Abstract
The oomycete pathogen causing root-rot and damping-off diseases in soybean (Glycine max) fields in the Huang-Huai region of China was identified as Pythium spinosum. To detect P. spinosum for disease diagnosis and control, we developed a loop-mediated isothermal amplification (LAMP) reaction with a primer set designed from the rDNA internal transcribed spacer 2 (ITS2) sequence of P. spinosum. The LAMP assay can efficiently amplify the target gene within 60 min at 63 °C. In specificity tests using 31 Pythium spp., 12 Phytophthora spp., 6 Phytopythium spp., and 9 other fungi strains, no cross-reactions were observed in the LAMP assay. The detection limit was 100 pg·μL−1 of genomic DNA per reaction. In cases of suspected disease, P. spinosum could be detected directly, using the LAMP assay, from soybean tissues and soil collected from fields in soybean production areas in the Huang-Huai region. This study provides a rapid method for diagnosing soybean root-rot and damping-off diseases caused by P. spinosum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There are more than 160 species in the genus Pythium (Chen et al. 2017), which includes plant pathogens found worldwide that frequently cause seed, seedling, and root rot in economically important crops such as peanut (Arachis hypogaea), wheat (Triticum spp.), and soybean (Glycine max) (Wang et al. 2003; Wrather and Koenning 2006). Most Pythium spp. are soil-borne, facultative plant pathogens with broad host ranges (Hendrix and Campbell 1973; Martin and Loper 2010).
Pythium spinosum infects several host species and occurs in both soil and water. P. spinosum often infects the seeds and young roots of plants (Botha 1993; Plaats-Niterink 1981), causing the plants to wilt, and also resulting in plant dieback, damping-off, and root rot (Hendricks and Roberts 2015; Ivimey and Collins 1937; Nawaz et al. 2016; Nzungize et al. 2011; Park et al. 2016; Takeuchi et al. 2002; Zhang and Yang 2000). For example, P. spinosum causes dieback in watermelon plants (Hendricks and Roberts 2015), damping-off in Cucumis melo (Park et al. 2016), root rot in chili plants (Nawaz et al. 2016), the common bean (Nzungize et al. 2011), and Sansevieria trifasciata (Takeuchi et al. 2002), and wilting in Primula spp. (Ivimey and Collins 1937). Based on our initial investigations in the Huang-Huai region of China, P. spinosum is also one of the major agents causing soybean root rot and damping-off.
Accurate identification of pathogens is the foundation of disease diagnosis and control. However, the symptoms of root rot and damping-off caused by P. spinosum are similar to those in diseases caused by other pathogens such as Fusarium graminearum, F. culmorum, Rhizoctonia solani, Phytophthora sojae, and Pythium ultimum (Lu et al. 2015; Zeng et al. 2017); thus, disease diagnosis in the field is difficult. Although many Pythium species can be identified using conventional laboratory methods, such as morphological and biological analyses, samples can display intraspecific phenotypic variation under different culture conditions (Pettitt et al. 2002), and may not always produce sexual reproductive structures in vitro for accurate identification (Botton et al. 2011). Therefore, the development of a rapid molecular assay that specifically detects P. spinosum would be useful for diagnosing soybean root rot caused by this pathogen in the field.
The loop-mediated isothermal amplification (LAMP) (Notomi et al. 2000) assay has high sensitivity and specificity because it uses a set of four primers with six binding sites that hybridise to the target gene sequence. The reaction results are visually confirmed by adding hydroxynaphthol blue (HNB) at the start of the assay (Goto et al. 2009). A positive result is indicated by a sky-blue colour, whereas a negative sample remains purple. LAMP assays have been developed to detect several Pythium pathogens at the species level, including P. aphanidermatum (Fukuta et al. 2013), P. inflatum (Cao et al. 2016), P. irregulare (Feng et al. 2015), P. myriotylum (Fukuta et al. 2014), and P. ultimum (Shen et al. 2017). However, no LAMP assay for detecting P. spinosum in soybean has been reported to date.
The objective of this study was to develop a simple LAMP detection method for specific identification of P. spinosum. We used the LAMP assay to diagnosis soybean root rot and damping-off caused by P. spinosum by directly detecting the pathogen in diseased host tissues and soil collected from three provinces.
Materials and methods
Strain isolation, identification, and DNA extraction
From 2016 to 2017, 142 soybean samples with symptoms of seed decay, seedling damping-off, and plant root rot were collected from Jiangsu, Shandong, and Anhui provinces in the Huang-Huai region of China. Pythium strains were isolated from the tissues of symptomatic soybean plants using selective V8 juice agar medium (V8A) containing rifampicin, ampicillin, and pentachloronitrobenzene. The isolation procedure followed methods described in Benard and Punja (1995).
The purified isolates were identified by morphological examination, and by sequencing the internal transcribed spacer 2 (ITS2) (White et al. 1990) and cytochrome oxidase I (COI) genes (Robideau et al. 2011). DNA was extracted from the pure cultures using the DNAsecure Plant Kit (Tiangen, Beijing, China) according to the manufacturer’s protocol. PCR to amplify the ITS2 and COI genes was then carried out as described previously (Chen et al. 2017). PCR products were sent to GenScript (Nanjing, China) for purification and sequencing. Amplicon sequences were compared with annotated sequences in the GenBank database hosted by the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov) using the BLASTN program. The species of each isolate was determined according to its best match in the database, with matches of at least 99% for both the ITS2 and COI regions.
The strains used in this study are maintained in a collection at the Department of Plant Pathology, Nanjing Agricultural University, China (Table 1). Pythium, Phytopythium, and Phytophthora strains were cultured on V8A, and the fungal strains isolated for our study were cultured on potato dextrose agar medium at 25 °C for at least 7 days before DNA extraction. Mycelia from each sample were harvested and then stored at −20 °C. DNA from mycelia was extracted using a modified hexadecyltrimethylammonium bromide (CTAB) method (Murray and Thompson 1980).
Pathogenicity assays
Pathogenicity was confirmed using six strains of P. spinosum isolated from Shandong, Anhui, and Jiangsu provinces to infect two soybean cultivars, Zhonghuang 13 and Xudou 18, which are commonly grown in these provinces. Pathogenicity in soybean seeds and seedlings was determined using assays similar to those previously described (Zhang and Yang 2000).
Pre-emergence damping-off assay: The six strains of P. spinosum were grown on V8A (Zheng 1997) for 3 days and then transferred to 2% water agar (WA) and incubated at room temperature (22–25 °C) for 48 h before inoculation. Six surface-sterilised soybean seeds (70% alcohol for 30 s and 2% sodium hypochlorite for 2 min, followed by three rinses with sterile water) were placed on the Pythium cultures. Seeds also were placed on WA plates without P. spinosum as controls. The plates were incubated at 25 °C in the dark for 7 days to allow seeds to germinate (i.e. primary root length equalled seed length) before disease levels were assessed. Three replicate plates were used for each strain. The disease levels were categorised as follows: 0 (no symptoms) = seed germinated without visible infection; 1 (weak) = seed germinated with short, light discoloured roots; 2 (strong) = seed died after germination; 3 (very strong) = seed died before germination.
Post-emergence damping-off assay: Three-day-old cultures of P. spinosum were cut into small pieces (5 × 5 mm), mixed with sterilised vermiculite (vermiculite subjected to moist heat at 121 °C for 20 min), and placed in Petri dishes (100 × 15 mm). One Petri dish was then buried at a 13-cm depth in each pot (500-mL plastic pots) used in this assay. Six soybean seeds were spread out evenly in each pot at a 2-cm depth. In control pots, seeds were planted in sterilised vermiculite mixed with sterile V8A. Three replicate pots were used for each isolate. All pots were incubated in a glasshouse at 25 °C for 14 days. After 14 days, the emerged seedlings were removed from pots, and germination status and plant height were measured.
All data were compared using one-way analysis of variance using the SPSS Statistics ver. 20 software for Windows (IBM, Armonk, NY, USA). Causal agents were isolated from symptomatic plants and identified based on morphological examination and comparisons of ITS and COI sequences.
LAMP primer design and screening
After comparing the sequences of different Pythium spp. from GenBank and the PCR products, ITS2 was selected to differentiate among Pythium spp. Sequences were aligned using BioEdit (Hall 1999) (Fig. 1). Multiple sets of primers were designed based on the polymorphic sequence regions of the ITS2 alignment, using PrimerExplorer ver. 4 (http://primerexplorer.jp/e), and screened using a series of specificity and sensitivity tests. One set of primers, listed in Table 2, was finally selected for use in our study.
LAMP assay reactions and pathogen detection
The LAMP assay was performed for each 26-μL sample containing 2.5 μL 10× ThermoPol Buffer (0.1% Triton X, 20 mM Tris-HCl, 10 mM KCl, 10 mM (NH4)2SO4, pH 8.8), 8 mmol·L−1 MgSO4, 0.8 μmol·L−1 betaine, 1.2 mmol·L−1 dNTPs, 1.6 μmol·L−1 each of the primers FIP and BIP, 0.4 μmol·L−1 each of the primers F3 and B3, 0.8 μmol·L−1 each of the primers LB, 180 mmol·L−1 HNB, 8 U·μL−1Bst DNA polymerase, and 2 μL DNA template. Amplification was conducted using the Gene Amp PCR system 2700 (ABI, Tokyo, Japan).
The LAMP assay was performed with the selected ITS2 primers using serial seven-fold dilutions (10 ng·μL−1 to 10 fg·μL−1) of pure P. spinosum genomic DNA to determine the detection limit of the LAMP assay. The optimal reaction conditions for distinguishing P. spinosum from other species were found to be 63 °C for 60 min. A positive control (standard P. spinosum DNA as the template) and negative control (double-distilled water in place of template) were included in each run, and each sample was analysed at least three times. After the reaction, the LAMP product was visually inspected to note any colour changes due to HNB. A change in colour of the reaction mixture from purple to sky blue denoted positive amplification, whereas nothing was amplified in reaction mixtures that remained purple.
DNA extraction from inoculated soybean tissues
After surface disinfection, the soybean seeds were planted in sterilised vermiculite and maintained in a glasshouse at 25 °C. When the first true leaf expanded, seedlings were inoculated by transferring culture discs (5 mm in diameter) from 3-day-old cultures of P. spinosum to the soybean hypocotyls using a sterilised metal needle. As a control, seedlings were inoculated with agar plugs from sterile V8A. The plants were incubated at 25 °C for 4 days with a 12-h photoperiod in the greenhouse. DNA was extracted from the tissues using the DNAsecure Plant Kit according to the manufacturer’s protocol.
DNA extraction from diseased tissues collected in the field
To assess the feasibility of using the LAMP assay to rapidly diagnose soybean disease caused by P. spinosum, samples of soybean plants suspected to be infected were collected, and DNA was extracted from the diseased root and basal stem tissues for LAMP assays using the DNAsecure Plant Kit. The same method was used to extract DNA from healthy soybean tissues, which served as negative controls.
DNA extraction from soil
Spore suspensions of P. spinosum containing 0, 10, 50, 100, 1000, and 10,000 oospores were added to 0.25-g replicate samples of sterilised soil. DNA was extracted from soil samples using a PowerSoil DNA Isolation Kit (MO BIO, Carlsbad, CA, USA), according to the manufacturer’s protocol. In addition, 20 soil samples were collected from soybean fields in Shandong, Anhui, and Jiangsu provinces to evaluate the effectiveness of the LAMP assay in directly detecting P. spinosum in soil.
Results
Isolation and identification of P. spinosum
From 2016 to 2017, a total of 108 Pythium and Phytopythium strains were isolated from 142 soybean plants that displayed root-rot and damping-off symptoms in the fields. Among the isolates, 29 strains (27%) were identified as P. spinosum using both morphological and molecular methods, including 21, 2, and 6 strains isolated from 92, 20, and 30 diseased soybean plants collected in Jiangsu, Anhui, and Shandong provinces, respectively (Table 3).
Morphological observations indicated that strains of P. spinosum were homothallic and could be distinguished from other Pythium spp. by the presence of hyphal swellings, oogonia with finger-like ornamentations, and monoclinous antheridia (Fig. S1). In addition, the cloned ITS2 and COI sequences of all isolates had the best matches (at least 99%) with GenBank sequences annotated as belonging to P. spinosum (GenBank IDs: ITS2, HQ643793; COI, HQ708834).
Pathogenicity of isolates of P. spinosum
We randomly selected two isolates of P. spinosum from each province for pathogenicity assays using the Zhonghuang 13 and Xudou 18 soybean cultivars. After growing on plates inoculated with P. spinosum for 7 days, most of the soybean seeds were dead before or after germination, whereas those grown on plates without P. spinosum did not display any visible disease symptoms (Fig. 2a–d). Zhonghuang 13 seeds displayed near-strong (close to Level 2) disease levels upon infection, whereas infected Xudou 18 seeds displayed strong (Level 2) and very strong (Level 3) disease levels (Fig. 2e). Conversely, almost all non-infected seeds exhibited no disease symptoms (Fig. 2e).
Pathogenicity of isolates ofP. spinosum. Examples of symptoms observed in the pre-emergence damping-off assay in the (a) non-inoculated and (b) inoculated Zhonghuang 13 seeds, and (c) non-inoculated and (d) inoculated Xudou 18 seeds. e Mean disease levels in the pre-emergence damping-off assay. Disease levels were classified as follows: 0, no symptoms; 1, weak; 2, strong; and 3, very strong. Examples of symptoms observed in the post-emergence damping-off assay in (f) Zhonghuang 13 and (g) Xudou 18 seedlings. The non-inoculated plant is indicated by CK. h Germination rates and (i) seed and seedling heights measured in the post-emergence damping-off assay. SD, JS, and AH refer to strains of P. spinosum isolated from samples collected from Shandong, Jiangsu, and Anhui provinces, respectively, TR indicates inoculation with P. spinosum. * P < 0.05
Similar results were obtained when seeds were grown in pots with soil mixed with hyphal plugs of P. spinosum or only with sterilised soil for two weeks. Compared with control seeds, seeds grown in P. spinosum-treated soil died before germination, or produced weaker seedlings with significantly shorter plant height, and shorter and fewer lateral roots (Fig. 2f–i). In addition, P. spinosum could be isolated from the diseased soybean seedlings and was identified based on morphological characteristics and comparisons of ITS2 and COI sequences. These results confirm that P. spinosum is a soybean pathogen.
Development of the LAMP assay for P. spinosum
The ITS2 region was selected as the target for developing the LAMP assay to distinguish P. spinosum from other congenerics (Fig. 1), and a set of LAMP primers was then selected after screening. The specificity of the LAMP assay to P. spinosum was tested using DNA from 20 randomly selected strains isolated from diseased soybean roots harvested from fields in Jiangsu, Anhui, and Shandong provinces. Positive and negative results could be easily distinguished based on visual detection using HNB; only samples containing P. spinosum displayed a sky-blue colour (Fig. 3a). From assays using decreasing concentrations of DNA from 10 ng·μL−1 to 10 fg·μL−1, the minimum DNA concentration required for detection by the LAMP assay was 100 pg·μL−1 (Fig. 3b).
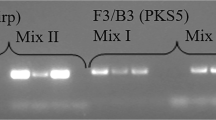
The ability of the LAMP assay to distinguish P. spinosum from other pathogens was tested using the DNA of various strains of P. spinosum, other Pythium spp., and non-Pythium species (Table 1). Only samples containing P. spinosum displayed a positive result (sky blue), whereas samples of related oomycete or fungi strains (many of which are also soybean root pathogens) all remained purple, indicating negative results (Fig. 4). Negative controls using double-distilled water as a template remained purple as well.
Detection of P. spinosum in diseased soybean tissues
DNA samples from artificially inoculated soybean seedlings produced positive reactions in the LAMP assay, whereas samples from non-inoculated soybean seedlings displayed no colour change, similar to the negative control (Fig. 5a). We also analysed 142 samples of diseased soybean tissues collected from fields in Jiangsu, Shandong, and Anhui provinces in 2016 and 2017. Using the LAMP assay, P. spinosum was detected in the tissues of 60 samples (Fig. 5b). The detection rate for P. spinosum using LAMP was higher than that using the isolation method (42% vs. 20%; Table 3). In addition, DNA of P. spinosum was isolated from some of the samples testing positive for disease using LAMP, but not from samples with negative LAMP results.
Ability of the LAMP assay to detectP. spinosumin infected soybean tissues and soil. a Results of using the LAMP assay to detect P. spinosum in inoculated soybean seedlings. b Analysis of diseased soybean tissues collected from fields. c Efficiency of the LAMP assay in detecting P. spinosum in soil containing different numbers of oospores. The detection limit for the assay was ten oospores per 0.25 g of soil
Detection of P. spinosum in soil
The LAMP assay detected the presence of P. spinosum when at least ten oospores of P. spinosum were present in 0.25 g of soil (Fig. 5c). We also analysed soil collected in different soybean fields. Using LAMP, P. spinosum was detected in some of the soil samples. Using a baiting method (Benson 1997), P. spinosum was isolated from some soil samples that tested positive in the LAMP assay, but not from any samples that tested negative (unpublished data).
Discussion
Root-rot and damping-off diseases have a substantial negative impact on soybean production. The soybean cultivars Zhonghuang 13 and Xudou 18, commonly grown in Jiangsu, Shandong, and Anhui provinces within the Huang-Huai region, the second largest soybean production area in China, were found to be susceptible to most isolated strains of P. spinosum. Here, we present the first report of P. spinosum causing soybean root rot and damping-off in China.
In recent years, molecular detection based on PCR has been applied to investigate P. spinosum (Feng et al. 2019; Nzungize et al. 2011; Toda et al. 2015). So far no research has been conducted to detect P. spinosum in soybean, and this is the first report of P. spinosum causing root rot and damping-off of soybean in China. Compared with PCR methods, LAMP assays are faster and easier to perform: a diagnosis can be obtained in only two to three hours, results can be scored visually by adding HNB, and the assay requires only a standard laboratory water bath or heat block to achieve isothermal conditions. Using the LAMP assay, P. spinosum can be detected directly from diseased soybean tissues and field soil without interference from impurities. Feng et al. (2019) applied the ITS-LAMP to detect P. spinosum in lettuce; however, their amplification products were separated on a 3% agarose gel, stained with GelRed, and photographed under UV light. HNB can be added before incubation so that amplification is completed in a closed tube system, and detection of the colour change requires no equipment (Duan et al. 2014; Ghosh et al. 2015). And our positive and negative results could be easily distinguished based on visual detection using HNB in this study.
The LAMP primer set designed from the ITS2 sequence specifically amplified DNA of P. spinosum only. The targeted ITS sequence is flanked by regions that are highly conserved within Pythium spp. (Le Floch et al. 2007), and has been used for the efficient detection and identification of Pythium spp. (Fukuta et al. 2013, 2014; Villa et al. 2006). A total of 142 samples of putatively diseased soybean plants were collected from the soybean fields. Our assay was able to detect P. spinosum successfully from samples containing the pathogen, indicating that the ITS2-based LAMP assay can be used to rapidly diagnose infections by P. spinosum in diseased soybean plants. The successful isolation of P. spinosum from the majority of tissue and soil samples testing positive in the LAMP assay, but not from the samples testing negative, indicates that the LAMP assay is robust and highly specific to P. spinosum. The oomycetes can remain viable for more than several years in soil as oospore, which makes it difficult to control (Zheng 1997). Residual pathogens in the soil can be used as primary infection sources to infect plants. Once plants are infected by Pythium, it is difficult to control. Our LAMP assay could provide an important preventive management tool for reducing loss by accurate soil and seedling testing. Therefore, the LAMP assay developed in our study provides a useful method for the rapid diagnosis of soybean root rot caused by P. spinosum.
References
Benard D, Punja ZK (1995) Role of Pythium species in cavity spot development on carrots in British Columbia. Can J Plant Pathol 17:31–45
Benson D (1997) Phytophthora diseases worldwide. 1st ed. Crop Protection 16(4): 399
Botha WJ (1993) Zoospore production in Pythium spinosum. Mycol Res 97:1495–1498
Botton SA, Pereira DI, Costa MM, Azevedo MI, Argenta JS, Jesus FP, Alves SH, Santurio JM (2011) Identification of Pythium insidiosum by nested PCR in cutaneous lesions of Brazilian horses and rabbits. Curr Microbiol 62(4):1225–1229
Cao YY, Li YQ, Li JJ, Wang LF, Cheng ZQ, Wang H et al (2016) Rapid and quantitative detection of Pythium inflatum by real-time fluorescence loop-mediated isothermal amplification assay. Eur J Plant Pathol 144:83–95
Chen JJ, LÜ L, Ye WW, Wang YC, Zheng XB (2017) Pythium cedri sp. nov. (Pythiaceae, Pythiales) from southern China based on morphological and molecular characters. Phytotaxa 309(2):135
Duan YB, Zhang XK, Ge CY, Wang Y, Cao JH, Jia XJ, Wang JX, Zhou MG (2014) Development and application of loop-mediated isothermal amplification for detection of the F167Y mutation of carbendazim-resistant isolates in Fusarium graminearum. Sci Report 4:7094
Feng W, Ishiguro Y, Hotta K, Watanabe H, Suga H, Kageyama K (2015) Simple detection of Pythium irregulare using loop-mediated isothermal amplification assay. FEMS Microbiol Lett 362(21)
Feng W, Hieno A, Kusunoki M, Suga H, Kageyama K (2019) LAMP detection of four plant-pathogenic oomycetes and its application in lettuce fields. Plant Dis 103(2):298–307
Fukuta S, Takahashi R, Kuroyanagi S, Miyake N, Nagai H, Suzuki H (2013) Detection of Pythium aphanidermatum in tomato using loop-mediated isothermal amplification (lamp) with species-specific primers. Eur J Plant Pathol 136:689–701
Fukuta S, Takahashi R, Kuroyanagi S, Ishiguro Y, Miyake N, Nagai H, Suzuki H et al (2014) Development of loop-mediated isothermal amplification assay for the detection of Pythium myriotylum. Lett Appl Microbiol 59(1):49–57
Ghosh R, Nagavardhini A, Sengupta A, Sharma M (2015) Development of Loop-Mediated Isothermal Amplification (LAMP) assay for rapid detection of Fusarium oxysporum f. sp. ciceris - wilt pathogen of chickpea BMC Research Notes (2015) 8:40
Goto M, Honda E, Ogura A, Nomoto A, Hanaki K (2009) Colorimetric detection of loop-mediated isothermal amplification reaction by using hydroxy naphthol blue. Biotechniques 46(3):167–172
Hall TA (1999) A user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hendricks KE, Roberts PD (2015) First report of Pythium spinosum as a pathogen of watermelon and in association with a dieback of watermelon in Southwest Florida. Plant Health Progress
Hendrix FF, Campbell WA (1973) Pythiums as plant pathogens. Annu Rev Phytopathol 11:77–98
Ivimey CW, Collins WB (1937) A pythium wilt of primula caused by Pythium spinosum, sawada. Trans Br Mycol Soc 21:29–33
Le Floch G, Tambong J, Vallance J, Tirilly Y, Levesque A, Rey P (2007) Rhizosphere persistence of three Pythium oligandrum strains in tomato soilless culture assessed by DNA macroarray and real-time PCR. FEMS Microbiol Ecol 61(2):317–326
Lu CC, Zhang HF, Wang YC, Zheng XB (2015) Rapid diagnosis of Fusarium root rot in soybean caused by Fusarium equiseti or Fusarium graminearum using loop-mediated isothermal amplification (LAMP) assays. Australas Plant Pathol 44:437–443
Martin FN, Loper JE (2010) Soilborne plant diseases caused by Pythium spp.: ecology, epidemiology, and prospects for biological control. Crit Rev Plant Sci 18(2):111–181
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nawaz K, Shahid AA, Subhani MN, Anwar W (2016) First report of Pythium spinosum causing root rot of chili (Capsicum annuum) in Pakistan. Plant Dis 100:526
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Amino N, Hase T (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28(12):e63–e663
Nzungize J, Gepts P, Buruchara R, Buah S, Ragama P, Busogoro JP, Baudoin JP (2011) Pathogenic and molecular characterization of Pythium species inducing root rot symptoms of common bean in Rwanda. Afr J Microbiol Res 5:1169–1181
Park MJ, Back CG, Han KS, Park JH (2016) Occurrence of damping-off caused by Pythium spinosum on Cucumis melo in Korea. Res Plant Dis 22(3):190–193
Pettitt TR, Wakeham AJ, Wainwright MF, White JG (2002) Comparison of serological, culture, and bait methods for detection of Pythium and Phytophthora zoospores in water. Plant Pathol 51:720–727
Plaats-Niterink AJVD (1981) Monograph of the genus Pythium. Stud Mycol 21
Robideau GP, De Cock AW, Coffey MD, Voglmayr H, Brouwer H, Bala K et al (2011) DNA barcoding of oomycetes with cytochrome coxidase subunit I and internal transcribed spacer. Mol Ecol Resour 11(6):1002–1011
Shen DY, Li QL, Yu J, Zhao YY, Zhu Y, Xu H, Dou DL (2017) Development of a loop-mediated isothermal amplification method for the rapid detection of Pythium ultimum. Australas Plant Pathol 46(6):571–576
Takeuchi J, Horie H, Nishimura S (2002) First report of Pythium rot of Sansevieria trifasciata caused by Pythium spinosum in Japan. Annu Rep Kanto-Tosan Soc Plant Prot 49:89–91
Toda T, Iwasa A, Fuji S, Furuya H (2015) Widespread occurrence of Pythium arrhenomanes pathogenic to Rice seedlings around Japanese Rice fields. Plant Dis 99(12):1823–1831
Villa NO, Kageyama K, Asano T, Suga H (2006) Phylogenetic relationships of Pythium and Phytophthora species based on ITS rDNA, cytochrome oxidase II and β-tubulin gene sequences. Mycologia 98(3):410–422
Wang PH, Chung CY, Lin YS, Yeh Y (2003) Use of polymerase chain reaction to detect the soft rot pathogen, Pythium myriotylum, in infected ginger rhizomes. Lett Appl Microbiol 36:116–120
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols:315–322
Wrather JA, Koenning SR (2006) Estimates of disease effects on soybean yields in the United States 2003 to 2005. J Nematol 38:173–180
Zeng DD, Ye WW, Xu M, Lu CC, Tian Q, Zheng XB (2017) Rapid diagnosis of soya bean root rot caused by Fusarium culmorum using a loop-mediated isothermal amplification assay. J Phytopathol 165(4):249–256
Zhang BD, Yang XB (2000) Pathogenicity of Pythium populations from corn–soybean rotation fields. Plant Dis 84:94–99
Zheng XB (1997) Methods in Phytophthora. China, Chinese Agriculture Press, Beijing
Acknowledgements
We thank Xiaoli Wang and Yue Yang for maintaining the strains used in this study. This work was supported by the China Agriculture Research System (CARS-004-PS14), the Fundamental Research Funds for the Central Universities (KJQN201738), the National Natural Science Foundation of China (31601618) and the Special Fund for Agro-scientific Research in the Public Interest of China (201303018).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: XBZ and YCW.
Performed the experiments: HF, JJC, ZY, and ZL.
Analyzed the data: HF, JJC, WWY, and XBZ.
Contributed reagents/materials/analysis tools: HF, JJC, ZY, ZL, WWY, and YCW.
Wrote the paper: HF, JJC, XBZ, and WWY.
Corresponding author
Electronic supplementary material
Figure S1
Asexual and sexual reproductive structures of P. spinosum. (A) Mycelia. (B, C) Hyphal swellings. (D) A terminal oogonium. (E) An intercalary oogonium. (F) Catenulate oogonia. (PNG 2048 kb)
Rights and permissions
About this article
Cite this article
Feng, H., Chen, J., Yu, Z. et al. A loop-mediated isothermal amplification assay can rapidly diagnose soybean root-rot and damping-off diseases caused by Pythium spinosum. Australasian Plant Pathol. 48, 553–562 (2019). https://doi.org/10.1007/s13313-019-00659-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-019-00659-7