Summary
Alzheimer’s disease (AD) is the most common cause of dementia and represents one of the highest unmet needs in medicine today. Drug development efforts for AD have been encumbered by largely unsuccessful clinical trials in the last decade. Drug repositioning, a process of discovering a new therapeutic use for existing drugs or drug candidates, is an attractive and timely drug development strategy especially for AD. Compared with traditional de novo drug development, time and cost are reduced as the safety and pharmacokinetic properties of most repositioning candidates have already been determined. A majority of drug repositioning efforts for AD have been based on positive clinical or epidemiological observations or in vivo efficacy found in mouse models of AD. More systematic, multidisciplinary approaches will further facilitate drug repositioning for AD. Some experimental approaches include unbiased phenotypic screening using the library of available drug collections in physiologically relevant model systems (e.g. stem cell-derived neurons or glial cells), computational prediction and selection approaches that leverage the accumulating data resulting from RNA expression profiles, and genome-wide association studies. This review will summarize several notable strategies and representative examples of drug repositioning for AD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For the last decade, pharmaceutical research and development productivity, measured by the relationship between investment input to develop new innovative drugs and the output/outcome (e.g., medical and financial benefits resulting from the drug), dramatically deceased and, at the time of writing, remains low [1–3]. A number of factors contribute to the productivity issues in drug development, including increasing development costs, generic competition due to the patent expiry of several blockbuster drugs, increasingly conservative regulatory policies, and insufficient breakthrough innovations. Increasing research and development expenditures are more apparent in the disease indications with the highest unmet need, such as Alzheimer’s disease (AD), where the attrition rate is high [1].
Given the growing elderly population, combined with no approved disease-modifying therapeutics and a weak pipeline of drugs under development, AD represents one of the biggest unmet needs in medicine [4, 5]. Recent analysis of historic trends revealed that AD drug discovery and development has been experiencing major setbacks for the last decade owing to the lowest success rate among all therapeutic areas (~0.4% during the period 2001–12) [6, 7]. The high attrition rate for AD drug development is mainly attributed to the failure of predicting the safety and/or efficacy of candidate drugs prior to human testing.
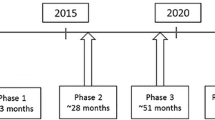
Drug repositioning strategies have been suggested as a solution to overcome some of these impending pharmaceutical industry and medical challenges [8–10]. “Drug repositioning”, also referred to as “drug repurposing”, is the process of identifying new indications for existing drugs (Fig. 1). In addition to approved drugs, this approach can also salvage drugs that have been shown to be safe but not effective for the indications for which they were originally developed. Drug repositioning has several advantages over the traditional de novo drug discovery approach, including reduced development costs and shorter time to approval and launch [8–10]. The drug repositioning approach can also substantially reduce the risk associated with conventional drug discovery by using existing pharmacokinetic, toxicology and safety data. For AD, drug repositioning is a very attractive strategy that should be more actively pursued [11–13].
A schematic representation of (A) traditional drug discovery and development versus (B) drug repositioning. Traditional de novo drug discovery and development involves a 10–17-year process. Drug repositioning decreases costs and time to launch, and reduces safety and pharmacokinetic uncertainty frequently associated with traditional drug development
Major advances in our understanding of AD disease biology have been made and a number of new resources have become available to facilitate drug discovery efforts through repositioning. These include abundant data from mechanistic, clinical and epidemiological research, development of biomarkers, and a number of well-validated models, both cell- and animal-based. Current drug repositioning approaches in AD are mainly based on ad hoc clinical and epidemiological risk assessment in human studies and preclinical changes in mouse models of AD [8]. However, more systematic approaches for both activity-based and computational methods should be implemented in AD drug repositioning efforts. Alongside rapid advances in disease biology, the availability of several established clinical drug libraries and sophisticated tools available in genomics and bioinformatics will greatly facilitate future endeavors in AD drug repositioning. For discovery of new AD indications, generally 2 alternative and complementary strategies can be employed: activity-based phenotypic screening or computational approaches.
Activity-Based Drug Repositioning Strategies
Existing Drug Collections
One of the prevailing strategies for drug repositioning is to screen, using phenotypic in vitro and in vivo assays relevant to a target disease(s), libraries of approved drugs and drug candidates. For instance, Chong et al. [14] assembled a library of 2687 existing drugs collected worldwide (known as John Hopkins University Clinical Compound Library) and performed phenotypic screening for parasite inhibition. A number of clinical library collections are now available through commercial sources: National Institutes of Health (NIH) Clinical Collections through Evotec (http://www.nihclinicalcollection.com), subsets of the compound collections from Microsource (e.g., Pharmakon1600, Spectrum, National Institute of Neurological Disorders and Stroke custom collection II and US and international drug collections), Sigma (e.g., LOPAC1280), Preswick, Selleckchem, Enzo Life Sciences, and Tocris (e.g., Tocriscreen) [15]. The NIH Chemical Genomics Center also built a collection of drugs approved in the USA and other foreign countries [16]. While these collections are ideal for first-line screening, a majority of these collections lack a number of clinical candidates that are actively being evaluated in human studies sponsored by major pharmaceutical and biotechnology companies. Public availability of the most up-to-date candidate drugs would greatly facilitate drug repositioning efforts in the drug discovery research community.
Phenotypic Approaches
Traditionally, identification of new drugs is mainly based on phenotypic changes in animal models or even serendipitous clinical observations made in humans [17, 18]. Starting in the 1980s, “rational” target-based drug discovery served as a dominant trend in the pharmaceutical industry [19]. In target-based drug discovery, small molecule screening of compound libraries consisting of structurally diverse small molecules is conduced to find compounds that bind specifically to the target with high affinity. In contrast, phenotypic drug discovery (PDD) is an experimental approach based on the discovery of bioactivities in model cells or organisms and does not rely on direct engagement of a specific cellular target. In fact, a seminal study attests that PDD led to substantially more successful first-in-class drugs compared with target-based drug discovery approaches, despite much greater resources invested in target-based approaches [19]. Once successful PDD is performed and identifies promising hit compounds, traditional target-directed or mechanism-based secondary assays are performed, followed by preclinical animal testing. For complex neurodegenerative disorders, such as AD, where molecular targets and disease mechanisms are far less clear, phenotypic assays can play an important role for selecting therapeutic leads that can effectively modify a disease-specific pathway [15, 20, 21].
A small number of studies have attempted to conduct phenotypic screening relevant to AD, mainly by examining amyloid precursor protein (APP) and APP metabolites, including amyloid β-peptide (Aβ), using libraries of a few hundred Food and Drug Administration (FDA)-approved drugs in neuronal cell lines [22–24]. Recently, screening has been conducted to identify compounds that can modulate additional AD-associated phenotypes such as Aβ-induced cytotoxicity, tau protein levels, apolipoprotein E (apoE) secretion or apoE4 conformation, Ca2+ signaling and neurogenesis in neuronal cell lines [22, 25–30]. Within industry-sponsored phenotypic screening initiatives, apoE stimulators for possible AD therapeutics have been pursued by Eli Lilly’s Phenotypic Drug Discovery Initiative (PD2) [31]. As screening data are not publically available, the initiative has made limited contributions to scientific knowledge in academia or to the public at large. The Lilly Initiative uses an astrocytoma cell line, CCF-STTG1 (vs primary astrocytes) [31, 32]. It is conceivable that the screen identified false positive hits or missed a number of hits (false negatives) that may be preferentially active in primary cells and more physiological cell models, as certain compounds have been found to exert biological activities only in physiologically relevant cells but not in transformed cell lines [33].
In neurodegenerative disease such as AD, protein aggregation could be also be considered an in vitro phenotype. Several studies screened small collections of FDA-approved drugs to identify new compounds harboring antiaggregation effects [34, 35]. One promising area of research will be to screen AD-associated phenotypes involving transcellular propagation of tau or Aβ aggregates in the manner shared with that of prion [36, 37]. Given increasing significance of cell-to-cell propagation of AD-associated protein aggregates [36, 37], modeling an emerging but potentially significant pathological phenotype is likely to yield new opportunities for discovering highly effective drug repositioning candidates.
As physiological context is crucial for establishing predictive in vitro models for AD phenotypic screening, there is an increasing need for more physiological models, such as primary brain cells or stem cell-derived neurons or glial cells, which are scalable and robust for high throughput screening (HTS) assays [15, 20, 33, 38]. Small molecule screenings for Aβ biogenesis and Aβ toxicity have been described in mouse embryonic stem cell-derived pyramidal neurons and commercial human induced pluripotent stem (iPS) cell-derived neurons [33, 39]. While human patient-derived stem cells (e.g., iPS cells) carry great promise [40, 41], their productive and reliable use for drug screening applications such as HTS remains challenging owing to laborious maintenance, which makes automation impractical [42–44]. Furthermore, genetic and epigenetic variability among patients may influence pharmacological responsiveness [45]. Disease modeling using iPS cells for AD has been actively pursued recently and has shown that pharmacological assays can be readily performed using these models [46–52]. Modeling and screening using glial–neuron co-culture and 3-dimensional cultures, both of which better resemble in vivo physiological states, should also be pursued [53, 54]. Thus, successful drug repositioning for AD can be further facilitated with the establishment of “AD-in-a-dish”, furnishing phenotypic assays suitable for robust and scalable HTS assays using physiological human neuron or glial cells [55].
As an alternative to the classical phenotypic screening platform using cultured cells relevant to AD, phenotypic screening using intact model organisms, such as yeast, nematode, Drosophila, and Zebrafish, substantially adds value [56]. Pharmacological effects of drug repositioning candidates can be systematically evaluated in molecular, cellular, and network contexts of intact model organisms, while offering reasonable throughput and allowing for screening of a relatively large number of compounds compared to mouse or other in vivo mammalian models [57, 58].
Target-Based Approaches
In addition to phenotypic screening, target-based screens can be used to discover a new activity of existing drugs. Advances in disease biology often lead to the identification of new druggable pharmaceutical targets, and the newly identified therapeutic target can be subjected to target-based screening such as enzymatic HTS assays [14, 59]. In AD, this approach has had limited success as the majority of target-based approaches have focused on development of high-affinity novel compounds that can potently inhibit known AD targets such as secretases. In contrast, the above-mentioned phenotypic screening paradigms could potentially reveal new drug pathway relationships, and specific molecular target(s) in the pathway could lead to new target-based project strategies.
Computational Drug Repositioning
Systematic computational analyses are powerful approaches for drug repositioning, allowing prediction and discovery of novel targets and therapeutic indications [60–63]. Owing to their ability to access virtually large numbers of compounds and data relating to the target disease, informatics-based approaches can complement and further facilitate drug repositioning efforts more systematically than the experimental approaches described above. Successful computational repositioning depends on several components, including the extent and quality of the data, appropriate analytical tools, and integration of the knowledge obtained from the analyses in order to effectively devise a new opportunity for drug repositioning. The key objective of the computational approach is to identify novel drug–disease connections using several computational methods.
Transcriptomic Approaches
Gene expression profiles serve as a quantitative tool to compare biological states associated with either a pharmacological response to a drug or a particular disease [64, 65]. This approach is based on the “connectivity map” project, which first built a database of gene expression profiles associated with a number of reference drugs [64, 65]. In silico analysis first establishes a disease-specific regulatory “signature” consisting of genes either up- or downregulated in the disease state compared with unaffected controls. By comparing these disease signatures with drug-specific signatures, new indications for a particular drug can be predicted.
Sirota et al. [66] demonstrated how computational analysis of public gene expression data led to the discovery of new indications for known drugs. The disease-associated gene expression data were first subjected to significance analysis by microarray to establish “disease signatures” for 100 diseases. In order to deduce possible new indications, each of the signatures was then compared against the reference drug signatures available through the connectivity map [64, 65]. Discovery of potential therapeutic drugs were identified based on the assumption that if a disease signature is opposite to a particular reference drug signature, the drug has a therapeutic potential for that specific disease. This concept has been validated experimentally. For instance, using this approach, topiramate (an epilepsy drug) was predicted to be a therapeutic for Crohn’s disease and ulcerative colitis [66], and, indeed, was found to exert potent efficacy in a rat model of colitis [61].
In the field of AD research, efforts to use the public databases have only just begun to immerge for integrating disease-linked gene expression profiles and gene expression data from drug exposure studies [67]. Currently, only limited gene expression profile data from neuronal tissues treated with small molecules are available for gene expression analysis. With increasing availability of the gene expression data set for central nervous system (CNS) tissues and brain cells from normal and AD models, this approach will gain further momentum in the field of AD drug discovery.
Genome-Wide Association Studies
The application of genomics technologies is also a powerful tool for drug repositioning. For the last several years, genome-wide association studies (GWAS) have revealed new information regarding the association of specific genomic variations with complex trait human diseases, such as AD. While these genetic studies provide new biological insights and pathobiological hypotheses, practical use of the vast amounts of GWAS data for clinical use, such as the development of personalized therapeutics, is still being established. Recently, new strategies have been proposed to use GWAS data for drug repositioning [68, 69].
Based on the list of GWAS genes associated with specific disease traits, a series of analyses can be performed to determine a subset of genes considered to be “drug targets” based on the druggability of each gene product [69]. Among these, further analysis can determine whether any of the gene products considered to be druggable have already been targeted by drugs currently in clinical development. Such analyses can identify 2 distinct groups among the GWAS genes linked to specific drug development project(s): one in which GWAS traits match the drug indication and the other that differs from the drug indication. For instance, the clinical candidate Biib-033, an antibody targeting the leucine-rich repeat and immunoglobulin domain-containing 1 (LINGO-1), was being developed for multiple sclerosis. Two GWAS studies reported that a marker in LINGO1 shows significant genome-wide association with essential tremor [70, 71]. These observations suggested that essential tremor could potentially be considered a new indication. As new GWAS data accumulate for AD, additional gene–disease associations and drug repositioning opportunities should be unveiled.
Drug Repositioning for AD Application
One of the FDA-approved medications for AD was, in fact, discovered through drug repositioning. Galanthamine, an acetylcholinesterase inhibitor that was originally used in Eastern Europe in the 1950s for poliomyelitis, was later developed as an AD drug [72]. Several major diseases are known to contribute to the risk of developing AD, such as hypertension and diabetes. Accordingly, some of the approved therapeutics for these disease conditions, especially those known to penetrate the blood–brain barrier (BBB), have been actively investigated to evaluate their potential as AD drugs [12, 13]. Drug repositioning projects are often supported by various experimental data such as preclinical efficacy of select drugs in models of AD. It has been shown that regardless of the target class, a number of FDA-approved drugs exert bioactivities perceived to be beneficial for AD, such as harboring Aβ-reducing activities in animal models. In contrast, some drugs show opposite effects, suggesting that both target biology and chemotypes (bioactivities associated with particular chemical structural features) contribute substantially to the AD-relevant pharmacological activities of the drugs being considered for drug repositioning [73]. Thus, it is important to evaluate the possibility of drug repositioning on a case-by-case basis with consideration of multiple parameters prior to engaging further drug development. This subsection will discuss some representative examples.
Selective Serotonin Reuptake Inhibitors
Many CNS drugs developed for non-AD indications such as antidepressants are ideal candidates for drug repositioning, as these compounds generally have been established to be safe for a chronic use and penetrable into the brain. Selective serotonin reuptake inhibitors (SSRIs) are a class of CNS drugs that are frequently used as antidepressants. Some SSRIs have been shown to increase cognitive function in clinical studies and have been prescribed alongside cholinesterase inhibitors for neuropsychiatric complications associated with AD [74, 75]. In addition to symptomatic benefits, much preclinical evidence suggest disease-modifying effects of select SSRIs, such as activities involving Aβ reduction and increased hippocampal neurogenesis [74]. Stimulation of 5-hydroxytryptamine receptors was shown to promote nonamyloidogenic processing of APP [76–79], and the SSRIs paroxetine and imipramine reduced Aβ levels in mouse models of AD [80]. Direct analysis of brain interstitial fluid using microdialysis revealed that citalopram, desvenlafaxine, and fluoxetine reduced the interstitial fluid Aβ levels in presenilin 1 APP double transgenic mice [81]. Chronic administration of citalopram also substantially reduced Aβ plaque burden. Interestingly, retrospective analysis revealed that amyloid loads measured by positron emission tomography imaging with Pittsburgh Compound B were found to be significantly lower in a patient group treated with citalopram compared with an untreated group, suggesting a disease-modifying activity of citalopram. The Aβ-lowering activity was also accessed directly in human cerebrospinal fluid (CSF) using stable isotope labeling kinetics. CSF sampling during acute dosing of citalopram, demonstrated decreased Aβ in the CSF of the citalopram-treated group [82]. These studies suggest that citalopram and potentially other SSRIs are promising candidate drugs for repositioning.
Antiepileptic Drugs
Both animal and human studies have demonstrated that epileptiform activity is associated with AD [83, 84]. Among several antiepileptic drugs tested, only levetiracetam reduced epileptiform activity and reversed cognitive deficits in human APP transgenic mice [85, 86]. Levetiracetam is currently undergoing a Phase IIa clinical trial for subclinical epileptiform activity and/or improved cognition in patients with AD (ClinicalTrials.gov identifier: NCT02002819). Valproic acid, a treatment for epilepsy and bipolar disorder, showed preclinical efficacy with diverse mechanisms [87–90]. However clinical studies showed no beneficial effects in AD [91, 92].
Antihypertensive Drugs
Mid-life hypertension increases the risk of AD in later life [93–95]. Accordingly, hypertension medications including angiotensin converting enzyme (ACE) inhibitors, calcium channel blockers (CCBs), and angiotensin-1 receptor blockers (ARBs) have been tested as potential therapeutics for AD. Some of these compounds exerted antiamyloid activities in cultured cells and mouse models of AD.
Perindopril, a brain penetrating ACE inhibitor, was shown to prevent memory impairments in the Aβ infusion model, as well as APP transgenic mouse models of AD [96, 97]. In contrast, imidapril and enalapril, ACE inhibitors that do not cross the BBB, failed to improve cognitive deficits in AD mouse models, suggesting that the observed benefit is likely due to ACE inhibition in the brain. While an epidemiological study demonstrated that the use of ACE inhibitors does not influence the incidence of AD [98], a clinical study known as Perindopril Protection Against Recurrent Stroke Study (PROGRESS) demonstrated that perindopril reduces cognitive decline associated with cardiovascular disease [99].
Nilvadipine is one of the dihydropyridine-type CCBs, shown to cross the BBB. Administration of nilvadipine reduced the levels of Aβ in the brain, likely by facilitating Aβ clearance, leading to improved learning and memory in mouse models of AD [100, 101]. In addition to its effects on Aβ, nilvadipine restored Aβ-triggered reduction in cerebral blood flow in an AD mouse model [102], as well as in patients with early-stage AD [103, 104]. Clinical investigations also showed that nilvadipine can prevent cognitive decline of patients with mild cognitive impairment [103, 105]. Among non-APOE4 carriers, nilvadipine also improved short-term cognitive functions [106]. Another dihydropyridine CCB, isradipine, conferred neuroprotective effects in vitro [107], and also attenuated the levels of hyperphosphorylated tau and suppressed autophagy of tau [108].
ARBs are another class of antihypertensive drugs that have been explored extensively for repositioning in AD. Medication with ARBs has been associated with decreased incidence of dementia [109, 110]. Epidemiological analyses revealed that a group of patients that took ARBs, displayed significantly reduced incidence and progression of AD and dementia than those treated with a comparable antihypertensive agent, ACE inhibitor [109]. In vivo animal studies revealed that oral or intranasal administration of some ARBs, including valsartan, olmesartan, and losartan, reduced Aβ levels and ameliorated spatial learning and memory deficits in mouse models of Aβ pathology [111, 112]. A few ARBs have also been clinically evaluated for their effect on cognitive function in a patient group with high risk of developing cardiovascular disease [113, 114]. Overall results were largely found to be negative, at least for ramipril, telmisartan, and candesartan. Thus, among the representative classes of antihypertensive drugs, there are “within-class” variations in preclinical efficacy, alleviation of cognitive impairment, and the risk of developing AD.
Carvedilol, a nonselective adrenergic receptor blocker, is another antihypertensive medication investigated for potential use in AD. Initial structural analysis revealed that carvedilol, along with rolitetracycline and daunomycin, has a structural motif essential for binding to Aβ [115]. Subsequently, carvedilol was found to interfere with the oligomerization of Aβ [116]. Consequently, carvedilol treatment resulted in reduced levels of Aβ oligomers, as well as improved learning and memory behaviors in a mouse model of AD [116]. Thus, the findings obtained with carvedilol represent an example of how structural modeling of pharmacophores of known compounds can lead to a mechanistic hypothesis that is subsequently validated experimentally. In this case it is critical to evaluate the structural properties of the repositioning candidate prior to engaging in further development to determine if specific structural moieties are associated with desired bioactivities in addition to drug-like properties, metabolism and toxicity profiles, and potential side effects through off-target interactions with unwanted cellular target proteins.
Anti-Diabetes Drugs
Ample clinical and epidemiological studies suggest that patients with type 2 diabetes have a much greater risk of developing AD [117, 118]. In animal models, both deficiency of insulin and insulin resistance lead to increased Aβ pathology. It has also been shown that Aβ disrupts insulin receptor signaling cascades in neurons [119]. Accordingly, some of the drugs developed for type 2 diabetes have been evaluated for their potential benefit in disease modification for AD.
Direct administration of insulin into the brain via intranasal injection showed beneficial effects on cognition, as well as improvement of biomarker profiles, such as glucose uptake in the brain, and CSF Aβ and tau levels [120]. In contrast, insulin deficiency has been shown to accelerate amyloidogenic pathology in a mouse model of AD [119]. In addition to direct insulin administration, compounds that can promote insulin secretion, such as glucagon-like peptide 1 (GLP-1) analogues, are being actively investigated., Liraglutide and Val(8)GLP-1, GLP-1 analogues, were found to be neuroprotective, rescue long-term potentiation and memory deficits, and reduce plaque loads in an AD mouse model [121–123]. Peripherally injected liraglutide and another GLP-1 analogue, lixisenatide, penetrated the BBB and promoted neurogenesis [124]. Concordently, clinical trials are being conducted to evaluate the potential therapeutic efficacy of liraglutide (ClinicalTrials.gov Identifier: NCT01843075).
A functionally and structurally distinct class of antidiabetic drugs, the biguanide class, which includes metformin, has also been under investigation for effects on AD pathogenesis. While initial reports indicate that metformin increases the levels of brain Aβ by upregulating β-site APP cleaving enzyme 1 (BACE1) [125], some studies show that metformin attenuated AD-like changes and also reduced tau phosphorylation [126–128].
Phosphodiesterase 5 Inhibitors
Sildenafil (Viagra), a phosphodiesterase 5 (PDE5) inhibitor, which was originally developed as a treatment for angina, has been repurposed as a therapeutic agent for erectile dysfunction, and more recently for idiopathic pulmonary arterial hypertension [129]. Puzzo et al. [130] first demonstrated the preclinical efficacy of PDE5 inhibitors in AD mouse models and the findings were later confirmed by other groups [131–133]. Among reported PDE5 inhibitors, tadalafil and sildenafil can penetrate into the brain and ameliorate synaptic, learning, and memory deficits by modulating the cyclic guanosine monophosphate intracellular signaling pathway [130, 134].
Private–Public Partnership for Drug Repositioning
Selected drug candidates from several major pharmaceutical companies are now available for drug repurposing projects through academia–industry partnership. The open innovation drug repositioning project was initiated by the Medical Research Council (MRC) and AstraZeneca. In addition to the MRC program, the National Center for Advancing Translational Sciences (NCATS) at the NIH created a collaborative drug repurposing initiative with 8 pharmaceutical companies that contributed 58 compounds initially. Nine have been selected for projects that are led by academic researchers [10, 135]. The initiatives of the NCATS and MRC help the pharmaceutical companies gain access to academic innovations with the hope of ultimately finding a new indication(s) and recovering investments made in failed drug candidates. To date, these collaborative projects only support “on-target” repositioning that deals with the discovery of new indications through mechanisms acting on the drugs’ original targets.
Among the 7 compounds to be investigated for repurposing under NCATS’s drug repositioning project, one under investigation for possible repurposing for AD therapeutic use is AZD0530 (Saracatinib), a Src tyrosine kinase inhibitor that was originally developed for various cancers but was discontinued owing to insufficient efficacy [136, 137]. The concept for repositioning of this agent was based on scientific discoveries that demonstrated that phosphorylation of Fyn tyrosine kinase is related to Aβ- and tau-associated synaptic dysfunction, as well as memory impairments in mouse models of AD [138–140].
Intellectual Property Considerations
Despite all of the positive virtues of drug repositioning, several obstacles and potential solutions have recently been discussed by Shineman et al. [141]. Notable challenges include the continued requirement for clinical trials that are still considered risky, especially in the case of AD. The commercialization process for repurposed drugs has also presented challenges as patent protection and market exclusivity are not as robust as drugs with a strong “composition-of-matter” patent. In the case of repurposed drugs, “use” patents serve as the primary means of protecting the intellectual property (IP) and market exclusivity once the drug becomes commercialized [142]. The best example is the multiple sclerosis drug BG-12 (dimethyl fumarate). BG-12 was repurposed from an older drug by Biogen Idec, who successfully developed and commercialized it using the use patent [141, 143]. There are several strategies that could be used to obtain adequate IP protection to maximize the market exclusivity of a drug. New doses and/or formulations can add modestly to the strength of the IP protection provided by the use patent for new indications. Drug combination is another protection strategy. If anticipated synergy in a drug combination is identified, such a combination could potentially be considered a new composition. One additional factor that contributes to the robustness of patent protection is related to the Orphan Drug Act of 1983. If the FDA approves the repurposed drug with an orphan drug status, 7 years of exclusivity is granted [144].
Conclusions
In this review, I have provided an overview of drug repositioning strategies with an emphasis on next-generation phenotypic approaches, as well as several representative drug repositioning activities in the AD field. Given the availability of translational biomarkers and drug libraries, advanced informatics tools and increasing AD genomics data, drug repositioning for AD provides a promising opportunity. Through drug repositioning, it will be possible to enrich AD drug development pipelines by introducing a greater number of drug candidates into clinical development and also by increasing overall probability of success of each candidate. Increased funding and collaborative activities among academia, government, and pharmaceutical companies, along with appropriate policy support will greatly facilitate drug repositioning opportunities for AD.
References
Pammolli F, Magazzini L, Riccaboni M. The productivity crisis in pharmaceutical R&D. Nat Rev Drug Discov 2011;10:428-438.
Schulze U, Baedeker M, Chen YT, Greber D. R&D productivity: on the comeback trail. Nat Rev Drug Discov 2014;13:331-332.
Scannell JW, Blanckley A, Boldon H, Warrington B. Diagnosing the decline in pharmaceutical R&D efficiency. Nat Rev Drug Discov 2012;11:191-200.
Citron M. Alzheimer’s disease: strategies for disease modification. Nat Rev Drug Discov 2010;9:387-398.
Lo AW, Ho C, Cummings J, Kosik KS. Parallel discovery of Alzheimer’s therapeutics. Sci Transl Med 2014;6:241cm5.
Cummings JL, Morstorf T, Zhong K. Alzheimer’s disease drug-development pipeline: few candidates, frequent failures. Alzheimers Res Ther 2014;6:37.
Hay M, Thomas DW, Craighead JL, Economides C, Rosenthal J. Clinical development success rates for investigational drugs. Nat Biotechnol 2014;32:40-51.
Ashburn TT, Thor KB. Drug repositioning: identifying and developing new uses for existing drugs. Nat Rev Drug Discov 2004;3:673-683.
Jin G, Wong ST. Toward better drug repositioning: prioritizing and integrating existing methods into efficient pipelines. Drug Discov Today 2014;19:637-644.
Mullard A. Drug repurposing programmes get lift off. Nat Rev Drug Discov. 2012;11:505-506.
Cummings JL, Zhong K. Repackaging FDA-approved drugs for degenerative diseases: promises and challenges. Expert Rev Clin Pharmacol 2014;7:161-165.
Appleby BS, Cummings JL. Discovering new treatments for Alzheimer’s disease by repurposing approved medications. Curr Top Med Chem 2013;13:2306-2327.
Corbett A, Pickett J, Burns A, et al. Drug repositioning for Alzheimer’s disease. Nat Rev Drug Discov 2012;11:833-846.
Chong CR, Chen X, Shi L, Liu JO, Sullivan DJ Jr. A clinical drug library screen identifies astemizole as an antimalarial agent. Nat Chem Biol 2006;2:415-416.
Zhang M, Luo G, Zhou Y, Wang S, Zhong Z. Phenotypic screens targeting neurodegenerative diseases. J Biomol Screen 2014;19:1-16.
Huang R, Southall N, Wang Y, et al. The NCGC pharmaceutical collection: a comprehensive resource of clinically approved drugs enabling repurposing and chemical genomics. Sci Transl Med 2011;3:80ps16.
Awouters FH, Lewi PJ. Forty years of antipsychotic Drug research—from haloperidol to paliperidone—with Dr. Paul Janssen. Arzneimittelforschung 2007;57:625-632.
Aronson JK. Old drugs—New uses. Br J Clin Pharmacol 2007;64:563-565.
Swinney DC, Anthony J. How were new medicines discovered? Nat Rev Drug Discov 2011;10:507-519.
Pruss RM. Phenotypic screening strategies for neurodegenerative diseases: a pathway to discover novel drug candidates and potential disease targets or mechanisms. CNS Neurol Disord Drug Targets 2010;9:693-700.
Lee JA, Uhlik MT, Moxham CM, Tomandl D, Sall DJ. Modern phenotypic drug discovery is a viable, neoclassic pharma strategy. J Med Chem 2012;55:4527-4538.
Chakrabarti E, Smith JD. Drug library screen to identify compounds that decrease secreted Abeta from a human cell line. Curr Alzheimer Res 2005;2:255-259.
Utsuki T, Yu QS, Davidson D, et al. Identification of novel small molecule inhibitors of amyloid precursor protein synthesis as a route to lower Alzheimer’s disease amyloid-beta peptide. J Pharmacol Exp Ther 2006;318:855-862.
Spilman P, Descamps O, Gorostiza O, et al. The multi-functional drug tropisetron binds APP and normalizes cognition in a murine Alzheimer’s model. Brain Res 2014;1551:25-44.
Seyb KI, Schuman ER, Ni J, Huang MM, Michaelis ML, Glicksman MA. Identification of small molecule inhibitors of beta-amyloid cytotoxicity through a cell-based high-throughput screening platform. J Biomol Screen 2008;13:870-878.
Dickey CA, Eriksen J, Kamal A, et al. Development of a high throughput drug screening assay for the detection of changes in tau levels—proof of concept with HSP90 inhibitors. Curr Alzheimer Res 2005;2:231-238.
Dehdashti SJ, Zheng W, Gever JR, et al. A high-throughput screening assay for determining cellular levels of total tau protein. Curr Alzheimer Res 2013;10:679-687.
Chen HK, Liu Z, Meyer-Franke A, et al. Small molecule structure correctors abolish detrimental effects of apolipoprotein E4 in cultured neurons. J Biol Chem 2012;287:5253-5266.
Honarnejad K, Kirsch AK, Daschner A, Szybinska A, Kuznicki J, Herms J. FRET-based calcium imaging: a tool for high-throughput/content phenotypic drug screening in Alzheimer disease. J Biomol Screen 2013;18:1309-1320.
Longo FM, Yang T, Xie Y, Massa SM. Small molecule approaches for promoting neurogenesis. Curr Alzheimer Res 2006;3:5-10.
Lee JA, Chu S, Willard FS, et al. Open innovation for phenotypic drug discovery: The PD2 assay panel. J Biomol Screen 2011;16:588-602.
Liang Y, Lin S, Beyer TP, et al. A Liver X receptor and retinoid X receptor heterodimer mediates apolipoprotein E expression, secretion and cholesterol homeostasis in astrocytes. J Neurochem 2004;88:623-634.
McIntire LB, Landman N, Kang MS, et al. Phenotypic assays for β-amyloid in mouse embryonic stem cell-derived neurons. Chem Biol 2013;20:956-967.
Blanchard BJ, Chen A, Rozeboom LM, Stafford KA, Weigele P, Ingram VM. Efficient reversal of Alzheimer’s disease fibril formation and elimination of neurotoxicity by a small molecule. Proc Natl Acad Sci U S A 2004;101:14326-14332.
McKoy AF, Chen J, Schupbach T, Hecht MH. A novel inhibitor of amyloid β (Aβ) peptide aggregation: from high throughput screening to efficacy in an animal model of Alzheimer disease. J Biol Chem 2012;287:38992-39000.
Walker LC, Diamond MI, Duff KE, Hyman BT. Mechanisms of protein seeding in neurodegenerative diseases. JAMA Neurol 2013;70:304-310.
Holmes BB, Diamond MI. Prion-like properties of tau protein: the importance of extracellular tau as a therapeutic target. J Biol Chem 2014;289:19855-19861.
Sharma P, Ando DM, Daub A, Kaye JA, Finkbeiner S. High-throughput screening in primary neurons. Methods Enzymol 2012;506:331-360.
Xu X, Lei Y, Luo J, et al. Prevention of β-amyloid induced toxicity in human iPS cell-derived neurons by inhibition of Cyclin-dependent kinases and associated cell cycle events. Stem Cell Res 2013;10:213-227.
Cundiff PE, Anderson SA. Impact of induced pluripotent stem cells on the study of central nervous system disease. Curr Opin Genet Dev 2011;21:354-361.
Marchetto MC, Gage FH. Modeling brain disease in a dish: really? Cell Stem Cell 2012;10:642-645.
Blanpain C, Daley GQ, Hochedlinger K, Passegué E, Rossant J, Yamanaka S. Stem cells assessed. Nat Rev Mol Cell Biol 2012;13:471-476.
Grskovic M, Javaherian A, Strulovici B, Daley GQ. Induced pluripotent stem cells—opportunities for disease modelling and drug discovery. Nat Rev Drug Discov 2011;10:915-929.
Ebert AD, Svendsen CN. Human stem cells and drug screening: opportunities and challenges. Nat Rev Drug Discov 2010;9:367-372.
Han SS, Williams LA, Eggan KC. Constructing and deconstructing stem cell models of neurological disease. Neuron 2011;70:626-644.
Yagi T, Ito D, Okada Y, et al. Modeling familial Alzheimer’s disease with induced pluripotent stem cells. Hum Mol Genet 2011;20:4530-4539.
Yahata N, Asai M, Kitaoka S, et al. Anti-Aβ drug screening platform using human iPS cell-derived neurons for the treatment of Alzheimer’s disease. PLoS One 2011;6:e25788.
Israel MA, Yuan SH, Bardy C, et al. Probing sporadic and familial Alzheimer’s disease using induced pluripotent stem cells. Nature 2012;482:216-220.
Shi Y, Kirwan P, Smith J, MacLean G, Orkin SH, Livesey FJ. A human stem cell model of early Alzheimer’s disease pathology in Down syndrome. Sci Transl Med 2012;4:124ra29.
Koch P, Tamboli IY, Mertens J, et al. Presenilin-1 L166P mutant human pluripotent stem cell-derived neurons exhibit partial loss of γ-secretase activity in endogenous amyloid-β generation. Am J Pathol 2012;180:2404-2416.
Kondo T, Asai M, Tsukita K, et al. Modeling Alzheimer’s disease with iPSCs reveals stress phenotypes associated with intracellular Aβ and differential drug responsiveness. Cell Stem Cell 2013;12:487-496.
Vazin T, Ball KA, Lu H, et al. Efficient derivation of cortical glutamatergic neurons from human pluripotent stem cells: a model system to study neurotoxicity in Alzheimer’s disease. Neurobiol Dis 2014;62:62-72.
Zhang D, Pekkanen-Mattila M, Shahsavani M, Falk A, Teixeira AI, Herland A. A 3D Alzheimer’s disease culture model and the induction of P21-activated kinase mediated sensing in iPSC derived neurons. Biomaterials 2014;35:1420-1428.
Choi SH, Kim YH, Hebisch M, et al. A three-dimensional human neural cell culture model of Alzheimer’s disease. Nature 2014;515:274-278.
Young JE, Goldstein LS. Alzheimer’s disease in a dish: promises and challenges of human stem cell models. Hum Mol Genet 2012;21:R82-R89.
Guo Q, Wang Z, Li H, Wiese M, Zheng H. APP physiological and pathophysiological functions: insights from animal models. Cell Res 2012;22:78-89.
McColl G, Roberts BR, Pukala TL, et al. Utility of an improved model of amyloid-beta (Aβ1–42) toxicity in Caenorhabditis elegans for drug screening for Alzheimer’s disease. Mol Neurodegener 2012;7:57.
Matlack KE, Tardiff DF, Narayan P, et al. Clioquinol promotes the degradation of metal-dependent amyloid-β (Aβ) oligomers to restore endocytosis and ameliorate Aβ toxicity. Proc Natl Acad Sci U S A 2014;111:4013-4018.
Singh N, Halliday AC, Thomas JM, et al. A safe lithium mimetic for bipolar disorder. Nat Commun 2013;4:1332.
Dubus E, Ijjaali I, Barberan O, Petitet F. Drug repositioning using in silico compound profiling. Future Med Chem 2009;1:1723-1736.
Dudley JT, Sirota M, Shenoy M, et al. Computational repositioning of the anticonvulsant topiramate for inflammatory bowel disease. Sci Transl Med 2011;3:96ra76.
Hurle MR, Yang L, Xie Q, Rajpal DK, Sanseau P, Agarwal P. Computational drug repositioning: from data to therapeutics. Clin Pharmacol Ther 2013;93:335-341.
Harrold JM, Ramanathan M, Mager DE. Network-based approaches in drug discovery and early development. Clin Pharmacol Ther 2013;94:651-658.
Lamb J, Crawford ED, Peck D, et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006;313:1929-1935.
Lamb J. The Connectivity Map: a new tool for biomedical research. Nat Rev Cancer 2007;7:54-60.
Sirota M, Dudley JT, Kim J, et al. Discovery and preclinical validation of drug indications using compendia of public gene expression data. Sci Transl Med 2011;3:96ra77.
Williams G. A searchable cross-platform gene expression database reveals connections between drug treatments and disease. BMC Genomics. 2012;13:12.
Sanseau P, Agarwal P, Barnes MR, Pastinen T, Richards JB, Cardon LR, Mooser V. Use of genome-wide association studies for drug repositioning. Nat Biotechnol 2012;30:317-320.
Cao C, Moult J. GWAS and drug targets. BMC Genomics 2014;15:S5.
Stefansson H, Steinberg S, Petursson H, et al. Variant in the sequence of the LINGO1 gene confers risk of essential tremor. Nat Genet 2009;41:277-279.
Clark LN, Park N, Kisselev S, Rios E, Lee JH, Louis ED. Replication of the LINGO1 gene association with essential tremor in a North American population. Eur J Hum Genet 2010;18:838-843.
Heinrich M, Lee Teoh H. Galanthamine from snowdrop—the development of a modern drug against Alzheimer’s disease from local Caucasian knowledge. J Ethnopharmacol 2004;92:147-162.
Wang J, Zhao Z, Lin E, et al. Unintended effects of cardiovascular drugs on the pathogenesis of Alzheimer’s disease. PLoS One 2013;8:e65232
Chow TW, Pollock BG, Milgram NW. Potential cognitive enhancing and disease modification effects of SSRIs for Alzheimer’s disease. Neuropsychiatr Dis Treat 2007;3:627-636.
Cummings JL, Zhong K. Treatments for behavioural disorders in neurodegenerative diseases: drug development strategies. Nat Rev Drug Discov 2006;5:64-74.
Nitsch RM, Deng M, Growdon JH, Wurtman RJ. Serotonin 5-HT2a and 5-HT2c receptors stimulate amyloid precursor protein ectodomain secretion. J Biol Chem 1996;271:4188-4194.
Arjona AA, Pooler AM, Lee RK, Wurtman RJ. Effect of a 5-HT(2C) serotonin agonist, dexnorfenfluramine, on amyloid precursor protein metabolism in guinea pigs. Brain Res 2002;951:135-140.
Shen F, Smith JA, Chang R, et al. 5-HT(4) receptor agonist mediated enhancement of cognitive function in vivo and amyloid precursor protein processing in vitro: a pharmacodynamic and pharmacokinetic assessment. Neuropharmacology 2011;61:69-79.
Pákáski M, Bjelik A, Hugyecz M, Kása P, Janka Z, Kálmán J. Imipramine and citalopram facilitate amyloid precursor protein secretion in vitro. Neurochem Int 2005;47:190-195.
Nelson RL, Guo Z, Halagappa VM, et al. Prophylactic treatment with paroxetine ameliorates behavioral deficits and retards the development of amyloid and tau pathologies in 3xTgAD mice. Exp Neurol 2007;205:166-176.
Cirrito JR, Disabato BM, Restivo JL, et al. Serotonin signaling is associated with lower amyloid-β levels and plaques in transgenic mice and humans. Proc Natl Acad Sci U S A 2011;108:14968-14973.
Sheline YI, West T, Yarasheski K, et al. An antidepressant decreases CSF Aβ production in healthy individuals and in transgenic AD mice. Sci Transl Med 2014;6:236re4.
Verret L, Mann EO, Hang GB, et al. Inhibitory interneuron deficit links altered network activity and cognitive dysfunction in Alzheimer model. Cell 2012;149:708-721.
Vossel KA, Beagle AJ, Rabinovici GD, et al. Seizures and epileptiform activity in the early stages of Alzheimer disease. JAMA Neurol 2013;70:1158-1166.
Sanchez PE, Zhu L, Verret L, et al. Levetiracetam suppresses neuronal network dysfunction and reverses synaptic and cognitive deficits in an Alzheimer’s disease model. Proc Natl Acad Sci U S A 2012;109:E2895-E2903.
Shi JQ, Wang BR, Tian YY, et al. Antiepileptics topiramate and levetiracetam alleviate behavioral deficits and reduce neuropathology in APPswe/PS1dE9 transgenic mice. CNS Neurosci Ther 2013;19:871-881.
Mark RJ, Ashford JW, Goodman Y, Mattson MP. Anticonvulsants attenuate amyloid beta-peptide neurotoxicity, Ca2+ deregulation, and cytoskeletal pathology. Neurobiol Aging 1995;16:187-198.
Qing H, He G, Ly PT, et al. Valproic acid inhibits Abeta production, neuritic plaque formation, and behavioral deficits in Alzheimer’s disease mouse models. J Exp Med 2008;205:2781-2789.
Smith AM, Gibbons HM, Dragunow M. Valproic acid enhances microglial phagocytosis of amyloid-beta(1-42). Neuroscience 2010;169:505-515.
Wang Z, Zhang XJ, Li T, Li J, Tang Y, Le W. Valproic acid reduces neuritic plaque formation and improves learning deficits in APP(Swe) /PS1(A246E) transgenic mice via preventing the prenatal hypoxia-induced down-regulation of neprilysin. CNS Neurosci Ther 2014;20:209-217.
Long Z, Zheng M, Zhao L, et al. Valproic acid attenuates neuronal loss in the brain of APP/PS1 double transgenic Alzheimer’s disease mice model. Curr Alzheimer Res 2013;10:261-269.
Tariot PN, Schneider LS, Cummings J, et al.; Alzheimer’s Disease Cooperative Study Group. Chronic divalproex sodium to attenuate agitation and clinical progression of Alzheimer disease. Arch Gen Psychiatry 2011;68:853-861.
Skoog I, Lernfelt B, Landahl S, et al. 15-year longitudinal study of blood pressure and dementia. Lancet 1996;347:1141-1145.
Posner HB, Tang MX, Luchsinger J, Lantigua R, Stern Y, Mayeux R. The relationship of hypertension in the elderly to AD, vascular dementia, and cognitive function. Neurology 2002;58:1175-1181.
Gottesman RF, Schneider AL, Albert M, et al. Midlife hypertension and 20-year cognitive change: the atherosclerosis risk in communities neurocognitive study. JAMA Neurol 2014;71:1218–1227.
Yamada K, Uchida S, Takahashi S, et al. Effect of a centrally active angiotensin-converting enzyme inhibitor, perindopril, on cognitive performance in a mouse model of Alzheimer’s disease. Brain Res 2010;1352:176-186.
Dong YF, Kataoka K, Tokutomi Y, et al. Perindopril, a centrally active angiotensin-converting enzyme inhibitor, prevents cognitive impairment in mouse models of Alzheimer’s disease. FASEB J 2011;25:2911-2920.
Khachaturian AS, Zandi PP, Lyketsos CG, et al. Antihypertensive medication use and incident Alzheimer disease: the Cache County Study. Arch. Neurol 2006;63:686-692.
Tzourio C, Anderson C, Chapman N, Woodward M, Neal B, MacMahon S, Chalmers J. Effects of blood pressure lowering with perindopril and indapamide therapy on dementia and cognitive decline in patients with cerebrovascular disease. Arch Intern Med 2003;163:1069-1075.
Paris D, Bachmeier C, Patel N, et al. Selective antihypertensive dihydropyridines lower Aβ accumulation by targeting both the production and the clearance of Aβ across the blood-brain barrier. Mol Med 2011;17:149-162.
Bachmeier C, Beaulieu-Abdelahad D, Mullan M, Paris D. Selective dihydropyiridine compounds facilitate the clearance of β-amyloid across the blood–brain barrier. Eur J Pharmacol 2011;659:124-129.
Paris D, Quadros A, Humphrey J, et al. Nilvadipine antagonizes both Abeta vasoactivity in isolated arteries, and the reduced cerebral blood flow in APPsw transgenic mice. Brain Res 2004;999:53-61.
Hanyu H, Hirao K, Shimizu S, Sato T, Kiuchi A, Iwamoto T. Nilvadipine prevents cognitive decline of patients with mild cognitive impairment. Int J Geriatr Psychiatry 2007;22:1264-1266.
Matsuda H, Araki N, Kuji I, Ohkubo T, Imabayashi E, Shimazu K. Effect of nilvadipine on regional cerebral blood flow in a patient with early Alzheimer disease. Clin Nucl Med 2008;33:34-35.
Hanyu H, Hirao K, Shimizu S, Iwamoto T, Koizumi K, Abe K. Favourable effects of nilvadipine on cognitive function and regional cerebral blood flow on SPECT in hypertensive patients with mild cognitive impairment. Nucl Med Commun 2007;28:281-287.
Kennelly S, Abdullah L, Kenny RA, et al. Apolipoprotein E genotype-specific short-term cognitive benefits of treatment with the antihypertensive nilvadipine in Alzheimer’s patients–an open-label trial. Int J Geriatr Psychiatry 2012;27:415-422.
Anekonda TS, Quinn JF, Harris C, Frahler K, Wadsworth TL, Woltjer RL. L-type voltage-gated calcium channel blockade with isradipine as a therapeutic strategy for Alzheimer’s disease. Neurobiol Dis 2011;41:62-70.
Anekonda TS, Quinn JF. Calcium channel blocking as a therapeutic strategy for Alzheimer’s disease: the case for isradipine. Biochim Biophys Acta 2011;1812:1584-1590.
Li NC, Lee A, Whitmer RA, et al. Use of angiotensin receptor blockers and risk of dementia in a predominantly male population: prospective cohort analysis. BMJ 2010;340:b5465.
Davies NM, Kehoe PG, Ben-Shlomo Y, Martin RM. Associations of anti-hypertensive treatments with Alzheimer’s disease, vascular dementia, and other dementias. J Alzheimers Dis 2011;26:699-708.
Wang J, Ho L, Chen L, et al. Valsartan lowers brain beta-amyloid protein levels and improves spatial learning in a mouse model of Alzheimer disease. J Clin Invest 2007;117:3393-3402.
Danielyan L, Klein R, Hanson LR, et al. Protective effects of intranasal losartan in the APP/PS1 transgenic mouse model of Alzheimer disease. Rejuvenation Res 2010;13:195-201.
Lithell H, Hansson L, Skoog I, et al; SCOPE Study Group. The Study on Cognition and Prognosis in the Elderly (SCOPE): principal results of a randomized double-blind intervention trial. J Hypertens 2003;21:875-886.
Anderson C, Teo K, Gao P, et al.; ONTARGET and TRANSCEND Investigators. Renin-angiotensin system blockade and cognitive function in patients at high risk of cardiovascular disease: analysis of data from the ONTARGET and TRANSCEND studies. Lancet Neurol 2011;10:43-53.
Howlett DR, George AR, Owen DE, Ward RV, Markwell RE. Common structural features determine the effectiveness of carvedilol, daunomycin and rolitetracycline as inhibitors of Alzheimer beta-amyloid fibril formation. Biochem J 1999;343:419-423.
Wang J, Ono K, Dickstein DL, et al. Carvedilol as a potential novel agent for the treatment of Alzheimer’s disease. Neurobiol Aging 2011;32:2321.e1-12.
Yang Y, Song W. Molecular links between Alzheimer’s disease and diabetes mellitus. Neuroscience 2013;250:140-150.
Butterfield DA, Di Domenico F, Barone E. Elevated risk of type 2 diabetes for development of Alzheimer disease: a key role for oxidative stress in brain. Biochim Biophys Acta 2014;1842:1693-1706.
Devi L, Alldred MJ, Ginsberg SD, Ohno M. Mechanisms underlying insulin deficiency-induced acceleration of β-amyloidosis in a mouse model of Alzheimer’s disease. PLoS One 2012;7:e32792.
Craft S, Baker LD, Montine TJ, et al. Intranasal insulin therapy for Alzheimer disease and amnestic mild cognitive impairment: a pilot clinical trial. Arch Neurol 2012;69:29-38.
Gengler S, McClean PL, McCurtin R, Gault VA, Hölscher C. Val(8)GLP-1 rescues synaptic plasticity and reduces dense core plaques in APP/PS1 mice. Neurobiol Aging 2012;33:265-276.
McClean PL, Parthsarathy V, Faivre E, Hölscher C. The diabetes drug liraglutide prevents degenerative processes in a mouse model of Alzheimer’s disease. J Neurosci 2011;31:6587-6594.
McClean PL, Hölscher C. Liraglutide can reverse memory impairment, synaptic loss and reduce plaque load in aged APP/PS1 mice, a model of Alzheimer’s disease. Neuropharmacology 2014;76:57-67.
Hunter K, Hölscher C. Drugs developed to treat diabetes, liraglutide and lixisenatide, cross the blood brain barrier and enhance neurogenesis. BMC Neurosci 2012;13:33.
Chen Y, Zhou K, Wang R, et al. Antidiabetic drug metformin (GlucophageR) increases biogenesis of Alzheimer’s amyloid peptides via up-regulating BACE1 transcription. Proc Natl Acad Sci U S A 2009;106:3907-3912.
Gupta A, Bisht B, Dey CS. Peripheral insulin-sensitizer drug metformin ameliorates neuronal insulin resistance and Alzheimer’s-like changes. Neuropharmacology 2011;60:910-920.
Li J, Deng J, Sheng W, Zuo Z. Metformin attenuates Alzheimer’s disease-like neuropathology in obese, leptin-resistant mice. Pharmacol Biochem Behav 2012;101:564-574.
Kickstein E, Krauss S, Thornhill P, et al. Biguanide metformin acts on tau phosphorylation via mTOR/protein phosphatase 2A (PP2A) signaling. Proc Natl Acad Sci U S A 2010;107:21830-21835.
Vlachopoulos C, Terentes-Printzios D, Ioakeimidis N, Rokkas K, Stefanadis C. PDE5 inhibitors in non-urological conditions. Curr Pharm Des 2009;15:3521-3539.
Puzzo D, Staniszewski A, Deng SX, et al. Phosphodiesterase 5 inhibition improves synaptic function, memory, and amyloid-beta load in an Alzheimer’s disease mouse model. J Neurosci 2009;29:8075-8086.
García-Barroso C, Ricobaraza A, Pascual-Lucas M, et al. Tadalafil crosses the blood-brain barrier and reverses cognitive dysfunction in a mouse model of AD. Neuropharmacology 2013;64:114-123.
Cuadrado-Tejedor M, Hervias I, Ricobaraza A, et al. Sildenafil restores cognitive function without affecting β-amyloid burden in a mouse model of Alzheimer’s disease. Br J Pharmacol 2011;164:2029-2041.
Zhang J, Guo J, Zhao X, et al. Phosphodiesterase-5 inhibitor sildenafil prevents neuroinflammation, lowers beta-amyloid levels and improves cognitive performance in APP/PS1 transgenic mice. Behav Brain Res 2013;250:230-237.
García-Osta A, Cuadrado-Tejedor M, García-Barroso C, Oyarzábal J, Franco R. Phosphodiesterases as therapeutic targets for Alzheimer’s disease. ACS Chem Neurosci 2012;3:832-844.
Allison M. NCATS launches drug repurposing program. Nat Biotechnol 2012;30:571-572.
Hennequin LF, Ballard P, Boyle FT, et al. Novel 4-anilinoquinazolines with C-6 carbon-linked side chains: synthesis and structure-activity relationship of a series of potent, orally active, EGF receptor tyrosine kinase inhibitors. Bioorg Med Chem Lett 2006;16:2672-2676.
Nygaard HB, van Dyck CH, Strittmatter SM. Fyn kinase inhibition as a novel therapy for Alzheimer’s disease. Alzheimers Res Ther 2014;6:8.
Roberson ED, Halabisky B, Yoo JW, et al. Amyloid-β/Fyn-induced synaptic, network, and cognitive impairments depend on tau levels in multiple mouse models of Alzheimer’s disease. J Neurosci 2011;31:700-711.
Larson M, Sherman MA, Amar F, et al. The complex PrP(c)-Fyn couples human oligomeric Aβ with pathological tau changes in Alzheimer’s disease. J Neurosci 2012;32:16857-16871a.
Um JW, Nygaard HB, Heiss JK, et al. Alzheimer amyloid-β oligomer bound to postsynaptic prion protein activates Fyn to impair neurons. Nat Neurosci 2012;15:1227-1235.
Shineman DW, Alam J, Anderson M, et al. Overcoming obstacles to repurposing for neurodegenerative disease. Ann Clin Transl Neurol 2014;1:512-518.
Rai AK, Rice G. Use patents can be useful: the case of rescued drugs. Sci Transl Med 2014;6:248fs30.
Burness CB, Deeks ED. Dimethyl fumarate: a review of its use in patients with relapsing-remitting multiple sclerosis. CNS Drugs 2014;28:373-387.
Sem DS. Repurposing - finding new uses for old (and patented) drugs: bridging the “valley of death,” to translate academic research into new medicines, 18 Marq. Intellectual Property L Rev 2014;18:139-166.
Acknowledgments
This work was supported by grants from National Institutes of Health (NS074536), BrightFocus Foundation, and Alzheimer’s Drug Discovery Foundation. I thank Laura Beth McIntire and Gina M. Finan for critical comments on the manuscript, and Meewon Park for editorial assistance.
Required Author Forms
Disclosure forms provided by the authors are available with the online version of this article.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1224 kb)
Rights and permissions
About this article
Cite this article
Kim, TW. Drug Repositioning Approaches for the Discovery of New Therapeutics for Alzheimer’s Disease. Neurotherapeutics 12, 132–142 (2015). https://doi.org/10.1007/s13311-014-0325-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13311-014-0325-7