Abstract
Bombyx mori nucleopolyhedrosis virus (BmNPV) is a highly pathogenic virus to domestic silkworm Bombyx mori, often causing severe economic losses to silk industry. Interestingly, there are no documented reports of NPV infection in the wild silkworm Antheraea mylitta. Analysis of gene expression datasets and comparative genomic analysis of sequence datasets from B. mori and A. mylitta was undertaken to unravel the potential immune-related proteins and immune pathways involved. The B. mori silkworm races Sarupat and CSR-2 which are resistant and susceptible to BmNPV respectively were selected and challenged with virus to study BmNPV resistance related genes and their expression profile. The genes were filtered to isolate membrane proteins, their sequences were retrieved from UniProt and were compared against A. mylitta using BLASTp to search for similarity. Major proteins were putative defence proteins. Further, KEGG database was used to check for the presence of differentially regulated proteins in certain metabolic pathways. The analysis of the pathways was carried out using cytoscape software based on betweeness centrality and stress. The PI3K-Akt pathway was found to be the hub of all signals triggered during the course of NPV infection. We analyzed how the NPV infection modulates PI3K-Akt signaling by gene expression studies of the key regulator of the pathway i.e. Akt using real-time PCR in A. mylitta. A significant upregulation of Akt expression from 72 h post infection reaching its peak with a 2.30 fold change at 120 h pi clearly indicates an enhanced level of immune response in host towards the viral infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The silkworm, Bombyx mori is one of the economically important insect which is being reared for the last 5000 years for the production of raw silk. Sericulture industry has a potential in generating both direct and indirect employment to various stakeholders and provides sustainable income at regular interval with minimal capital investment with sufficient returns.
India is the second largest producer of silk in the world and also the largest consumer of silk in the world [1]. Sericulture faces biological challenges from pathogenic bacteria, fungi and viruses which causes loss of almost 20–30% of the potential cocoon production every year [2]. Among the various silkworm diseases viral diseases alone accounts for 80% of total cocoon loss of which B. mori nucleopolyhedrovirus (BmNPV) is the most serious one which calls for intensive research efforts towards this disease control. Till date, no effective treatment measures are available to combat BmNPV infection [3]. Also traditional strain breeding has been tried to improve BmNPV resistance in silkworms, but it takes considerable time of over years to achieve the same and also results in low cocoon production which is not economically feasible. Therefore, improving silkworm’s resistance to NPV by an effective and simple method is the need of hour.
The silkworm relies on different innate immune responses to act against invading pathogens. Some proteins from silkworm have shown various levels of anti BmNPV activity. The proteins purified from midgut juice of NPV resistant silkworms, such as serine protease [4], Bm-lipase [5] or a NADH- oxidoreductase like protein named BmNOX [6] have shown to impair the virulence of NPV, suggesting these protein’s antiviral activity. The Janus kinase/signal transducer and activator of transcription (JAK/STAT) signalling pathway is an evolutionarily conserved innate immune pathway in the insect immune response mechanism [7, 8]. After the infection of Autographa californica nucleopolyhedrovirus (AcMNPV) in Spodoptera frugiperda Sf9 cells, the key gene stat in the JAK/STAT signalling pathway is activated to mediate the immune response against AcMNPV [9].
Though the domestic silkworm B. mori and wild silkworm A. mylitta both belong to the order Lepidoptera and being sericigenous insects of economic importance they still differ in their susceptibilities to viral infections. The NPV infection and its crop loss is well established in B. mori whereas, there are no reports of NPV infection in A. mylitta [10]. The microarray datasets of the B. mori silkworm race CSR-2 and sarupat which are susceptible and resistant to BmNPV respectively were selected for this study [11]. To find out the BmNPV resistance related proteins/pathways involved in resistant silkworm, the differentially expressed membrane proteins in the midgut of sarupat and CSR-2 were considered as it is the first site of virus host interaction. The differential regulation of genes in resistant race after BmNPV infection would indicate their possible role in antiviral immune response.
In the present study in order to unravel the potential immune-related genes and pathways we naturally challenged A. mylitta with BmNPV to check for the validation of the data driven hypothesis which would prove a step ahead of the TiO2 nanoparticles induced immunity in silkworms. Immune signalling pathway analysis revealed that TiO2 nanoparticles inhibited the proliferation of silkworm BmNPV, while promoting the expression of BmAkt to improve immunity reporting the role of PI3K-Akt signalling pathway in B. mori in response to BmNPV infection [12]. Our results may provide some reliable mechanism associated with the interaction of virus and the host cell by using A. mylitta as the invertebrate insect model for the first time.
Materials and methods
In-silico studies
Analysis of gene expression datasets and comparative genomic analysis of sequence datasets from B. mori and A. mylitta to identify the key proteins/signalling pathway involved was carried out. Datasets which consisted of microarray data of two sets of experiments, SET 2A (sarupat resistant race) and SET 2B (CSR-2 susceptible race) of B. mori were downloaded which consisted of supplementary data in the form of excel sheets. It gave a list of all the upregulated and downregulated proteins along with their UniProt accession numbers, names, fold change, etc. The data was processed for the identification of significantly modulated in their expression specially belonging to membrane proteins i.e. receptors, transmembrane proteins, transporter proteins, exchange proteins, recognition proteins and co-transporters since the focus was on the host–pathogen interactions. The sequence of the filtered proteins was downloaded from Uniprot in FASTA format.
Comparison against Antheraea mylitta
Sequence alignment of the sequences was carried out using BLASTp server. All the protein sequences were compared against the genome of A. mylitta and the percentage similarity of the sequences was noted down.
KEGG database
The KEGG database was exploited to search for involvement of the proteins in certain pathways [13]. This was done by using the links provided on the UniProt database for the specific proteins [14]. The database was also used to create pathway network of the following pathways: mTOR signaling pathway, FOXO signaling pathway and Longevity regulating pathway.
Pathway reconstruction and cytoscape analysis
From the results obtained for the KEGG database analysis, three pathways were found whose analysis was carried out using cytoscape based on betweenness centrality and stress while treating the network as directed [15].
In-vivo studies: relative gene expression analysis of Akt gene
Fourth instar silkworm A. mylitta larvae were procured from Mudimiyal farm, Chevala dist., Telangana. Indoor rearing of the same on Terminalia arjuna branches at 25 ± 1 °C and 75 ± 2% relative humidity was carried out in our laboratory. Fifth instar 1st day larvae of the experimental zone (30 larvae) were fed with leaves containing BmNPV (4.5 × 108 polyhedra/ml) and the control zone ones (30 larvae) with leaves smeared with sterile water. All treated leaves were air dried before feeding. Silkworms in both control zone and experimental zone were dissected to isolate midgut once every 24 h, flash frozen in liquid nitrogen and stored at − 80 °C till further use.
PCR amplification and sequencing of novel gene (Akt)
As the Akt gene in A. mylitta was not reported before and also the unavailability of the sequence for further studies drove us to check for its presence in the genome using PCR. Primers were designed for Akt gene using the conserved region from the Clustal W alignment of the genes in different silkworm species. The PCR amplicon product was further sequenced. PCR amplification was carried out whose conditions were 94 °C for 5 min, 35 cycles of 94 °C for 5 s, 55 °C for 10 s, 72 °C for 10 s and 72 °C for 5 min. Primers used for the Akt gene are: 5′-GACATGGCACGGCTCGGCACCAGC-3′ (Forward) and 5′-CTTCAATATCTTCATAGCGTACA-3′ (Reverse).
Real time expression analysis of Akt gene
Primers were designed and synthesized for the housekeeping tubulin gene (5′-TCGTCGAGCCCTACAACTCT- 3′ (Forward) 5′-ACTCGGTGAGGTCCACATTC-3′ (Reverse) and for the Akt gene as above.
Total RNA was extracted from A. mylitta midgut samples using Trizol reagent (Invitrogen), reverse transcribed into cDNA using oligo dT primer (Chromous kit). PCR was standardized to obtain amplification for the gene. A Real-time run was taken on ABI Step-one Real Time PCR machine. The PCR conditions were 94 °C for 5 min, 40 cycles of 94 °C for 5 s, 55 °C for 10 s, 72 °C for 10 s and 72 °C for 5 min. Results were calculated using − ΔΔCt method.
Results and discussion
Genome wide expression profiles associated with BmNPV resistance and susceptibility
The microarray data of differentially regulated genes in response to BmNPV infection showed a total of 1324 genes of which 735 were upregulated and 589 were downregulated in the resistant Sarupat race. Whereas in case of susceptible CSR-2 race out of 4298 genes 2183 were upregulated and 2115 were downregulated.
The disparity of differentially regulated proteins seen between the two races of B. mori is evident as shown in Table 1. The number of upregulated receptor proteins in CSR-2 susceptible race was almost thrice than that found in the resistant Sarupat race. Similarly, there was a fivefold increase in the number of transmembrane proteins from resistant to susceptible race. Likewise, the number of downregulated receptors doubled in the susceptible CSR-2 race from the resistant Sarupat race and although there was no significant increase in the number of transmembrane proteins, the transporters showed a quintuple rise.
Comparison against Antheraea mylitta
The differentially regulated genes compared with A. mylitta genome showed the following ortholog proteins (Tables 2, 3, 4, 5) Since the availability of either the genomic sequence source or expressed sequence tag data is limited for A. mylitta, the differential gene expression analysis was done with resistant and susceptible strains of B. mori. However, the differentially regulated genes were searched for their presence in A. mylitta through reciprocal blast search which confirm the presence of orthologous proteins in A. mylitta, genome. The reciprocal blast resulted in identification of 47 orthologs out of 227 differentially expressed genes. For which the UNIPROT IDs were mined and their details are given in Tables 2 and 4 which consists of upregulated proteins found in two different races, resistant Sarupat and susceptible CSR-2 race of B. mori that have orthologous proteins present in A. mylitta. The table distinctly shows the reduction in number of upregulated proteins found in both the races from the result obtained upon plain filtering of the raw data. Out of 33 upregulated membrane proteins, only 11 had orthologs in A. mylitta in the resistant race whereas in the susceptible race, the number reduced to a mere 15 proteins from 82. Even though the number of upregulated proteins in both the races appears almost closer to each other, upon further examination it can be seen that with the exception of 3 proteins, the rest are completely different. This indicates that there is an upregulation of completely different set of proteins in the resistant race as compared to susceptible race.
Out of the three proteins that were similarly upregulated in both the races, accession numbers, Q9U5A8, B7FF34 and A0SY07, only Q9U5A8 which was Tyrosine protein kinase receptor was found to have a role in FOXO signaling pathway. The protein when checked against KEGG database showed that it was in fact involved in not just FOXO signaling pathway, but also in Longevity regulating pathway and mTOR signaling pathway. Tables 3 and 5 consist of downregulated proteins present in the resistant Sarupat race and susceptible CSR-2 race of B. mori. As with the upregulated proteins, it can be seen that the number of proteins present drastically decreased when compared to that of the filtered proteins. It shows that not all the proteins present in B. mori had orthologs present in A. mylitta. Out of 26 membrane proteins in the resistant race, only 5 showed similarity with A. mylitta whereas in the susceptible race, the number declined down from 81 membrane proteins to 17.
However, unlike the upregulated proteins, there was a significant difference in the number of downregulated proteins between the two races. The susceptible race had almost thrice the number of proteins as compared to the resistant race. Yet, at the same time, the set of proteins differentially regulated in the resistant variety was different than those in susceptible except for two further supporting the conclusion derived from the analysis of the upregulated proteins that proteins differentially regulated in the susceptible and resistant races during NPV infection are completely different. These results indicate that, it is not just a different set of proteins that is differentially regulated during the infection, there is a difference in the entire biological mechanism that takes place during infection.
Cytoscape analysis
Cytoscape analysis of pathways FOXO signaling, Longevity regulating, mTOR signaling, showed that PI3K played a major role connecting all the three pathways (Fig. 1), getting affected even due to a slight change, thereby affecting all three pathways, the receptor of which turned out to be Tyrosine protein Kinase receptor. This was also shown in one of the previous reports which determined the involvement of PI3K-Akt signaling pathway in the event of BmNPV infection [12].
Expression characteristics of key gene Akt in PI3K-Akt signaling pathway
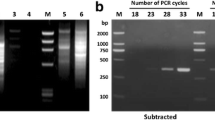
The resistance of silkworms against BmNPV is not only associated with resistance genes but also immune signalling pathways [12]. Gene Akt amplified from A. mylitta midgut (Fig. 2) was sequenced and deposited in NCBI with Gene accession no. MK263735. In this study, the transcription levels of a key gene in the PI3K-Akt pathway was measured. Time course analysis of gene expression of Akt gene by real-time PCR clearly shows an upregulation of Akt gene in BmNPV infected midgut samples. As shown (Fig. 3) the transcription levels of PI3K-Akt pathway marker gene Akt showed significant increase in Akt transcription level from 72 h pi with a fold change of 1.47 followed by an increase of 2.24 fold change at 96 hpi which reached a peak value thereafter with a fold change of 2.30 at 120 h pi indicating that the upregulation of Akt was induced by BmNPV infection aiding in increased host immune response. So, overall Akt gene expression was upregulated 0.47 times, 1.24 times and 1.30 times respectively at 72 h pi, 96 h pi and 120 h pi. Relative expression of Akt gene among the control and infected group at different time intervals was found to be statistically insignificant (P < 0.05) upon ANOVA analysis. These data suggest that activation of the PI3K-Akt pathway may be at least partially responsible for the antiviral response in A. mylitta upon BmNPV infection restricting a productive infection in a non-permissive host.
Immune response in insects is quite dynamic and different effector genes are likely expressed at variable time points during infection, contributing efficiently to the ability of the insects to ward off infections inspite of the absence of adaptive immunity. Wild silkworms have hardly been the subject of detailed scientific investigations delinquent largely to non-availability of molecular and genetic data on these species. As a first step in the present study we explored the differentially regulated membrane proteins during virus host negotiation in A. mylitta upon NPV infection by retrieving the gene expression data for susceptible and resistant variety of B. mori as complete genomic data for A. mylitta is not available. Doing this we also propose a new strategy for further comparative gene expression studies in other organisms whose genomic data is not available.
This report focuses on combining functional and subtractive genomic approach with special reference to the membrane proteins that were differentially regulated during BmNPV infection which has not been previously done according to the accessible information. Similarly, in spite of countless microarray assay reports being available for antiviral proteins, to this end no reports deal with the ability of membrane proteins to play an important role in BmNPV infection. All the accessed previous studies dealt with the situation that takes place upon the entry of the virus into the cell, like the differential regulation of the proteins, production of certain enzymes, etc. Shifting the focus from differential regulation of extracellular or intracellular proteins during a viral infection to membrane proteins is important as the first interaction of the virus takes place with that of the membrane proteins. Thus, any differential regulation in the membrane proteins determines the ability of the organism to fend off the virus since it is the site of virus host negotiation. Moreover, many membrane proteins play a significant role in various signaling mechanisms to communicate the external stimulus. Thus, any up or downregulation of membrane proteins is bound to affect the entire transcriptome and in turn the cellular behavior and response.
In silico analysis data has shown the involvement of several membrane proteins which do modulate differentially in a race dependent manner. Many proteins show conserved orthologs in A. mylitta further suggesting PI3K-Akt signaling pathway to be the hub of all the signals that is triggered during the NPV infection and can be exploited to understand the immunological consequences. On one hand, PI3K-Akt signalling is critical for many viruses to complete their life cycle, including cellular entry, replication and egress. On the other hand, the same PI3K-Akt pathway also induces antiviral immunity suggesting that PI3K-Akt pathway represents a two-faced player in interactions between the virus and cell [16]. However, the exact role (favourable or unfavourable for virus infection) of the PI3K-Akt pathway seems to be virus-specific. Time course analysis of gene expression of Akt by real-time PCR clearly shows an upregulation of Akt gene in infected midgut samples from 72 h onwards reaching a peak at 120 h pi. This study is the first one to reveal the activation of Akt in PI3K-Akt pathway that lead to improved resistance of wild silkworm A. mylitta to NPV infection. Furthermore, the observed bivalent action of the PI3K-Akt module represents a perfect example of a signalling pathway that is part of the antiviral response, however, is misused by the virus to support its own replication.
However, the biological reasons for NPV activating PI3K-Akt signalling pathway remain unclear, from both a virus and a host perspective which need further investigation. A better understanding of immune signalling pathways is required that could be activated by NPV infection to further delineate the cellular signalling pathways involved in baculovirus infection and finding novel therapeutic targets for nucleopolyhedrosis virus (NPV).
References
Bairoch A, Apweiler R, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ. The universal protein resource (UniProt). Nucleic Acids Res. 2005;33(suppl_1):D154–9. https://doi.org/10.1093/nar/gki070.
Dostert C, Jouanguy E, Irving P, Troxler L, Galiana-Arnoux D, Hetru C, Hoffmann JA, Imler JL. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of drosophila. Nat Immunol. 2005;6(9):946. https://doi.org/10.1038/ni1237.
Dunn EF, Connor JH. HijAkt: the PI3K/Akt pathway in virus replication and pathogenesis. In Progress in molecular biology and translational science, vol. 106, pp. 223–250. Academic Press; 2012. https://doi.org/10.1016/B978-0-12-396456-4.00002-X.
Jiang L, Cheng T, Zhao P, Yang Q, Wang G, Jin S, Lin P, Xiao Y, Xia Q. Resistance to BmNPV via overexpression of an exogenous gene controlled by an inducible promoter and enhancer in transgenic silkworm, Bombyx mori. PLoS ONE. 2012;7(8):e41838. https://doi.org/10.1371/journal.pone.0041838.
Jiang L, Zhao P, Cheng T, Sun Q, Peng Z, Dang Y, Wu X, Wang G, Jin S, Lin P, Xia Q. A transgenic animal with antiviral properties that might inhibit multiple stages of infection. Antivir Res. 2013;98(2):171–3. https://doi.org/10.1016/j.antiviral.2013.02.015.
Leclerc V, Reichhart JM. The immune response of Drosophila melanogaster. Immunol Rev. 2004;198(1):59–71. https://doi.org/10.1111/j.0105-2896.2004.0130.x.
Lekha G, Gupta T, Awasthi AK, Murthy GN, Trivedy K, Ponnuvel KM. Genome wide microarray based expression profiles associated with BmNPV resistance and susceptibility in Indian silkworm races of Bombyx mori. Genomics. 2015;106(6):393–403. https://doi.org/10.1371/journal.pone.0165865.
Nakazawa H, Tsuneishi E, Ponnuvel KM, Furukawa S, Asaoka A, Tanaka H, Ishibashi J, Yamakawa M. Antiviral activity of a serine protease from the digestive juice of Bombyx mori larvae against nucleopolyhedrovirus. Virology. 2004;321(1):154–62. https://doi.org/10.1016/j.virol.2003.12.011.
Narayanan S. The betweenness centrality of biological networks a study of betweenness centrality. Doctoral dissertation, PhD thesis, Virginia Polytechnic Institute. 2005. http://hdl.handle.net/10919/35405.
Ogata H, Goto S, Sato K, Fujibuchi W, Bono H, Kanehisa M. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 1999;27(1):29–34. https://doi.org/10.1093/nar/27.1.29.
Ponnuvel KM, Kangayam and Rao GP. Recent trends in seribiotechnology. In: Proceedings of UGC national symposium on modern biotechnology: prospects & challenges, Bengaluru, January; 2013. pp. 22–23.
Ponnuvel KM, Nakazawa H, Furukawa S, Asoka A, Ishibashi J, Tanaka H, Yamakawa M. A lipase isolated from the silkworm Bombyx mori shows antiviral activity against nucleopolyhedrovirus. J Virol. 2003;77(19):10725–9. https://doi.org/10.1016/j.virol.2003.12.011.
Selot R, Kumar V, Sekhar SC, Kumar PG. Molecular characterization and expression analysis of BmNOX in two strains of Bombyx mori with contrasting viral resistance phenotype. Arch Insect Biochem Physiol Publ Collab Entomol Soc Am. 2010;73(3):163–75. https://doi.org/10.1673/031.014.76.
Xu K, Li F, Ma L, Wang B, Zhang H, Ni M, Hong F, Shen W, Li B. Mechanism of enhanced Bombyx mori nucleopolyhedrovirus-resistance by titanium dioxide nanoparticles in silkworm. PLoS ONE. 2015;10(2):e0118222. https://doi.org/10.1371/journal.pone.0118222.
Yaseen S. Indian sericulture industry: its importance, problems and prospects. 2013. Acme Intellects International Journal of Research in Management. ISSN 2320 –2939 (Print) ISSN 2320-2793 (online) Let your Research be Global search—an ultimate search of truth-reforms through research, vol 2, no. 2; 2013.
Yeh MS, Cheng CH, Chou CM, Hsu YL, Chu CY, Chen GD, Chen ST, Chen GC, Huang C. Expression and characterization of two STAT isoforms from Sf9 cells. Dev Comp Immunol. 2008;32(7):814–24. https://doi.org/10.1016/j.dci.2007.12.001.
Acknowledgements
We thank University Grants Commission of National Fellowship for Higher Education with Award letter number: F1-17.1/2015-16/NFST-2015-17-ST-KAR-1674/(SA-III/Website) Dated: April, 2016 for awarding the fellowship enabling to carry out the present research work.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nallabothula, T., Avabhrath, N.K., Hulikal, S.K.S. et al. PI3K-Akt pathway mediated antiviral mechanism in silkworm Antheraea mylitta. VirusDis. 31, 349–356 (2020). https://doi.org/10.1007/s13337-020-00578-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13337-020-00578-y