (P) and Thr (T) constituted 42.85%, 41.50% in P. koreensis and P. syringae while the same was 40.3% and 39% in P. aeruginosa and P. stutzeri. Although the difference in amount of tiny/small amino acids present in cold adaptive and mesophilic species was not much, it was interesting to note that the out of the three amino acids explaining the variation in CspA of P. koreensis and P. syringae two (P and T) were tiny/small amino acids. While out of the five amino acids explaining the variation in CspA of mesophilic Pseudomonas, only one (S) was tiny/small. CspA of P. koreensis P2 was found to be localized in cytoplasm. Secondary structure predicted by PDBSUM server showed 1 sheet, 3 β-hairpins,1 Ψ loop, 4 β-beta bulges,5 strands, 2 helices and 3 β-turns (Supplementary figure 2). Multiple sequence alignment of CspA proteins (Supplementary figure 3) showed presence of two motifs viz. RNP1and RNP2 while 16 of 70 residues were found to be 100% conserved. Similar structure and composition of RNA binding domains (RNP1 and RNP2) were reported in E. coli and Bacillus subtilis (Schindelin et al. 1993, 1994). Schröder et al. (1995) showed through mutational analysis that RNP1 and RNP2 were essential for ssDNA-binding activity. RNA molecules typically form stable secondary structures under low temperature and may cause premature transcription termination. Hence, the RNA chaperones are essential to transcriptional process under low temperature. The presence of these motifs in CspA of P. koreensis P2 indicated the conservation of the nucleic acid binding capacity and, possibly, of the functional role as RNA chaperone observed in other bacteria.
Phylogeny of CspA protein
CspA is reported in a large number of bacteria (Phadtare et al. 1999; Kortmann and Narberhaus 2012). Phylogenetic analysis of CspA protein showed close evolutionary relationship with CspA of P. fluorescens WH6 (100%), P. agarici (98.4%) and P. rhizosphaerae (98.4%) forming a monophyletic clade (Fig. 4). P. fluorescens belonged to P. fluorescens subgroup which was phylogenetically distant from P. koreensis subgroup (Gomila et al. 2015). Other close neighbours like P. rhizospharae, P. japonica, P. cremoricolorata belonged to the P. putida subgroup which was also distant from P. koreensis group while P. agarici belonged to unclassified group of Pseudomonas (Anzai et al. 2000). Phylogenetic analysis revelaed that the cold shock proteins (CspA) were highly diverse in Pseudomonads as the similarity ranged from 35 to 100%. The evolutionary relationship among different CspA proteins indicated probable gene transfer among different subgroups of Pseudomonas. The AT content of the cspA gene was 51.24% which was higher than the genome average (39.55%, Unpublished). Jensen et al. (2004) reported that high AT genomic regions in Pseudomonas were more prone to gene transfer. Moreover is it is well accepted that the laterally transferred genes are generally AT rich (Moszer et al. 1999; Lawrence and Ochman 1997; Médigue et al. 1991). It was quite possible that interspecies transfer of genes among the pseudomonads might have resulted in the cspA of P. koreensis which was also supported by the phylogenetic analyses as the most closest cspA (of P. fluorescens WH6) was from different sub-group of Pseudomonas. Such gene transfer might have enabled cold adaptive P. koreensis P2 to survive under cold climatic conditions of eastern Himalayas. Earlier, Bisht et al. (2014) reported possible gene transfer between Pseudomonads and Bacillus cereus under cold Himalayan environment.
Prediction and evaluation of 3D structure of cold shock protein
As mentioned earlier that the 3D structure of CspA from Pseudomonas koreensis P2 was not available in PDB and in any other structural database, the structure was modeled by comparative modeling and a composite approach using threading and ab initio modeling for 3D structure prediction.
For comparative modeling Modweb, Swiss Model and Phyre2 were used. Best models obtained from these methods were evaluated. Five models were predicted and retrieved for CspA with threading and ab initio modeling at I-TASSER. Model1 predicted by I-TASSER was found the best among the predicted model. The best model of CspA had an acceptable C-score (correlation scoring) of 1.11 as it lied within the range of reliable models C-score, i.e., − 0.5 to + 2.0. C-score is a “confidence score” for estimating the quality of a computed model. The C-score in I-TASSER can range from − 5 to 2. 3D structure of CspA of P. koreensis P2 with the highest C score is depicted in Fig. 5. The refined 3D model was deposited in Protein Model database (PMDB) with PMDB id: PM0081019.
The results of stereo-chemical estimation of backbone Ψ and Φ dihedral angles of the CspA revealed that 87.5, 10.7, and 1.8% of residues were falling within the most favored regions, additionally allowed regions and disallowed regions respectively (Supplementary figure 4). Q-mean-Z-score of the model was found to be − 0.90 indicating reliable structure and fell within the range of scores characteristically found for similar size proteins (Supplementary figure 5).Verify-3D checks the compatibility of the model with its own amino acid sequence. The Verify 3-D predicted the model which lied within the range 0.19–0.71 (Supplementary figure 6). About 97% of the residues had an average 3D–1D score ≥ 0.2, confirming good quality model. The ERRAT results indicated that the CspA model’s overall quality factor was 95.161%, supporting the robust nature of the developed model (Supplementary figure 7). The qualitative evaluation of the model revealed that the generated model was reliable and of good quality.
Structural analysis of CspA model
Protein 3D structure and its surface topography are able to provide vital information for understanding the protein function. Structural details of surface regions of protein enables detailed studies of the relationship of protein structure and function. In CspA from P. koreensis P2, seven binding pockets were identified with volume ranging from 14.7 to 41.9 (Supplementary figure 8). During analysis of NABPs in CspA model, three largest positive patches were detected viz Patch 1MET1 SER2 LYS4 MET5 GLN38 SER52 PHE53 THR54 GLU56 GLY65 ASN66 THR68 LEU70 Patch 2:PHE31 HIS33 PHE34 GLU56 GLY58 LYS60 PRO62 ALA63 ALA64 Patch 3: HIS33 SER35 GLY41 LYS43 (Supplementary figure 9-10), which predicted CspA as nucleic acid (NA) binding protein. The results suggested the role of CspA protein of P. koreensis P2 as chaperone.
In the present study, 213 bp long cspA gene was identified in cold adaptive Pseudomonas koreensis which was isolated from the Eastern Himalayas. Expression analyses of the gene indicated its role in survival during moderately low temperature. In silico analysis revealed that tiny/small and neutral amino acids characteristic to psychrophilic proteomes were predominant in CspA protein. This protein contained two highly conserved motifs viz. RNP1 and RNP2 involved in nucleic acid binding. Phylogenetic analyses revealed that the CspA protein of P. koreensis P2 was more close to CspA of distant subgroups of Pseudomonas like P. fluorescens and P. putida subgroup indicating a possible intra-specific gene transfer. The cold shock proteins are of significant biotechnological importance as they can be useful for engineering crop plants for tolerating abiotic stresses, particularly cold and drought.
Abbreviations
- CSPs:
-
Cold shock proteins
- GRAVY:
-
Grand average hydropathy
- MEGA:
-
Molecular evolutionary genetics analysis
- PDB:
-
Protein Data Bank
- NCBI:
-
National Center for Biotechnology Information
- NABPs:
-
Nucleic acid binding proteins
- DDBJ:
-
DNA Data Bank of Japan
- PMDB:
-
Protein model database
- C-score:
-
Correlation scoring
- MIQE:
-
Minimum information for publication of quantitative RT PCR experiments
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Anzai Y, Kim H, Park JY, Wakabayashi H, Oyaizu H (2000) Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence. Int J Syst Evol Microbiol 50(4):1563–1589
Artimo P (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40(W1):W597–W603
Atsushi IKAI (1980) Thermostability and aliphatic index of globular proteins. J Biochem 88(6):1895–1898
Ausubel FM, Brent R, Moore RE, Seidman JG, Smith JA (2003) Short protocols in molecular biology: a compendium of methods from current protocols in molecular biology. Wiley, New York
Benkert P, Michael K, Torsten S (2009) QMEAN server for protein model quality estimation. Nucleic Acids Res 37:W510–W514
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat VH, Weissig SI, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Binkowski TA, Naghibzadeh S, Liang J (2003) CASTp: computed atlas of surface topography of proteins. Nucleic Acids Res 31(13):3352–3355
Bisht SC, Joshi GK, Mishra PK (2014) cspA encodes a major cold shock protein in Himalayan psychrotolerant Pseudomonas strains. Interdiscip Sci Comput Life Sci 6(2):140–148
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55(4):611–22
Casanueva A, Tuffin M, Cary C, Cowan DA (2010) Molecular adaptations to psychrophily: the impact of ‘omic’ technologies. Trends Microbiol 18:374–381
Colovos C, Yeates TO (1997) Verification of protein structures:patterns of non bonded atomic interactions. Protein Sci 2:1511–1519
D’Amico S, Collins T, Marx JC, Feller G, Gerday C (2006) Psychrophilic microorganisms: challenges for life. EMBO Rep 7:385–389
Eisenberg D, Lüthy R, Bowie JU (1997) VERIFY3D: assessment of protein models with three-dimensional profiles. Methods Enzymol 277:396–404
Fang SH, Chiang SH, Hsu SY, Chou CC (2012) Cold shock treatments affect the viability of Streptococcus thermophilus BCRC 14085 in various adverse conditions. J Food Drug Anal 20(1):117–124
Gasteiger E (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press, Springer, New York, pp 571–607
Gill SC, Peter H, Hippel V (1989) Calculation of protein extinction coefficients from amino acid sequence data. Anal Biochem 182(2):319–326
Goldenberg D, Azar I, Oppenheim AB (1996) Differential mRNA stability of the cspA gene in the cold-shock response of Escherichia coli. Mol Microbiol 19:241–248
Goldstein J, Pollitt NS, Inouye M (1990) Major cold shock protein of Escherichia coli. Proc Natl Acad Sci USA 87:283–287
Gomila M, Peña A, Mulet M, Lalucat J, García-Valdés E (2015) Phylogenomics and systematics in Pseudomonas. Front Microbiol 6:214
Graumann P, Wendrich TM, Weber MH, Schroder K, Marahiel MA (1997) A family of cold shock proteins in Bacillus subtilis is essential for cellular growth and for efficient protein synthesis at optimal and low temperatures. Mol Microbiol 25:741–756
Guruprasad K, Reddy Bhasker BV, Pandit Madhusudan W (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng 4(2):155–161
Hoffmann T, Tych KM, Brockwell DJ, Dougan L (2013) Single-molecule force spectroscopy identifies a small cold shock protein as being mechanically robust. J Phys Chem B 117:1819–1826
Ivancic T, Jamnik P, Stopar D (2013) Cold shock CspA and CspB protein production during periodic temperature cycling in Escherichia coli. BMC Res Notes 6:248
Jensen LJ, Skovgaard M, Sicheritz-Pontén T, Hansen NT, Johansson H, Jørgensen MK, Ussery D (2004) Comparative genomics of four Pseudomonas species. In: Ramos J-L (ed) Pseudomonas, Volume 1; Genomics, life style and molecular architecture. Springer, New York, pp 139–164
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10(6):845–858
Kortmann J, Narberhaus F (2012) Bacterial RNA thermometers: molecular zippers and switches. Nat Rev Microbiol 10:255–265
Kyte J, Russell FD (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1):105–132
Laskowski RA (2001) PDBsum: summaries and analyses of PDB structures. Nucleic Acids Res 29(1):221–222
Laskowski RA, MacArthur MW, Thornton JM (2001) PROCHECK: validation of protein structure coordinates. International tables of crystallography, Vol. F. Crystallography of biological macromolecules. Kluwer Academic Publishers, Dordrecht, pp 722–725
Lawrence JG, Ochman H (1997) Amelioration of bacterial genomes: rates of change and exchange. J Mol Evol 44(4):383–397
Lee J, Jeong KW, Jin B, Ryu KS, Kim EH, Ahn JH, Kim Y (2013) Structural and dynamic features of cold-shock proteins of Listeria monocytogenes, a psychrophilic bacterium. Biochemistry 52:2492–2504
Lee SK, Park SH, Lee JW, Lim HM, Jung SY, Park IC, Park SC (2014) A putative cold shock protein-encoding gene isolated from Arthrobacter sp. A2-5 confers cold stress tolerance in yeast and plants. J Korean Soc Appl Biol Chem 57(6):775–782
Mazzon RR, Lang EAS, Silva CAPT, Marques MV (2012) Cold shock genes CspA and CspB from Caulobacter crescentus are post transcriptionally regulated and important for cold adaptation. J Bacteriol 194:6507–6517
Médigue C, Rouxel T, Vigier P, Hénaut A, Danchin A (1991) Evidence for horizontal gene transfer in Escherichia coli speciation. J Mol Biol 222(4):851–856
Metpally RPR, Reddy BVB (2009) Comparative proteome analysis of psychrophilic versus mesophilic bacterial species: insights into the molecular basis of cold adaptation of proteins. BMC Genom 10(1):11
Moon C, Jeong K, Kim HJ, Heo Y, Kim Y (2009) Recombinant expression, isotope labeling and purification of cold shock protein from Colwellia psychrerythraea for NMR study. Bull Korean Chem Soc 30:2647–2650
Moszer I, Rochaa EP, Danchin A (1999) Codon usage and lateral gene transfer in Bacillus subtilis. Curr Opin Microbiol 2(5):524–528
Nancy YY, Wagner JR, Laird MR, Melli G, Rey S, Lo R, Brinkman FS (2010) PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26(13):1608–1615
Panicker G, Jackie A, David S, Asim KB (2002) Cold tolerance of Pseudomonas sp. 30-3 isolated from oil-contaminated soil, Antarctica. Polar Biol 25(1):5–11
Paz I, Kligun E, Bengad B, Mandel-Gutfreund Y (2016) BindUP: a web server for non-homology-based prediction of DNA and RNA binding proteins. Nucleic Acids Res 44(W1):W568–W574
Phadtare S (2004) Recent developments in bacterial cold shock response. Curr Issues Mol Biol 6:125–136
Phadtare S, Alsina J, Inouye M (1999) Cold-shock response and cold-shock proteins. Curr Opin Microbiol 2:175–180
Piette A, D’Amico S, Mazzucchelli G, Danchin A, Leprince P, Feller G (2011) Life in the cold: a proteomic study of cold-repressed proteins in the Antarctic bacterium Pseudoalteromonashaloplanktis TAC125. Appl Environ Microbiol 77:3881–3883
Polissi A, DeLaurentis W, Zangrossi S, Briani F, Longhi V, Pesole G, Deho G (2003) Changes in Escherichia coli transcriptome during acclimatization at low temperature. Res Microbiol 154:573–580
Rai P, Sharma A, Saxena P, Soni A, Chakdar H, Kashyap PL, Srivastava A, Sharma AK (2015) Comparison of molecular and phenetic typing methods to assess diversity of selected members of the genus Bacillus. Microbiology 84(2):236–246
Ray MK, Sitaramamma T, Ghandhi S, Shivaji S (1994) Occurrence and expression of cspA, a cold shock gene, Antarctic psychrotropic bacteria. FEMS Microbiol Lett 116(1):55–60
Rodrigues DF, Tiedje JM (2008) Coping with our cold planet. Appl Environ Microbiol 74:1677–1686
Roy A, Alper K, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738
Schindelin H, Marahiel MA, Heinemann U (1993) Universal nucleic acid-binding domain revealed by crystal structure of the B. subtilis major cold-shock protein. Nature 364:164–168
Schindelin H, Jiang W, Inouye M, Heinemann U (1994) Crystal structure of CspA, the major cold shock protein of Escherichia coli. PNAS 91(11):5119–5123
Schröder K, Graumann P, Schnuchel A, Holak TA, Marahiel MA (1995) Mutational analysis of the putative nucleic acid-binding surface of the cold-shock domain, CspB, revealed an essential role of aromatic and basic residues in binding of single-stranded DNA containing the Y-box motif. Mol Microbiol 16(4):699–708
Schwede T (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
Shazman S, Mandel-Gutfreund Y (2008) Classifying RNA-binding proteins based on electrostatic properties. PLoS Comput Biol 4:1000–1146
Song W, Lin X, Huang X (2012) Characterization and expression analysis of three cold shock protein (CSP) genes under different stress conditions in the Antarctic bacterium Psychrobacter sp. G. Polar Biol 35:1515–1524
Stawiski EW, Gregoret LM, Mandel-Gutfreund Y (2003) Annotating nucleic acid-binding function based on protein structure. J Mol Biol 326:1065–1079
Tamura K (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Thieringer HA, Jones PG, Inouye M (1998) Cold shock and adaptation. BioEssays 20:49–57
Thompson JD, Gibson T, Higgins DG (2002) Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinform. https://doi.org/10.1002/0471250953.bi0203s00
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins Struct Funct Bioinform 64:643–651
Acknowledgements
The authors gratefully acknowledge the financial assistance under network project ‘Application of Microorganisms in Agriculture and Allied Sectors (AMAAS)’ and “CRP Genomics” from Indian Council of Agricultural Research (ICAR), India.
Author information
Authors and Affiliations
Contributions
HC conceptualized the study. Primers were designed by KM. Molecular works were carried out by ASh and PS. SA performed all computational analyses. JY performed the gene expression experiment. AB performed statistical analyses. HC, KM, KP and ASi drafted and revised the manuscript. AKS, PLK and AKS helped in execution of the experiments.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

Supplementary Figure 1
Multiple sequence alignment of cspA sequences from different Pseudomonas strains (JPEG 374 kb)

Supplementary Figure 2
The secondary structure of CSP from Pseudomonas koreensis P2 (JPEG 75 kb)

Supplementary Figure 3
Multiple alignment of the deduced amino acid sequences of CspA of Pseudomonas koreensisP2 (JPEG 467 kb)

Supplementary Figure 4
Ramchandran plot of the CspA model. The most favored regions are colored red, additional allowed, generously allowed and disallowed regions are indicated as yellow, light yellow and white fields, respectively (JPEG 84 kb)

Supplementary Figure 5
Model quality estimation plot obtained by QMEAN server. The area built by the circles colored in different shades of gray in the plot represents the QMEAN scores of the reference structures from the PDB (JPEG 70 kb)

Supplementary Figure 6
Verify 3D score of predicted CspA model (JPEG 77 kb)

Supplementary Figure 7
Overall quality factor for CspA protein model obtained from ERRAT server (JPEG 14 kb)

Supplementary Figure 8
Binding pockets (shown in different colors) of CspA protein from Psedomonas koreensis P2 (JPEG 58 kb)

Supplementary Figure 9
Figure showing electrostatic potential on CspA model (JPEG 54 kb)

Supplementary Figure 10
Three largest positive patches (in different blue color balls), calculated on a structural model of CspA. The model is predicted to be NA-binding (JPEG 49 kb)
Rights and permissions
About this article
Cite this article
Awasthi, S., Sharma, A., Saxena, P. et al. Molecular detection and in silico characterization of cold shock protein coding gene (cspA) from cold adaptive Pseudomonas koreensis. J. Plant Biochem. Biotechnol. 28, 405–413 (2019). https://doi.org/10.1007/s13562-019-00500-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-019-00500-8