Abstract
Background
Marine mammals, which have evolved independently into three distinct lineages, share common physiological features that contribute to their adaptation to the marine environment.
Objective
To identify positively selected genes (PSGs) for adaptation to the marine environment using available genomic data from three taxonomic orders: cetaceans, pinnipeds, and sirenians.
Methods
Based on the genomes within each group of Artiodactyla, Carnivora and Afrotheria, we performed selection analysis using the branch-site model in CODEML.
Results
Based on the branch-site model, 460, 614, and 359 PSGs were predicted for the cetaceans, pinnipeds, and sirenians, respectively. Functional enrichment analysis indicated that genes associated with hemostasis were positively selected across all lineages of marine mammals. We observed positive selection signals for the hemostasis and coagulation-related genes plasminogen activator, urokinase (PLAU), multimerin 1 (MMRN1), gamma-glutamyl carboxylase (GGCX), and platelet endothelial aggregation receptor 1 (PEAR1). Additionally, we found out that the sodium voltage-gated channel alpha subunit 9 (SCN9A), serine/arginine repetitive matrix 4 (SRRM4), and Ki-ras-induced actin-interacting protein (KRAP) are under positive selection pressure and are associated with cognition, neurite outgrowth, and IP3-mediated Ca2 + release, respectively.
Conclusion
This study will contribute to our understanding of the adaptive evolution of marine mammals by providing information on a group of candidate genes that are predicted to influence adaptation to aquatic environments, as well as their functional characteristics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Marine mammals, which migrated from terrestrial to aquatic environments, are fascinating subjects in the study of evolutionary adaptations to aquatic life. They are classified into several groups under three orders: Cetartiodactyla (cetaceans), Carnivora (pinnipeds and marine fissipeds), and Afrotheria (sirenians) (Berta et al. 2006). Cetaceans, pinnipeds, and sirenians share several common physiological and anatomical features that facilitated their adaptation to aquatic life, despite independently evolving from different ancestors. Marine mammals have unique adaptations to their hemostasis systems, which enable them to survive in aquatic environments. For instance, the deletion of certain blood coagulation factors (F12 and KLKB1) (Yim et al. 2014; Huelsmann et al. 2019) and an increase in activated partial thromboplastin time (aPTT) have been reported in cetaceans (Robinson et al. 1969). This delay in clotting prevents thrombosis and hypertension, which occur when a clot blocks the pulmonary artery. In contrast, the manatee possesses all blood coagulation factors (Barratclough et al. 2016) and tends to experience hypercoagulation compared to other land animals (Gerlach et al. 2015).
Positive selection analysis of genes can be performed to study the evolutionary changes resulting from adaptation. Several studies on the selective pressure of genes in marine mammals (Foote et al. 2015; Zhou et al. 2015; Chikina et al. 2016; Yuan et al. 2021) have used terrestrial mammals as background groups and performed positive selection studies on marine mammal groups. For instance, Chikina et al. (2016) found that marine-accelerated genes are involved in functions related to marine adaptation, including muscle physiology, lipid metabolism, and sensory systems. Yuan et al. (2021) reported analytical results for cetaceans, pinnipeds, and sirenians using 16 terrestrial mammalian species from diverse taxonomic orders. This study identified positively selected genes (PSGs) involved in thermoregulation and vision in marine mammals.
To determine the genetic distinctions between terrestrial and adaptive aquatic mammals within their respective orders, we conducted a comprehensive analysis of positive selection across all available genomes and genes in the three marine mammal groups. Moreover, we aimed to identify the commonly evolved characteristics across diverse groups of marine mammals. A total of 56 high-quality genomes from three orders were used. To determine the evolutionary differences between marine and terrestrial mammals, we performed a branch-site model analysis, revealing 460, 614, and 359 PSGs in cetaceans, pinnipeds, and sirenians, respectively. Through gene enrichment, network, and comparative analyses of the predicted PSGs, common genes and biological functions were identified as the underlying molecular bases for aquatic adaptations. Our primary objective was to broaden our understanding of aquatic adaptations in marine mammals and derive relevant genes.
Materials and methods
Data collection
From the NCBI RefSeq database, we retrieved 24 genomes of Cetartiodactyla (including twelve cetaceans), 24 genomes of Carnivora (including eight pinnipeds), and 6 genomes of Afrotheria (including one sirenian). In addition, two Xenarthra genomes, the closest relatives of Afrotheria, were added to the Afrotheria genome for statistical significance. Based on data from the gene ortholog database (ftp://ftp.ncbi.nlm.nih.gov/gene/DATA/gene_orthologs.gz), we extracted 17,898, 17,883, and 17,759 orthologous gene groups for Cetartiodactyla, Carnivora, and Afrotheria + Xenarthra, respectively, from the 56 genomes (Supplementary Fig. S1, Table S1).
Quality control pipeline for multiple sequence alignment
A multiple sequence alignment (MSA) with adequate accuracy is required for positive selection analysis. Due to the relatively low accuracy of MSA using the extracted coding sequences (CDSs) of the orthologous genes, multiple calculation methods were used (Supplementary Fig. S2). CDSs with frameshift exhibit poor MSA quality. To reduce frameshift errors, CDSs were converted into protein sequences and aligned using Clustal Omega. Subsequently, protein alignment was converted back into a CDS alignment using previous translation information, and CDSs with premature stop codons were excluded. The Clustal Omega guide tree option was used for the alignment. The NCBI taxonomy and TimeTree database were used to obtain the taxonomic information (Kumar et al. 2017). A guide tree for each MSA was generated in advance by replacing the pairwise alignment step with a faster and more accurate MSA step, for the clustering sequences. Using in-house programs to scan each CDS MSA, sequences with a low alignment quality were detected and excluded from the MSA. In addition, the front and back areas of the MSA with low alignment accuracy were trimmed to produce a high-quality multi-alignment region, which we termed Clean Segment Block (CSB). After manual inspection, low-quality sequences or alignment sites that were not detected by the scan programs were identified and adjusted for realignment. The final MSA, obtained after completing the aforementioned steps, was used as the input for positive selection analysis (Supplementary Fig. S2).
Positive selection analysis
The branch-site model of the CODEML program in the PAML 4.9j (Yang 2007) package was used to predict PSGs. An ancestral branch of cetaceans, pinnipeds, and sirenians was set as the independent foreground in each of the Artiodactyla, Carnivora, and Afrotheria + Xenarthra datasets, using a branch-site model. To ensure statistical reliability, analysis was performed only when seven or more species were included in the CSB for each orthologous gene dataset.
The significance of the likelihood difference (2ΔL) between the null hypothesis model (ω = 1) and the alternative hypothesis model was evaluated using a chi-square distribution with np2-np1 degrees of freedom. Using empirical Bayes analysis, all genes that rejected the null hypothesis and adopted the alternative hypothesis presented positively selected codons in the foreground branch. After performing false discovery rate (FDR) correction for multiple testing, the PSGs used for analysis were filtered using a cutoff value of FDR < 0.05.
Prediction of biological functions of PSGs
To investigate the functional distribution of PSGs in each group of marine mammals, we used clusterProfiler (Yu et al. 2012) to perform gene enrichment analysis.
For a more integrated view of the functions related to PSGs, hub genes were predicted for each group by the protein–protein interaction network using the STRING V11 database and 11 topological analysis methods (Degree, Edge Percolated Component [EPC], Maximum Neighborhood Component [MNC], Density of Maximum Neighborhood Component [DMNC], Maximal Clique Centrality [MCC], Bottleneck, EcCentricity, Closeness, Radiality, Betweenness, and Stress) of CytoHubba 0.1 (Chin et al. 2014), provided in Cytoscape 3.8.2 (Shannon et al. 2003). The top 10 genes ranked by each method were selected, and the genes shared by at least five methods were identified as “hub genes.”
In addition, common PSGs from each group pair, including, cetaceans + pinnipeds, pinnipeds + sirenians, and sirenians + cetaceans, were investigated to identify common genes that were subjected to selection pressure to adapt to marine environments.
Results
Identification of PSGs and enrichment analysis
We selected 11,449 orthologous genes from Cetartiodactyla, 14,838 from Carnivora, and 5,270 from Afrotheria + Xenarthra for positive selection calculation using CODEML after MSA. Using the branch-site model, 460, 614, and 359 PSGs (adjusted p-value < 0.05) were detected in cetaceans, pinnipeds, and sirenians, respectively (Supplementary Tables S2–S5).
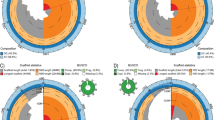
Functional enrichment analysis using PSGs for each group of marine mammals found gene ontology (GO) terms related to hemostasis across all groups, with a significance threshold of p < 0.05. These terms included "coagulation,” “regulation of hemostasis,” and “negative regulation of hemostasis” (Fig. 1A). For hemostasis systems, we identified six PSGs (PLAU, PRKG1, TSPAN8, COL3A1, MMRN1, and SYK) in cetaceans, one (TPH1) in pinnipeds, and four PSGs (GGCX, PLA2G4A, DGKE, and PEAR1) in sirenians (Table 1). Specifically, four genes (PLAU, MMRN1, GGCX, and PEAR1), related to various aspects of hemostasis and coagulation, were predicted to cause protein damage due to amino acid substitutions in at least one program in the analysis results (Table 1).
Association between PSGs in marine mammals and the hemostatic system. A Gene enrichment results using clusterProfiler (p < 0.05). This study included 460 genes from cetaceans, 614 genes from pinnipeds, and 359 genes from sirenians. B The major components of hemostasis. C The number of PSGs in marine mammalian groups related to each function was obtained from the GeneCards database (https://www.genecards.org/)
Hemostasis is a complex physiological process involving three major steps: vasoconstriction, platelet activation, and the coagulation cascade (Fig. 1B). Blood clots formed during this process are typically removed from the body through a mechanism that dissolves them. Thus, we identified genes related to hemostasis or coagulation using the GeneCards database based on five keywords: “hemostasis,” “coagulation,” “thrombosis,” “embolism,” and “fibrinolysis.” The results of matching the PSGs of each group of marine mammals to the derived genes are shown in Fig. 1C. Our findings indicate that aquatic adaptations in marine mammals are more strongly associated with blood coagulation than with clot dissolution in the hemostatic system.
Investigation of common PSGs and hub genes
Common genes were investigated for each pair of PSGs in the cetaceans + pinnipeds, pinnipeds + sirenians, and sirenians + cetaceans groups (Fig. 2A, Supplementary Table S6). Seven genes (C2orf73, PAX5, SCN9A, ITPRID2, PMFBP1, MYCBPAP, and SRRM4) were identified as common PSGs in the cetaceans + pinnipeds group. Both groups had the same amino acid substitution (I990T) in SCN9A, which encodes the voltage-gated sodium channel Nav1.7 (Fig. 2B). Previous studies analyzing the SCN9A gene in sperm whale and baleen whale lineages also found positive selection sites located in critical functional regions of the gene. Congenital insensitivity, or diminished response to pain, is caused by mutations in SCN9A (Raouf et al. 2010; Ding et al. 2022). To investigate the effect of this amino acid change on protein structure and function, we performed an analysis using the PolyPhen program (Adzhubei et al. 2013). This analysis predicted that amino acid alterations would have detrimental effects on protein function.
Common genes for each group of marine mammals. A Venn diagram illustrating the shared genes in cetaceans, pinnipeds, and sirenians. Seven PSGs were detected in cetaceans and pinnipeds. B Identical amino acid substitution in the SCN9A gene of cetaceans and pinnipeds. Evolutionary changes led to isoleucine (I) to threonine (T) amino acid substitution at site 990 of the SCN9A protein in cetaceans and pinnipeds, based on the human protein sequence. The silhouette images were collected from PhyloPic (http://www.phylopic.org/). C Schematic diagram of how the KRAP gene affects calcium ion regulation. The positive selection sites for KRAP and the predicted effects of amino acid substitutions on protein function can be found in Supplementary Table 7. CREB binding protein (CBP); E1A binding protein P300 (EP300); Endoplasmic reticulum (ER)
SRRM4 encodes serine/arginine repetitive matrix protein 4 (Safran et al. 2010), and its mutations are associated with deafness (Nakano et al. 2012). Marcovitz et al. (2019) studied the G310E mutation of SRRM4 in toothed whales, dolphins, and bats that underwent convergent evolution of echolocation, and a clear protein-coding signal was observed in echolocating bats and whales. Pinnipeds do not echolocate (Schusterman et al. 2000) but require auditory adaptations in aquatic and terrestrial environments for an amphibious lifestyle (Hanke and Dehnhardt 2013). Consequently, amino acid changes in SRRM4 may help pinnipeds adapt to aquatic environments (Supplementary Table S7).
ITPR Interacting Domain Containing 2 (ITPRID2), also known as KRAP or SSFA2, plays a crucial role in the localization and function of inositol trisphosphate receptors (IP3Rs) that mediate the release of Ca2+ from the endoplasmic reticulum (Arige and Yule 2021; Vorontsova et al. 2022). The effects of amino acid changes at positively selected positions on protein function and structure were predicted in each group of aquatic mammals using PROVEAN (Choi and Chan 2015), SIFT (Ng and Henikoff 2001), and PolyPhen-2 (Adzhubei et al. 2013). At least one selection site for each gene was identified as potentially damaging (Supplementary Table S7).
The MAT2B gene, observed in both pinnipeds and sirenians, encodes a member of the MAT family. Studies have linked this gene to tumorigenesis in hepatocellular carcinoma, colon cancer, and malignant melanomas. Additionally, SPA17 is found in sirenians and cetaceans (Peng et al. 2013; Tang et al. 2017; Yuan et al. 2019). Positive selection for this gene has been observed in high-altitude frogs. However, this gene is newly discovered in marine mammals (Sun et al. 2018). SPA17 encodes a cell surface transmembrane protein that plays a role in male reproductive functions, such as spermatogenesis and sperm-egg interactions (Chiriva-Internati et al. 2009). For the first time, we present this gene as a positive selection candidate for aquatic reproduction in fully aquatic mammals, including cetaceans and sirenians.
Using hub gene analysis, we identified five genes (DKC1, CCT4, PA2G4, CCT2, and RRM1) in cetaceans, four genes (APOA1, CD4, EP300, and SOX9) in pinnipeds, and seven genes (TMED1, COX15, DERL2, STYK1, MAP2K5, RB1CC1, and RBX1) in sirenians (Table 2).
Discussion
Previous studies identified PSGs by comparing marine mammal branches across the entire mammalian tree. In contrast, our study focused on comparing each marine mammal with terrestrial mammals of the same order. Through an extensive analysis using the branch-site model in CODEML, we detected 460, 614, and 359 PSGs in cetaceans, pinnipeds, and sirenians, respectively.
We performed a functional enrichment analysis of PSGs from three groups of marine mammals and found that genes related to hemostasis were positively selected across all lineages. Hemostasis is the process by which bleeding from the blood vessels is stopped (LaPelusa and Dave 2023). This process includes three main stages: vascular contraction, platelet clotting, and coagulation cascade (Robinson et al. 1969; LaPelusa and Dave 2023). Previous studies have reported the loss or selection of blood coagulation factors in Cetacea (Yim et al. 2014; Huelsmann et al. 2019). Under hypoxia, extended pulmonary arteries have been observed in sea lions (Olson et al. 2010).
We identified four genes (GGCX, PLA2G4A, DGKE, and PEAR1) that may cause protein damage owing to amino acid substitutions at positively selected sites. To our knowledge, this is the first study to identify MMRN1, GGCX, and PEAR1 as PSGs in marine mammals. PLAU, also known as urokinase-type plasminogen activator, encodes a secreted serine protease that converts plasminogen to plasmin. This gene is associated with diseases such as Quebec platelet disorder, which causes bleeding disorders (Paterson et al. 2010). This disease is characterized by the overexpression of PLAU (Liang et al. 2020). MMRN1, also known as multimerin-1, encodes a protein found in platelets and blood vessel walls that may have platelet adhesive functions (Tasneem et al. 2009). MMRN1 deficiency is associated with Quebec platelet disorder, and it plays a role in thrombus formation (Leatherdale et al. 2021). GGCX, also known as gamma-glutamyl carboxylase, encodes an enzyme required for the activation of vitamin K-dependent clotting factors. Mutations in this gene are associated with bleeding disorders (Hao et al. 2021; Rishavy et al. 2022). The PEAR1 (platelet endothelial aggregation receptor 1) gene activates platelet cells by encoding a protein in the platelet (Nanda et al. 2005; Johnson 2016). Therefore, these genes may be linked to delayed blood coagulation in cetaceans (Lohman et al. 1998). These results are consistent with those of previous studies. In contrast to other marine mammals, manatees tend to exhibit higher coagulation (hypercoagulation) than terrestrial mammals (Gerlach et al. 2015; Barratclough et al. 2016). An herbivorous diet comprising vitamin-K-rich seaweed and seagrass reduces blood clotting time (Shiau and Liu 1994; Wei et al. 2023). The exact mechanisms require further research; however, their dietary habits and genes (such as GGCX) might be related.
Although no PSG related to blood coagulation factors or platelets was found in the pinnipeds, we identified TPH1 as a PSG. The TPH genes, including TPH1 (expressed in the periphery) and TPH2 (expressed in the brain), are responsible for the production of serotonin (5-HT), a potent vasoconstrictor. TPH1 is involved in blood vessel regulation. In a TPH1 null mouse experiment, symptoms such as reduced cardiac output, increased heart rate, and abnormal blood circulation were observed (Côté et al. 2003). This gene is also associated with pulmonary hypertension through serotonin regulation. Although pulmonary hypertension typically occurs in oxygen-deficient mammals, it is absent in diving mammals such as pinnipeds (MacLean 2007; Olson et al. 2010). Therefore, TPH1 may have enabled pinnipeds to adapt to land and sea habitats by affecting their blood circulation-related functions.
For the first time, we reported positive selection results for KRAP, which encodes a protein involved in the function of IP3Rs in Ca2 + release, in marine mammals (Vorontsova et al. 2022). Calcium ions in the cytosol control various cellular activities, such as cell death, survival, secretion, gene transcription, and metabolism. Moreover, calcium regulation is associated with anti-obesity (Berridge et al. 2000; Zemel 2002; La Rovere et al. 2016; Kania et al. 2017; Zhang et al. 2019). IP3Rs regulate calcium ion levels by releasing them from the endoplasmic reticulum (Arige and Yule 2021; Thillaiappan et al. 2021; Vorontsova et al. 2022). KRAP is necessary for IP3Rs to function correctly and allow for an accurate location (Fig. 2). KRAP (-/-) mice have been reported to exhibit a higher basal metabolic rate, glucose tolerance, and hypoinsulinemia than wild-type mice (Fujimoto et al. 2007; Chou et al. 2010; Hägg et al. 2015). A recent study has reported that cAMP response element-binding proteins (CREB) regulate KRAP expression (Arige et al. 2021). In previous studies, positive selection for the CREB-binding protein (CBP) and EP300 (known coactivators of CREB) has been reported in cetaceans and pinnipeds, respectively (Noh et al. 2022). Our study identified EP300 as a PSG and hub gene in pinnipeds, and there is evidence linking CBP and EP300 to fat metabolism and obesity (Nishimura et al. 2015; Namwanje et al. 2019). These results suggest that the regulation of calcium signaling may also affect metabolism and fat accumulation in marine mammals.
In summary, we identified PSGs that have undergone selection pressure for adaptation to the aquatic environment in marine mammals, including the hemostatic system. These selected genes provide insight into the physiological characteristics of marine mammals and help address questions regarding their adaptation to aquatic life.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article or its supplementary materials. The data that support the findings of this study are available from the corresponding author, Lee or Park, upon reasonable request.
Abbreviations
- PSGs:
-
Positively selected genes
- aPTT:
-
Activated partial thromboplastin time
- MSA:
-
Multiple sequence alignment
- CDSs:
-
Coding sequences
- CSB:
-
Clean Segment Block
- EPC:
-
Edge Percolated Component
- MNC:
-
Maximum Neighborhood Component
- DMNC:
-
Density of Maximum Neighborhood Component
- MCC:
-
Maximal Clique Centrality
- BN:
-
Bottleneck
- GO:
-
Gene ontology
- CREB:
-
CAMP response element-binding proteins
- CBP:
-
CREB-binding protein
References
Adzhubei I, Jordan DM, Sunyaev SR (2013) Predicting functional effect of human missense mutations using PolyPhen-2. Curr Protoc Hum Genet 76:21–27. https://doi.org/10.1002/0471142905.hg0720s76
Arige V, Yule DI (2021) A KRAP (y) job: Defining the localization of active IP3R. Cell Calcium 100:102470. https://doi.org/10.1016/j.ceca.2021.102470
Arige V, Terry LE, Malik S, Knebel TR, Wagner LE II, Yule DI (2021) CREB regulates the expression of type 1 inositol 1, 4, 5-trisphosphate receptors. J Cell Sci 134:jcs258875. https://doi.org/10.1242/jcs.258875
Barratclough A, Floyd RF, Conner B, Reep R, Ball R, Stacy N (2016) Normal hemostatic profiles and coagulation factors in healthy free-living Florida manatees (Trichechus manatus latirostris). J Wildl Dis 52:907–911. https://doi.org/10.7589/2015-12-328
Berridge MJ, Lipp P, Bootman MD (2000) The versatility and universality of calcium signalling. Nat Rev Mol Cell Biol 1:11–21. https://doi.org/10.1038/35036035
Berta A, Sumich JL, Kovacs KM (2006) Marine mammals: evolutionary biology, 2nd edn. Academic Press, San Diego, California
Chikina M, Robinson JD, Clark NL (2016) Hundreds of genes experienced convergent shifts in selective pressure in marine mammals. Mol Biol Evol 33:2182–2192. https://doi.org/10.1093/molbev/msw112
Chin C-H, Chen S-H, Wu H-H, Ho C-W, Ko M-T, Lin C-Y (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8:1–7. https://doi.org/10.1186/1752-0509-8-S4-S11
Chiriva-Internati M, Gagliano N, Donetti E, Costa F, Grizzi F, Franceschini B, Albani E, Levi-Setti PE, Gioia M, Jenkins M (2009) Sperm protein 17 is expressed in the sperm fibrous sheath. J Transl Med 7:1–5. https://doi.org/10.1186/1479-5876-7-61
Choi Y, Chan AP (2015) PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31:2745–2747. https://doi.org/10.1093/bioinformatics/btv195
Chou JY, Jun HS, Mansfield BC (2010) Glycogen storage disease type I and G6Pase-β deficiency: etiology and therapy. Nat Rev Endocrinol 6:676–688. https://doi.org/10.1038/nrendo.2010.189
Côté F, Thévenot E, Fligny C, Fromes Y, Darmon M, Ripoche M-A, Bayard E, Hanoun N, Saurini F, Lechat P (2003) Disruption of the nonneuronal tph1 gene demonstrates the importance of peripheral serotonin in cardiac function. PNAS 100:13525–13530. https://doi.org/10.1073/pnas.2233056100
Ding X, Yu F, He X, Xu S, Yang G, Ren W (2022) Rubbing Salt in the Wound: Molecular Evolutionary Analysis of Pain-Related Genes Reveals the Pain Adaptation of Cetaceans in Seawater. Animals 12:3571. https://doi.org/10.3390/ani12243571
Foote AD, Liu Y, Thomas GW, Vinař T, Alföldi J, Deng J, Dugan S, van Elk CE, Hunter ME, Joshi V (2015) Convergent evolution of the genomes of marine mammals. Nat Genet 47:272–275. https://doi.org/10.1038/ng.3198
Fujimoto T, Koyanagi M, Baba I, Nakabayashi K, Kato N, Sasazuki T, Shirasawa S (2007) Analysis of KRAP expression and localization, and genes regulated by KRAP in a human colon cancer cell line. J Hum Genet 52:978–984. https://doi.org/10.1007/s10038-007-0204-8
Gerlach TJ, Bandt C, Conner B, Ball RL (2015) Establishment of reference values for various coagulation tests in healthy Florida manatees (Trichechus manatus latirostris) and evaluation of coagulation in debilitated manatees during rehabilitation. J Am Vet Med Assoc 247:1048–1055. https://doi.org/10.2460/javma.247.9.1048
Hägg S, Ganna A, Van Der Laan SW, Esko T, Pers TH, Locke AE, Berndt SI, Justice AE, Kahali B, Siemelink MA (2015) Gene-based meta-analysis of genome-wide association studies implicates new loci involved in obesity. Hum Mol Genet 24:6849–6860. https://doi.org/10.1093/hmg/ddv379
Hanke W, Dehnhardt G (2013) Sensory biology of aquatic mammals. J Comp Physiol, A 199:417–420. https://doi.org/10.1007/s00359-013-0823-9
Hao Z, Jin D-Y, Chen X, Schurgers LJ, Stafford DW, Tie J-K (2021) γ-Glutamyl carboxylase mutations differentially affect the biological function of vitamin K–dependent proteins. Blood 137:533–543. https://doi.org/10.1182/blood.2020006329
Huelsmann M, Hecker N, Springer MS, Gatesy J, Sharma V, Hiller M (2019) Genes lost during the transition from land to water in cetaceans highlight genomic changes associated with aquatic adaptations. Sci Adv 5:eqw6671. https://doi.org/10.1126/sciadv.aaw6671
Johnson AD (2016) Pairing megakaryopoiesis methylation with PEAR1. Blood 128:890–892. https://doi.org/10.1182/blood-2016-06-723940
Kania E, Roest G, Vervliet T, Parys JB, Bultynck G (2017) IP3 receptor-mediated calcium signaling and its role in autophagy in cancer. Front Oncol 7:140. https://doi.org/10.3389/fonc.2017.00140
Kumar S, Stecher G, Suleski M, Hedges SB (2017) TimeTree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol 34:1812–1819. https://doi.org/10.1093/molbev/msx116
La Rovere RM, Roest G, Bultynck G, Parys JB (2016) Intracellular Ca2+ signaling and Ca2+ microdomains in the control of cell survival, apoptosis and autophagy. Cell Calcium 60:74–87. https://doi.org/10.1016/j.ceca.2016.04.005
LaPelusa A, Dave HD (2023) Physiology, hemostasis. StatPearls Publishing, Stat Pearls
Leatherdale A, Parker DA, Tasneem S, Wang Y, Bihan D, Bonna A, Hamaia SW, Gross PL, Ni H, Doble BW (2021) Multimerin 1 supports platelet function in vivo and binds to specific GPAGPOGPX motifs in fibrillar collagens that enhance platelet adhesion. J Thromb Haemost 19:547–561. https://doi.org/10.1111/jth.15171
Liang M, Soomro A, Tasneem S, Abatti LE, Alizada A, Yuan X, Uusküla-Reimand L, Antounians L, Alvi SA, Paterson AD (2020) Enhancer-gene rewiring in the pathogenesis of Quebec platelet disorder. Blood 136:2679–2690. https://doi.org/10.1182/blood.2020005394
Lohman S, Folkow LP, Osterud B, Sager G (1998) Changes in fibrinolytic activity in diving grey seals. Comp Biochem Physiol A Mol Integr Physiol 120:693–698. https://doi.org/10.1016/s1095-6433(98)10088-0
MacLean M (2007) Pulmonary hypertension and the serotonin hypothesis: where are we now? Int J Clin Pract 61:27–31. https://doi.org/10.1111/j.1742-1241.2007.01497.x
Marcovitz A, Turakhia Y, Chen HI, Gloudemans M, Braun BA, Wang H, Bejerano G (2019) A functional enrichment test for molecular convergent evolution finds a clear protein-coding signal in echolocating bats and whales. PNAS 116:21094–21103. https://doi.org/10.1073/pnas.1818532116
Nakano Y, Jahan I, Bonde G, Sun X, Hildebrand MS, Engelhardt JF, Smith RJ, Cornell RA, Fritzsch B, Bánfi B (2012) A mutation in the Srrm4 gene causes alternative splicing defects and deafness in the Bronx waltzer mouse. PLoS Genet 8:e1002966. https://doi.org/10.1371/journal.pgen.1002966
Namwanje M, Liu L, Chan M, Aaron N, Kraakman MJ, Qiang L (2019) The depot-specific and essential roles of CBP/p300 in regulating adipose plasticity. J Endocrinol 240:257–269. https://doi.org/10.1530/JOE-18-0361
Nanda N, Bao M, Lin H, Clauser K, Komuves L, Quertermous T, Conley PB, Phillips DR, Hart MJ (2005) Platelet endothelial aggregation receptor 1 (PEAR1), a novel epidermal growth factor repeat-containing transmembrane receptor, participates in platelet contact-induced activation. J Biol Chem 280:24680–24689. https://doi.org/10.1074/jbc.M413411200
Ng PC, Henikoff S (2001) Predicting deleterious amino acid substitutions. Genome Res 11:863–874. https://doi.org/10.1101/gr.176601
Nishimura Y, Sasagawa S, Ariyoshi M, Ichikawa S, Shimada Y, Kawaguchi K, Kawase R, Yamamoto R, Uehara T, Yanai T (2015) Systems pharmacology of adiposity reveals inhibition of EP300 as a common therapeutic mechanism of caloric restriction and resveratrol for obesity. Front Pharmacol 6:199. https://doi.org/10.3389/fphar.2015.00199
Noh HJ, Turner-Maier J, Schulberg SA, Fitzgerald ML, Johnson J, Allen KN, Hückstädt LA, Batten AJ, Alfoldi J, Costa DP (2022) The Antarctic Weddell seal genome reveals evidence of selection on cardiovascular phenotype and lipid handling. Commun Biol 5:140. https://doi.org/10.1038/s42003-022-03089-2
Olson KR, Whitfield NL, Bearden SE, Leger J, Nilson E, Gao Y, Madden JA (2010) Hypoxic pulmonary vasodilation: a paradigm shift with a hydrogen sulfide mechanism. Am J Physiol Regul Integr Comp Physiol 298:R51–R60. https://doi.org/10.1152/ajpregu.00576.2009
Paterson AD, Rommens JM, Bharaj B, Blavignac J, Wong I, Diamandis M, Waye JS, Rivard GE, Hayward CP (2010) Persons with Quebec platelet disorder have a tandem duplication of PLAU, the urokinase plasminogen activator gene. Blood 115:1264–1266. https://doi.org/10.1182/blood-2009-07-233965
Peng H, Dara L, Li TW, Zheng Y, Yang H, Tomasi ML, Tomasi I, Giordano P, Mato JM, Lu SC (2013) MAT2B-GIT1 interplay activates MEK1/ERK 1 and 2 to induce growth in human liver and colon cancer. Hepatology 57:2299–2313. https://doi.org/10.1002/hep.26258
Raouf R, Quick K, Wood JN (2010) Pain as a channelopathy. J Clin Invest 120:3745–3752. https://doi.org/10.1172/JCI43158
Rishavy MA, Hallgren KW, Wilson LA, Hiznay JM, Runge KW, Berkner KL (2022) GGCX mutants that impair hemostasis reveal the importance of processivity and full carboxylation to VKD protein function. Blood 140:1710–1722. https://doi.org/10.1182/blood.2021014275
Robinson AJ, Kropatkin M, Aggeler PM (1969) Hageman factor (factor XII) deficiency in marine mammals. Science 166:1420–1422. https://doi.org/10.1126/science.166.3911.1420
Safran M, Dalah I, Alexander J, Rosen N, Iny Stein T, Shmoish M, Nativ N, Bahir I, Doniger T, Krug H (2010) GeneCards Version 3: the human gene integrator. Database (oxford) 2010:baq020. https://doi.org/10.1093/database/baq020
Schusterman RJ, Kastak D, Levenson DH, Reichmuth CJ, Southall BL (2000) Why pinnipeds don’t echolocate. J Acoust Soc Am 107:2256–2264. https://doi.org/10.1121/1.428506
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Shiau S-Y, Liu J-S (1994) Estimation of the dietary vitamin K requirement of juvenile Penaeus chinensis using menadione. Aquaculture 126:129–135. https://doi.org/10.1016/0044-8486(94)90254-2
Sun Y-B, Fu T-T, Jin J-Q, Murphy RW, Hillis DM, Zhang Y-P, Che J (2018) Species groups distributed across elevational gradients reveal convergent and continuous genetic adaptation to high elevations. PNAS 115:E10634–E10641. https://doi.org/10.1073/pnas.1813593115
Tang Z, He G, Xu J, Zhongfu L (2017) Knockdown of Cbp/P300-interacting transactivator with Glu/Asp-rich carboxy-terminal domain 2 inhibits cell division and increases apoptosis in gastric cancer. J Surg Res 211:1–7. https://doi.org/10.1016/j.jss.2016.11.049
Tasneem S, Adam F, Minullina I, Pawlikowska M, Hui S, Zheng S, Miller J, Hayward C (2009) Platelet adhesion to multimerin 1 in vitro: influences of platelet membrane receptors, von Willebrand factor and shear. J Thromb Haemost 7:685–692. https://doi.org/10.1111/j.1538-7836.2009.03284.x
Thillaiappan NB, Smith HA, Atakpa-Adaji P, Taylor CW (2021) KRAP tethers IP3 receptors to actin and licenses them to evoke cytosolic Ca2+ signals. Nat Commun 12:4514. https://doi.org/10.1038/s41467-021-24739-9
Vorontsova I, Lock JT, Parker I (2022) KRAP is required for diffuse and punctate IP3-mediated Ca2+ liberation and determines the number of functional IP3R channels within clusters. Cell Calcium 107:102638. https://doi.org/10.1016/j.ceca.2022.102638
Wei X, Hang Y, Li X, Hua X, Cong X, Yi W, Guo X (2023) Effects of dietary vitamin K3 levels on growth, coagulation, calcium content, and antioxidant capacity in largemouth bass, Micropterus salmoides. Aquac Fish 8:159–165. https://doi.org/10.1016/j.aaf.2021.08.004
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591. https://doi.org/10.1093/molbev/msm088
Yim H-S, Cho YS, Guang X, Kang SG, Jeong J-Y, Cha S-S, Oh H-M, Lee J-H, Yang EC, Kwon KK (2014) Minke whale genome and aquatic adaptation in cetaceans. Nat Genet 46:88–92. https://doi.org/10.1038/ng.2835
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–287. https://doi.org/10.1089/omi.2011.0118
Yuan Y, Wang Y, Liu Z, Sun Y, Yao Y, Yu W, Shen Z (2019) MAT2B promotes proliferation and inhibits apoptosis in osteosarcoma by targeting epidermal growth factor receptor and proliferating cell nuclear antigen. Int J Oncol 54:2019–2029. https://doi.org/10.3892/ijo.2019.4764
Yuan Y, Zhang Y, Zhang P, Liu C, Wang J, Gao H, Hoelzel AR, Seim I, Lv M, Lin M (2021) Comparative genomics provides insights into the aquatic adaptations of mammals. PNAS 118:e2106080118. https://doi.org/10.1073/pnas.2106080118
Zemel MB (2002) Regulation of adiposity and obesity risk by dietary calcium: mechanisms and implications. J Am Coll Nutr 21:146S-151S. https://doi.org/10.1080/07315724.2002.10719212
Zhang F, Ye J, Zhu X, Wang L, Gao P, Shu G, Jiang Q, Wang S (2019) Anti-obesity effects of dietary calcium: the evidence and possible mechanisms. Int J Mol Sci 20:3072. https://doi.org/10.3390/ijms20123072
Zhou X, Seim I, Gladyshev VN (2015) Convergent evolution of marine mammals is associated with distinct substitutions in common genes. Sci Rep 5:16550. https://doi.org/10.1038/srep16550
Acknowledgements
This research was part of the “Development of biomedical materials based on marine proteins” project (20170305) funded by the Ministry of Oceans and Fisheries, Korea, and an in-house program (EA0122) from the Korea Institute of Ocean Science & Technology (KIOST).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Roh, YR., Yim, HS., Park, K. et al. Molecular characterization of positively selected genes contributing aquatic adaptation in marine mammals. Genes Genom 46, 775–783 (2024). https://doi.org/10.1007/s13258-023-01487-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01487-2