Abstract
Given that only a small proportion of women infected by high-risk human papillomavirus (hrHPV) develop cervical cancer, it’s important to identify biomarkers for distinguishing women with hrHPV positivity who might develop cervical cancer from the transient infections. In this study, we hypothesized that human leukocyte antigens (HLA) susceptibility alleles might contribute to cervical cancer risk among females infected by hrHPV, and interact with hrHPV types. A case-control study with 593 cervical cancer cases and 407 controls (all hrHPV positive) was conducted to evaluate the effect of eight HLA-related single-nucleotide polymorphisms (SNPs) and their interactions with hrHPV types on the risk of cervical cancer. Three HLA-DP SNPs (rs4282438, rs3117027, and rs3077) were found to be significantly associated with risk of cervical cancer (rs4282438: odds ratio (OR) = 0.72, 95 % confidence interval (CI) = 0.56–0.93; rs3117027: OR = 1.41, 95 % CI = 1.10–1.83; and rs3077: OR = 1.37, 95 % CI = 1.04–1.80) among women infected with hrHPV. An additive interaction between HPV16 and rs4282438 for cervical cancer risk was also found (P for interaction = 0.002). Compared with subjects carrying variant genotypes (GG/TG) and non-HPV16 infections, those carrying wild-type genotype (TT) of rs4282438 and HPV16 positive had a 5.22-fold increased risk of cervical cancer (95 % CI = 3.39–8.04). Our study supported that certain HLA-DP alleles in concert with HPV16 could have a predisposition for cervical cancer development, which may be translated for triage of hrHPV-positive women.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cervical cancer is the third most commonly diagnosed cancer and the fourth leading cause of cancer deaths among women worldwide, and nearly 85 % of cervical cancer cases and deaths occurred in developing countries [1]. Human papillomavirus (HPV) infections, very common among general females, are necessary [2, 3] but not sufficient for the development of cervical cancer. Studies have shown that up to 22 % of general population worldwide are HPV positive [4], less than 10 % HPV infections become persistent [5, 6], and fewer than 4 % of individuals with HPV positivity develop premalignant lesion, and even fewer develop invasive cancer [7, 8]. Therefore, although high-risk HPV (hrHPV) testing has been incorporated into screening programs in many countries for its high sensitivity, the mediocre specificity and positive predictive value for cervical cancer identification resulting from the resolve of most HPV infections in a few months or years [9], makes it necessary to look for a better way of the triage of hrHPV-positive women. Cytology is the widely preferred tool for the triage of hrHPV-positive women but is restricted by the subjective diagnosis of pathologists mostly as a morphology-based method [10]. Identification of effective and objective biomarkers to make up for the deficiency of hrHPV test is needed to improve the efficiency of cervical cancer prevention.
The pathogenesis of cervical cancer is not entirely clear but is closely related with the virus-specific host immunity [11]. It has been reported that the human leukocyte antigens (HLA) system plays a pivotal role in presenting foreign peptides to antigen-specific T cell receptors that are responsible for clearance of virus-infected cells and tumor cells [12]. Our recent study found two single-nucleotide polymorphisms (SNPs) of HLA-DP (rs3077 and rs9277535) associated with increased risk of cervical cancer among general Chinese women using a candidate gene approach [13]. Meanwhile, two genome-wide association studies (GWAS) of cervical cancer also identified susceptibility locus on HLA genes in both the Chinese and European populations [14, 15]. The previous studies suggested that genetic variations of HLA genes could be used as candidate susceptibility biomarkers for cervical cancer among Chinese females [13]. However, whether these host genetic variations could be potentially used to distinguish the hrHPV-positive women who would develop cervical cancer from that who would regress, and whether these SNPs could have interaction with HPV types, still needs to be studied.
In this study, we performed a case-control study to explore association between candidate SNPs of HLA genes and the risk of cervical cancer after hrHPV acquisition, and to investigate the possible interaction with certain HPV types. The study was designed to enroll both cases and controls with hrHPV positivity and the findings would add important value for decision-making among the hrHPV-positive women for further evaluation or treatment [16].
Materials and methods
Study population and design
This study was approved by the ethics committee of Cancer Institute and Hospital, Chinese Academy of Medical Sciences and Nanjing Medical University. Written consent was obtained from each study subject. Cervical cancer cases were recruited from Cancer Institute and Hospital, Chinese Academy of Medical Sciences, between January 2010 and July 2013. All the cervical cancer cases in the study were newly diagnosed and histologically confirmed. The control subjects were selected from the women who participated in the physical examination in the Department of Cancer Prevention, Cancer Institute and Hospital, Chinese Academy of Medical Sciences, between January 2010 and July 2013 and were frequency matched to the cases by age (±5 years). All controls have any liquid-based cytologically (LBC) confirmed abnormal intraepithelial lesions (atypical squamous cells of undetermined significance, ASCUS, or worse) and self-reported cancer history. In addition, hrHPV positivity was one of the most important criteria for both cases and controls. Finally, a total of 593 cancer cases and 407 controls were included in the study. The hrHPV prevalence in cases and controls was 97.69 % (593/607) and 8.03 % (407/5066), respectively.
Data collection
Peripheral blood samples and exfoliated cervical cells were collected. Special-trained physicians and nurses conducted in-person interview using a standardized questionnaire.
SNP and HPV genotyping
Eight recently reported HLA-related SNPs (HLA-DP rs3117027, HLA-DP rs9277535, HLA-DP rs4282438, HLA-DP rs3077, HLA-DP rs9277535, HLA-DQ rs2856718, HLA-DQ rs7453920, and HLA-B/MICA rs2516488) were derived from recently published GWAS and candidate gene-based study [17–19] (Supplemental Fig. 1). All of the SNPs were genotyped by TaqMan assays except for one by direct sequence (primers and probes shown in Supplemental Table 1).
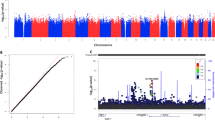
HPV DNA was extracted from exfoliated cervical cells. Genotyping was performed twice for both cases and controls. Firstly, 21 HPV types (including 15 hrHPV types: HPV16, 18, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, and 68; and 6 low-risk HPV types: HPV6, 11, 42, 43, 44, and 81) were tested in the Department of Clinical Laboratory, Cancer Institute and Hospital, Chinese Academy of Medical Sciences, by HPV GenoArray Test Kit (Hybribio company) preliminarily. Then, positive hrHPV DNA was re-evaluated by type-specific RCR (primers shown in Supplemental Table 2) in the Department of Epidemiology, Jiangsu Key Lab of the Cancer Biomarkers, Prevention and Treatment, Nanjing Medical University (Fig. 1 and Supplemental Fig. 2). Subjects with consistent results of hrHPV types were included in the present study. Each hrHPV type was confirmed by direct sequencing (Supplemental Fig. 3).
All the SNPs and HPV DNA were genotyped by experienced researchers, blinded to the status of cases and controls. For the purpose of quality control, two blank (water) samples were added in each 384-well plate. Ten percent repeated samples were randomly selected from both cases and controls for SNP genotyping.
Statistical analyses
Student’s t test or χ 2 tests were used to evaluate differences in the distributions of age and hrHPV types between the cases and controls. Hardy-Weinberg equilibrium (HWE) was tested by a goodness-of-fit χ 2 test to compare the observed genotype frequencies to the expected ones among the control subjects. Unconditional logistic regression model was used to calculate the odds ratios (ORs) and 95 % confidence intervals (CIs). The dominant genetic models compared the variant allele homozygote and heterozygote to the common allele which served as the reference group. A linear trend test assuming an additive genetic model was conducted by assigning an ordinal value of 1, 2, or 3 corresponding to the homozygous wild-type, heterozygote, and homozygous variant genotype, respectively. Interactions between SNPs of genes and HPV genotypes were determined if it followed an additive or multiplicative model. Statistical analyses were conducted using R package and Stata Version 10.0 software (Stata, College Station, TX). P < 0.05 was the criterion of statistical significance, and all statistical tests were two sided.
Results
Of the 593 cervical cancer cases, 544 (91.74 %) were squamous cell carcinoma, 24 (4.04 %) were adenocarcinoma, and 5 (0.84 %) were adenosquamous carcinoma. Cases and controls had similar mean age (48.09 ± 0.46 years for cases and 48.81 ± 0.34 years for controls, P = 0.197).
Among 15 hrHPV types, 14 types (including HPV16, 18, 31, 33, 35, 39, 45, 51, 52, 56, 58, 59, 66, and 68) were detected in the study population (Fig. 1 and Supplemental Figs. 1 and 2). HPV16, 18, 52, and 58 were four common types in both cases and controls. More than half of the cases were infected by HPV16 (54.81 %) while only one fourth of the controls were infected by HPV16 (25.80 %) (Table 1). HPV16 was the unique hrHPV type in which prevalence was significantly higher in cases than that in controls (P < 0.001).
Of the observed genotypes frequencies for eight SNPs in control subjects, five SNPs were in Hardy-Weinberg equilibrium (HWE, P = 0.740 for rs4282438; P = 0.200 for rs3117027; P = 0.756 for rs3077; P = 1.00 for rs9277535, and P = 0.486 for rs2856718) except the other three SNPs (P = 0.016 for rs2516448; P = 0.013 for rs9272143; and P = 0.009 for rs7453920). Associations between these SNPs and cervical cancer risk among women infected by hrHPV were presented in Table 2. Three SNPs (rs4282438, rs3117027, and rs3077) in HLA-DP gene were significantly associated with the risk of cervical cancer in dominant model (rs4282438: adjusted OR = 0.72, 95 % CI = 0.56–0.93; rs3117027: adjusted OR = 1.41, 95 % CI = 1.10–1.83; rs3077: adjusted OR = 1.37, 95 % CI = 1.04–1.80). HLA-DP rs9277535 also showed suggestive association with cervical cancer risk (Table 2).
We further analyzed the combined effect between HPV16 infection and the three SNPs (rs4282438, rs3117027, and rs3077) in HLA-DP since HPV16 was the unique hrHPV type which prevalence was higher in cervical cancer cases than that in controls. The interaction between HPV16 infection and HLA-DP rs4282438 was found in additive model. Compared with G allele carriers of rs4382438 in the HLA-DP gene infected by non-HPV 16 types, TT carriers with HPV16 positivity had 5.22-fold increased risk of cervical cancer (OR = 5.22, 95 % CI = 3.39–8.04; P for interaction = 0.002) (Table 3).
Haplotype analyses did not show highly linkage disequilibrium (LD) for the three HLA-DP SNPs with all R 2 less than 0.1 (Supplemental Table 3).
Discussion
Several studies on HLA genetic variants and the risk of cervical cancer have been published, and one of the studies was performed by us [13]. Although these studies reported the risk of cervical cancer related to the genetic variants of HLA-DP, none of them considered the HPV status, either in the cases and controls. Thus, it was difficult to identify individuals who have higher risk of cervical cancer after hrHPV acquisitions, not to mention to explore the interaction between HLA genetic variant and hrHPV genotypes. Unlike previous studies (including those mentioned by the reviewer), we investigated cases and controls infected with hrHPV. We found that three SNPs of HLA-DP (rs3077, rs3117027, and rs4282438) were susceptible markers of cervical cancer for women with hrHPV infection. More importantly, a significant interaction was observed between HLA-DP rs4282438 and HPV16 infection. To the best of our knowledge, this is the first study to explore the interaction between genetic variations of HLA genes and HPV genotype on the risk of cervical cancer
Current studies widely believed that cervical cancer development after HPV infection largely depends on the genetic differences [17, 18]. Recently, Chen D conducted a systematic genome-wide association study to investigate the association between genetic variants in the HLA-DP region and cervical cancer risk. Their findings provided further evidence about the contribution of polymorphisms in the HLA-DP region to risk of cervical cancer [19]. In the present study, three HLA-DP SNPs (rs4282438, rs3117027, and rs3077) were found to be significantly associated with risk of cervical cancer (rs4282438: OR = 0.72, 95 % CI = 0.56–0.93; rs3117027: OR = 1.41, 95 % CI = 1.10–1.83; and rs3077: OR = 1.37, 95 % CI = 1.04–1.80) among women infected with hrHPV. An additive interaction between HPV16 and rs4282438 for cervical cancer risk (P for interaction = 0.002) was also found. Compared with subjects carrying variant genotypes (GG/TG) and non-HPV16 infections, those carrying wild-type genotype (TT) of rs4282438 and HPV16 positivity had a 5.22-fold increased risk of cervical cancer (95 % CI = 3.39–8.04). Our study supported that certain HLA-DP alleles in concert with HPV16 could have a predisposition for cervical cancer development, and the potential to be translated for triage of hrHPV-positive women. Above all, our results supported that those three SNPs may be associated with cervical cancer risk.
To explore the potential function of those three SNPs, we use the public database RegulomeDB (http://www.regulomedb.org) for predicting. Interestingly, we noticed that the rs3077 was the eQTL of HLA-DPA1. Furthermore, the rs3077 was located in the transcript factor binding site. The binding proteins include POLR2A, NT-ATC1, and ZNF263. Dai, J. et al. found that genetic variants in transcript factor binding site could influence the susceptibility to cancer [20].
The HLA system is the name of the loci of genes which encode for major histocompatibility complex (MHC) in humans including HLA-I, HLA-II, and HLA-III genes. There are three main HLA-II genes in humans (HLA-DP, HLA-DQ, and HLA-DR) encoding particular antigens that stimulate the multiplication of T helper cells, which in turn stimulate antibody-producing B cells to produce antibodies to that specific antigen [21, 22]. Series of studies have found that genetic variations of HLA-II genes were involved in neoplastic cervical disease and cancer development [23–26]. However, as we mentioned earlier, all the association between the SNPs of HLA-DP and the risk of cervical cancer were evaluated without the consideration of HPV infection status in these previous studies. For better understanding of the association between the host genetic variants and the risk of cervical cancer, the typical virus infection-related cancer, we did HPV DNA genotype in this study based on our previous research findings and further validated that HLA-DP rs4282438, rs3077, and rs3117027 increased the risk of cervical cancer among women infected by hrHPV and might play role in the progress of invasive cancer development, especially after hrHPV acquisition. More importantly, the interaction between rs4282438 and HPV16 was found in this study, which further validated the genotype-infection correlation in the development of cervical cancer and the explained partly why hrHPV infection is only a necessary but not a sufficient cause of cervical cancer. HLA-DP, composed of 2 subunits (DPα and DPβ), belongs to HLA class II molecules. HLA-DP was expressed as cell surface glycoproteins that bind and present foreign peptides such as virus antigens to CD4+ T cells [27]. The rs4282438 are located at 3ʹUTR of HLA-DPA1, intron of HLA-DPB1 and in the upstream of HLA-DPB/2, respectively. An in vitro study found that HPV16 E5 could decrease immune recognition of infected keratinocytes through function disruption of HLA class II proteins [28], which could partly explain the interaction between HLA-DP rs4282438 and HPV16 found in the present study.
The major strength of our study was that all controls were genotyped for HPV positivity with double check, which allowed us to investigate the association between related SNPs and cervical cancer especially among women with hrHPV positivity, and analysis interaction between host genetic variation and certain HPV genotype. Additionally, all of the controls in our study were diagnosed to be with normal cervix by LBC in hospital when they participated in physical check during the study period. Exclusion of subjects with cervical lesions by LBC diagnosis assured the minimum bias of misclassification. The prevalence of hrHPV in the controls of this study was 8.03 % (407/5066), which was similar to the prevalence reported in other Chinese populations [29, 30], validated, in some extent, the precise classification in the study. However, there are several limitations of the study which need to be solved in further research. Firstly, we conducted the case-control study for only one stage without validation. Further large-scale studies are required for validation. Secondly, the biological function of these three SNPs in HLA-DP was not further investigated in this study. The associations between the SNPs and the risk of cervical cancer among women infected by hrHPV should be interpreted with caution.
In conclusion, based on the findings of our previous study, the present study further shown that certain HLA-DP alleles could have a predisposition for cervical cancer development especially among women with hrHPV positivity. HLA-DP alleles (rs3077) could be potential biomarkers for the triage of hrHPV-positive women. Meanwhile, HLA-DP rs4282438, in concert with HPV16 infection, could be more important in the identification of high-risk population of cervical cancer among hrHPV-positive women. Further studies with different ethnic background, biological functional researches, and large sample size will help to further understand the role of immune regulation in HPV infection-related cervical cancer carcinogenesis.
Abbreviations
- hrHPV:
-
High-risk human papillomavirus
- SNPs:
-
Single-nucleotide polymorphisms
- HLA:
-
Human leukocyte antigens
- GWAS:
-
Genome-wide association studies
- LBC:
-
Liquid-based cytologically
- ASCUS:
-
Atypical squamous cells of undetermined significance
- Ors:
-
Odds ratios
- CIs:
-
Confidence intervals
- HWE:
-
Hardy-Weinberg equilibrium
- MHC:
-
Major histocompatibility complex
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61(2):69–90.
Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol. 1999;189(1):12–9.
zur Hausen H. Papillomaviruses and cancer: from basic studies to clinical application. Nat Rev Cancer. 2002;2(5):342–50.
de Sanjose S, Diaz M, Castellsague X, Clifford G, Bruni L, Munoz N, et al. Worldwide prevalence and genotype distribution of cervical human papillomavirus DNA in women with normal cytology: a meta-analysis. Lancet Infect Dis. 2007;7(7):453–9.
Ho GY, Bierman R, Beardsley L, Chang CJ, Burk RD. Natural history of cervicovaginal papillomavirus infection in young women. N Engl J Med. 1998;338(7):423–8.
Moscicki AB, Shiboski S, Broering J, Powell K, Clayton L, Jay N, et al. The natural history of human papillomavirus infection as measured by repeated DNA testing in adolescent and young women. J Pediatr. 1998;132(2):277–84.
Dalstein V, Riethmuller D, Pretet JL, Le Bail CK, Sautiere JL, Carbillet JP, et al. Persistence and load of high-risk HPV are predictors for development of high-grade cervical lesions: a longitudinal French cohort study. Int J Cancer. 2003;106(3):396–403.
Kulasingam SL, Hughes JP, Kiviat NB, Mao C, Weiss NS, Kuypers JM, et al. Evaluation of human papillomavirus testing in primary screening for cervical abnormalities: comparison of sensitivity, specificity, and frequency of referral. JAMA. 2002;288(14):1749–57.
Schiffman M, Wentzensen N, Wacholder S, Kinney W, Gage JC, Castle PE. Human papillomavirus testing in the prevention of cervical cancer. J Natl Cancer Inst. 2011;103(5):368–83.
Li N, Shi JF, Franceschi S, Zhang WH, Dai M, Liu B, et al. Different cervical cancer screening approaches in a Chinese multicentre study. Br J Cancer. 2009;100(3):532–7.
Hibma MH. The immune response to papillomavirus during infection persistence and regression. Open Virol J. 2012;6:241–8.
Horton R, Wilming L, Rand V, Lovering RC, Bruford EA, Khodiyar VK, et al. Gene map of the extended human MHC. Nat Rev Genet. 2004;5(12):889–99.
Jiang J, Li N, Shen Y, Liu J, Liu L, Du J, et al. Genetic variants in HLA-DP/DQ contribute to risk of cervical cancer: a two-stage study in Chinese women. Gynecol Oncol. 2013;129(2):401–5.
Chen D, Juko-Pecirep I, Hammer J, Ivansson E, Enroth S, Gustavsson I, et al. Genome-wide association study of susceptibility loci for cervical cancer. J Natl Cancer Inst. 2013;105(9):624–33.
Shi Y, Li L, Hu Z, Li S, Wang S, Liu J, et al. A genome-wide association study identifies two new cervical cancer susceptibility loci at 4q12 and 17q12. Nat Genet. 2013;45(8):918–22.
Sahasrabuddhe VV, Luhn P, Wentzensen N. Human papillomavirus and cervical cancer: biomarkers for improved prevention efforts. Future Microbiol. 2011;6(9):1083–98.
Chen D, Gyllensten U. Lessons and implications from association studies and post-GWAS analyses of cervical cancer. Trends Genet. 2015;31(1):41–54.
Chen D, Gyllensten U. Systematic investigation of contribution of genetic variation in the HLA-DP region to cervical cancer susceptibility. Carcinogenesis. 2014;35(8):1765–9.
Dai J, Zhu M, Wang C, Shen W, Zhou W, Sun J, et al. Systematical analyses of variants in CTCF-binding sites identified a novel lung cancer susceptibility locus among Chinese population. Sci Rep. 2015;5:7833.
Magnusson PK, Lichtenstein P, Gyllensten UB. Heritability of cervical tumours. Int J Cancer. 2000;88(5):698–701.
Shiina T, Inoko H, Kulski JK. An update of the HLA genomic region, locus information and disease associations: 2004. Tissue Antigens. 2004;64(6):631–49.
Biochemistry. Major-histocompatibility-complex proteins present peptide antigens on cell surfaces for recognition by T-cell receptors. This link leads to a site outside Genetics Home Reference. fifth edition. US National Library of Medicine. 2002.
Kohaar I, Hussain S, Thakur N, Tiwari P, Nasare V, Batra S, et al. Association between human leukocyte antigen class II alleles and human papillomavirus-mediated cervical cancer in Indian women. Hum Immunol. 2009;70(4):222–9.
Liang J, Xu A, Xie Y, Awonuga AO, Lin Z. Some but not all of HLA-II alleles are associated with cervical cancer in Chinese women. Cancer Genet Cytogenet. 2008;187(2):95–100.
Wu Y, Liu B, Lin W, Xu Y, Li L, Zhang Y, et al. Human leukocyte antigen class II alleles and risk of cervical cancer in China. Hum Immunol. 2007;68(3):192–200.
Zoodsma M, Nolte IM, Te Meerman GJ, De Vries EG, Van der Zee AG. HLA genes and other candidate genes involved in susceptibility for (pre)neoplastic cervical disease. Int J Oncol. 2005;26(3):769–84.
Diaz G, Amicosante M, Jaraquemada D, Butler RH, Guillen MV, Sanchez M, et al. Functional analysis of HLA-DP polymorphism: a crucial role for DPbeta residues 9, 11, 35, 55, 56, 69 and 84–87 in T cell allorecognition and peptide binding. Int Immunol. 2003;15(5):565–76.
Zhang B, Li P, Wang E, Brahmi Z, Dunn KW, Blum JS, et al. The E5 protein of human papillomavirus type 16 perturbs MHC class II antigen maturation in human foreskin keratinocytes treated with interferon-gamma. Virology. 2003;310(1):100–8.
Dai M, Bao YP, Li N, Clifford GM, Vaccarella S, Snijders PJ, et al. Human papillomavirus infection in Shanxi Province, People’s Republic of China: a population-based study. Br J Cancer. 2006;95(1):96–101.
Li LK, Dai M, Clifford GM, Yao WQ, Arslan A, Li N, et al. Human papillomavirus infection in Shenyang City, People’s Republic of China: a population-based study. Br J Cancer. 2006;95(11):1593–7.
Acknowledgments
This work was supported in part by the National Natural Science Fund from the National Natural Science Foundation of China (grant nos. 81172757, 81373079, and 81502873); Beijing Natural Science Foundation (grant nos. 7123225), Beijing Nova Program (grant nos. xx2012067); National Science Foundation for Distinguished Young Scholars of China (81225020), National Key Basic Research Program For Youth (2013CB911400), National Program for Support of Top-notch Young Professionals, and Natural Science Foundation of Jiangsu Province for Youth (grant no. BK20150997). We also acknowledge the support from Collaborative Innovation Center for Cancer Medicine.
Authors’ contributions
Dai Min, Li Ni, and Hu Zhibin directed the study, obtained financial support, and were responsible for the study design. Li Ni drafted the initial manuscript. Jia Meiqun, Han Jing, Hang Dong, Jiang Jie, Wang Minjie, Wei Baojun, Guo Lanwei, Zhang Kai, Qi Ju, and Ma Hongxia were responsible for samples processing and managed the genotyping data. Li Ni, Han Jing, Dai Juncheng, Jufang Shi, and Jiansong Ren were responsible for data management and statistically analysis. Dai Min and Hu Zhibin supervised the study design, and contributed to interpretation of results. All authors approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
None
Additional information
Meiqun Jia and Jing Han contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 1524 kb)
Rights and permissions
About this article
Cite this article
Jia, M., Han, J., Hang, D. et al. HLA-DP is the cervical cancer susceptibility loci among women infected by high-risk human papillomavirus: potential implication for triage of human papillomavirus-positive women. Tumor Biol. 37, 8019–8025 (2016). https://doi.org/10.1007/s13277-015-4673-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4673-7