Abstract
Mass spectrometry-based methods have made significant progress in characterizing post-translational modifications in peptides and proteins; however, certain aspects regarding fragmentation methods must still be improved. A good technique is expected to provide excellent sequence information, locate PTM sites, and retain the labile PTM groups. To address these issues, we investigate 10.6 μm IRMPD, 213 nm UVPD, and combined UV and IR photodissociation, known as HiLoPD (high-low photodissociation), for phospho-, sulfo-, and glyco-peptide cations. IRMPD shows excellent backbone fragmentation and produces equal numbers of N- and C-terminal ions. The results reveal that 213 nm UVPD and HiLoPD methods can provide diverse backbone fragmentation producing a/x, b/y, and c/z ions with excellent sequence coverage, locate PTM sites, and offer reasonable retention efficiency for phospho- and glyco-peptides. Excellent sequence coverage is achieved for sulfo-peptides and the position of the SO3 group can be pinpointed; however, widespread SO3 losses are detected irrespective of the methods used herein. Based on the overall performance achieved, we believe that 213 nm UVPD and HiLoPD can serve as alternative options to collision activation and electron transfer dissociations for phospho- and glyco-proteomics.

ᅟ
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The identification and mapping of post-translational modifications (PTMs) in peptides and proteins is challenging because of their low abundance, lability, and unique chemical properties [1, 2]. Mass spectrometry-based analysis of phosphorylation [3], sulfonation [4], and glycosylation [5] plays an important role in understanding their diverse biological functions. Phosphorylation by protein kinases regulates signal transduction for diverse intracellular processes [6, 7]. Many diseases such as cancer, inflammation, metabolic disorders, and neurodegenerative diseases are also linked to kinase protein phosphorylation [8]. The key functions of tyrosine sulfation are protein–protein interaction regulation, hormonal regulation, and hemostasis [9, 10]. It is difficult to characterize the sulfo-proteome because of its very acidic nature and labile sulfo-ester bond [11]. In glycoproteins, an oligosaccharide chain (glycan) is covalently attached to the polypeptide side-chain [12]. Glycosylation is associated with plasma-membrane and secretory proteins [13]. Moreover, proteins that have an extracellular segment are often glycosylated. Glycosylation has been linked with several human diseases such as inflammation [14], cancer [15], genetic disorders [16], and neurodegenerative disorders [17, 18]. Glycoproteins are difficult to characterize because of the low-abundance, complexity, and heterogeneity of glycan structures [19].
Tandem mass spectrometry (MS/MS) has emerged as an indispensable tool for analyzing the PTMs of proteins as it can provide structural information with high accuracy, relative speed, and sensitivity [20, 21]. Fragmentation methods are crucial to obtain precise structural information. Collision induced dissociation (CID) is frequently applied for fragmenting peptide ions. Although CID can recognize the presence of phosphate (especially from pSer and pThr) in a peptide or protein, by identifying the loss of a phosphate (–80 HPO3 or –98 H3PO4) group from the precursor ion, identifying the exact site is a challenging problem [22]. The neutral loss of phosphate groups from tyrosine is not always observed because of the strong phosphate-tyrosine binding energy and lower abundance of pTyr phosphorylation compared with pSer and pThr [23, 24]. Moreover, as sulfonation (SO3) and phosphorylation (HPO3) both result in the loss of 80 Da, it makes PTM identification even more challenging. One inherent problem with CID is that the excitation of precursor ions requires increasing internal energy, which increases neutral loss and in turn provides limited structural information [22]. However, metastable atom-activated dissociation (MAD) and higher-energy collision dissociation (HCD) experiments on phosphorylated and sulfonated peptides in negative ion mode have led to significant improvements [25, 26]. Recently, using dual spray ion/ion reactions, traditional CID underwent significant improvement in terms of phosphate group fragmentation and retention [27].
Electron-driven methods based on ‘ion–electron’ activation in electron capture dissociation (ECD) [28] and ‘ion–ion’ activation in electron transfer dissociation (ETD) [29] have been developed as an alternative to CID. In ECD and ETD, low energy electrons (~1 eV) are captured (or transferred) by precursor ions [30]. After receiving an electron, the activated precursor ions specifically break the N–Cα bonds and yield c and z ions without abundant side-chain loss, making it possible to identify the locations of PTM sites [31, 32]. However, ECD and ETD methods require multiply charged ions, which is difficult to form for the acidic phosphate and sulfonate groups in PTMs [30, 33, 34]. Incorporating metal ions in phospho and sulfo sites can improve localization and fragmentation by generating multiple charge states [35, 36]. Owing to the acidic nature of phospho- and sulfo- groups in the PTM peptide, they can present improved ionization when negative polarity is used in ESI (electrospray ionization) and provide good fragmentation while retaining the PTM groups [37,38,39]. However, irrespective of activation methods, peptide anions produce more complex MS/MS spectra caused by manifold fragmentation events with widespread side chain losses, making it difficult to process, interpret, and analyze the resulting complex data sets [40,41,42,43,44,45].
Alternatives to ‘ion–ion’ activation technique are also available, such as various UV photon-based methods, including 157 nm [46], 193 nm [47,48,49,50], 220 nm [51], 266 nm [52] ultraviolet photodissociation (UVPD), and electron detachment dissociations (EDD) [53]. Kim and Reilly observed series of a/x ions in 157 nm VUVPD on phospho-peptides and noticed the retention of the phosphate group [46]. They also found that phospho-tyrosine is more stable compared with phosphoserine or phosphothreonine. 193 nm UVPD with negative polarity is capable of providing interesting features, such as excellent sequence coverage and the retention of H3PO4 and SO3 groups from product ions of phospho- and sulfo-peptides [47, 48, 50]. 220 nm UVPD on protonated tyrosine containing phospho-peptides showed characteristic aromatic side chain losses of the tyrosine residue [51]. Aromatic side chain loss was also observed at 266 nm for electron detachment dissociation (EDD) for peptide anions [53]; however, this loss was suppressed for phospho-peptide cations [52]. Compared with high-energy UV photodissociation, few studies have been performed using 10.6 μm infrared multiphoton dissociation (IRMPD) [22, 54, 55, 56] for PTM characterization. Despite several challenges and difficulties, the potential of IRMPD is similar to certain other photoexcitation techniques [57, 58] since this method (1) requires no alternation of the stable trajectory or kinetic energy of the trapped ion for excitation; (2) is not associated with low cutoff m/z; (3) can provide reasonable fragmentation efficiency; (4) can operate without collision gas; (5) it is compatible with the vibrational modes of PO4 3– and SO3 groups present in PTMs.
Although various wavelengths between 157 and 266 nm were employed in ultraviolet photodissociation for characterizing post-translational modifications, 213 nm UVPD [41, 59] has never been used before for this purpose. A single high energy UV photon is sufficient to promote the dissociation of a peptide and protein, cleaving mainly Cα–C bonds and producing abundant a/x ions with some y and z ions. On the other hand, the multiple absorption of low energy IR photons is required to increase the internal energy and cleave the labile C–N bonds, generating mainly b/y ions. In general, the key advantages of coupling high and low energy photon-based activation are that they provide a balanced and diverse array of fragment ions and an even distribution of the fragment ions across the entire m/z range. Although 213 nm UVPD and other methods (157 nm and 193 nm UVPD) provide large numbers of fragment ions, the latter are usually highly charged and are thus observed at m/z very close to that of the precursor ion, hence crowding the MS/MS spectra. In this study, we employ a new method called HiLoPD (high-low photodissociation) [60], which combines high-energy UV and low-energy IR lasers with a high resolution Q-Exactive mass spectrometer encompassing high and low photoactivation channels for PTM characterization. We also evaluate the performance of 10.6 μm IRMPD, 213 nm UVPD, and HiLoPD for phospho-, sulfo- and glyco-peptide cation characterization in view of achieving three goals: (1) obtain adequate backbone fragmentation with good sequence coverage; (2) identify the exact position of PTMs groups; and (3) compare the loss and retain events of the labile PTMs groups in the fragment ions.
Materials and Methods
Sample Preparation
Phospho-, sulfo-, and glyco-peptides such as RRLIEDAEY(H2PO4) AARG from tyrosine kinase peptide, FFKNIVTPRT(H2PO4)PPPSQGK, RDY(SO3)TGWLDF and EAISPPDAAS(GalNAc)AAPLR from GalNAc-Ser Erythropoietin (177-131) were obtained from GeneCust Europe. All the peptides were used without any further purification. All the peptide samples were prepared at 2 μM concentration in 50/49/1 (v/v/v) methanol/water/acetic acid.
Mass Spectrometry
All the experiments were performed on a hybrid quadrupole-Orbitrap Q-Exactive mass spectrometer (Thermo Fisher Scientific, San Jose, CA, USA) equipped with an HESI ion source. Positive polarity was used for all the peptides. All the mass spectra were acquired using a mass range of 200–2000 m/z and resolving power of 140,000 at m/z 400. Spray voltage, capillary temperature, and sheath gas flow rate were set to 3.5–4.0 kV, 320 °C, and 5–10, respectively. The AGC (automatic gain control) target was set to 5 × 106 and the maximum injection time was set at 250 ms. The isolation width was 1–2 Th. To avoid collisions and CID contamination, the HCD collision energy was set to the minimum 2 eV. All the experiments were performed for three microscans with averaging for 50 scans.
Photodissociation
IRMPD experiments were performed with a 50 W continuous-wave CO2 laser (model ULR-50; Universal Laser System, Scottsdale, AZ, USA). To compare backbone fragmentation and retaining PTM groups, various nominal laser powers from 10% to 60% were used. However, we noticed that 10%–30% laser power is enough to achieve excellent fragmentation and comparable PTM loss and retention events. Although high laser power can provide a significant number of fragment ions, it is difficult to retain the PTM groups. Compared with protein samples, peptides generally required shorter irradiation times from 50 to 500 ms. For the IRMPD experiment, N2 gas pressure in the HCD cell had to be lowered to reduce collisional cooling and obtain fragmentation [60]. For the PTM peptides, the pressure controller was set to ~0.1–0.15 MPa to obtain reasonable trapping and good signals.
For the UVPD experiments, which were similar to the previous experiment [41, 59], the fifth harmonic (λ=213 nm, ~1 mJ/pulse) of a 20 Hz BrillantB solid-state Nd:YAG laser (Quantel, Les Ulis, France) was used. In brief, to generate 213 nm UV light, the fundamental 1064 nm light is passed through the non-linear crystal to generate the second harmonic at 532 nm. Then the second harmonic is sent through the second crystal where the fourth harmonic at 266 nm is generated. The remaining fundamental light is then allowed to interact with the fourth harmonic (266 nm) to produce the fifth harmonic at 213 nm by sum-frequency generation (SFG). A mechanical shutter (SH05/TSC001; Thorslab) was used to control the UV beam in the HCD cell. For PTM peptides, the optimal shutter opening time used here was 50–100 ms (1–2 laser shots), as these peptides require few UV laser shots.
HiLoPD experiments were performed with combined IRMPD and UVPD irradiation in the HCD cell of a hybrid quadrupole Orbitrap mass spectrometer. The detailed set up was described elsewhere [60]. In brief, the generated fifth harmonic 213 nm laser beam passes through two dichroic mirrors, lenses, optical mirrors, and is then introduced into the HCD cell. In addition, the IR beam is directed at the HCD cell using gold mirrors and a half-moon (D-shaped) mirror. The IR beam is gated on an external TTL signal. To combine the irradiating UV and IR beams, a BaF2 window (wavelength range 0.2–12 μm, Ø 25.4 mm, thickness 5 mm) is placed at the rear of the HCD cell. This window can transmit both IR (10.6 μm) and UV (213 nm) beams with an efficiency of 90% and 85%, respectively. Coupling schemes between IR and UV are implemented by simultaneous irradiation of the CO2 laser (50–200 ms) with 10%–30% of nominal laser power and 1–2 shots of UV laser.
Data Analysis
Manual interpretation of the IRMPD, UVPD, and HiLoPD data was performed with the assistance of Protein Prospector v5.14.4. (http://prospector.ucsf.edu/prospector/mshome.htm). All the major ion types (a, a+1, a+2, b–1, b, b+1, b+2, c–1, c, c+1, x–1, x, x+1, x+2, y, y–1, y–2, z–1, z, z+1) were considered. To identify PTM loss, the exact masses of the labile groups were subtracted from the precursor and fragment ions and a mass list was created in Excel manually. These values were then searched throughout the spectra. H2O and NH3 losses from the fragment ions were also considered.
For the PTM loss and retention assessments, the position of the PTM sites relative to the N- and C-terminal ions of a/b/c and x/y/z, respectively, were considered (Scheme 1). Only fragment ions that contained the modified amino acid were taken into account. The losses of similar groups (such as H3PO4 and HPO3) from the same fragment ion were counted as a ‘one loss’ event. For instance, if both H3PO4 and HPO3 were lost from the y5 ion, this was counted as a ‘one loss’ event and not ‘two losses’. In addition, the detection of several y–1, y, y+1 or a, a+2 ions from the same backbone position was counted as a ‘one retain’ event rather than several events. PTM retention specificity was calculated by the following equation:
Results and Discussion
IRMPD, UVPD, and HiLoPD on RRLIEDAEY(H2PO4)AARG
The IRMPD, UVPD, and HiLoPD photodissociation spectra of the triply protonated [M+3H]3+ (m/z 533.9346) of peptide RRLIEDAEY(H2PO4)AARG are presented in Figure 1. Theoretical m/z, observed m/z, and assignments for fragment ions detected in the IRMPD, UVPD and HiLoPD experiments on this peptide are summarized in Supplementary Table S1. The abundance of fragment ions, excluding phosphate losses, in all three methods is compared in Figure 2a.
IRMPD, UVPD and HiLoPD spectra of the +3 charge state precursor ion (m/z 533.9356) of RRLIEDAEY(H2PO4)AARG peptide. The precursor ions are indicated by star (*) sign. Sequence coverage (%) presented in the bracket. Detailed assignments of the fragment ions are summarized in Supplementary Table S1
Number of fragment ions (without PTM loss) detected in IRMPD, UVPD, and HiLoPD spectra of (a) the +3 charge state precursor ion (m/z 533.9356) of RRLIEDAEY(H2PO4)AARG peptide. (b) the +3 charge state precursor ion (m/z 665.3544) of FFKNIVTPRT(H2PO4)PPPSQGK peptide. (c) the +2 charge state precursor ion (m/z 626.7492) of RDY(SO3)TGWLDF peptide. (d) the +3 charge state precursor ion (m/z 556.9529) of EAISPPDAAS(GalNAc)AAPLR peptide
Since the stretching of the P–O bond (9.6–11 μm or 1042-909 cm–1) is in resonance with the 10.6 μm wavelength, the phosphate group can stimulate chromophore-driven efficient dissociation [61]. The IRMPD spectrum provided good overall sequence coverage of 75%, including sequence information in the low m/z region. IRMPD showed similar sequence coverage with N-terminal ions and C-terminal ions (66%). The neural losses of 98.0118 and 79.9986 Da, which corresponded to the elimination of H3PO4 and HPO3 groups, were observed from the precursor ions at m/z 501.2637 and 507.2681, respectively. The neutral loss of H2O was detected at m/z 527.9238. In addition, IRMPD exhibited substantial backbone fragmentation (excluding phosphate losses) producing 30 b ions and 13 y ions (Figure 2a). Regarding site-specific PTM losses, five were detected from yn ions (n = 5–9), whereas only two were identified from bn ions (n = 9, 10) (Table 1). However, phosphate groups were retained in 4 yn ions (n = 5, 6, 7, 8). The overall phosphate retention efficiency in IRMPD was 36.4%.
The UVPD experiment on the +3 ion of this peptide allowed the detection of a total of 87 fragment ions (excluding phosphate losses) with one laser pulse, which is nearly twice the number of fragment ions detected with IRMPD (Figure 2a and Supplementary Table S1). The neutral loss peaks at m/z 527.9256, 506.9291, and 501.2650 correspond to the elimination of H2O, HPO3 and H3PO4 from the precursor ion [M+3H]3+. The neutral loss of CH3CH2 noticed at m/z 524.2445 represents the side chain of Ile [62]. The peak at m/z 498.5760 corresponds to the loss of O=C6H4=CH2 (106.0836 Da) from tyrosine [63]. However, in comparison to IRMPD, only 14 b ions were identified in UVPD. Besides the traditional a/x, y, and c/z ions, a+1/x+1, x+2, y–1, y–2, c–1, and c+1 ions of this peptide were also detected. Despite the absence of a proline residue, we observed y–1 and y–2 ions from the secondary detachment of the x+1 radical [64, 38]. The UVPD spectrum provided 83% overall sequence coverage, with a significant number of fragment ions. However, the same sequence coverage (66%) was observed with N- and C-terminal ions.
For UVPD, the neutral losses of H3PO4 and HPO3 groups from the fragment ions were observed from yn ions at positions n = 5–9. Only two such losses were identified for zn ions (n = 5, 7). Surprisingly, no such phosphate loss was detected for a, b, c, or x ions. A significant number of preserved phosphate groups were observed for xn (n = 5, 6, 7), yn (n = 5, 6, 7, 8), zn (n = 5, 6), and an (n = 9) ions (Table 1). An overall phosphate retention efficiency of 58.8% was obtained for UVPD.
HiLoPD on the same peptide produced a wider range of fragmentation types, a, b, c, x, y, and z, owing to its high and low activation channels combining UV and IR photodissociation. While UVPD and HiLoPD had the same sequence coverages (83%), HiLoPD presented a diverse array of fragment ions from the N- and C-terminals. “UV-type” fragment ions (i.e., a/c and x/z) allowed increasing the sequence coverage in HiLoPD compared to IRMPD. A significant number of b ions (31 fragments) were identified in HiLoPD, similar to IRMPD, whereas many of them were absent in UVPD (Figure 2a). However, compared with UVPD, the number of a/x ions was lower in HiLoPD. In addition to the traditional ion types, the spectrum also contained highly abundant ions corresponding to neutral losses of water and ammonia from the fragment ions (Supplementary Table S1). As with UVPD, phosphate losses were observed for yn (n = 5–9) and zn (n = 5, 7) ions (Table 1). Interestingly, it appeared that the loss of HPO3 from y ions was more frequent in HiLoPD and UVPD compared with IRMPD. In general, the activation of phospho-peptide molecular ion by collision induced the cleavage of the C–O–P ester bridge. If the cleavage of C–O bond occurred with hydrogen transfer, this led to phosphoric acid (H3PO4) loss, whereas breaking the O–P bond promoted the loss of the HPO3 group. In the collision activation of tyrosine phosphorylated peptide, neutral loss of the HPO3 (79.9657 Da) group is usually observed [65] and the loss of the H3PO4 group is less likely to occur, since the bond dissociation energy of a C–O bond adjacent to an aromatic ring is quite high compared with that of a P–O bond, and the second aromatic group does not promote E2-elimination or SN2-neighbouring group participation reaction [66]. Previous studies reported that the H3PO4 group can be removed from phosphorylated tyrosine through the concurrent or sequential loss of HPO3 from the tyrosine residue and of H2O from elsewhere in the peptide [67, 68]. Moreover, six x-ions and seven y-ions, still containing the phosphate groups, were also detected (Table 1). The overall phosphate retention efficiency in HiLoPD was 50.0%.
Although significant losses of phosphate groups from the product ions are not desirable, some of these losses along with high sequence coverage can certainly confirm the phosphate location on a phospho-peptide. In IRMPD, the elimination of the H3PO4 group was identified from b9-10 as well as from y5-9 ions (Table 1 ). In UVPD and HiLoPD, the neutral losses of H3PO4 and HPO3 groups were observed only from y5-9, z5, and z7 fragment ions. As evidenced by these results, no phosphate loss was detected from y1-4 and b1-8 ions, confirming that the phosphate group was attached to the tyrosine at position 9 from the N-terminal.
IRMPD, UVPD, and HiLoPD on FFKNIVTPRT(H2PO4)PPPSQGK
The IRMPD, UVPD, and HiLoPD photodissociation spectra of the triply-protonated [M+3H]3+ (m/z 665.3544) of peptide FFKNIVTPRT(H2PO4)PPPSQGK is presented in Figure 3. Theoretical m/z, observed m/z, and assignments for fragment ions detected in IRMPD, UVPD, and HiLoPD experiments on this peptide are summarized in Supplementary Table S2. Losses of HPO3 and H3PO4 groups from the precursor ion were detected at m/z 638.6534 and 632.6923, respectively. Neutral losses of water and ammonia were also observed in these spectra.
IRMPD, UVPD, and HiLoPD spectra of the +3 charge state precursor ion (m/z 665.3544) of FFKNIVTPRT(H2PO4)PPPSQGK peptide. The precursor ions are indicated by a star (*) sign. Sequence coverage (%) presented in the bracket. Detailed assignments of the fragment ions are summarized in Supplementary Table S2
The abundance of fragment ions excluding phosphate losses in all three methods is compared in Figure 2b.The IRMPD, UVPD, and HiLoPD spectra provided excellent sequence coverage (94%–100%) and with a substantial number of fragment ions. Nearly equal numbers of b (27) and y (25) ions were detected in IRMPD (Figure 2b). In UVPD, a significant number of fragment ions were observed compared with IRMPD and HiLoPD.
With IRMPD, the losses of H3PO4/HPO3 groups were detected from b10-12, b15, and b16 ions as well as from y8-12, y14, and y16 ions (Table 2). However, bn (n = 10, 12–16) and yn (n = 8–14) fragment ions retained the phosphate groups. The overall phosphate retention efficiency achieved with IRMPD is 52%. In UVPD, significant numbers (42) of fragment ions such as an (n = 10–16), bn (n = 10–16), cn ( n = 13–16), xn (n = 8–16), yn (n = 8–16), and zn (n = 11–16) retain the phosphate groups (Table 2). However, phosphate losses occurred only from 29 fragment ions. The phosphate retention efficiency attained in UVPD was higher (59%) than that for IRMPD (52%) and HiLoPD (52%). For this phospho-threonine peptide, no phosphate loss was detected from x1-7/y1-7/z1-7 and a1-9/b1-9/c1-9 ions, which confirmed that the phosphate group is attached to the threonine at position 10 from the N-terminal.
IRMPD, UVPD, and HiLoPD on RDY(SO3)TGWLDF
The IRMPD, UVPD, and HiLoPD photodissociation spectra of the doubly protonated [M+2H]2+ (m/z 626.7492) of peptide RDY(SO3)TGWLDF are presented in Figure 4. The observed m/z and assignments of fragment ions of this peptide are summarized in Supplementary Table S3. The numbers of fragment ions (excluding sulfonate loss) detected by IRMPD, UVPD, and HiLoPD are summarized in Figure 2c. Singly and doubly protonated precursor ions provided nearly the same fragment ions in UVPD; however, the singly protonated ion was not stable and was difficult to isolate prior to MS/MS activation in IRMPD and HiLoPD methods. Similar events were also witnessed in a previous study [47]. In contrast, the doubly protonated ion of this peptide was easier to analyze with all three methods. Moreover, earlier studies reported that higher charge states can increase the sulfonate retention [39]. In all cases, the neutral loss of SO3 (79.9573 Da) was observed at m/z 1172.5388 from the [M+H]+ ions and at m/z 586.7710 from the precursor ion, respectively. Moreover, the sequential loss of SO3 and H2O was detected at m/z 577.7677 from the precursor ion.
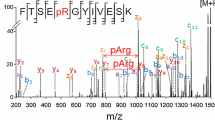
IRMPD, UVPD, and HiLoPD spectra of the +2 charge state precursor ion (m/z 626.7492) of RDY(SO3)TGWLDF peptide. The precursor ions are indicated by a star (*) sign. Sequence coverage (%) presented in the bracket. Detailed assignment of the fragment ions are summarized in Supplementary Table S3
The vibrational frequencies of C–O(SO3) and symmetrical O=S=O were in the range 9.4–10 μm [69], which is very close to the wavelength of the CO2 laser (10.6 μm). The IRMPD spectrum provided 87% sequence coverage and was dominated by series of y-ions, as well as minor contributions from b-ions (Figure 2c). Although more fragment ions were generated from the C-terminus, higher sequence coverage was observed for the N-terminal peptides (87.5%) compared with the C-terminal peptides (62.5%). Most of the y ions were formed close to the aspartic acid at position 2. Previous CID studies have shown that acidic residues near the C-terminus may promote the formation of y ions in sulfonated peptides [70]. Some y ions in IRMPD eliminated a molecule of water (18 Da) in secondary fragmentation. Interestingly, the loss of SO3 was seen predominantly from b3-8 ions but no such loss was detected from y ions (Table 3), probably because of the location of the PTM close to the N-terminal.
UVPD and HiLoPD of the doubly protonated precursor ion provided 100% sequence coverage with a/x, b/y, and c/z ions; 20 and 28 fragment ions (excluding SO3 loss) were detected in UVPD and HiLoPD, respectively. In addition to the typical fragment ions types, the UVPD and HiLoPD spectra also showed abundant ions corresponding to consecutive neutral losses of water and ammonia. The losses of 171.0099 and 129.0141 Da from the precursor ions corresponded to the removal of the related ion of tryptophan at m/z 541.2400 and the immonium ion of arginine at m/z 562.2380, respectively (Supplementary Table S3). Hydrogen-deficient and hydrogen-rich fragment ions were prevalent with UVPD and HiLoPD. The loss of SO3 from the backbone a/b/c fragment ions in UVPD and HiLoPD could be observed (Table 3). Only few a and c ions still containing the SO3 group were detected. As with IRMPD, the overall SO3 retention efficiency obtained by these two methods was very poor (12.5%).
In addition to sequence coverage and retention efficiency, the actual site of sulfation had to be pinpointed. Although the CID method has routinely been utilized to confirm the presence of sulfo groups detecting the neural loss of SO3 from the precursor ion, backbone fragment ions are required to confirm the position [47]. In IRMPD, UVPD, and HiLoPD, no SO3 loss was detected from a1 - 2/b1-2/c1-2, and this loss was noticed only from a3/b3/c3 and onward. In addition, no loss of SO3 was witnessed from x1-6/y1-6/z1-6 ions, and such losses began to occur only from y7 and x8/z8 ions, which confirmed the presence of the SO3 group on tyrosine at position 3 from the N-terminal.
IRMPD, UVPD, and HiLoPD on EAISPPDAAS (GalNAc) AAPLR
High throughput and residue-specific investigation of the O-glycosylation is challenging since the O-glycan core structure is very heterogeneous compared with that of N-glycan, and there is no straightforward protein sequence available for O-glycan [5, 71, 72]. In a given protein, O-glycan can be found with several serine/threonine residues. CID technique is routinely used for deducing glycan composition; however, determining the exact position of glycosylation and the peptide sequence is difficult to achieve. Although IRMPD provides quite similar fragment ions to CID, a previous study demonstrated that a low photon energy-based method can detect informative side chain losses from non-glycosylated serine and threonine residues, which indirectly implicates glycan attachment sites [71].
The IRMPD photodissociation spectrum obtained for the triply protonated [M+3H]3+ (m/z 556.9529) of peptide EAISPPDAAS(GalNAc)AAPLR is presented in Figure 5. Theoretical and observed m/z values of fragment ions of this peptide are summarized in Supplementary Table S4. The abundance of fragment ions excluding glycan losses in all three methods is compared in Figure 2d. The sequence coverage obtained by IRMPD is 86%. The neutral losses of GalNAc (221.0999 Da) and GalNAc-H2O (203.0899 Da) from the precursor ion were observed at m/z 483.2556 and 489.2591, respectively. For all the methods, the sequential losses of GalNAc and H2O were observed specifically from positions b10-b12. In IRMPD, nearly equal numbers of b and y ions could be seen (Figure 2d). The neutral losses of GalNAc and GalNAc-H2O from b10-12 ions are apparent whereas for yn ions these occurred from n = 7, 8, 10, and 12 (Table 4). No such neutral losses were observed for b1-9 and y1-5, which unambiguously confirmed that the GalNAc group is attached to serine at position 10 from the N-terminal. Some GalNAc groups were also preserved in y/b ions (Table 4). Overall, IRMPD showed 41.6% PTM retention efficiency, which was better than the sulfo-peptide.
IRMPD, UVPD, and HiLoPD spectra of the +3 charge state precursor ion (m/z 556.9529) of EAISPPDAAS(GalNAc)AAPLR peptide. The precursor ions are indicated by a star (*) sign. Sequence coverage (%) presented in the bracket. Detailed assignment of the fragment ions are summarized in Supplementary Table S4
The UVPD spectrum provided a wealth of fragment ions with sequence coverage of 86%. A significant number of a/x and b/y ions retaining the GalNAc group were detected. Compared with IRMPD (23 ions), the UVPD spectrum provided more fragment ions (63 ions) (Figure 2d). In addition, some c and z ions were also observed. Moreover, the neutral losses of GalNAc (221.0999Da) and GalNAc-H2O (203.0899 Da) groups were observed from all the series of ions (Supplementary Table S4). More losses were detected from y/b ions compared with other fragment ions. However, several a/x, b/y, and z fragment ions containing the PTM were detected (Table 4). The overall PTM retention efficiency was 46.6%.
In HiLoPD, a significant number of a/x, b/y, and c/z ions were identified with similar sequence coverage (86%) from the N- and C-terminals. The number of b ions detected in HiLoPD was higher than in UVPD and IRMPD. As in our previous studies, the number of a ions was lower in comparison to UVPD, possibly because the secondary fragmentation of these ions [60]. Interestingly, very few x ions were generated in HiLoPD or in UVPD. Compared with UVPD, the neutral losses of GalNAc and GalNAC-H2O groups were significantly reduced in HiLoPD. Such losses were only observed for b/y and c ions (Table 4). All a and z ions retained glycan groups. As with UVPD and IRMPD, neutral losses started to occur from positions 10 and 7 from the N- and C-terminals, respectively, which confirms the position of the glycan group (in serine at position 10 from the N-terminal) in this peptide. The retention efficiency of GalNAc in HiLoPD was 55.0%, which was the highest efficiency compared with IRMPD (41.6%) and UVPD (46.6%).
Conclusion
In this work, we reported the use of IRMPD, UVPD, and HiLoPD to characterize phospho-, sulfo-, and glyco-peptides in the gas phase. These results showed the proof-of-principal of 213 nm UVPD and HiLoPD methods for PTM characterization. Compared with a whole protein, the characterization of these PTM peptides requires lower CO2 laser power and fewer UV laser shots. Controlled and tunable parameters can improve the performance of these techniques. The IRMPD results demonstrated that sufficient backbone fragmentation and sequence coverage can be obtained. The IRMPD sequence coverage for phospho-tyrosine, phospho-threonine, sulfo-, and glyco-peptides was 75%, 94%, 87% and 86%, respectively. The exact location of the PTM groups in a peptide can be pinpointed. However, fragment-specific and overall PTM retention efficiency in IRMPD was somewhat reduced for all peptides. Compared with phospho- and glyco-peptides, the SO3 group was very prompt to dissociate in IRMPD, which may have been attributable to the low bond dissociation energy associated with the O–S bond or strong absorption of 10.64 μm IR photons by the SO3 group. This is the first study of 213 nm UVPD and HiLoPD used to characterize different PTM peptides. UVPD and HiLoPD gave excellent sequence coverage of 83%, 100%, 100%, and 86%, for phospho-tyrosine, phospho-threonine, sulfo-, and glyco-peptides. PTM retention efficiencies were better than in IRMPD (up to 59% for the phospho-peptides). Photodissociation at 213 nm UVPD and HiLoPD on peptide cations offers several promising benefits, including: (1) the production of more arrays of fragment ions with excellent sequence coverage; (2) the identification of the exact PTM position; (3) balanced PTM loss and retention events; and (4) no widespread side-chain losses. Our first set of results show that UVPD and HiLoPD prove to be promising methods for characterizing phospho- and glyco-proteomics.
References
Mann, M., Jensen, O.N.: Proteomic analysis of post-translational modifications. Nat. Biotechnol. 21, 255–261 (2003)
Zhao, Y., Jensen, O.N.: Modification-specific proteomics: strategies for characterization of post-translational modifications using enrichment techniques. Proteomics. 9, 4632–4641 (2009)
Macek, B., Mann, M., Olsen, J.: V: Global and site-specific quantitative phosphoproteomics: principles and applications. Annu. Rev. Pharmacol. Toxicol. 49, 199–221 (2009)
Önnerfjord, P., Heathfield, T.F., Heinegård, D.: Identification of tyrosine sulfation in extracellular leucine-rich repeat proteins using mass spectrometry. J. Biol. Chem. 279, 26–33 (2004)
Christiansen, M.N., Kolarich, D., Nevalainen, H., Packer, N.H., Jensen, P.H.: Challenges of determining o-glycopeptide heterogeneity: a fungal glucanase model system. Anal. Chem. 82, 3500–3509 (2010)
Cohen, P.: The origins of protein phosphorylation. Nat. Cell. Biol. 4, E127–E130 (2002)
Ubersax, J.A., Ferrell Jr., J.E.: Mechanisms of specificity in protein phosphorylation. Nat. Rev. Mol. Cell. Biol. 8, 530–541 (2007)
Pearce, L.R., Komander, D., Alessi, D.R.: The nuts and bolts of AGC protein kinases. Nat. Rev. Mol. Cell. Biol. 11, 9–22 (2010)
Bettelheim, F.R.: Tyrosine-O-sulfate in a peptide from fibrinogen. J. Am. Chem. Soc. 76, 2838–2839 (1954)
Moore, K.L.: Protein tyrosine sulfation: a critical posttranslation modification in plants and animals. Proc. Natl. Acad. Sci. 106, 14741–14742 (2009)
Yu, Y., Hoffhines, A.J., Moore, K.L., Leary, J.A.: Determination of the sites of tyrosine O-sulfation in peptides and proteins. Nat. Methods. 4, 583–588 (2007)
Steen, P. Van den, Rudd, P.M., Dwek, R.A., Opdenakker, G.: Concepts and principles of O-linked glycosylation. Crit. Rev. Biochem. Mol. Biol. 33, 151–208 (1998)
Gill, D.J., Clausen, H., Bard, F.: Location, location, location: new insights into O-GalNAc protein glycosylation. Trends Cell. Biol. 21, 149–158 (2017)
Kreisman, L.S.C., Cobb, B.A.: Infection, inflammation, and host carbohydrates: a glyco-evasion hypothesis. Glycobiology. 22, 1019–1030 (2012)
Pinho, S.S., Reis, C.A.: Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer. 15, 540–555 (2015)
Freeze, H.H., Eklund, E.A., Ng, B.G., Patterson, M.C.: Neurological aspects of human glycosylation disorders. Annu. Rev. Neurosci. 38, 105–125 (2015)
Schedin-Weiss, S., Winblad, B., Tjernberg, L.O.: The role of protein glycosylation in Alzheimer disease. FEBS J. 281, 46–62 (2014)
Hwang, H., Zhang, J., Chung, K.A., Leverenz, J.B., Zabetian, C.P., Peskind, E.R., Jankovic, J., Su, Z., Hancock, A.M., Pan, C., Montine, T.J., Pan, S., Nutt, J., Albin, R., Gearing, M., Beyer, R.P., Shi, M., Zhang, J.: Glycoproteomics in neurodegenerative diseases. Mass Spectrom. Rev. 29, 79–125 (2010)
Pan, S., Chen, R., Aebersold, R., Brentnall, T.A.: Mass spectrometry based glycoproteomics—from a proteomics perspective. Mol. Cell. Proteom. 10, (2011)
Masselon, C., Anderson, G.A., Harkewicz, R., Bruce, J.E., Pasa-Tolic, L., Smith, R.D.: Accurate mass multiplexed tandem mass spectrometry for high-throughput polypeptide identification from mixtures. Anal. Chem. 72, 1918–1924 (2000)
Pandey, A., Mann, M.: Proteomics to study genes and genomes. Nature. 405, 837–846 (2000)
Crowe, M.C., Brodbelt, J.S.: Infrared multiphoton dissociation (IRMPD) and collisionally activated dissociationof peptides in a quadrupole ion trapwith selective IRMPD of phosphopeptides. J. Am. Soc. Mass Spectrom. 15, 1581–1592 (2004)
Zolodz, M.D., Wood, K.V.: Detection of tyrosine phosphorylated peptides via skimmer collision-induced dissociation/ion trap mass spectrometry. J. Mass Spectrom. 38, 257–264 (2003)
Affolter, M., Watts, J.D., Krebs, D.L., Aebersold, R.: Evaluation of two-dimensional phosphopeptide maps by electrospray ionization mass spectrometry of recovered peptides. Anal. Biochem. 223, 74–81 (1994)
Cook, S.L., Jackson, G.P.: Metastable atom-activated dissociation mass spectrometry of phosphorylated and sulfonated peptides in negative ion mode. J. Am. Soc. Mass Spectrom. 22, 1088–1099 (2011)
Edelson-Averbukh, M., Shevchenko, A., Pipkorn, R., Lehmann, W.D.: Discrimination between peptide O-sulfo- and O-phosphotyrosine residues by negative ion mode electrospray tandem mass spectrometry. J. Am. Soc. Mass Spectrom. 22, 2256–2268 (2011)
Cotham, V.C., McGee, W.M., Brodbelt, J.S.: Modulation of phosphopeptide fragmentation via dual spray ion/ion reactions using a sulfonate-incorporating reagent. Anal. Chem. 88, 8158–8165 (2016)
Zubarev, R.A., Kelleher, N.L., McLafferty, F.W.: Electron capture dissociation of multiply charged protein cations. A nonergodic process. J. Am. Chem. Soc. 120, 3265–3266 (1998)
Syka, J.E.P., Coon, J.J., Schroeder, M.J., Shabanowitz, J., Hunt, D.F.: Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc. Natl. Acad. Sci. USA. 101, 9528–9533 (2004)
Zhurov, K.O., Fornelli, L., Wodrich, M.D., Laskay, U.A., Tsybin, Y.O.: Principles of electron capture and transfer dissociation mass spectrometry applied to peptide and protein structure analysis. Chem. Soc. Rev. 42, 5014–5030 (2013)
Coon, J.J.: Collision or electron? Protein sequence analysis on the 21st century. Anal. Chem. 81, 3208–3215 (2009)
Shi, S.D.H., Hemling, M.E., Carr, S.A., Horn, D.M., Lindh, I., McLafferty, F.W.: Phosphopeptide/phosphoprotein mapping by electron capture dissociation mass spectrometry. Anal. Chem. 73, 19–22 (2000)
Good, D.M., Wirtala, M., McAlister, G.C., Coon, J.J.: Performance characteristics of electron transfer dissociation mass spectrometry. Mol. Cell. Proteom. 6, 1942–1951 (2007)
Frese, C.K., Altelaar, A.F.M., Hennrich, M.L., Nolting, D., Zeller, M., Griep-Raming, J., Heck, A.J.R., Mohammed, S.: Improved peptide identification by targeted fragmentation using CID, HCD, and ETD on an LTQ-Orbitrap Velos. J. Proteome Res. 10, 2377–2388 (2011)
Asakawa, D., Takeuchi, T., Yamashita, A., Wada, Y.: Influence of metal–peptide complexation on fragmentation and inter-fragment hydrogen migration in electron transfer dissociation. J. Am. Soc. Mass Spectrom. 25, 1029–1039 (2014)
Asakawa, D., Osaka, I.: High-confidence sequencing of phosphopeptides by electron transfer dissociation mass spectrometry using dinuclear zinc(II) complex. Anal. Chem. 88, 12393–12402 (2016)
Loo, J.A., Loo, R.R.O., Light, K.J., Edmonds, C.G., Smith, R.D.: Multiply charged negative ions by electrospray ionization of polypeptides and proteins. Anal. Chem. 64, 81–88 (1992)
Huzarska, M., Ugalde, I., Kaplan, D.A., Hartmer, R., Easterling, M.L., Polfer, N.C.: Negative electron transfer dissociation of deprotonated phosphopeptide anions: choice of radical cation reagent and competition between electron and proton transfer. Anal. Chem. 82, 2873–2878 (2010)
Hersberger, K.E., Håkansson, K.: Characterization of O-sulfopeptides by negative ion mode tandem mass spectrometry: superior performance of negative ion electron capture dissociation. Anal. Chem. 84, 6370–6377 (2012)
Madsen, J.A., Xu, H., Robinson, M.R., Horton, A.P., Shaw, J.B., Giles, D.K., Kaoud, T.S., Dalby, K.N., Trent, M.S., Brodbelt, J.S.: High-throughput database search and large-scale negative polarity liquid chromatography-tandem mass spectrometry with ultraviolet photodissociation for complex proteomic samples. Mol. Cell. Proteom. 12, 2604–2614 (2013)
Halim, M., Girod, M., MacAleese, L., Lemoine, J., Antoine, R., Dugourd, P.: 213 nm ultraviolet photodissociation on peptide anions: radical-directed fragmentation patterns. J. Am. Soc. Mass Spctrom. 27, 474–486 (2016)
Riley, N.M., Rush, M.J.P., Rose, C.M., Richards, A.L., Kwiecien, N.W., Bailey, D.J., Hebert, A.S., Westphall, M.S., Coon, J.J.: The negative mode proteome with activated ion negative electron transfer dissociation (AI-NETD). Mol. Cell. Proteom. 14, 2644–2660 (2015)
Riley, N.M., Bern, M., Westphall, M.S., Coon, J.J.: Full-featured search algorithm for negative electron-transfer dissociation. J. Proteome Res. 15, 2768–2776 (2016)
Bowie, J.H., Brinkworth, C.S., Dua, S.: Collision-induced fragmentations of the (M-H)− parent anions of underivatized peptides: an aid to structure determination and some unusual negative ion cleavages. Mass Spectrom. Rev. 21, 87–107 (2002)
Shaw, J.B., Kaplan, D.A., Brodbelt, J.S.: Activated ion negative electron transfer dissociation of multiply charged peptide anions. Anal. Chem. 85, 4721–4728 (2013)
Kim, T.-Y., Reilly, J.P.: Time-resolved observation of product ions generated by 157 nm photodissociation of singly protonated phosphopeptides. J. Am. Soc. Mass Spectrom. 20, 2334–2341 (2009)
Robinson, M.R., Moore, K.L., Brodbelt, J.S.: Direct identification of tyrosine sulfation by using ultraviolet photodissociation mass spectrometry. J. Am. Soc. Mass Spectrom. 25, 1461–1471 (2014)
Madsen, J.A., Kaoud, T.S., Dalby, K.N., Brodbelt, J.S.: 193-nm Photodissociation of singly and multiply charged peptide anions for acidic proteome characterization. Proteomics. 11, 1329–1334 (2011)
Fort, K.L., Dyachenko, A., Potel, C.M., Corradini, E., Marino, F., Barendregt, A., Makarov, A.A., Scheltema, R.A., Heck, A.J.R.: Implementation of ultraviolet photodissociation on a benchtop Q Exactive mass spectrometer and its application to phosphoproteomics. Anal. Chem. 88, 2303–2310 (2016)
Robinson, M.R., Taliaferro, J.M., Dalby, K.N., Brodbelt, J.S.: 193 nm Ultraviolet photodissociation mass spectrometry for phosphopeptide characterization in the positive and negative ion modes. J. Proteome Res. 15, 2739–2748 (2016)
Lemoine, J., Tabarin, T., Antoine, R., Broyer, M., Dugourd, P.: UV photodissociation of phospho-seryl-containing peptides: Laser stabilization of the phospho-seryl bond with multistage mass spectrometry. Rapid Commun. Mass Spectrom. 20, 507–511 (2006)
Park, S., Ahn, W.-K., Lee, S., Han, S.Y., Rhee, B.K., Oh, H.: Bin: Ultraviolet photodissociation at 266 nm of phosphorylated peptide cations. Rapid Commun. Mass Spectrom. 23, 3609–3620 (2009)
Antoine, R., Joly, L., Tabarin, T., Broyer, M., Dugourd, P., Lemoine, J.: Photo-induced formation of radical anion peptides. Electron photodetachment dissociation experiments. Rapid Commun. Mass Spectrom. 21, 265–268 (2007)
Wilson, J.J., Brodbelt, J.S.: Infrared multiphoton dissociation for enhanced de novo sequence interpretation of N-terminal sulfonated peptides in a quadrupole ion trap. Anal. Chem. 78, 6855–6862 (2006)
Pikulski, M., Hargrove, A., Shabbir, S.H., Anslyn, E.V., Brodbelt, J.S.: Sequencing and characterization of oligosaccharides using infrared multiphoton dissociation and boronic acid derivatization in a quadrupole ion trap. J. Am. Soc. Mass Spectrom. 18, 2094–2106 (2007)
Li, B., Russell, S.C., Zhang, J., Hedrick, J.L., Lebrilla, C.B.: Structure determination by MALDI-IRMPD mass spectrometry and exoglycosidase digestions of O-linked oligosaccharides from Xenopus borealis egg jelly. Glycobiology. 21, 877–894 (2011)
Brodbelt, J.S., Wilson, J.J.: Infrared multiphoton dissociation in quadrupole ion traps. Mass Spectrom. Rev. 28, 390–424 (2009)
Zhou, W., Hakansson, K.: Structural characterization of carbohydrates by Fourier transform tandem mass spectrometry. Curr. Proteom. 8, 297–308 (2011)
Girod, M., Sanader, Z., Vojkovic, M., Antoine, R., MacAleese, L., Lemoine, J., Bonacic-Koutecky, V., Dugourd, P.: UV photodissociation of proline-containing peptide ions: insights from molecular dynamics. J. Am. Soc. Mass Spctrom. 26, 432–443 (2014)
Halim, M.A., Girod, M., MacAleese, L., Lemoine, J., Antoine, R., Dugourd, P.: Combined infrared multiphoton dissociation with ultraviolet photodissociation for ubiquitin characterization. J. Am. Soc. Mass Spectrom. 27, 1435–1442 (2016)
Flora, J.W., Muddiman, D.C.: Determination of the relative energies of activation for the dissociation of aromatic versus aliphatic phosphopeptides by ESI-FTICR-MS and IRMPD. J. Am. Soc. Mass Spectrom. 15, 121–127 (2004)
Han, H., Xia, Y., McLuckey, S.A.: Ion trap collisional activation of c and z• ions formed via gas-phase ion/ion electron-transfer dissociation. J. Proteome Res. 6, 3062–3069 (2007)
Antoine, R., Lemoine, J., Dugourd, P.: Electron photodetachment dissociation for structural characterization of synthetic and bio-polymer anions. Mass Spectrom. Rev. 33, 501–522 (2014)
Madsen, J., Cheng, R.R., Kaoud, T.S., Dalby, K., Makarov, D.E., Brodbelt, J.: Charge-site-dependent dissociation of hydrogen-rich radical peptide cations upon vacuum UV photoexcitation. Chem. Eur. J. 18, 5374–5383 (2012)
DeGnore, J., Qin, J.: Fragmentation of phosphopeptides in an ion trap mass spectrometer. J. Am. Soc. Mass Spectrom. 9, 1175–1188 (1998)
Tholey, A., Reed, J., Lehmann, W.D.: Electrospray tandem mass spectrometric studies of phosphopeptides and phosphopeptide analogues. J. Mass Spectrom. 34, 117–123 (1999)
Moyer, S.C., VonSeggern, C.E., Cotter, R.J.: Fragmentation of cationized phosphotyrosine containing peptides by atmospheric pressure MALDI/ion trap mass spectrometry. J. Am. Soc. Mass Spectrom. 14, 581–592 (2003)
Annan, R.S., Carr, S.A.: Phosphopeptide analysis by matrix-assisted laser desorption time-of-flight mass spectrometry. Anal. Chem. 68, 3413–3421 (1996)
Paciotti, R., Coletti, C., Re, N., Scuderi, D., Chiavarino, B., Fornarini, S., Crestoni, M.E.: Serine O-sulfation probed by IRMPD spectroscopy. Phys. Chem., Chem. Phys. 17, 25891–25904 (2015)
Budnik, B.A., Haselmann, K.F., Zubarev, R.A.: Electron detachment dissociation of peptide di-anions: an electron-hole recombination phenomenon. Chem. Phys. Lett. 342, 299–302 (2001)
Seipert, R.R., Dodds, E.D., Lebrilla, C.B.: Exploiting differential dissociation chemistries of O-linked glycopeptide ions for the localization of mucin-type protein glycosylation. J. Proteome Res. 8, 493–501 (2009)
Zhu, Z., Su, X., Clark, D.F., Go, E.P., Desaire, H.: Characterizing O-linked glycopeptides by electron transfer dissociation: fragmentation rules and applications in data analysis. Anal. Chem. 85, 8403–8411 (2013)
Acknowledgements
The research leading to these results has received funding from the European Research Council under the European Union’s Seventh Framework Program (FP7/2007-2013 grant agreement no. 320659).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 61 kb)
Rights and permissions
About this article
Cite this article
Halim, M.A., MacAleese, L., Lemoine, J. et al. Ultraviolet, Infrared, and High-Low Energy Photodissociation of Post-Translationally Modified Peptides. J. Am. Soc. Mass Spectrom. 29, 270–283 (2018). https://doi.org/10.1007/s13361-017-1794-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-017-1794-9