Abstract
Background and Objectives
Previous studies reported that tapentadol-sulfate represented one of the major metabolites of tapentadol excreted in urine. The current study aimed to identify the human cytosolic sulfotransferases (SULTs) that is(are) capable of sulfating tapentadol and to examine whether human cells and human organ specimens are capable of sulfating tapentadol.
Methods
Thirteen human SULTs, previously expressed and purified, as well as human organ cytosols, were analyzed for tapentadol-sulfating activity using an established sulfotransferase assay. Cultured HepG2 human hepatoma cells and Caco-2 human colon carcinoma cells were labeled with [35S]sulfate in the presence of different concentrations of tapentadol.
Results
Three of the thirteen human SULTs, SULT1A1, SULT1A3, and SULT1C4, were found to display sulfating activity toward tapentadol. Kinetic analysis revealed that SULT1A3 displayed the highest catalytic efficiency in mediating the sulfation of tapentadol, followed by SULT1A1 and SULT1C4. Using cultured HepG2 and Caco-2 cells, the generation and release of sulfated tapentadol under metabolic conditions was demonstrated. Moreover, of the four human organ specimens (kidney, liver, lung, and small intestine) tested, the cytosols prepared from small intestine and liver showed significant tapentadol-sulfating capacity (at 0.0203 and 0.0054 nmol/min/mg, respectively).
Conclusion
Taken together, the results derived from the current study provided a molecular basis underlying the sulfation of tapentadol in humans.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Human SULT1A1, SULT1A3, and SULT1C4 were found to be capable of sulfating tapentadol. |
SULT1A3 showed the strongest sulfating efficiency with tapentadol. |
Sulfated tapentadol was shown to be produced by cultured HepG2 and Caco-2 cells and by human organ cytosols. |
1 Introduction
Tapentadol is an analgesic typically taken for the alleviation of moderate to severe chronic pain such as that associated with diabetic peripheral neuropathy [1]. It is a centrally acting opioid, exerting its analgesic activity by acting both as a μ-opioid receptor agonist and as a norepinephrine reuptake inhibitor [1, 2]. In regard to its pharmacokinetics, conjugation reactions have been shown to be the main pathways for the biotransformation of tapentadol, being responsible for the deactivation of about 70% of the oral dose [3]. Main metabolites have been demonstrated to be tapentadol-glucuronide and tapentadol-sulfate, with only about 3% of the drug eliminated unchanged by the kidneys. It is noted that tapentadol metabolites have been shown to be free of analgesic activity [3–6].
In vertebrates including humans, the cytosolic SULT-mediated sulfation plays a significant role in the metabolism of both endogenous compounds, such as dopamine (DA) and other catecholamines, as well as xenobiotics including a variety of drugs [7–9]. Sulfation is generally believed to result in the inactivation of substrate compounds [10, 11], and sulfated metabolites, due to the negatively charged sulfate group, tend to be more water-soluble and therefore are more easily excreted from the body [12]. There are, however, occasional exceptions. For example, morphine-6-sulfate and morphine-6-glucuronide have been reported to exhibit greater analgesic potency than the parent compound and appear to play an important role in the pharmacological effect of morphine [13–15]. All SULTs found in vertebrates constitute a gene superfamily. Within the SULT gene superfamily, members are categorized into SULT families, which are further divided into subfamilies [16, 17]. In humans, there are thirteen SULTs that are classified into four families; SULT1, SULT2, SULT4, and SULT6 [18]. SULT1 family, previously called the phenol sulfotransferase family, comprises eight members of which all have been shown to exhibit sulfating activity toward phenolic substrate compounds [17]. Of the three members of the SULT2 family, SULT2A1 has been described as a dehydroepiandrosterone SULT [19], whereas SULT2B1a and SULT2B1b have been shown to be capable of catalyzing the sulfation of pregnenolone and cholesterol, respectively [20, 21]. The sole members of the SULT4 and SULT6 families, on the other hand, have not been characterized in regard to their physiological substrates [22–24].
We report in this communication a detailed analysis of the tapentadol-sulfating activity of all thirteen known human SULTs. In addition, the pH-dependence and kinetic characteristics of the three SULTs (SULT1A3, SULT1A1, and SULT1C4) that displayed sulfating activity toward tapentadol were analyzed. Furthermore, tapentadol sulfation by cultured human cells and by human organ cytosols was investigated.
2 Materials and Methods
2.1 Materials
Tapentadol HCL was purchased from Cerilliant. Adenosine 5′-triphosphate (ATP), dithiothreitol, dimethyl sulfoxide, p-nitrophenol (pNP), DA, N-2-hydroxylpiperazine-N′-2-ethanesulfonic acid (HEPES), 2-morpholinoethanesulfonic acid (MES), 2-(cyclohexylamino)ethanesulfonic acid (CHES), 3-[N-tris-(hydroxymethyl)methylamino]-propanesulfonic acid (TAPS), 3-(cyclohexylamino)-1-propanesulfonic acid (CAPS), minimum essential medium (MEM), and silica gel thin-layer chromatography (TLC) plates, these plates are products of Sigma Chemical Company (St. Louis, MO, USA). Adenosine-3′-phosphate-5′-phospho[35S]sulfate (PAP[35S]) was prepared using ATP and carrier-free [35S]sulfate based on a previously established protocol [25]. Purified human SULTs, SULT1A1 (GenBank Accession # AAI10888.1), SULT1A2 (GenBank Accession # NP_001045.1), SULT1A3 (GenBank Accession # AAH78144.1), SULT1B1 (GenBank Accession # NP_055280.2), SULT1C2 (GenBank Accession # AAH05353.1), SULT1C3 (GenBank Accession # NP_001307807.1), SULT1C4 (GenBank Accession # AAI25044.1), SULT1E1 (GenBank Accession # EAX05597.1), SULT2A1 (GenBank Accession # AAH20755.1), SULT2B1a (GenBank Accession # AAC78498.1), SULT2B1b (GenBank Accession # AAC78499.1), SULT4A1 (GenBank Accession # CAG30474.1), and SULT6B1 (GenBank Accession # AAI40798.1), were prepared as previously described [26–28]. Cellulose TLC plates were from EMD Millipore. Ecolume scintillation cocktail was purchased from MP Biomedical. HepG2 human hepatoma cells (ATCC HB-8065) and Caco-2 human colon carcinoma cells (ATCC HTB-37) were obtained from American Type Culture Collection. Pooled human lung, liver, small intestine (duodenum and jejunum), and kidney cytosols were purchased from XenoTech, LLC. Other chemicals were of the highest brand commercially available.
2.2 Sulfotransferase Assay
The sulfating activity of purified recombinant human SULTs toward tapentadol was assayed based on a previously established protocol [29], with radioactive PAP[35S] as the sulfuryl donor. The assay mixture, following a 10 min reaction at 37 °C, was separated by TLC and analyzed for the [35S]sulfated tapentadol as previously described [29]. The same protocol was employed for the analysis of pH-dependence of the tapentadol-sulfating activity of SULT1A1, SULT1A3, and SULT1C4 using different buffers (sodium acetate at 4.5, 5.0, and 5.5; MES at 6.0, and 6.5; HEPES at 7.0, 7.5, and 8.0; TAPS at 8.5; CHES at 9.0, 9.5, and 10.0; and CAPS at 10.5, 11.0, and 11.5) in different assay mixtures. To determine the kinetic constants of the sulfation of tapentadol by SULT1A1, SULT1A3, and SULT1C4, the assays were performed using varying concentrations (10, 12.5, 16.67, 25, 50, and 100 µM) of tapentadol as substrates at their respective optimum pH, as well as at neutral pH and pH 7.4 (the pH inside the cell [30–34]). The activity data obtained were analyzed based on Michaelis–Menten kinetics using nonlinear regression (GraphPad Prism). To assay for tapentadol-sulfating activity of human organ cytosols, the assay mixture was supplemented with 50 mM NaF (a phosphatase inhibitor that prevents PAPS degradation by the phosphatases that may be present in the human organ cytosols [35]), and the reaction time was 30 min.
2.3 Examination of the Sulfation of Tapentadol by Cultured HepG2 and Caco-2 Cells
A previously established metabolic labeling procedure was employed to examine the metabolism of tapentadol by sulfation in HepG2 cells and Caco-2 cells. The cells were labeled in sulfate-free Minimum Essential Media (MEM) containing [35S]sulfate and varying concentrations (1, 2, 4, 10, 20, 40, and 100 µM) of tapentadol. The labeling media, collected following an 18-h incubation, were collected and analyzed by TLC as previously described [29].
3 Results
3.1 Differential Sulfating Activities of the Human SULTs Toward Tapentadol
In an initial study, all thirteen known human SULTs previously prepared were evaluated for sulfating activity with tapentadol as a substrate. Activity data compiled in Table 1 indicated that three (SULT1A1, SULT1A3, and SULT1C4) of the thirteen human SULTs (SULT1A1, SULT1A2, SULT1A3, SULT1B1, SULT1C2, SULT1C3, SULT1C4, SULT1E1, SULT2A1, SULT2B1a, SULT2B1b, SULT4A1, and SULT6B1) displayed significant sulfating activities toward tapentadol. Of the three, SULT1A3 showed the strongest tapentadol-sulfating activity (2.74 nmol/min/mg enzyme) at neutral pH, while SULT1A1 and SULT1C4 displayed considerably lower activities (0.88 and 0.66 nmol/min/mg enzyme, respectively). At pH 7.4, the presumable pH inside the cell [30–34], the sulfating activity of SULT1A3 toward tapentadol (4.32 nmol/min/mg enzyme) was also much higher than those of SULT1A1 and SULT1C4 (1.71 and 0.79 nmol/min/mg enzyme, respectively). Whereas at their respective optimal pH, SULT1A1 and SULT1A3 showed comparable and much higher tapentadol-sulfating activities (22.87 and 21.92 nmol/min/mg enzyme, respectively) than did SULT1C4 (at 2.37 nmol/min/mg enzyme).
3.2 pH-dependence of the Sulfation of Tapentadol by Human SULT1A1, SULT1A3, and SULT1C4
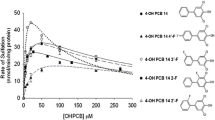
In an effort to characterize further the sulfation of tapentadol by SULT1A1, SULT1A3, and SULT1C4, a pH-dependence experiment was performed. As shown in Fig. 1, SULT1A1, SULT1A3, and SULT1C4 displayed optimal tapentadol-sulfating activity at, respectively, pH 10.5, 9.0, and 9.5.
pH-dependence of tapentadol-sulfating activity of the human SULT1A1 (a), SULT1A3 (b), and SULT1C4 (c). Enzymatic assays were carried out under standard assay conditions as described in Sect. 2 using different buffer systems as indicated. Data shown represent calculated mean ± standard deviation derived from three independent (separate) experiments using the same batch of enzymes. Symbols used are closed diamond for the acetate buffer at pH 4.5–5.5; closed triangle for MES buffer at pH 6.0 and 6.5; closed circle for HEPES buffer at pH 7.0–8.0; open diamond for TAPS buffer at pH 8.5; open circle for CHES buffer at pH 9.0–10.0; and open triangle for CAPS buffer at pH 10.5–11.5. MES 2-morpholinoethanesulfonic acid, HEPES N-2-hydroxylpiperazine-N′-2-ethanesulfonic acid, TAPAS 3-[N-tris-(hydroxymethyl)methylamino]-propane-sulfonic acid, CHES 2-(cyclohexylamino)ethanesulfonic acid, CAPS 3-(cyclohexylamino)-1-propanesul-fonic acid, SULT sulfotransferase
3.3 Kinetics of the Sulfation of Tapentadol by Human SULT1A1, SULT1A3, and SULT1C4
The kinetics of the sulfation of tapentadol by human SULT1A1, SULT1A3, and SULT1C4 were analyzed using varying concentrations of tapentadol as substrate at their respective optimum pH (10.5 for SULT1A1, 9.0 for SULT1A3, and 9.5 for SULT1C4), as well as neutral pH (7.0) and pH 7.4 (the presumable pH inside the cell) [30–34]. Kinetic constants calculated based on the results obtained are compiled in Table 2. Of the three SULT enzymes, SULT1C4 showed the lowest Michaelis constant (k m) (20.20 μM), followed by SULT1A1 (38.17 μM) and SULT1A3 (49.93 μM), at their respective optimal pH. In regard to maximum velocity (V max), SULT1A3 showed the highest values at all three pHs tested.
3.4 Metabolic Sulfation of Tapentadol in Cultured Cells and Sulfation of Tapentadol by Human Organ Specimens
As shown in Fig. 2, upon incubation in sulfate-free media containing [35S]sulfate and different concentrations of tapentadol, both HepG2 cells and Caco-2 cells were capable of producing [35S]sulfated tapentadol and releasing it into the labeling media in a concentration-dependent manner. Furthermore, as shown in Table 3, the cytosols prepared from human small intestine and liver were capable of mediating the sulfation of tapentadol (at 0.0203 and 0.0054 nmol/min/mg, respectively); whereas the cytosols of human lung and kidney showed no detectable activity toward tapentadol.
Production and release of [35S]sulfated tapentadol by HepG2 human hepatoma cells and Caco-2 human colon carcinoma cells labeled with [35S]sulfate in the presence of tapentadol. The figure shows the autoradiographs taken from the plates at the end of the thin-layer chromatography (TLC) analysis. Confluent HepG2 or Caco-2 cells were incubated in labeling media containing, respectively, 0, 1, 2, 4, 10, 20, 40, and 100 µM (corresponding to lanes 1–7) of tapentadol for 18 h. C refers to the control labeling medium without added drug compounds. E refers to [35S]sulfated tapentadol enzymatically synthesized using human SULT1A3. The arrow indicates the radioactive spot corresponding to [35S]sulfated tapentadol. SULT sulfotransferase
4 Discussion
To better understand the efficacy and the underlying basis for the adverse effects of tapentadol, it is necessary to find out more about its pharmacokinetics. As mentioned earlier, a significant portion of the oral dose of tapentadol was found to be directly metabolized through sulfate conjugation [3]. The current study aimed to first identify the SULT enzyme(s) that is(are) responsible for the sulfate conjugation of tapentadol. An initial survey revealed that three of the thirteen known human SULTs, SULT1A1, SULT1A3, and SULT1C4, could catalyze the sulfation of tapentadol. Previous studies have demonstrated the expression of SULT1A3 at high levels in the small intestine and platelets and at low levels in the liver [36, 37]. In contrast, SULT1A1 was found to be mainly expressed in the liver [38, 39], whereas SULT1C4 was found to be expressed at high levels in fetal lung, kidney and at low levels in fetal heart, adult kidney, ovary, and spinal cord [26]. The expression of SULT1A3 and SULT1A1 in small intestine and liver implicates the possibility of the first-pass metabolism of tapentadol, and may explain, in part at least, the relatively low oral bioavailability (32%) of this drug [2, 4]. It is noted that other drugs such as isoprenaline had previously been reported to undergo extensive first-pass metabolism by SULT-mediated sulfation upon oral administration [40].
An important factor that may affect the bioavailability of tapentadol is the developmental expression of tapentadol-sulfating SULTs [41, 42]. Many drug-metabolizing enzymes are known to be poorly expressed during the fetal period [43, 44], whereas SULT enzymes have been reported to be exceptions [45]. Both SULT1A1 and SULT1A3 have been reported to be expressed even at fetal stages [42]. At the beginning of postnatal life, there appears to be an elevation in the expression of SULT1A1 in the liver and a concomitant decrease in SULT1A3 expression in the liver and other tissues [41, 46]. On the other hand, SULT1C4 has been shown to be expressed at higher levels in fetuses than in adults [26]. These dynamic changes of tapentadol-sulfating SULTs during fetal development, through neonatal/child stages, onto adulthood may impact on the bioavailability and pharmacokinetics when the drug is used during pregnancy or for different age groups of patients.
pH-dependence analysis of the sulfation of tapentadol by human SULT1A3, SULT1A1, and SULT1C4 (cf. Fig. 1) revealed that the pH optima of the tested SULTs were all basic and close to the pKa values of the two ionizable groups of tapentadol (9.34 for the –N(CH3)2 group and 10.45 for the −OH group [4, 47]). It is possible that the protonated or deprotonated state of these functional groups may affect the interaction between the tapentadol molecule and corresponding amino acid residues of each of the three SULT enzymes in regard to substrate binding and/or catalysis and, therefore, their sulfating activity under different pH environments. Kinetic analysis of the sulfation of tapentadol by SULT1A1, SULT1A3, and SULT1C4 (cf. Table 2) revealed that the catalytic efficiency, as reflected by V max/K m, for the three enzymes is in the order SULT1A3 (0.63 ml/min/mg) > SULT1A1 (0.47 ml/min/mg) > SULT1C4 (0.08 ml/min/mg) at their optimal pH. Similar trend was also found for the catalytic efficiency determined for the three SULTs at neutral pH or pH 7.4. It is noted that a previous study employing 14C-labeled tapentadol indicated that the maximum plasma concentration in tested human subjects reached was 2.45 µg/ml [6], an equivalent of 11.1 µM. While this plasma concentration is considerably lower than the K m values determined for each of the three tapentadol-sulfating SULTs as mentioned above, it is possible that the tapentadol may be more highly concentrated in cells that contain SULT1A1, SULT1A3, and/or SULT1C4 and, therefore, allow for their metabolism through sulfation. As described earlier, studies using animal models and human subjects had revealed the production and urinary excretion of sulfated tapentadol [3, 48].
To gather evidence that sulfation of tapentadol may occur in cells of human organs, we first performed a metabolic labeling experiment using cultured cells. Two cell lines, HepG2 and Caco-2, were employed. Previous studies have demonstrated that HepG2 cells express a number of SULTs including SULT1A subfamily (SULT1A1, SULT1A2, SULT1A3), SULT1E1, and SULT2A1, while Caco-2 cells express SULT1A1, SULT1A2, SULT1A3, SULT1B1, SULT1C2, SULT1C4, and SULT2A1 [49–51]. Both these two cell lines therefore are equipped with enzymes that are capable of sulfating tapentadol. That these liver- or intestine-derived cells were capable of metabolizing tapentadol through sulfation may underscore the first-pass metabolism of tapentadol upon oral administration. To obtain further evidence for the sulfation of tapentadol in human organs, cytosols prepared from four human organs (small intestine, liver, lung, and kidney) were tested for tapentadol-sulfating activity. Of the four cytosols tested, only cytosols prepared from liver and small intestine showed tapentadol-sulfating activity. Notably, the tapentadol-sulfating activity detected for the intestine cytosol was nearly four times that detected for the liver cytosol. These results are in line with the much higher level of expression of SULT1A3 in intestine than in liver [36, 37, 52]. Both intestine and liver have been reported to express considerable levels of SULT1A1, another tapentadol-sulfating SULT [52].
5 Conclusions
The current study revealed that of the thirteen known human SULTs, SULT1A1, SULT1A3, and SULT1C4 were capable of sulfating tapentadol. Of the three, SULT1A3 showed considerably higher catalytic efficiency in catalyzing the sulfation of tapentadol, suggesting that SULT1A3 is likely the major enzyme responsible for tapentadol sulfation in human body, followed by SULT1A1 and SULT1C4. That both cultured HepG2 and Caco-2 cells, as well as small intestine and liver cytosols, were capable of mediating the sulfation of tapentadol implied that small intestine and liver may represent major human organs responsible for the metabolism of tapentadol through sulfation. Collectively, these results contribute to a better understanding about the molecular basis underlying the pharmacokinetics of tapentadol. While it remains to be clarified, the differential expression of tapentadol-sulfating SULT1A1, SULT1A3, and SULT1C4 during fetal, neonatal, and child development onto adulthood may dictate the differential metabolism of tapentadol administered during pregnancy or at different stages during neonatal/child development and in adults.
References
Kress HG. Tapentadol and its two mechanisms of action: is there a new pharmacological class of centrally-acting analgesics on the horizon? Eur J Pain. 2010;14:781–3.
Tzschentke TM, Christoph T, Kögel B, Schiene K, Hennies HH, Englberger W, Haurand M, Jahnel U, Cremers TI, Friderichs E, De Vry J. (−)-(1R,2R)-3-(3-dimethylamino-1-ethyl-2-methyl-propyl)-phenol hydrochloride (tapentadol HCl): a novel mu-opioid receptor agonist/norepinephrine reuptake inhibitor with broadspectrum analgesic properties. J Pharmacol Exp Ther. 2007;323:265–76.
Coulter C, Taruc M, Tuyay J, Moore C. Determination of tapentadol and its metabolite N-desmethyltapentadol in urine and oral fluid using liquid chromatography with tandem mass spectral detection. J Anal Toxicol. 2007;34:458–63.
Nucynta® ER (tapentadol) extended-release oral tablets C-II [package insert]. Janssen Pharmaceuticals, Inc. NJ: Raritan; 2014.
Terlinden R, Kogel BY, Englberger W, Tzschentke TM. In vitro and in vivo characterization of tapentadol metabolites. Methods Find Exp Clin Pharmacol. 2012;32:31–8.
Terlinden R, Ossig J, Fliegert F, Lange C, Göhler K. Absorption, metabolism, and excretion of 14C-labeled tapentadol HCl in healthy male subjects. Eur J Drug Metab Pharmacokinet. 2007;32:163–9.
Kurogi K, Chepak A, Hanrahan MT, Liu M-Y, Sakakibara Y, Suiko M, Liu M-C. Sulfation of opioid drugs by human cytosolic sulfotransferases: metabolic labeling study and enzymatic analysis. Eur J Pharm Sci. 2014;62:40–8.
Ko K, Kurogi K, Davidson G, Liu M-Y, Sakakibara Y, Suiko M, Liu M-C. Sulfation of ractopamine and salbutamol by the human cytosolic sulfotransferases. J Biochem. 2012;152:275–83.
Clements J, Critchley JA, Prescott LF. The role of sulphate conjugation in the metabolism and disposition of oral and intravenous paracetamol in man. Br J Clin Pharmacol. 1984;18:481–5.
Falany CN, Roth JA. Properties of human cytosolic sulfotransferases involved in the drug metabolism. In: Jeffery HE, editor. Human drug metabolism. From molecular biology to man. Boca Raton: CRC Press; 1993. p. 101–15.
Weinshilboum RM, Otterness DM. Sulfotransferase enzymes in conjugation–deconjugation reactions. In: Kaufman FC, editor. Drug metabolism and toxicology. Berlin: Springer; 1994. p. 45–78.
Falany CN, Ström P, Swedmark S. Sulphation of O-desmethyl naproxen and related compounds by human cytosolic sulfotransferases. Br J Clin Pharmacol. 2005;60:632–40.
Zuckerman A, Bolan E, de Paulis T, Schmidt D, Spector S, Pasternak GW. Pharmacological characterization of morphine-6-sulfate and codeine-6-sulfate. Brain Res. 1999;842:1–5.
Houdi AA, Kottayil S, Crooks PA, Butterfield DA. 3-0-Acetylmorphine-6-O-sulfate: a potent, centrally acting morphine derivative. Pharmacol Biochem Behav. 1996;53:665–71.
Pasternak GW, Pan YX. Mu opioids and their receptors: evolution of a concept. Pharmacol Rev. 2013;65:1257–317.
Glatt H. Sulfotransferases in the bioactivation of xenobiotics. Chem Biol Interact. 2000;129:141–70.
Blanchard RL, Freimuth RR, Buck J, Weinshilboum RM, Coughtrie MW. A proposed nomenclature system for the cytosolic sulfotransferase (SULT) superfamily. Pharmacogenetics. 2004;14:199–211.
Hui Y, Liu MC. Sulfation of ritodrine by the human cytosolic sulfotransferases (SULTs): effects of SULT1A3 genetic polymorphism. Eur J Pharmacol. 2015;761:125–9.
Cook IT, Leyh TS, Kadlubar SA, Falany CN. Structural rearrangement of SULT2A1: effects on dehydroepiandrosterone and raloxifene sulfation. Horm Mol Biol Clin Investig. 2010;1:81–7.
Javitt NB, Lee YC, Shimizu C, Fuda H, Strott CA. Cholesterol and hydroxycholesterol sulfotransferases: identification, distinction from dehydroepiandrosterone sulfotransferase, and differential tissue expression. Endocrinology. 2001;142:2978–84.
Falany CN, He D, Dumas N, Frost AR, Falany JL. Human cytosolic sulfotransferase 2b1: isoform expression, tissue specificity and subcellular localization. J Steroid Biochem Mol Biol. 2006;102:214–21.
Falany CN, Xie X, Wang J, Ferrer J, Falany JL. Molecular cloning and expression of novel sulphotransferase-like cDNAs from human and rat brain. Biochem J. 2000;346:857–64.
Lindsay J, Wang LL, Li Y, Zhou SF. Structure, function and polymorphism of human cytosolic sulfotransferases. Curr Drug Metab. 2008;9:99–105.
Runge-Morris M, Kocarek TA, Falany CN. Regulation of the cytosolic sulfotransferases by nuclear receptors. Drug Metab Rev. 2013;45:15–33.
Yanagisawa K, Sakakibara Y, Suiko M, Takami Y, Nakayama T, Nakajima H, Takayanagi K, Natori Y, Liu M-C. cDNA cloning, expression, and characterization of the human bifunctional ATP sulfurylase/adenosine 5′-phosphosulfate kinase enzyme. Biosci Biotechnol Biochem. 1998;62:1037–40.
Sakakibara Y, Yanagisawa K, Katafuchi J, Ringer DP, Takami Y, Nakayama T, Suiko M, Liu M-C. Molecular cloning, expression, and characterization of novel human SULT1C sulfotransferases that catalyze the sulfonation of N-hydroxy-2-acetylaminofluorene. J Biol Chem. 1998;273:33929–35.
Sakakibara Y, Takami Y, Nakayama T, Suiko M, Liu M-C. Localization and functional analysis of the substrate specificity/catalytic domains of human M-form and P-form phenol sulfotransferases. J Biol Chem. 1998;273:6242–7.
Suiko M, Sakakibara Y, Liu M-C. Sulfation of environmental estrogen-like chemicals by human cytosolic sulfotransferases. Biochem Biophys Res Commun. 2000;267:80–4.
Liu MC, Lipmann F. Decrease of tyrosine-O-sulfate-containing proteins found in rat fibroblasts infected with Rous sarcoma virus or Fujinami sarcoma virus. Proc Natl Acad Sci USA. 1984;81:3695–8.
Roos A, Boron WF. Intracellular pH. Physiol Rev. 1981;61:296–434.
Bright GR, Fisher GW, Rogowska J, Taylor DL. Fluorescence ratio imaging microscopy: temporal and spatial measurements of cytoplasmic pH. J Cell Biol. 1987;104:1019–33.
Madshus IH. Regulation of intracellular pH in eukaryotic cells. Biochem J. 1988;250:1–8.
Gerweck LE, Seetharaman K. Cellular pH gradient in tumor versus normal tissue: potential exploitation for the treatment of cancer. Cancer Res. 1996;56:1194–8.
Boron WF. Regulation of intracellular pH. Adv Physiol Educ. 2004;28:160–79.
Rens-Domiano S, Roth JA. Characterization of tyrosylprotein sulfotransferase from rat liver and other tissues. J Biol Chem. 1989;264:899–905.
Weinshilboum RM. Phenol sulfotransferase in humans: properties, regulation, and function. Fed Proc. 1986;45:2223–8.
Ganguly TC, Krasnykh V, Falany CN. Bacterial expression and kinetic characterization of the human monoamine-sulfating form of phenol sulfotransferase. Drug Metab Dispos. 1995;23:945–50.
Butler PR, Anderson RJ, Venton DL. Human platelet phenol sulfotransferase: partial purification and detection of two forms of the enzyme. J Neurochem. 1983;41:630–9.
Stanley EL, Hume R, Visser TJ, Coughtrie MWH. Differential expression of sulfotransferase enzymes involved in thyroid hormone metabolism during human placental development. J Clin Endocrinol Metab. 2001;86:5944–55.
Conolly E, Davies DS, Dollery CT, Morgan CD, Paterson JW, Sandler M. Metabolism of isoprenaline in dog and man. Br J Pharmacol. 1972;46:458–72.
Richard K, Hume R, Kaptein E, Stanley EL, Visser TJ, Coughtrie MW. Sulfation of thyroid hormone and dopamine during human development: ontogeny of phenol sulfotransferases and arylsulfatase in liver, lung and brain. J Clin Endocrinol Metab. 2001;86:2734–42.
Stanley EL, Hume R, Coughtrie MW. Expression profiling of human fetal cytosolic sulfotransferases involved in steroid and thyroid hormone metabolism and in detoxification. Mol Cell Endocrinol. 2005;240:32–42.
McCarver DG, Hines RN. The ontogeny of human drug-metabolizing enzymes: phase II conjugation enzymes and regulatory mechanisms. J Pharmacol Exp Ther. 2002;300:361–6.
Coughtrie MWH, Burchell B, Leaky JEA, Hume R. The inadequacy of perinatal glucuronidation-immunoblot analysis of the developmental expression of individual UDP-glucuronosyltransferase isoenzymes in rat and human-liver microsomes. Mol Pharmacol. 1988;34:729–35.
Coughtrie MWH. Sulfation through the looking glass-recent advances in sulfotransferase research for the curious. Pharmacogenomics J. 2002;2:297–308.
Vietri M, Pietrabissa A, Mosca F, Rane A, Pacific GM. Human adult and foetal liver suphotransferases: inhibition by mefenamic acid and salicylic acid. Xenobiotica. 2001;31:153–61.
Schneider J, Jahnel U, Linz K. Neutral effects of the novel analgesic tapentadol on cardiac repolarization due to mixed ion channel inhibitory activities. Drug Dev Res. 2010;71:197–208.
Wade WE, Spruill WJ. Tapentadol hydrochloride: a centrally acting oral analgesic. Clin Ther. 2009;31:2804–18.
Teubner W, Meinl W, Florian S, Kretzschmar M, Glatt H. Identification and localization of soluble sulfotransferases in the human gastrointestinal tract. Biochem J. 2007;404:207–15.
Westerink WM, Schoonen WG. Phase II enzyme levels in HepG2 cells and cryopreserved primary human hepatocytes and their induction in HepG2 cells. Toxicol In Vitro. 2007;21:1592–602.
Meinl W, Ebert B, Glatt H, Lampen A. Sulfotransferase forms expressed in human intestinal Caco-2 and TC7 cells at varying stages of differentiation and role in benzo[a]pyrene metabolism. Drug Metab Dispos. 2008;36:276–83.
Riches Z, Stanley EL, Bloomer JC, Coughtrie MW. Quantitative evaluation of the expression and activity of five major sulfotransferases (SULTs) in human tissues: the SULT “pie”. Drug Metab Dispos. 2009;37:2255–61.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was supported in part by a startup fund from the College of Pharmacy and Pharmaceutical Sciences, the University of Toledo.
Conflict of interest
Ahsan F. Bairam, Mohammed I. Rasool, Katsuhisa Kurogi, and Ming-Cheh Liu declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Bairam, A.F., Rasool, M.I., Kurogi, K. et al. On the Molecular Basis Underlying the Metabolism of Tapentadol Through Sulfation. Eur J Drug Metab Pharmacokinet 42, 793–800 (2017). https://doi.org/10.1007/s13318-016-0392-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13318-016-0392-8