Abstract
Background
Immunoglobulin A nephropathy (IgAN) is one of the most common primary forms of glomerulonephritis, while IgA vasculitis (IgAV) is the most common systemic vasculitis in children.
Objective
Herein we aimed to uncover single nucleotide polymorphism (SNP) markers associated with these two related diseases by applying association tests and Sanger sequencing.
Methods
Within the discovery stage, genomic DNA in blood samples from 101 enrolled patients were genotyped by the Korean Biobank Array. Association tests were performed with 397 Korean reference genomes. In the validation stage, 26 independent samples were genotyped by Sanger sequencing.
Results
Four SNPs were identified (P < 5 × 10–8) in the discovery stage. To determine whether the genotypes determined by SNP array were accurate, additional genotyping via Sanger sequencing was performed. As a result, only one SNP, rs9428555, was properly genotyped. In the validation stage, the minor allele (A > G) was found in as many as 15 out of 26 samples (minor allele frequency = 0.288), even though this minor allele is rare in East Asians (< 3%).
Conclusions
We found rs9428555 as a novel susceptible locus associated with the development of both IgAN and IgAV in Koreans. Though we cannot conclude rs9428555 is the unique susceptible locus of IgAN and IgAV, it is likely a good marker as the minor allele of this SNP occurred much more often in the patient group here versus within East Asians as a whole.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Immunoglobulin A nephropathy (IgAN) is one of the most common primary forms of glomerulonephritis in children and adolescents worldwide, and is characterized by predominant IgA-containing immune complex deposits within the glomerular mesangium upon renal biopsy (Noel et al. 1987). IgA vasculitis (IgAV), formerly known as Henoch-Schönlein purpura, is the most common form of systemic vasculitis in children and is also characterized by IgA-containing immune complex in small vessels (Trnka 2013). Renal involvement of IgAV (IgAV nephritis) occurs in about 30% of IgAV patients and is histologically indistinguishable from IgAN (Davin and Coppo 2014).
Though the pathogeneses of IgAN and IgAV and/or nephritis remain unclear, the multi-hit hypothesis, including production of galactose-deficient IgA, autoantibodies that recognize abnormal IgA1, their subsequent immune complexes formation and glomerular deposition has been widely supported by many studies (Davin and Coppo 2014; Suzuki 2019). IgAN and IgAV and/or nephritis are considered to be related diseases due to their similar histological features and IgA abnormalities, as well as the occurrence of IgAV and IgAN among identical twins or within the same patient (Kamei et al. 2016; Suzuki et al. 2018). Notably, serum levels of galactose-deficient IgA are high in both groups of patients—those with IgAN and IgAV nephritis—and their asymptomatic first-degree relatives (Hastings et al. 2010; Kiryluk et al. 2011).
Genetic background is considered important for the development or progression of disease (Yeo et al. 2018). The prevalence of both the aforementioned diseases varies between different ethnicities, and is higher within Asian populations (Oni and Sampath 2019; Schena and Nistor 2018). However, the causative genes of IgAN/IgAV have not yet been identified, probably because these diseases are polygenic or highly influenced by environment (Kiryluk et al. 2014; Yeo et al. 2018). In an effort to explore the genetic susceptibility markers of these diseases, several large-scale genome-wide association studies (GWAS) have been performed in IgAN patients, mainly with cohorts of European and East Asian ancestry, and nearly 20 risk variants were identified (Li and Yu 2018; Neugut and Kiryluk 2018). In respect to IgAV and/or nephritis, to date there is only one GWAS study with 308 IgAV patients and 1018 controls from Spain, reported by Lopez-Mejias et al. (2017). Within a Korean cohort of IgAN patients, Jeong et al. reported a new novel susceptible locus rs2296136 within the gene ANKRD16 as a candidate marker (Jeong et al. 2019).
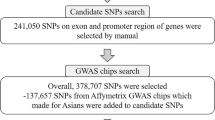
However, there has been no report regarding genome-wide association of pediatric-onset IgAN and IgAV patients. Thus, the aim of this study was to identify novel genetic susceptibility loci for the two related diseases IgAN and IgAV and/or nephritis, particularly in Korean children and adolescents (Fig. 1).
Methods
Subject
A total of 127 individuals were enrolled for this study, of whom all provided informed consent. We conducted a two-stage analysis: the first stage (i.e. discovery cohort) consisted of 101 cases, and the second (i.e. validation cohort) involved a replication analysis of the top single nucleotide polymorphism (SNP) signals that were identified during the discovery phase for 26 additional cases. Whole genome sequencing data (depth > 30x×for 397 average Koreans accessed from the Korean Reference Genome Database (KRGDB; http://coda.nih.go.kr/coda/KRGDB/) were used with approval as the normal control set. One hundred and twenty-seven cases included IgAN patients (n = 57) or those of IgAV with or without nephritis (n = 70), who were diagnosed before the age of 18 and followed at two pediatric nephrology centers (Seoul National University Children’s Hospital and Bucheon St. Mary’s Hospital of the Catholic University of Korea). All IgAN patients were diagnosed via renal biopsy. IgAV diagnoses were clinically determined according to the European League Against Rheumatism/Pediatric Rheumatology International Trials Organisation/Pediatric Rheumatology European Society criteria, including purpura or petechial with lower limb predominance and at least one of the four following features: acute onset abdominal pain, histopathology exhibiting leukocytoclastic vasculitis or proliferative glomerulonephritis with predominant IgA deposition, arthralgia or arthritis, and renal involvement. Renal involvement, also known as IgA nephritis, is defined as when IgAV patients have proteinuria or hematuria during the disease course (Ozen et al. 2010). IgAN or IgAV secondary to other conditions—including systemic lupus erythematosus, chronic hepatitis, diabetes, or cancer—were excluded. The first stage (Genomic DNA samples collected before August 2019) included 48 patients with IgAN, 10 with IgAV and 43 having IgAV nephritis, while the second stage (Genomic DNA samples collected after August 2019) included 9 IgAN patients and 12 with IgAV and 5 patients of IgAV nephritis. Study procedures were carried out in accordance with the Declaration of Helsinki. The Institutional Review Boards of Bucheon St. Mary’s Hospital of the Catholic University of Korea (IRB No. HC18TNDI0012) and Seoul National University Hospital (No. 1808-157-967) approved this study. All participants and their parents gave written informed consent. Patient demographics and clinical information are summarized in Table 1.
Genotyping
Genomic DNA was extracted from peripheral blood samples collected in tubes containing EDTA. We genotyped these samples via Korea Biobank Array (DNA Link Inc., Seoul, Korea), which consists of more than 800,000 markers, including > 247,000 rare-frequency variants within Koreans (Moon et al. 2019). To validate four SNP markers, genomic DNA of 14 samples previously genotyped during the discovery stage and 26 which were not used in the discovery stage, were separately prepared and genotyped via Sanger sequencing (Cosmo GeneTech, Seoul, Korea). Four primer pairs, (GAGCCAGATGCCTTACACCA and TAAGACATTTCAATCCAGCACAGC; TTGGCAGCTGGACTTACTGTT and CTTGGGATTCCTTCCAGGCT; ACTGAAAGCAATGGCTCAAAC and GTGTCTGACTTACTTCACTTAATATGC; CATGGCCTTGATTCAATCCT and TTGGAACTTGGTGTAAATGAGA) were used for Sanger sequencing of rs4926802, rs147294199, rs9428555, and rs11660485, respectively.
Identifying candidate SNP markers and statistical analysis
Out of 827,783 SNP markers within the Korean Biobank Array, only autosomal SNPs that were assigned as ‘Recommended’ by the SNPolisher package (Affymetrix) were selected. In addition, only SNPs with marker call rate < 0.05, Hardy–Weinberg Equilibrium p-value > 10–6, and minor allele frequency (MAF) < 0.01 were selected. After applying these filters, an association test was carried out for 509,500 SNP markers using PLINK 1.9 (Purcell et al. 2007). SNPs whose p-value for the association test was < 5 × 10–8 were considered as candidate SNPs.
Results
Four candidate loci identified in array-based discovery stages
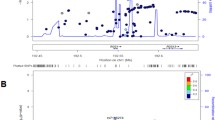
In the discovery stage, an association test was carried out for 101 test subjects and 397 healthy controls based on the genotypes of > 500,000 SNPs identified using the Korean Biobank Array. Based on p-values < 5 × 10–8 (See Methods), four loci (rs4926802 in CYP4Z1, rs147294199 in FAM151A, rs9428555 in intergenic region, and rs11660485 in SLC14A2) were selected and considered as candidate markers for further validation (Table 2). As shown in Quantile–Quantile plot and Manhattan plot in Fig. 2, p-values of the four SNPs were significantly different from those of other SNPs. More detailed regional Manhattan plots and plots of signal intensities for the four loci in the Korean Biobank Array were presented in Supplementary Figs. S1 and S2.
Resequencing by Sanger methods validated one SNP, rs9428555
All of the MAF values of candidate loci in control group were less than 0.05 (Table 2) and HWE p-values were relatively small. This implies that the list of candidates may include false positives or incorrectly genotyped. To verify whether the genotypes identified by SNP array were accurate, Sanger sequencing was carried out for the four above-mentioned SNPs using genomic DNA samples that were already used in the discovery stage. Eight samples having minor allele variant(s) were used for each SNP, with a resulting 14 samples used in total (Supplementary Table S1). Only one SNP, rs9428555, was validated with Sanger sequencing, while variant alleles were not identified in three out of four SNPs (Supplementary Table S1). Because we tested only four loci, we could not conclude what made this inconsistency, but additional verification seems necessary when using the result of Korean Biobank Array.
Sanger sequencing validation for independent samples
In the validation stage, we utilized blood samples from 26 patients that were not included in the discovery stage. Although this sample size is relatively small, we estimated that validation with this number of cases would be relevant because the determined variant (rs9428555) is known to be very rare in East Asians (Supplementary Table S2). Interestingly, 15 out of 26 patients harbored the minor allele (G) of the SNP marker (Supplementary Table S3; Supplementary Fig. S3), resulting in the MAF of the discovery cohort being 0.2885, which is significantly higher than those in all populations with the exception of Africans (MAF = 0.0278 in East Asians; Supplementary Table S2).
Minor allele frequencies in subgroups
As summarized in Table 1, our samples could be divided into three subgroups, IgAN, IgAVN, and IgAV. To check whether there were differences of allele frequencies among subgroups, we compared MAFs in all subgroups (Table 3) for the four loci identified in the discovery stage. For rs9428555, since we genotyped the locus in validation stage by Sanger sequencing, we also calculated MAFs for each subgroup in the validation samples. Without any exception, all of MAFs in subgroups were much higher than those in control set. Although we compared all pairwise combination of subgroups and calculated p-values based on chi-square test, we could not find any significant differences among subgroups in the three types of diseases (data not shown).
Investigation of functional roles of rs9428555
To the best of our knowledge, no reports or literature exist concerning the functional role of this variant, which is located within an intergenic region between PLD5 and CEP170. We additionally carried out literature search about PLD5 and CEP170. PLD5 is a gene encoding a protein, Phospholipase D (PLD) Family Member 5. We could not find any direct evidences refereeing relation between PLD5 and any disorders. Knockout study in a mouse could not detect any abnormalities (Karp et al. 2010). We could find only a weak evidence from the fact that both PLD and diacylglycerol kinase (DGK) are known to be involved in generation of phosphatidic acid (PA). In the previous two studies, mutations in DGKe, one of DGK members, were shown to lead hemolytic uremic syndrome, a type of Nephrotic syndrome (Lemaire et al. 2013; Ozaltin et al. 2013). CEP170 encodes a CEntrosomal Protein of 170 kDa. Unfortunately, we could not find any evidence that CEP170 is associated with IgAN or any neurological disorders. Even though we found a weak clue about association with PLD5 and a nephrotic syndrome, the distance from the rs9428555 to PLD5 are 383 kb. Thus, it is unlikely that a genotype of rs9428555 directly affect the functions of PLD5.
Then, we searched for related expression quantitative trait loci (eQTL) information in Genotype-Tissue Expression (GTEx) (Baran et al. 2015). However, we could not find any gene expression linked with rs9428555. Also, we tried to find epigenetic involvements by exploring ENCODE data using UCSC genome browser (Rosenbloom et al. 2013). Unfortunately, we could not detect any related known findings of epigenetic role of the locus.
We have investigated related variants using LDproxy, which is a part of LDlink (Machiela and Chanock 2015) and provides proxy and putatively functional variants for a query variant. We found three alleles highly related (D′ = 1, R2 > 0.99) to the identified variant rs9428555 (Supplementary Table S4) from the result of global population in 1,000 genomes (Genomes Project et al. 2015). All of these were within intergenic regions, but we ascertained that rs2491835, one of the three SNPs contains a probable sequence of binding motif of vitamin D receptor (VDR) based on RegulomeDB (Boyle et al. 2012).
Comparison to loci previously reported to be associated with IgAN
Recently, another GWAS study for Korean IgAN cohort was reported using customized DNA chip containing gene regions which were selected manually (Jeong et al. 2019). In that study, twelve previously reported IgAN loci were compared to other GWAS based on other populations. Out of the twelve ones, five loci were genotyped in our study. We compared the p-values of previously reported susceptible loci in our study and Jeong et al.’s result (Table 4). As Jeong et al. failed to show susceptible loci reported in other populations were also susceptible in Korean population, we also failed to common susceptible loci. One locus (rs660895 in HLA-DRB1) showed moderate association with IgAN (p < 0.05), which was identified in Han Chinese study (Yu et al. 2011).
Discussion
The present study represents the first GWAS of pediatric-onset IgAN and IgAV in a Korean population. We identified one novel susceptible SNP, rs9428555 in chromosome 1. Although we tried to find any direct pathological association between the locus and the disease, we only find an indirect and weak evidence about VDR binding motif. VDR binding had known to be significantly enriched in alleles associated with various diseases and phenotypes identified by GWAS (Ramagopalan et al. 2010). VDR binding sites in the human genome have been reported to be related to the evolution hypothesis regarding skin color (Jablonski and Chaplin 2000; Ramagopalan et al. 2010). Although we could not identify a detailed functional mechanism linked to this variant, its notable frequency provides some evidence that the variant may affect vitamin D-related pathway(s) and renal functions. In support, association of the VDR with nephropathies, including IgAN, have been elucidated in many studies (Yang et al. 2018). Several studies reported that VDR gene polymorphisms are associated with increased susceptibility to glomerulonephropathies, such as: diabetic nephropathy, lupus nephritis, and chronic renal failure (Azab et al. 2016; Li et al. 2018; Yang et al. 2017). Recently, Mo et al. reported that a VDR gene FokI polymorphism is associated with an increased risk of renal dysfunction in IgAN patients within a Chinese population (Mo et al. 2019). In addition, treatments with VDR agonist exhibited therapeutic effects in IgAN patients and an IgAN rat model via regulation of immune responses (Deng et al. 2017; Yuan et al. 2017). Additional studies could help to elucidate the association of rs9428555 with the VDR gene and its functional role for the development or prognosis of IgAN.
Interestingly, the MAF of rs9428555 has previously been shown to be significantly higher only in African populations (Supplementary Table S2), even considering that risk allele frequencies were known to be higher than those of non-African populations (+ 1.15% on average) (Kim et al. 2018). However, in the present study this MAF was also found to be higher in IgAN and IgAV patients. Therefore, theoretically the incidence of IgAN or IgAV would be expected to be high in African populations. Nevertheless, the incidence of IgAN in Africa is known to be lower than that of other populations (Schena and Nistor 2018). The causes of this phenomenon could not be determined here, but we may speculate that other important variants or environmental factors, such as local pathogen variation, may affect the occurrence of these diseases because IgAN and IgAV are polygenic (Kiryluk et al. 2011).
To date, several large-scale GWAS for IgAN patients within East Asian populations have been conducted, primarily within the Han Chinese (Li and Yu 2018). These showed that several loci encoding proteins playing roles in innate immunity, complement activation, regulation of mucosal IgA production and intestinal mucosal barrier maintenance are associated with the development of IgAN, and these identified loci are also related with other immune-mediated diseases such as inflammatory bowel disease (Kiryluk et al. 2014; Li and Yu 2018). Among these loci, one variant in complement factor H (rs6677604) identified as an IgAN-susceptible variant by GWAS, was also associated with the development of IgAV nephritis in a Chinese population (Jia et al. 2020). Herein rs9428555 was associated with the development of both IgAN and IgAV. These data may support that IgAN and IgAV are related diseases. However, affecting alleles were not consistently found within other East Asian populations. The results for pediatric-onset IgAN and IgAV in the Korean population described here were also not consistent with those for adult IgAN patients within the same ethnicity (Jeong et al. 2019). Furthermore, age at disease onset may be affected by the cumulative burden of risk-associated alleles for the development of IgAN or IgAV (Kiryluk et al. 2014). Thus, large-scale cohort studies including pediatric-onset IgAN and IgAV patients should be conducted to elucidate the genetic effects for occurrence or progression of these diseases.
The present study has some limitations in that this association study was conducted for a relatively small number of patients, therefore sensitivity is not guaranteed. In other words, other loci may be identified as IgAN susceptible loci if subsequent study is conducted using larger cohort. Nevertheless, this study is the first to identify rs9428555, a SNP marker associated with pediatric-onset IgAN and IgAV by applying an association test, and the SNP was successfully validated by Sanger sequencing in an independent cohort. In other words, there may be other SNP markers but at least the SNP (rs9428555) is not a false positive and is an apparent marker associated with IgAN and IgAV. Unfortunately, however, the SNP was located in intergenic region, and we had difficultly to find reliable functional link between the locus and the disease. Thus, one limitation is that we could not clearly explain why A > G in rs9428555 is associated with IgAN. However, we still believe the SNP can be a possible marker for IgAN according to our observations although we could not answer a mechanism.
In summary, we identified one candidate SNP marker, rs9428555 for both pediatric-onset IgAN and IgAV. We could not find any previous studies or directly related annotations in multiple types of databases. Although it is unclear whether the marker is functionally and directly relevant to the disease or not, it could be utilized as a genetic marker diagnosing IgAN or IgAV as its MAF is quite rare in East Asians.
References
Azab SF, Ali YF, Farghaly MA, Hamed ME, Allah MA, Emam AA, Abdelsalam NI, Hashem MI, Gawish HH, Nabil RM et al (2016) Vitamin D receptor gene BsmI polymorphisms in Egyptian children and adolescents with systemic lupus erythematosus: a case-control study. Medicine (baltimore) 95:e5233
Baran Y, Subramaniam M, Biton A, Tukiainen T, Tsang EK, Rivas MA, Pirinen M, Gutierrez-Arcelus M, Smith KS, Kukurba KR et al (2015) The landscape of genomic imprinting across diverse adult human tissues. Genome Res 25:927–936
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, Karczewski KJ, Park J, Hitz BC, Weng S et al (2012) Annotation of functional variation in personal genomes using RegulomeDB. Genome Res 22:1790–1797
Davin JC, Coppo R (2014) Henoch-Schonlein purpura nephritis in children. Nat Rev Nephrol 10:563–573
Deng J, Zheng X, Xie H, Chen L (2017) Calcitriol in the treatment of IgA nephropathy with non-nephrotic range proteinuria: a meta-analysis of randomized controlled trials. Clin Nephrol 87:21–27
Ferreira RC, Pan-Hammarstrom Q, Graham RR, Gateva V, Fontan G, Lee AT, Ortmann W, Urcelay E, Fernandez-Arquero M, Nunez C et al (2010) Association of IFIH1 and other autoimmunity risk alleles with selective IgA deficiency. Nat Genet 42:777–780
Genomes Project C, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, Marchini JL, McCarthy S, McVean GA et al (2015) A global reference for human genetic variation. Nature 526:68–74
Hastings MC, Moldoveanu Z, Julian BA, Novak J, Sanders JT, McGlothan KR, Gharavi AG, Wyatt RJ (2010) Galactose-deficient IgA1 in African Americans with IgA nephropathy: serum levels and heritability. Clin J Am Soc Nephrol 5:2069–2074
Jablonski NG, Chaplin G (2000) The evolution of human skin coloration. J Hum Evol 39:57–106
Jeong KH, Kim JS, Lee YH, Kim YG, Moon JY, Kim SK, Kang SW, Kim TH, Lee SH, Kim YH et al (2019) Genome-wide association study identifies new susceptible loci of IgA nephropathy in Koreans. BMC Med Genom 12:122
Jia M, Zhu L, Zhai YL, Chen P, Xu BY, Guo WY, Shi SF, Liu LJ, Lv JC, Zhang H (2020) Variation in complement factor H affects complement activation in immunoglobulin A vasculitis with nephritis. Nephrology (carlton) 25:40–47
Kamei K, Ogura M, Sato M, Ito S, Ishikura K (2016) Evolution of IgA nephropathy into anaphylactoid purpura in six cases–further evidence that IgA nephropathy and Henoch-Schonlein purpura nephritis share common pathogenesis. Pediatr Nephrol 31:779–785
Karp NA, Baker LA, Gerdin AK, Adams NC, Ramirez-Solis R, White JK (2010) Optimising experimental design for high-throughput phenotyping in mice: a case study. Mamm Genome 21:467–476
Kim MS, Patel KP, Teng AK, Berens AJ, Lachance J (2018) Genetic disease risks can be misestimated across global populations. Genome Biol 19:179
Kiryluk K, Moldoveanu Z, Sanders JT, Eison TM, Suzuki H, Julian BA, Novak J, Gharavi AG, Wyatt RJ (2011) Aberrant glycosylation of IgA1 is inherited in both pediatric IgA nephropathy and Henoch-Schonlein purpura nephritis. Kidney Int 80:79–87
Kiryluk K, Li Y, Scolari F, Sanna-Cherchi S, Choi M, Verbitsky M, Fasel D, Lata S, Prakash S, Shapiro S et al (2014) Discovery of new risk loci for IgA nephropathy implicates genes involved in immunity against intestinal pathogens. Nat Genet 46:1187–1196
Lemaire M, Fremeaux-Bacchi V, Schaefer F, Choi M, Tang WH, Le Quintrec M, Fakhouri F, Taque S, Nobili F, Martinez F et al (2013) Recessive mutations in DGKE cause atypical hemolytic-uremic syndrome. Nat Genet 45:531–536
Li M, Yu XQ (2018) Genetic determinants of IgA nephropathy: eastern perspective. Semin Nephrol 38:455–460
Li L, Wan Q, Yang S, Zhao S (2018) Impact of vitamin D receptor gene polymorphism on chronic renal failure susceptibility. Ther Apher Dial 22:575–587
Lopez-Mejias R, Carmona FD, Castaneda S, Genre F, Remuzgo-Martinez S, Sevilla-Perez B, Ortego-Centeno N, Llorca J, Ubilla B, Mijares V et al (2017) A genome-wide association study suggests the HLA Class II region as the major susceptibility locus for IgA vasculitis. Sci Rep 7:5088
Machiela MJ, Chanock SJ (2015) LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 31:3555–3557
Mo MQ, Pan L, Tan L, Jiang L, Pan YQ, Li FJ, Yang ZH, Liao YH (2019) Association between VDR gene FokI polymorphism and renal function in patients with IgA nephropathy. PeerJ 7:e7092
Moon S, Kim YJ, Han S, Hwang MY, Shin DM, Park MY, Lu Y, Yoon K, Jang HM, Kim YK et al (2019) The Korea Biobank array: design and identification of coding variants associated with blood biochemical traits. Sci Rep 9:1382
Neugut YD, Kiryluk K (2018) Genetic determinants of IgA nephropathy: western perspective. Semin Nephrol 38:443–454
Noel LH, Droz D, Gascon M, Berger J (1987) Primary IgA nephropathy: from the first-described cases to the present. Semin Nephrol 7:351–354
Oni L, Sampath S (2019) Childhood IgA vasculitis (Henoch Schonlein Purpura)—advances and knowledge gaps. Front Pediatr 7:257
Ozaltin F, Li B, Rauhauser A, An SW, Soylemezoglu O, Gonul II, Taskiran EZ, Ibsirlioglu T, Korkmaz E, Bilginer Y et al (2013) DGKE variants cause a glomerular microangiopathy that mimics membranoproliferative GN. J Am Soc Nephrol 24:377–384
Ozen S, Pistorio A, Iusan SM, Bakkaloglu A, Herlin T, Brik R, Buoncompagni A, Lazar C, Bilge I, Uziel Y et al (2010) EULAR/PRINTO/PRES criteria for Henoch-Schonlein purpura, childhood polyarteritis nodosa, childhood Wegener granulomatosis and childhood Takayasu arteritis: Ankara 2008. Part II: Final classification criteria. Ann Rheum Dis 69:798–806
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Ramagopalan SV, Heger A, Berlanga AJ, Maugeri NJ, Lincoln MR, Burrell A, Handunnetthi L, Handel AE, Disanto G, Orton SM et al (2010) A ChIP-seq defined genome-wide map of vitamin D receptor binding: associations with disease and evolution. Genome Res 20:1352–1360
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, Wong MC, Maddren M, Fang R, Heitner SG et al (2013) ENCODE data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res 41:D56-63
Schena FP, Nistor I (2018) Epidemiology of IgA nephropathy: a global perspective. Semin Nephrol 38:435–442
Suzuki H (2019) Biomarkers for IgA nephropathy on the basis of multi-hit pathogenesis. Clin Exp Nephrol 23:26–31
Suzuki H, Yasutake J, Makita Y, Tanbo Y, Yamasaki K, Sofue T, Kano T, Suzuki Y (2018) IgA nephropathy and IgA vasculitis with nephritis have a shared feature involving galactose-deficient IgA1-oriented pathogenesis. Kidney Int 93:700–705
Trnka P (2013) Henoch-Schonlein purpura in children. J Paediatr Child Health 49:995–1003
Yang L, Wu L, Fan Y, Ma J (2017) Vitamin D receptor gene polymorphisms in association with diabetic nephropathy: a systematic review and meta-analysis. BMC Med Genet 18:95
Yang S, Li A, Wang J, Liu J, Han Y, Zhang W, Li YC, Zhang H (2018) Vitamin D receptor: a novel therapeutic target for kidney diseases. Curr Med Chem 25:3256–3271
Yeo SC, Cheung CK, Barratt J (2018) New insights into the pathogenesis of IgA nephropathy. Pediatr Nephrol 33:763–777
Yu XQ, Li M, Zhang H, Low HQ, Wei X, Wang JQ, Sun LD, Sim KS, Li Y, Foo JN et al (2011) A genome-wide association study in Han Chinese identifies multiple susceptibility loci for IgA nephropathy. Nat Genet 44:178–182
Yuan D, Fang Z, Sun F, Chang J, Teng J, Lin S, Liu X (2017) Effect of vitamin D and tacrolimus combination therapy on IgA nephropathy. Med Sci Monit 23:3170–3177
Zhu L, Zhai YL, Wang FM, Hou P, Lv JC, Xu DM, Shi SF, Liu LJ, Yu F, Zhao MH et al (2015) Variants in complement factor H and complement factor H-related protein genes, CFHR3 and CFHR1, affect complement activation in IgA nephropathy. J Am Soc Nephrol 26:1195–1204
Acknowledgements
This work was supported by National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT (NRF-2017R1C1B5076675 and NRF-2017M3A9B6061511); and the Catholic Medical Center Research Foundation from the program year of 2017. Some of the Biospecimens and data used in this study were provided by the Biobank of Seoul National University Hospital, a member of Korea Biobank Network.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Minho Lee, Gunhee Lee, Hee Gyung Kang, and Jin-Soon Suh declare that they have no conflict of interest.
Ethical statement
The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The trial was conducted in accordance with the Declaration of Helsinki and the Harmonized Tripartite Guideline for Good Clinical Practice from the International Conference on Harmonization. This study was reviewed and approved by the Institutional Review Boards of Bucheon St. Mary’s Hospital of the Catholic University of Korea (IRB No. HC18TNDI0012) and Seoul National University Hospital (No. 1808–157-967). All patients enrolled completed the informed consent form.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, M., Lee, G., Kang, H.G. et al. New susceptible locus, rs9428555, is associated with pediatric-onset immunoglobulin A nephropathy and immunoglobulin A vasculitis in Koreans. Genes Genom 43, 1049–1057 (2021). https://doi.org/10.1007/s13258-021-01120-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01120-0