Abstract
Background
Cyclin-dependent kinase 10 (CDK10) has recently been identified as a tumor suppressor and, concordantly, its encoding gene has frequently been found to be inactivated in various human cancers. Here, we examined the expression status of CDK10 in a panel of primary human breast cancers and evaluated its correlation with clinicopathological parameters and clinical outcome.
Methods
Western blotting was used to assess CDK10 protein levels in 20 paired breast cancer tissues and adjacent noncancerous tissues. In addition, immunohistochemistry was performed in 128 formalin-fixed, paraffin-embedded tumor tissues. Associations of CDK10 expression with various clinicopathological parameters were evaluated and Kaplan-Meier survival analyses and Cox proportional hazards models were used to estimate its effect on patient survival.
Results
We found that CDK10 protein expression was markedly decreased in cancer tissues compared to adjacent noncancerous tissues. Immunohistochemistry revealed decreased CDK10 levels in 65/128 (50.8 %) of the primary breast cancer tissues tested. These decreased levels were found to be significantly associated with lymph node metastasis (P = 0.003), advanced tumor stage (P < 0.001) and unfavorable overall survival (P < 0.001). Furthermore, multivariate analyses indicated that CDK10 expression may serve as an independent prognostic factor for survival (P = 0.001).
Conclusion
Down-regulated CDK10 expression frequently occurs in breast cancers and correlates with disease progression and poor survival. CDK10 may serve as a prognostic biomarker for breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Breast cancer is a heterogeneous disease encompassing a variety of pathological entities. These entities exhibit variable clinical characteristics [1, 2]. During the past decade we have witnessed substantial advances in the comprehensive molecular profiling of different breast cancer entities and the maturation of our understanding of their biology [3–7]. Such advances allow a more precise molecular stratification of patients, allow a better understanding of their clinical behavior and provide tools to identify novel targets for therapy [4, 8, 9]. Despite these improvements, however, numerous problems remain. For example, endocrine therapies and newly developed targeted drugs may lead to a better survival, but still a number of patients encounters either de novo or acquired resistance [10, 11]. Additionally, current molecular biomarkers and gene expression signatures for prognostics and therapy are still inadequate. Thus, there is a need to identify more and/or better biomarkers to improve a personalized clinical management of these patients [10].
Cyclin-dependent kinase 10 (CDK10; also called PISSLRE), whose encoding gene is located at 16q24.3, has in the past emerged as a tumor suppressor in breast cancer [12–14]. The CDK10/ETS2/RAF-1 signaling pathway has been found to act as an important determinant of resistance to endocrine therapy for breast cancer, as well as for neural progenitor survival [14–17]. Furthermore, it has been found that CDK10 over-expression results in enhanced chemosensitivity of hepatocellular carcinoma and biliary tract cancer cells [18, 19]. Previously, we found that the CDK10 gene is frequently silenced via promoter hypermethylation in nasopharyngeal carcinoma cells and that restoration of its expression strongly suppresses the malignant behavior of these cells [20]. More recently, we found that CDK10 expression, both at the mRNA and protein level, is positively correlated with C1orf63 expression, which may be involved in cell cycle exit and, as such, may act as a cellular quiescence-controlling gene [21]. These findings add new clues to our understanding of the role of CDK10 in breast cancer and signify its clinical value. Nevertheless, to the best of our knowledge, the prognostic value of CDK10 expression in patients with primary breast cancer has not been reported yet. Therefore, the specific objective in this study was to better characterize the clinical relevance of CDK10 expression in breast cancer. To do this, we first set out to determine CDK10 protein expression levels in 20 paired breast cancers and its adjacent noncancerous tissues by Western blotting. Next, we examined CDK10 expression in 128 primary breast cancer samples by immunohistochemistry and investigated its association with clinicopathologic parameters and clinical outcome.
2 Materials and methods
2.1 Patients and tissue samples
All patient samples were randomly collected from the Cancer Hospital of Shantou University Medical College. We collected 128 formalin-fixed paraffin-embedded specimens from patients with breast cancer undergoing curative surgery between 2000 and 2005 (median age, 51 years; range, 29–88 years). The patients were followed up for a mean period of 42 months (range, 1–77 months) from the date of surgery. In addition, cancer tissues (n = 20) and paired noncancerous tissues (n = 20) were collected for Western blot analysis. These latter tissues, independently obtained from another cohort of patients with breast cancer who underwent surgery at the same institution between May 2010 and July 2012, were immediately snap frozen in liquid nitrogen and kept at - 80 °C until further use.
Tumor grade and stage were classified in accordance with the American Joint Committee on Cancer (AJCC) pathologic tumor-node-metastasis (TNM) classification, sixth edition. The clinicopathologic characteristics of the patients, including the expression status of the estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor 2 (HER-2), are listed in Table 1. All cases were confirmed by two pathologists. No patients had undergone preoperative radiotherapy or chemotherapy. Signed informed consents were obtained. This study was approved by the Institution Review Board (# 04–070) of the Cancer Hospital of Shantou University Medical College.
2.2 Western blot analysis
CDK10 protein expression was assessed by Western blot analysis in human tissue lysates (60 mg of protein in RIPA lysis buffer). Proteins were separated by SDS-PAGE and transferred to PVDF membranes. The membranes were incubated in blocking buffer (Tris-buffered saline with 0.1 % Tween and 5 % nonfat dry milk) for 1 h and then incubated with a rabbit anti-CDK10 antibody (Abgent, San Diego, USA) at a dilution of 1:500 in blocking buffer, followed by a horseradish peroxidase-conjugated secondary antibody directed against rabbit IgG. Signals were visualized using an ECL chemiluminescence system according to the manufacturer’s instructions (Amersham Pharmacia, Piscataway, NJ, USA). Blots were re-probed with an anti-β-actin monoclonal antibody (Abcam) to confirm equal loading of the different samples. Quantification of CDK10 on the Western blots was performed using Bio-Rad Quantity One quantification software [22], and a ratio cancer versus paired non-cancer of less than two-fold was considered as down-regulated expression.
2.3 Immunohistochemistry and staining evaluation
Immunohistochemical staining for CDK10 was carried out using a standard EnVision complex method as reported previously [21]. Sections (4-μm) were cut from resected specimens, fixed in 10 % buffered formalin and embedded in paraffin. After deparaffinization and rehydration, endogenous peroxidase activity was blocked with 0.3 % hydrogen peroxide for 30 min at room temperature. For antigen retrieval, tissue sections were autoclaved at 121 °C in citrate buffer (10-mmol/L concentration, pH 6.0) for 10 min. Next, the specimens were incubated with a rabbit polyclonal anti-CDK10 antibody (1:300 dilution, Abgent). Immunohistochemical staining was carried out by an EnVision antibody complex (anti-Mouse/Rabbit) method using an EnvisionTM Detection kit (ZSGB-BIO, Beijing, China) and 3,3′-diaminobenzidine as the chromogen substrate. Nuclei were counterstained with hematoxylin. Negative controls were composed of identically treated histological sections with normal rabbit IgG to replace the primary antibody.
Staining evaluations were performed as follows: 10 random 400× microscopic fields per slide were evaluated by two independent observers who were blinded to the clinical information. CDK10 immunostaining was assessed using a semi-quantitative approach, which combines the percentage of positive cells and the staining intensity. The mean percentage of positively stained cells was scored as follows: 0 % (0); 1–25 % (1); 26–50 % (2); 51–75 % (3); 76–100 % (4). The staining intensities were categorized as follows: absent (0); weak staining exhibited as light-yellow (1); moderate staining exhibited as yellow-brown (2); and strong staining exhibited as brown (3). The sum of the intensity and percentage positive cells scores was used as the final staining score. For the purpose of statistical evaluation, tumors having a final staining score of < 3 were grouped into negative CDK10 expression and those with scores ≥ 3 were grouped into positive CDK10 expression.
2.4 Statistical analyses
Statistical analyses were performed using the SPSS 17.0 statistical software package (SPSS Inc., IL, USA). Differences in CDK10 expression between tumors and adjacent normal tissues were assessed by Student’s t test. Correlations between CDK10 expression and other molecular and clinicopathologic parameters were assessed using the Pearson χ 2 test or Fisher’s exact test. The correlation coefficient between variables was obtained using the Spearman’s rank method. Overall survival (OS) was defined as the time from diagnosis to the date of last contact or of death from any cause. Survival curves were calculated using the Kaplan-Meier method with a log rank test. The impact of clinicopathological variables on patient survival was determined by univariate analysis, and further examined by multivariate regression analysis using the Cox hazards model. P < 0.05 (two-tailed) was considered as statistically significant.
3 Results and discussion
Based on our previous results indicating that CDK10 may act as a tumor suppressor [22], we first set out to determine whether CDK10 is down-regulated in primary breast cancer tissues versus noncancerous tissues. In Fig. 1 representative Western blot results are shown from a cohort of 20 cases. We found that 15/20 (75 %) of the cancer tissues (C) exhibited a lower CDK10 protein expression compared to the paired adjacent noncancerous (N) tissues (P < 0.001).
Next, we investigated the CDK10 expression profiles in a cohort of 168 primary breast cancer specimens using immunohistochemistry. By doing so, positive CDK10 immunostaining was observed in the nucleus of neoplastic cells in 65/128 (50.8 %) of the primary tumors (Fig. 2). To further explore the significance of CDK10 expression in breast cancer, we next examined its relationship with various clinicopathological features. We found that CDK10 expression was associated with earlier tumor stages (stageI/II versus stage III/IV; P < 0.001), a lack of lymph node metastasis (P = 0.003), positive HER-2 expression (P = 0.004) and negative ER expression (P = 0.003) (Table 1). No significant association was found between CDK10 expression and the other clinicopathologic parameters examined, including age, tumor depth, histological type and PR expression. Spearman correlation analysis showed that the CDK10 expression levels were correlated with clinical tumor stage (r =−0.395; P < 0.001), lymph node metastasis (r =−0.359; P < 0.001), HER-2 expression (r = 0.245; P < 0.001) and ER expression (r =−0.177; P = 0.012). Together, these data indicate that loss of CDK10 expression may be associated with malignancy, mainly related to lymph node metastasis, and support the notion that deregulation of CDK10 expression may contribute to breast cancer progression.
Immunohistochemiscal CDK10 staining in primary breast cancer tissues and adjacent noncancerous tissues. a Strong CDK10 staining in breast cancer tissue. b Moderate CDK10 staining in breast cancer tissue. c Weak CDK10 staining in breast cancer tissue. d Positive CDK10 staining in adjacent noncancerous tissue. Original magnification, 200×
In order to assess the prognostic impact of CDK10 expression on the outcome of breast cancer patients, Kaplan-Meier survival analyses were performed. As shown in Fig. 3a, the overall survival (OS) rate of patients with a positive CDK10 score was significantly higher than that of patients with a negative score (P < 0.001). The relationship between CDK10 expression and patient survival was independently assessed in patients with lymph node metastasis (N1-N3) and in patients without lymph node metastasis (N0). In both groups, the OS rate was higher in patients carrying tumors with a positive CDK10 expression score than in patients carrying tumors with a negative CDK10 expression score (P = 0.019 for N0 group and P < 0.001 for N1-N3 group) (Fig. 3b and c).
Kaplan-Meier survival curves with univariate analyses (log-rank) according to the expression statues of CDK10 in patients with breast cancer. a The overall survival rate of patients with CDK10-positive tumors was significantly higher than that of patients with CDK10-negative tumors (P < 0.001). b Among the patients without lymph node metastasis (N0), the overall survival was significantly better in patients with CDK10-positive tumors than in patients with CDK10-negative tumors (P = 0.019). c Among the patients with lymph node metastasis (N1-N3), the overall survival was significantly better in patients with CDK10-positive tumors than in patients with CDK10-negative tumors (P < 0.001)
To test the possibility that CDK10 expression may serve as a prognostic biomarker for breast cancer patients, we applied univariate and multivariate Cox-regression models. Upon univariate analyses, we found that older patient age (P = 0.026), negative CDK10 expression score (P < 0.001), lymph node metastasis (N1- N3 versus N0; P < 0.001), advanced tumor stages (III/IV versus I/II; P = 0.001), tumor histological type (ductal versus non-ductal; P = 0.001) and HER2 expression (P < 0.006) were significantly correlated with an unfavorable OS (Table 2). Multivariate analyses, however, indicated that age (P = 0.029), tumor stage (P = 0.045) and decreased CDK10 expression (P = 0.001) acted as independent prognostic indicators for an unfavorable OS in our study. These results indicate that CDK10 expression may serve as a predictor for the survival of patients with breast cancer.
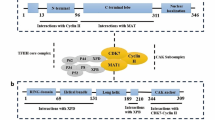
Orderly progression through the cell cycle requires sequential activation and inactivation of cyclin-dependent kinases (CDKs). This is achieved in part through the association of CDKs with positive regulators (i.e., cyclins) and inactivation of cyclin-CDK complexes by cyclin-CDK inhibitors [23]. The role of cell cycle control proteins, both as primary effectors and as mediators of tumorigenesis, has gained increasing interest [24–26]. As a member of the family of Cdc2-related kinases, CDK10 has been found to act as a cyclin-dependent kinase (CDK) through the identification of cyclin M as activating cyclin [27]. Functional studies have revealed that CDK10 silencing increases ETS2-driven activation of the MAPK pathway, which confers tamoxifen resistance to breast cancer cells [14–16], raising the possibility that patients carrying tumors that express low levels of CDK10 may have a worse clinical outcome.
The present work is part of our effort to unravel the link between CDK10 expression profiles and clinical outcome, with the aim to identify new diagnostic and prognostic biomarkers, as well as new therapeutic targets. CDK10 down-regulation has been reported to occur in several earlier cancer-related studies [14, 18, 19]. Recently, we found that the CDK10 gene was frequently silenced via promoter hypermethylation in nasopharyngeal carcinomas [20], but in breast cancer this mechanism is still controversial [14, 28]. A reduced CDK10 expression was previously found to be associated with lymph node metastasis and tumor stage in biliary tract cancer [18] and with alpha-fetoprotein level, tumor size and tumor stage in hepatocellular carcinoma [19]. Here, we found that loss of CDK10 expression was associated with advanced disease stage, lymph node metastasis, HER-2 expression and ER expression in breast cancer. Of these factors, metastasis appears to be of major prognostic significance [29]. Our results suggest that CDK10 silencing affects breast cancer progression and, more directly, correlates with an aggressive biological behavior. Our survival analyses revealed a prognostic value of CDK10 expression in breast cancer patients in general, whereas our multivariate analyses identified CDK10 expression as a strong independent prognostic factor for a favorable OS. These observations suggest that CDK10 may serve as an important node in tumor progression. The exact downstream effects of CDK10 in breast cancer progression require further exploration. Since patients usually present with locally advanced disease, biomarkers other than lymph node status may be needed to predict patient outcome [30]. In this study, a lack of CDK10 expression was also associated with a poor OS in patients without lymph node metastasis. This result suggests that CDK10 may be of value in predicting the outcomes of patients with early-stage disease.
The estrogen receptor (ER)-alpha is a key regulatory molecule in mammary epithelial cell development. Therapies that target estrogen signaling have transformed the treatment of breast cancer. Loss of ER-alpha in breast cancer is correlated with poor prognosis, increased recurrence after treatment, and an elevated incidence of metastasis [29]. It has been shown that patients carrying ER-alpha positive tumors that express low levels of CDK10 relapse early upon tamoxifen treatment [14]. CDK10 silencing leads to ETS2-driven activation of the promoter of the RAF1 gene, thereby enhancing ERK/MAPK kinase pathway activity and relieving tamoxifen-induced G1 cell cycle arrest in cancer cells [14–16]. We, however, failed to observe any significant difference in OS between patients with or without CDK10 expression in the subgroup of ER-positive patients (data not shown).
There are some limitations to our current study. First, to reveal more significant data an extension of our sample size may be required. Second, to substantiate the results obtained from primary patient samples, validation under well-controlled conditions in cell and/or animal models may be required. Nevertheless, collectively our results indicate that decreased CDK10 expression serves as a molecular signature of breast cancer progression. CDK10 down-regulation was found to be associated with malignant properties mainly relevant to metastasis and to correlate with a poor overall survival. We conclude that CDK10 expression may serve as an independent prognostic predictor that holds therapeutic promise for breast cancer.
References
L.A. Torre, F. Bray, R.L. Siegel, J. Ferlay, J. Lortet-Tieulent, A. Jemal, Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015)
C.M. Perou, T. Sørlie, M.B. Eisen, M. van de Rijn, S.S. Jeffrey, C.A. Rees, J.R. Pollack, D.T. Ross, H. Johnsen, L.A. Akslen, O. Fluge, A. Pergamenschikov, C. Williams, S.X. Zhu, P.E. Lønning, A.L. Børresen-Dale, P.O. Brown, D. Botstein, Molecular portraits of human breast tumours. Nature 406, 747–752 (2000)
S. Di Cosimo, J. Baselga, Management of breast cancer with targeted agents: importance of heterogeneity. Nat. Rev. Clin. Oncol. 7, 139–147 (2010)
J. Peppercorn, C.M. Perou, L.A. Carey, Molecular subtypes in breast cancer evaluation and management: divide and conquer. Cancer Investig. 26, 1–10 (2008)
S. Tabarestani, S.M. Ghaderian, H. Rezvani, R. Mirfakhraie, A. Ebrahimi, H. Attarian, J. Rafat, M. Ghadyani, H.A. Alavi, N. Kamalian, A. Rakhsha, E. Azargashb, Prognostic and predictive value of copy number alterations in invasive breast cancer as determined by multiplex ligation-dependent probe amplification. Cell. Oncol. 37, 107–118 (2014)
A.H. Verschuur-Maes, C.B. Moelans, P.C. de Bruin, P.J. van Diest, Analysis of gene copy number alterations by multiplex ligation-dependent probe amplification in columnarcell lesions of the breast. Cell. Oncol. 37(147–154) (2014)
E. Yiannakopoulou, Etiology of familial breast cancer with undetected BRCA1 and BRCA2 mutations: clinical implications. Cell. Oncol. 37(1–8) (2014)
L.J. van’t Veer, H. Dai, M.J. van de Vijver, Y.D. He, A.A. Hart, M. Mao, H.L. Peterse, K. van der Kooy, M.J. Marton, A.T. Witteveen, G.J. Schreiber, R.M. Kerkhoven, C. Roberts, P.S. Linsley, R. Bernards, S.H. Friend, Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002)
Cancer Genome Atlas Network, Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012)
E.A. Musgrove, R.L. Sutherland, Biological determinants of endocrine resistance in breast cancer. Nat. Rev. Cancer 9, 631–643 (2009)
C.A. Hudis, Trastuzumab-mechanism of action and use in clinical practice. N. Engl. J. Med. 357, 39–51 (2007)
F. Bullrich, T.K. MacLachlan, N. Sang, T. Druck, M.L. Veronese, S.L. Allen, N. Chiorazzi, A. Koff, K. Heubner, C.M. Croce et al., Chromosomal mapping of members of the cdc2 family of protein kinases, cdk3, cdk6, PISSLRE, and PITALRE, and a cdk inhibitor, p27Kip1, to regions involved in human cancer. Cancer Res. 55, 1199–1205 (1995)
J. Crawford, L. Ianzano, M. Savino, S. Whitmore, A.M. Cleton-Jansen, C. Settasatian, M. D’apolito, R. Seshadri, J.C. Pronk, A.D. Auerbach, P.C. Verlander, C.G. Mathew, A.J. Tipping, N.A. Doggett, L. Zelante, D.F. Callen, A. Savoia, The PISSLRE gene: structure, exon skipping, and exclusion as tumor suppressor in breast cancer. Genomics 56, 90–97 (1999)
E. Iorns, N.C. Turner, R. Elliott, N. Syed, O. Garrone, M. Gasco, A.N. Tutt, T. Crook, C.J. Lord, A. Ashworth, Identification of CDK10 as an important determinant of resistance to endocrine therapy for breast cancer. Cancer Cell 13, 91–104 (2008)
C.J. Lord, E. Iorns, A. Ashworth, Dissecting resistance to endocrine therapy in breast cancer. Cell Cycle 7, 1895–1898 (2008)
P. Khanal, H.J. Yun, S.C. Lim, S.G. Ahn, H.E. Yoon, K.W. Kang, R. Hong, H.S. Choi, Proyl isomerase Pin1 facilitates ubiquitin-mediated degradation of cyclin-dependent kinase 10 to induce tamoxifen resistance in breast cancer cells. Oncogene 31, 3845–3856 (2012)
C.W. Yeh, S.H. Kao, Y.C. Cheng, L.S. Hsu, Knockdown of cyclin-dependent kinase 10 (cdk10) gene impairs neural progenitor survival via modulation of raf1a gene expression. J. Biol. Chem. 288, 27927–27939 (2013)
J.H. Yu, X.Y. Zhong, W.G. Zhang, Z.D. Wang, Q. Dong, S. Tai, H. Li, Y.F. Cui, CDK10 functions as a tumor suppressor gene and regulates survivability of biliary tract cancer cells. Oncol. Rep. 27, 1266–1276 (2012)
X.Y. Zhong, X.X. Xu, J.H. Yu, G.X. Jiang, Y. Yu, S. Tai, Z.D. Wang, Y.F. Cui, Clinical and biological significance of Cdk10 in hepatocellular carcinoma. Gene 498, 68–74 (2012)
Y. You, W. Yang, Z. Wang, H. Zhu, H. Li, C. Lin, Y. Ran, Promoter hypermethylation contributes to the frequent suppression of the CDK10 gene in human nasopharyngeal carcinomas. Cell. Oncol. 36, 323–331 (2013)
C.Q. Hong, F. Zhang, Y.J. You, W.L. Qiu, A.E. Giuliano, X.J. Cui, G.J. Zhang, Y.K. Cui, Elevated C1orf63 expression is correlated with CDK10 and predicts better outcome for advanced breast cancers: a retrospective study. BMC Cancer 15, 548 (2015)
Y. You, W. Yang, X. Qin, F. Wang, H. Li, C. Lin, W. Li, C. Gu, Y. Zhang, Y. Ran, ECRG4 acts as a tumor suppressor and as a determinant of chemotherapy resistance in human nasopharyngeal carcinoma. Cell. Oncol. 38, 205–14 (2015)
A.S. Howell, D.J. Lew, Morphogenesis and the cell cycle. Genetics 190(1), 51–77 (2012)
S. Mahale, S.B. Bharate, S. Manda, P. Joshi, P.R. Jenkins, R.A. Vishwakarma, B. Chaudhuri, Antitumour potential of BPT: a dual inhibitor of cdk4 and tubulin polymerization. Cell Death Dis. 6, e1743 (2015)
T. VanArsdale, C. Boshoff, K.T. Arndt, R.T. Abraham, Molecular pathways: targeting the cyclin D-CDK4/6 axis for cancer treatment. Clin. Cancer Res. 21, 2905–2910 (2015)
Z. Feng, S. Xu, M. Liu, Y.X. Zeng, T. Kang, Chk1 inhibitor Gö6976 enhances the sensitivity of nasopharyngeal carcinoma cells to radiotherapy and chemotherapy in vitro and in vivo. Cancer Lett. 297, 190–197 (2010)
V.J. Guen, C. Gamble, M. Flajolet, S. Unger, A. Thollet, Y. Ferandin, A. Superti-Furga, P.A. Cohen, L. Meijer, P. Colas, CDK10/cyclin M is a protein kinase that controls ETS2 degradation and is deficient in STAR syndrome. Proc. Natl. Acad. Sci. U. S. A. 110, 19525–19530 (2013)
G. Heller, B. Ziegler, A. Brandstetter, S. Novak, M. Rudas, G. Hennig, M. Gehrmann, T. Acht, S. Zöchbauer-Müller, M. Filipits, CDK10 is not a target for aberrant DNA methylation in breast cancer. Anticancer Res. 29, 3939–3944 (2009)
A. Dhasarathy, M. Kajita, P.A. Wade, The transcription factor snail mediates epithelial to mesenchymal transitions by repression of estrogen receptor-alpha. Mol. Endocrinol. 21, 2907–2918 (2007)
Y. Ran, S. Wu, Y. You, Demethylation of E-cadherin gene in nasopharyngeal carcinoma could serve as a potential therapeutic strategy. J. Biochem. 149, 49–54 (2011)
Acknowledgments
This work was supported in part by the Science and Technology Planning Project of Henan Province, China (142102310464), the Key Research Foundation of Higher Education of Henan Province, China (15B320003), the Annual Natural Science Foundation of Luohe Medical College (2015-S-LMC02), the Natural Science Foundation of Hubei Province (2014CFC1154), the Foundation of Medical College of Hubei University of Arts and Science (YXKY 201402) and the Scientific Research Foundation for Doctoral Program of Hubei University of Arts and Science (X. Qin).
Conflicts of interest statement
No potential conflicts of interest were disclosed.
Author information
Authors and Affiliations
Corresponding author
Additional information
Yanjie You and Haijun Li are co-first authors for this manuscript. These authors have contributed equally to this work.
Rights and permissions
About this article
Cite this article
You, Y., Li, H., Qin, X. et al. Decreased CDK10 expression correlates with lymph node metastasis and predicts poor outcome in breast cancer patients - a short report. Cell Oncol. 38, 485–491 (2015). https://doi.org/10.1007/s13402-015-0246-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-015-0246-4