Abstract
Background
Current advances in the molecular biology of multiple myeloma (MM) are not sufficient to fully delineate the genesis and development of this disease.
Objective
This study aimed to identify molecular targets underlying MM pathogenesis.
Methods
mRNA expression profiling for 29 samples (19 MM samples, 7 MM cell lines and 3 controls) were obtained using microarray. We evaluated the in vitro effects of RAD54L gene silencing on the proliferation, apoptosis and cell cycle distribution in KMS-28BM human MM cells using siRNA approach. Cell proliferation was determined by MTS assay while apoptosis and cell cycle distribution were analysed with flow cytometry. Gene and protein expression was evaluated using RT-qPCR and ELISA, respectively.
Results
Microarray results revealed a total of 5124 differentially expressed genes (DEGs), in which 2696 and 2428 genes were up-regulated and down-regulated in MM compared to the normal controls, respectively (fold change ≥ 2.0; P < 0.05). Up-regulated genes (RAD54L, DIAPH3, SHCBP1, SKA3 and ANLN) and down-regulated genes (HKDC1, RASGRF2, CYSLTR2) have never been reported in association with MM. Up-regulation of RAD54L was further verified by RT-qPCR (P < 0.001). In vitro functional studies revealed that RAD54L gene silencing significantly induced growth inhibition, apoptosis (small changes) and cell cycle arrest in G0/G1 phase in KMS-28BM (P < 0.05). Silencing of RAD54L also decreased its protein level (P < 0.05).
Conclusions
This study has identified possible molecular targets underlying the pathogenesis of MM. For the first time, we reveal RAD54L as a potential therapeutic target in MM, possibly functioning in the cell cycle and checkpoint control.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multiple myeloma (MM) is a cancer of plasma cells. It is a highly heterogeneous and genetically complex form of blood cancer (Corre et al. 2015). MM is the second leading cause of haematological malignancy in the world (de Mel et al. 2014). Development and progression of MM required multiple primary and secondary oncogenic events. Hyperdiploidy and translocations concerning the IgH locus are frequent primary genetic events in the initiation of MM disease. IgH translocations frequently lead to the activation of proto-oncogenes for instance FGFR3/MMSET/t(4;14), CCND1/t(11;14), c-MAF/CCND2/t(14;16), MAF-B/t(14;20) and CCND3/t(6;14) (Colombo et al. 2015; Pinto et al. 2020). Acquired mutations (K-RAS, N-RAS, BRAF, DIS3, FAM46C and TP53 genes), copy number aberrations (del(17p)/TP53, del(13) and 1q gain), NF-κB pathway mutation and MYC aberrations are recurrent secondary oncogenic events in MM, which are associated with disease progression (Colombo et al. 2015; Roy et al. 2018; Pinto et al. 2020).
Current advances in research have dramatically expanded our understanding of the genomic landscape, tumour heterogeneity and clonal evolution in MM (Furukawa and Kikuchi 2020). However, MM remains incurable with a poor prognosis due to a lack of suitable tumour markers for early diagnosis and treatment. Therefore, identifying new tumour biomarkers and therapeutic targets are essential steps to improve the prognostic and outcomes of patients. The aims of this study are to identify molecular targets underlying the pathogenesis of MM by mRNA expression profiling and functional target validation by using siRNA and cell-based assays.
Materials and methods
Patient samples
Bone marrow or whole blood were taken from MM patients (N = 19: MM1-MM19) and healthy donors (N = 3: NB1, NB2, and NB3) and stored at − 80 °C. The average age of the patient was 57 years while the median age was 61 years. The age of the patients was in the range of 28–74 years. Seventeen patients were newly diagnosed MM (NDMM), 2 were from relapsed MM cases. Patients were with > 10% plasma cell infiltration at the time of sample recruitment. Cytogenetic analysis indicated 13 patients with normal karyotype, 1 patient with hypodiploidy and 5 patients with unknown karyotype. Patients’ clinical parameters are shown in Table 1.
Cell lines
The U-266, RPMI-8226, IM-9 and MM.1S cells were obtained from American Type Culture Collection (ATCC, USA). The KMS-28BM, KMS-12BM and KMS-20 were purchased from the Japanese Collection of Research Bioresources (JCRB) cell bank. Multiple myeloma cells were cultured and maintained in RPMI-1640 medium (PAN-Biotech, Germany) with 10% fetal bovine serum (Sigma-Aldrich, Germany) in an incubator at 37 ℃ with 5% CO2. Cells were sub-cultured when achieved 70–75% confluence. Cells at the logarithmic phase were used for transfection.
Isolation of total RNAs
Total RNAs were isolated from the bone marrow/peripheral blood/cells according to manufacturer’s recommendation (Qiagen RNeasy mini kit). DNA digestion was performed to ensure that the RNA was free of DNA contamination (Qiagen DNase I, Hilden, Germany). The integrity of the total RNAs used for gene expression microarray were checked with Bioanalyser (RNA Nano Chip, Agilent 2100 Bioanalyser). All samples included in the gene expression microarray had an RNA integrity number (RIN) of at least 8.0. The purity of the isolated Total RNAs was also checked with NanoDrop ND-1000 UV–VIS spectrophotometer to ensure that the purity was within the range of 1.80–2.10.
Gene expression microarray assay
Sample processing, labelling, and hybridisation were performed following the standard protocol recommended by Agilent’s manufacturer. Approximately 60,000 probes were contained on each array. Briefly, 100 ng of total RNAs were labelled using one colour Agilent’s Low Input Quick Amp Labelling kit and purified with spin column (Qiagen RNeasy mini kit, Hilden, Germany). Hybridisation was performed using 600 ng of labelled cRNAs onto SurePrint G3 Human GE 8 × 60 K V2 Microarray Kit (Agilent Technologies, USA). The microarray slide was then put into an incubator with rotation at 65 °C for 17 h. Microarray images were scanned and data from raw microarray image files were extracted with Agilent Feature Extraction Software Version 10.7.3.1 to prepare for analysis. Only samples that passed the raw data quality control metrics as described by Agilent’s recommended procedure were proceeded to data analysis. Pre-processing of the data files in this study was performed by using GeneSpring software version 14.9. All the raw data were thresholded to 1 and normalised to the 75th percentile. This was followed by a baseline transformation set to the median of all samples.
Significantly differentially expressed probes in MM vs. normal controls were identified by unpaired unequal variance t-test (Welch) (P < 0.05). The Benjamini Hochberg false discovery rate (FDR) multiple testing corrections was used to identify differentially expressed probes. The resulting list was further refined by analysing it to a second filter, which specified a 2.0-fold change between MM vs. controls. Only probes that passed a p-value cut-off of 0.05 and fold change ≥ 2.0 were considered significant. Unsupervised hierarchical clustering analysis was carried out for the up-regulated and down-regulated probes, respectively, with a p-value cut-off of 0.05 and fold change ≥ 2.0. Unsupervised hierarchical clustering was generated using Euclidean distance metric and average linkage statistical methods.
In vitro functional study of RAD54L in KMS-28BM human MM cell line
siRNA transfection
Three target-specific siRNA oligo duplexes (Cohesion Biosciences, Catalog No.: CRH5528) of the human RAD54L gene were pooled together to knockdown the target gene in KMS-28BM MM cells. Briefly, 2 × 106 cells were resuspended in 100 μl of 4D-Nucleofector™ solution (Lonza, USA) and mixed with 500 nM of RAD54L siRNA or negative control siRNA. The mixture was transferred to a cuvette. Transfection was then carried out using program DY-100 in a 4D-Nucleofector™ system (Lonza, USA). Then, cells in the cuvette were gently transferred to warm medium in 24 well plates. Two or three independent experiments were carried out for each transfection.
RT-qPCR analysis
Total RNAs were converted to first strand cDNA following manufacturer’s protocol (High Capacity RNA-to-cDNA kit, Applied Biosystems, USA). The RT-qPCR was performed using the TaqMan gene expression assay (Applied Biosystems, USA) in the StepOnePlus™ Real-time PCR System (Applied Biosystems, USA). Pre-designed TaqMan gene expression assays for RAD54L (Hs00936473_m1, ThermoFisher Scientific, USA) and internal control GAPDH (Hs02758991-g1, ThermoFisher Scientific, USA) were used. Thermal cycling conditions consisted of the following: initial denaturation at 95 °C for 20 s, followed by 40 cycles of 95 °C for 1 s and 60 °C for 20 s. The relative expression of genes was calculated and quantified based on 2−ΔΔCt method.
MTS assay
RAD54L siRNA or negative control siRNA was transfected into KMS-28BM. Cells were seeded onto 96-well plates at a density of 2.0 × 104 cells/well in 100 μl of culture medium. Cells were cultured for 24 h, 48 h and 72 h. At 0 h and after 24 h, 48 h and 72 h post-transfection, 20 μl of CellTiter 96 AQueous One Solution Reagent (Promega, USA) was added into each sample well. The plate was then incubated at 37 ℃ for 3–4 h in a humidified 5% CO2 incubator. Subsequently, the plate was measured at absorbances of 490 nm and 630 nm using Synergy HTX microplate reader (BioTek®, USA). The 630 nm reading was then subtracted from the 490 nm reading.
Apoptosis assay
After 48 h of transfection, RAD54L siRNA and control siRNA-treated cells were harvested by centrifugation at 1000 rpm. Cells were washed with ice-cold PBS and then resuspended in Annexin binding buffer at 5.0 × 105 cells/mL. This was followed by staining the cells with 5 μl of Annexin-V-FITC and 5 μl of propidium iodide (PI) (Elabscience, USA). Finally, the cells were analysed with the FACSCANTO II flow cytometer (BD BioScience, USA) in which 10,000 events were recorded for each analysis.
Cell-cycle analysis
Briefly, transfection was performed according to the procedure described above. Cells transfected with RAD54L siRNA or control siRNA were then transferred to a 24-well cell culture plate containing pre-warm medium. The plate was then incubated at 37 °C CO2 cell incubator for 48 h. After 48 h of transfection, cells were harvested and washed with ice-cold PBS. This was followed by fixing the cells with 70% ethanol overnight. Approximately 1 × 105 cells were stained with 500 μl of PI solution (BD BioScience, USA). The mixture was incubated for 15 min at room temperature. Cell cycle distribution was then analysed with FACSCANTO II flow cytometer (BD BioScience, USA) and ModFit LT (Verity Software House).
The determination of RAD54L protein level with ELISA
Standards were diluted and prepared according to manufacturer’s protocol (SUNLONG Human RAD54L ELISA kit). Forty microliters of sample dilution buffer and 10 μl of sample were added into sample wells and then incubated for 30 min at 37 °C. Then, the wash buffer stock solution was diluted 1:30 in distilled water. Microtiter plate was washed for 30 s, repeatedly for 5 times. This was followed by adding 50 μl of HRP-conjugate reagent to each sample well. The mixture was then incubated and washed as described above. Then, 50 μl of each Chromogen Solution A and B were added into the sample well. Subsequently, the plate was incubated for 15 min at 37 °C. This was followed by adding 50 μl of stop solution to terminate the reaction. Lastly, absorbance was measured at 450 nm with a microplate reader (Tecan Infinite® M1000).
Statistical analysis
Student’s t-test was used to assess statistical significance between the means of the 2 groups. Those having P-value lower than 0.05 were considered significant.
Results
mRNA expression profiling and RT-qPCR verification
mRNA expression profiling was performed for 19 MM samples, 7 MM cell lines and 3 normal controls. A total of 50739 probes were retained after normalising and filtering at selected thresholds. Out of 50739 probes, 5888 probes were significantly differentially expressed (P < 0.05; fold change ≥ 2.0). The results are depicted in a volcano plot (Fig. 1A). These 5888 probes were consisted of 5124 genes (including long intergenic non-protein coding RNAs, lincRNAs and novel transcripts). Among 5124 genes, 2696 and 2428 genes were up-regulated and down-regulated in MM vs. the control group, respectively. Genes showing up-regulation and down-regulation in MM vs. controls are listed in Online Resource 1 and Online Resource 2, respectively. Unsupervised hierarchical clustering was performed on up-regulated and down-regulated genes for all samples, respectively. Genes which expressed at similar patterns were clustered together and joined by a sequence of branches or dendrogram. The genes input list contained DEGs with a corrected P < 0.05 and a fold change of ≥ 2.0. Online Resources 3 & 4 showing the heatmaps of three distinct sub-clusters classified according to the degree of similarity in gene expression: MM cell lines, MM clinical samples (except MM9 for Online Resource 3; MM5 and MM9 for Online Resource 4) and controls.
A Volcano plot identified 5888 differentially expressed probes in multiple myeloma vs. normal controls (fold change ≥ 2.0; P < 0.05). The x-axis represents the log2-fold change of probes, while the y-axis represents the − log10 of the corrected P values for each probe. Each dot represents a probe and the red- and blue-coloured areas represent the up-regulated and down-regulated probes, respectively, that met the selection criteria of a fold change of at least 2 (fold change ≥ 2.0 or ≤ 2.0) and a P < 0.05. Orange and light blue dots represent up-regulated and down-regulated probes that failed to pass the fold change cut-off, respectively. Green dots represent probes for RAD54L, DIAPH3, SHCBP1, SKA3, ANLN, HKDC1, RASGRF2 and CYSLTR2. B Graph showing the mean expression of RAD54L detected in microarray and RT-qPCR. RAD54L is up-regulated in 23 MM samples as measured by RT-qPCR, consistent with microarray findings. C Graph showing mean expression of RAD54L in relapsed MM (N = 14) is higher than NDMM (N = 2) as determined by RT-qPCR. ***P < 0.001
The up-regulation of RAD54L as detected with microarray analysis was further verified by using RT-qPCR in 23 MM samples from the same cohort (except MM2, MM5 and MM9, where the RNA concentration was insufficient for RT-qPCR). The RT-qPCR results showed that RAD54L was significantly up-regulated in the MM vs. control group, consistent with microarray findings (P < 0.001) (Fig. 1B). Additionally, when we compared the relative expression of RAD54L in NDMM (N = 14) with the relapsed MM (N = 2), the RT-qPCR findings showed that RAD54L expression in relapsed MM was higher than NDMM, although the result was not significant (Fig. 1C).
The effects of RAD54L gene silencing on cell proliferation, apoptosis, cell cycle distribution and protein level in KMS-28BM MM cells
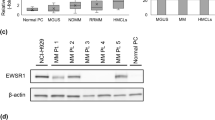
We have previously integrated the gene expression profiles with the miRNA expression profiles from the matched MM samples and revealed an inverse correlation between 5 putative target genes (RAD54L, CCNA2, CYSLTR2, HKDC1 and RASGRF2) and 15 dysregulated miRNAs (Bong et al. 2017). Among the 5 genes listed, RAD54L is one of the new genes that has never been reported in association with MM. This prompted us to explore the in vitro function of RAD54L in MM cells. Since the MM clinical samples used in this study were derived from the Asian population, we would like to study the function of RAD54L in MM cell lines derived from the same origin. Thus, we have selected KMS-28BM as a model for functional analysis: a cell line which showed the highest RAD54L expression level among the 3 cell lines derived from Asian patients included for the gene expression profiling (Fig. 2A).
A Graph showing relative RAD54L expression in KMS-20, KMS-12BM and KMS-28BM MM cell lines as measured by RT-qPCR. B Relative RAD54L expression was reduced by 71% in RAD54L siRNA-treated vs. control siRNA-treated KMS-28BM cells at 24 h post-transfection as measured by RT-qPCR. C Cell proliferation was decreased in KMS-28BM cells transfected with RAD54L siRNA compared to control siRNA at 48 h and 72 h post-transfection as analysed by MTS assay. D Number of early apoptotic cells showing small but significant increase in KMS-28BM cells transfected with RAD54L siRNA compared to control siRNA at 48 h post-transfection as analysed by flow cytometry. E RAD54L gene silencing significantly induced cell cycle arrest at G0/G1 phase while reduced S phase in RAD54L siRNA-treated cells compared to control at 48 h post-transfection as determined by flow cytometry. F RAD54L protein expression level was decreased by approximately 19% in KMS-28BM cells transfected with RAD54L siRNA compared to control siRNA at 48 h post-transfection as measured by ELISA assay. *P < 0.05; **P < 0.01
Silencing of RAD54L gene with siRNA duplexes successfully knockdown RAD54L expression in KMS-28BM by 71% at 24 h post-transfection as measured by RT-qPCR (P < 0.05) (Fig. 2B). The MTS assay revealed significant decreased in proliferation of MM cells treated with RAD54L siRNA compared to the control siRNA at 48 h (P < 0.01) and 72 h (P < 0.05) post-transfection (Fig. 2C). Flow cytometry results showed that silencing of RAD54L led to a small but significant increase in the numbers of early apoptotic cells in siRNA-treated cells compared to the control siRNA-treated cells (P < 0.01) (Fig. 2D). Additionally, cell cycle analysis showed that gene silencing of RAD54L significantly induced cell cycle arrest at G0/G1 phase (P < 0.01) while reduced S phase (P < 0.05) in KMS-28BM (Fig. 2E). ELISA results showed that RAD54L protein expression diminished by approximately 19% with RAD54L knockdown at 48 h post-transfection (P < 0.05) (Fig. 2F).
Discussion
MM is a highly heterogeneous and complex disease which develops via a stepwise process involving multiple genetic aberrations. Numerous genetic aberrations have been identified in association with MM development and malignancy. However, to date MM is still an incurable disease mainly because most of the patients eventually relapse or are refractory to the available treatments (Davis and Sherbenou 2021). Thus, identification of new molecular targets is urgently needed to solve this problem.
Herein, we used mRNA expression profiling to identify potential genes involved in the molecular pathogenesis of MM. Unsupervised hierarchical clustering analysis of the up-regulated and down-regulated probes clearly clustered the samples into MM cell lines, MM clinical samples (except MM9 for the up-regulated probes and MM5 & MM9 for the down-regulated probes) and control groups (Online Resources 3 & 4). This scenario depicts that gene expression changes occur during the transition from normal to malignant plasma cells (Szalat et al. 2016). This study identified a total of 5124 DEGs in the MM vs. control group. Our findings revealed up-regulation of recurrent genes involved in primary oncogenic events: CCND1 and FGFR3 (Kuehl and Bergsagel 2012). Prominent and potential genes involved in the secondary oncogenic event such as NRAS, IRF4, IDH2, PSMB5 and APOBEC2 were found to be up-regulated whereas SP140, LTB and ATM were down-regulated in MM (Kuehl and Bergsagel 2012; Bolli et al. 2014; Walker et al. 2018; Allmeroth et al. 2021). Apart from that, differential expression of genes involved in NF-κB pathway were identified (NFKBIB, IKBKB, CARD11, TNFRSF1A, MAP3K1, MAP3K14 and TLR4) suggesting the important role of NF-κB pathway in myelomagenesis (Chapman et al. 2011). Aberrant expression of apoptosis-related genes such as BCL2, BIK and BAX were also detected in the current study (Gupta et al. 2021). Additionally, the most established growth factors and cytokines involved in bone marrow microenvironment, namely IL6 and IGF1, were found to be over-expressed in the MM vs. control group in this study (Birmann et al. 2009). Other DEGs, namely BIRC5, CENPA, CCNB1, CHEK1, AURKB, BUB1, BUB1B, NEK2, ASPM, TOP2A and EZH2 have been described elsewhere and they play a role, at least in part, in the molecular pathogenesis of MM (Chng and Fonseca 2009; Broyl et al. 2010; Chung et al. 2013). Our findings revealed that most of the DEGs are involved in DNA repair, cell proliferation, cell cycle and mitotic/spindle checkpoints, mismatch repair pathway, kinetochore and microtubule attachment and NF-κB pathway (Chng et al. 2006; Ueki et al. 2008; Chng and Fonseca 2009; Ruiz et al. 2009; Broyl et al. 2010; Agarwal et al. 2011; Chapman et al. 2011; Asano et al. 2013; Bengtsson et al. 2013; Chung et al. 2013; Jiao et al. 2013; Li and Huang 2013; Chuang and Ou 2014; Morley et al. 2015; Perez-Peña et al. 2017; Contreras et al. 2019; Ji et al. 2019; Xia et al. 2020; Borah and Reddy 2021; Li et al. 2021). DEGs and their functions are shown in Table 2.
Interestingly, our study reveals significant DEGs, which have never been reported in association with myelomagenesis. They are RAD54L, DIAPH3, SHCBP1, SKA3, ANLN, HKDC1, RASGRF2 and CYSLTR2. The RAD54L, DIAPH3, SHCBP1, SKA3 and ANLN were up-regulated while HKDC1, RASGRF2 and CYSLTR2 were down-regulated in MM vs. controls. Over-expression of RAD54L was detected in the MM vs. control group in this study by 18.3-folds. We then verified the expression of RAD54L in 23 samples by using RT-qPCR and the results are consistent with microarray analysis. RAD54L is involved in homologous recombination repairing of DNA double-strand breaks to facilitate human genomic integrity and genetic diversity (Andriuskevicius et al. 2018). Defects in homologous recombination pathway-related genes RAD54, RAD51 and RAD52 could lead to tumour development (Mun et al. 2020). RAD54 interacts with RAD51 nucleoprotein filament to form RAD54-RAD51-ssDNA nucleoprotein complex to stimulate homology search and DNA strand exchange (Zohud et al. 2020). RAD54 also removes RAD51 from heterodimeric DNA in an ATP-dependent manner after DNA strand exchange (Andriuskevicius et al. 2018; Rosenbaum et al. 2019). In G2 phase of the cell cycle, RAD54 is phosphorylated to remove RAD51 to facilitate homologous recombination while it is not phosphorylated in S phase to allow RAD51 stabilizing and protecting the stalled replication forks from nucleolytic degradation (Spies et al. 2016). Thus, RAD54 deficiency reduces homologous recombination efficiency. In addition to RAD54L, our results revealed up-regulation of RAD51 (13.8-folds), implicating that the deficiency in the homologous recombination pathway plays a critical role, at least in part, in the pathogenesis of MM.
Elevated expression of RAD54L is detected in carcinomas of the breast, colon, lymphoma and meningioma; however, its role in MM pathogenesis is unknown (Leone et al. 2003). In colorectal carcinoma, RAD54L taking part in maintaining chromosomal stability via DNA homologous recombination and p53 signalling pathway by interacting with POLE (Zohud et al. 2020). Apart from p53 pathway, RAD54L also interacting with other genes such as E2F1 and NEK2 in tumourigenesis (Mun et al. 2020; Pavan et al. 2021). Interestingly, E2F1 and NEK2 were found to be up-regulated in the MM vs. control group in this study by 14.2 and 21.8-folds, respectively. E2F1 is a well-recognized transcription factor that regulates cell cycle progression in the suppression of tumourigenesis, whereas NEK2 is involved in maintaining the stability of replication forks (Karras et al. 2016; Pavan et al. 2021).
The p53-DREAM pathway is a newly described p53-mediated cell cycle arrest pathway. The p53-DREAM pathway regulates not only cell cycle associated genes essential for cell cycle progression from G1 phase to the end of mitosis, but also DNA repair and telomere maintenance genes (Fischer et al. 2014; Engeland 2018). Thus, defects in this pathway lead to the loss of checkpoint control and uncontrolled cell division (Engeland 2018). Apart from that, aberrant expression of genes in this pathway induces chromosomal instability and aneuploidy in cancer cells (Thompson and Compton 2010). To date, almost 250 target genes regulated by the p53-DREAM pathway have been identified (Fischer et al. 2014; Engeland 2018). Interestingly, RAD54L and many other DEGs that function in cell cycle or checkpoint control identified in the current study are components of the p53-DREAM pathway (CCNA2, BIRC5, CENPA, CENPF, CCNB1, CCNB2, CDC25C, KIF14, DEPDC1 and CDK1) (Table 2) (Fischer et al. 2014; Engeland 2018). CCNA2, BIRC5, CCNB1, CCNB2, CDC25C and CDK1 are key regulators in the p53-DREAM pathway (Fischer et al. 2015, 2016). It was postulated that RAD54L, CCNA2 and CCNB1 are p53-dependent repression of the cell cycle genes, in which they contain cell cycle genes homology region (CHR) elements that allow them to bind with DREAM transcriptional repressor (Fischer et al. 2014, 2016). Most of these target genes are involved in G2/M checkpoint control and progression through mitosis (Fischer et al. 2016). Our results indicate that disruption of the p53-DREAM pathway plays a critical role in MM pathogenesis as many genes involved in this pathway were differentially expressed in MM.
In terms of epigenetic aspect, our previous findings on the integrative analysis of the mRNA and miRNA expression profiling for matched samples revealed an inverse correlation between RAD54L and cancer-associated miR-150 (Bong et al. 2017). Based on computational predictions, our previous findings revealed that differential expression of RAD54L might be correlated with aberrant expression of miR-150-5p, which regulates the gene. Thus, according to our observation, we suggest that RAD54L possibly triggers MM either independently or by interacting with other genes/miRNA via impairing DNA repair mechanism and cell cycle progression.
To validate the function of RAD54L in MM, we transiently knockdown RAD54L in KMS-28BM human MM cell line using siRNA method. The effects of RAD54L-mediated gene silencing on cell proliferation, apoptosis and cell cycle distribution in KMS-28BM were determined. Our results reveal that gene silencing of RAD54L inhibits cell proliferation in KMS-28BM (Fig. 2C). In addition, RAD54L knockdown induces small but significant changes in early apoptotic cells (Fig. 2D). Moreover, cell cycle analysis demonstrates that depletion of RAD54L induces cell cycle arrest at G0/G1 phase and inhibits cells at S phase suggesting that RAD54L is involved in cell cycle progression (Fig. 2E). The ELISA assay shows that silencing of RAD54L not only knockdown the gene expression at the mRNA level but also decreases its protein expression level (Fig. 2F). For the first time, our results demonstrate that up-regulation of RAD54L might play a key role, at least in part, in MM pathogenesis by activating myeloma cell growth, inhibiting apoptosis and impairing cell cycle progression.
The limitation of the present study is the small number of samples used for gene expression profiling, which may reduce the statistical power of the study. Our future direction is to validate the results with larger sample sizes by using a more advanced technology such as RNA-sequencing (RNA-seq).
Conclusions
This study has identified possible molecular targets underlying MM pathogenesis. RAD54L might be a potential therapeutic target in MM, possibly functioning in the cell cycle and checkpoint control.
Data availability
The microarray-generated gene expression data can be retrieved from NCBI Gene Expression Omnibus (GEO) repository as series accession identifier GSE72213.
References
Agarwal S, van Cappellen WA, Guénolé A, Eppink B, Linsen SEV, Meijering E (2011) ATP-dependent and independent functions of Rad54 in genome maintenance. J Cell Biol 192(5):735–750. https://doi.org/10.1083/jcb.201011025
Allmeroth K, Horn M, Kroef V, Miethe S, Müller RU, Denzel MS (2021) Bortezomib resistance mutations in PSMB5 determine response to second-generation proteasome inhibitors in multiple myeloma. Leukemia 35(3):887–892. https://doi.org/10.1038/s41375-020-0989-4
Andriuskevicius T, Kotenko O, Makovets S (2018) Putting together and taking apart: assembly and disassembly of the Rad51 nucleoprotein filament in DNA repair and genome stability. Cell Stress 2(5):96–112. https://doi.org/10.15698/cst2018.05.134
Asano E, Hasegawa H, Hyodo T, Ito S, Maeda M, Takahashi M, Hamaquchi M, Senga T (2013) The Aurora B-mediated phosphorylation of SHCBP1 regulates cytokinetic furrow ingression. J Cell Sci 126(Pt 15):3263–3270. https://doi.org/10.1242/jcs.124875
Bengtsson AM, Jönsson G, Magnusson C, Salim T, Axelsson C, Sjölander A (2013) The cysteinyl leukotriene 2 receptor contributes to all-trans retinoic acid-induced differentiation of colon cancer cells. BMC Cancer 13:336. https://doi.org/10.1186/1471-2407-13-336
Birmann BM, Tamimi RM, Giovannucci E, Rosner B, Hunter DJ, Kraft P, Mitsiades C, Anderson KC, Colditz GA (2009) Insulin-like growth factor-1- and interleukin-6-related gene variation and risk of multiple myeloma. Cancer Epidemiol Biomarkers Prev 18(1):282–288. https://doi.org/10.1158/1055-9965
Bolli N, Avet-Loiseau H, Wedge DC, Van Loo P, Alexandrov LB, Martincorena I, Dawson KJ, Iorio F, Nik-Zainal S, Bignell GR et al (2014) Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat Commun 5:2997. https://doi.org/10.1038/ncomms3997
Bong IPN, Ng CC, Baharuddin P, Zakaria Z (2017) MicroRNA expression patterns and target prediction in multiple myeloma development and malignancy. Genes Genomics 39(5):533–540. https://doi.org/10.1007/s13258-017-0518-7
Borah NA, Reddy MM (2021) Aurora kinase B inhibition: a potential therapeutic strategy for cancer. Molecules 26(7):1981. https://doi.org/10.3390/molecules26071981
Broyl A, Hose D, Lokhorst H, Peeters J, Jauch A, Bertsch U, Buijs A, Stevens-Kroef M, Beverloo HB, Vellenga E et al (2010) Gene expression profiling for molecular classification of multiple myeloma in newly diagnosed patients. Blood 116(14):2543–2553. https://doi.org/10.1182/blood-2009-12-261032
Chapman MA, Lawrence MS, Keats JJ, Cibulskis K, Sougnez C, Schinzel AC, Harview CL, Brunet JP, Ahmann GJ, Adli M et al (2011) Initial genome sequencing and analysis of multiple myeloma. Nature 471(7339):467–472. https://doi.org/10.1038/nature09837
Chng WJ, Fonseca R (2009) Centrosomes and myeloma; aneuploidy and proliferation. Environ Mol Mutagen 50(8):697–707. https://doi.org/10.1002/em.20528
Chng W-J, Baker A, Henry T, Price-Troska T, van Wier S, Chung T-H (2006) Combined high resolution array comparative genomic hybridization and gene expression profiling reveal rb1 haploinsufficiency as a possibile tumorigenic mechanism in myeloma. Blood 108(11):113. https://doi.org/10.1182/blood.V108.11.113.113
Chuang HY, Ou YH (2014) Overexpression of anillin in colorectal cancer promotes the cell proliferation, cell mobility and cell invasion. Cancer Res 74(19):4068. https://doi.org/10.1158/1538-7445.AM2014-4068
Chung T-H, Mulligan G, Fonseca R, Chng WJ (2013) A novel measure of chromosome instability can account for prognostic difference in multiple myeloma. PLoS ONE 8(6):e66361. https://doi.org/10.1371/journal.pone.0066361
Colombo M, Galletti S, Garavelli S, Platonova N, Paoli A, Basile A, Taiana E, Neri A, Chiaramonte R (2015) Notch signaling deregulation in multiple myeloma: a rational molecular target. Oncotarget 6(29):26826–26840. https://doi.org/10.18632/oncotarget.5025
Contreras J, Caldas-Lopes E, Jayabalan D, Monge J, Lombardi N, Li H-y, Gunaganti N, Niesvizky R, Guzman N (2019) Targeting Aurora-kinase B with the novel Aurora Kinase B inhibitor MK7 in multiple myeloma. Clin Lymphoma Myeloma Leuk 19(10):e128. https://doi.org/10.1016/j.clml.2019.09.212
Corre J, Munshi N, Avet-Loiseau H (2015) Genetics of multiple myeloma: another heterogeneity level? Blood 125(12):1870–1876. https://doi.org/10.1038/sj.leu.2404621
Davis LN, Sherbenou DW (2021) Emerging therapeutic strategies to overcome drug resistance in multiple myeloma. Cancers (basel) 13(7):1686. https://doi.org/10.3390/cancers13071686
de Mel S, Lim SH, Tung ML, Chng W-J (2014) Implications of heterogeneity in multiple myeloma. Biomed Res Int 2014:232546. https://doi.org/10.3390/ijms20051248
Engeland K (2018) Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ 25(1):114–132. https://doi.org/10.1038/cdd.2017.172
Fischer M, Steiner L, Engeland K (2014) The transcription factor p53: not a repressor, solely an activator. Cell Cycle 13(19):3037–3058. https://doi.org/10.4161/15384101.2014.949083
Fischer M, Quaas M, Nickel A, Engeland K (2015) Indirect p53-dependent transcriptional repression of Survivin, CDC25C, and PLK1 genes requires the cyclin-dependent kinase inhibitor p21/CDKN1A and CDE/CHR promoter sites binding the DREAM complex. Oncotarget 6(39):41402–41417. https://doi.org/10.18632/oncotarget.6356
Fischer M, Quaas M, Steiner L, Engeland K (2016) The p53–p21-DREAM-CDE/CHR pathway regulates G2/M cell cycle genes. Nucleic Acids Res 44(1):164–174. https://doi.org/10.1093/nar/gkv927
Furukawa Y, Kikuchi J (2020) Molecular basis of clonal evolution in multiple myeloma. Int J Hematol 111(4):496–511. https://doi.org/10.1007/s12185-020-02829-6
Gupta VA, Ackley J, Kaufman JL, Boise LH (2021) BCL2 family inhibitors in the biology and treatment of multiple myeloma. Blood Lymphat Cancer 11:11–24. https://doi.org/10.2147/BLCTT.S245191
Ji C, Ma J, Shu P (2019) Effect of dentideless e3 ubiquitin protein ligase on proliferation and colone formation of multiple myeloma cells. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 41(2):194–202. https://doi.org/10.3881/j.issn.1000-503X.10674
Jiao X, Hooper SD, Djureinovic T, Larsson C, Wärnberg F, Tellgren-Roth C, Botling J, Sjoblom T (2013) Gene rearrangements in hormone receptor negative breast cancers revealed by mate pair sequencing. BMC Genomics 14:165. https://doi.org/10.1186/1471-2164-14-165
Karras JR, Schrock MS, Batar B, Huebner K (2016) Fragile genes that are frequently altered in cancer: players not passengers. Cytogenet Genome Res 150(3–4):208–216. https://doi.org/10.1159/000455753
Kuehl WM, Bergsagel PL (2012) Molecular pathogenesis of multiple myeloma and its premalignant precursor. J Clin Invest 122(10):3456–3463. https://doi.org/10.1172/JCI61188
Leone PE, Mendiola M, Alonso J, Paz-y-Miño C, Pestaña A (2003) Implications of a RAD54L polymorphism (2290C/T) in human meningiomas as a risk factor and/or a genetic marker. BMC Cancer 3:6. https://doi.org/10.1186/1471-2407-3-6
Li G-H, Huang J-F (2013) Inferring therapeutic targets from heterogeneous data: HKDC1 is a novel potential therapeutic target for cancer. Bioinformatics 30(6):748–752. https://doi.org/10.1093/bioinformatics/btt606
Li C, Wendlandt EB, Darbro B, Xu H, Thomas GS, Tricot G, Chen F, Shaughnessy JD Jr, Zhan F (2021) Genetic analysis of multiple myeloma identifies cytogenetic alterations implicated in disease complexity and progression. Cancers (basel) 13(3):517. https://doi.org/10.3390/cancers13030517
Morley S, You S, Pollan S, Choi J, Zhou B, Hager MH, Steadman K, Spinelli C, Rajendran K, Gertych A et al (2015) Regulation of microtubule dynamics by DIAPH3 influences amoeboid tumor cell mechanics and sensitivity to taxanes. Sci Rep 5:12136. https://doi.org/10.1038/srep12136
Mun J-Y, Baek S-W, Park WY, Kim W-T, Kim S-K, Roh Y-G, Jeong M-S, Yang G-E, Lee J-H, Chung JW et al (2020) E2F1 promotes progression of bladder cancer by modulating RAD54L involved in homologous recombination repair. Int J Mol Sci 21(23):9025. https://doi.org/10.3390/ijms21239025
Pavan ICB, Peres de Oliveira A, Dias PRF, Basei FL, Issayama LK, Ferezin C, Silva FR, Rodrigues de Oliveira AL, Dos Reis A, Moura L, Martins MB et al (2021) On broken ne(c)ks and broken DNA: the role of human NEKs in the DNA damage response. Cells 10(3):507. https://doi.org/10.3390/cells10030507
Perez-Peña J, Corrales-Sánchez V, Amir E, Pandiella A, Ocana A (2017) Ubiquitin-conjugating enzyme E2T (UBE2T) and denticleless protein homolog (DTL) are linked to poor outcome in breast and lung cancers. Sci Rep 7:17530. https://doi.org/10.1038/s41598-017-17836-7
Pinto V, Bergantim R, Caires HR, Seca H, Guimarães JE, Vasconcelos MH (2020) Multiple myeloma: available therapies and causes of drug resistance. Cancers (basel) 12(2):407. https://doi.org/10.3390/cancers12020407
Rosenbaum JC, Bonilla B, Hengel SR, Mertz TM, Herken BW, Kazemier HG, Pressimone CA, Ratterman TC, MacNary E, De Magis A et al (2019) The Rad51 paralogs facilitate a novel DNA strand specific damage tolerance pathway. Nat Commun 10:3515. https://doi.org/10.1038/s41467-019-11374-8
Roy P, Sarkar UA, Basak S (2018) The NF-κB activating pathways in multiple myeloma. Biomedicines 6(2):59. https://doi.org/10.3390/biomedicines6020059
Ruiz S, Santos E, Bustelo XR (2009) The use of knockout mice reveals a synergistic role of the Vav1 and Rasgrf2 gene deficiencies in lymphomagenesis and metastasis. PLoS ONE 4(12):e8229. https://doi.org/10.1371/journal.pone.0008229
Spies J, Waizenegger A, Barton O, Sürder M, Wright WD, Heyer W-D, Löbrich M (2016) Nek1 regulates Rad54 to orchestrate homologous recombination and replication fork stability. Mol Cell 62(6):903–917. https://doi.org/10.1016/j.molcel.2016.04.032
Szalat R, Avet-Loiseau H, Munshi NC (2016) Gene expression profiles in myeloma: ready for the real world? Clin Cancer Res 22(22):5434–5442. https://doi.org/10.1158/1078-0432.CCR-16-0867
Thompson SL, Compton DA (2010) Proliferation of aneuploid human cells is limited by a p53-dependent mechanism. J Cell Biol 188(3):369–381. https://doi.org/10.1083/jcb.200905057
Ueki T, Nishidate T, Park JH, Lin ML, Shimo A, Hirata K, Nakamura Y, Katagiri T (2008) Involvement of elevated expression of multiple cell-cycle regulator, DTL/RAMP (denticleless/RA-regulated nuclear matrix associated protein), in the growth of breast cancer cells. Oncogene 27(43):5672–5683. https://doi.org/10.1038/onc.2008.186
Walker BA, Mavrommatis K, Wardell CP, Ashby TC, Bauer M, Davies FE, Rosenthal A, Wang H, Qu P, Hoering A et al (2018) Identification of novel mutational drivers reveals oncogene dependencies in multiple myeloma. Blood 132(6):587–597. https://doi.org/10.1182/blood-2018-03-840132
Xia J, He Y, Meng B, Chen S, Zhang J, Wu X, Zhu Y, Shen Y, Feng X, Guan Y et al (2020) NEK2 induces autophagy-mediated bortezomib resistance by stabilizing Beclin-1 in multiple myeloma. Mol Oncol 14(4):763–778. https://doi.org/10.1002/1878-0261
Zohud BA, Wang M, Cai X (2020) Germline RAD54L with somatic POLE defect implicated in Hypermutation phenotype: case report. BMC Gastroenterol 20:255. https://doi.org/10.1186/s12876-020-01403-y
Acknowledgements
The authors would like to thank the Director General of Health, Ministry of Health Malaysia for approval to publish this research paper. We would also like to thank the Deputy Director General of Health (Research and Technical Support) and the Director of the Institute for Medical Research (IMR) for their support in the publication of this paper.
Funding
This study was funded by a research grant from the Ministry of Health Malaysia (NMRR-18-729-41414), awarded to Bong IPN.
Author information
Authors and Affiliations
Contributions
Conceptualisation: IPNB; formal analysis: IPNB; methodology: IPNB, NO; funding acquisition: IPNB; writing—original draft: IPNB; writing—review and editing: IPNB, CCN; final approval of the version to be published: EE.
Corresponding author
Ethics declarations
Conflict of interest
Bong IPN, Ng CC, Othman N and Esa E declare that they have no conflict of interest.
Ethical approval
This study was approved by the Medical Research & Ethics Committee (MREC), Ministry of Health, Malaysia. Informed consent was obtained from all subjects recruited in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13258_2022_1272_MOESM1_ESM.xlsx
Supplementary file1 Significant up-regulated probes identified by mRNA expression profiling (≥2.0 fold change, P < 0.05) (XLSX 243 KB)
13258_2022_1272_MOESM2_ESM.xlsx
Supplementary file2 Significant down-regulated probes identified by mRNA expression profiling (≥2.0 fold change, P < 0.05) (XLSX 225 KB)
13258_2022_1272_MOESM3_ESM.tif
Supplementary file3 Heatmap showing up-regulated probes and detailed view of the newly identified probes RAD54L, DIAPH3, SHCBP1, SKA3, ANLN (fold change ≥2.0; P < 0.05) (TIF 1037 KB)
13258_2022_1272_MOESM4_ESM.tif
Supplementary file4 Heatmap showing down-regulated probes and detailed view of the newly identified probes HKDC1, RASGRF2 and CYSLTR2 (fold change ≥2.0; P < 0.05) (TIF 1035 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bong, I.P.N., Ng, C.C., Othman, N. et al. Gene expression profiling and in vitro functional studies reveal RAD54L as a potential therapeutic target in multiple myeloma. Genes Genom 44, 957–966 (2022). https://doi.org/10.1007/s13258-022-01272-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-022-01272-7