Abstract
Background
Acute coronary syndrome (ACS) is a complex cardiovascular disease whose development involves the dysregulation of adaptive immune responses. Though it has been proven that T cells associate with inflammation in the development of ACS, the function of B cells in disease remains unclear.
Objective
The aim of this study was to reveal the diversity of the B cell receptor (BCR) repertoire of patients with ACS.
Methods
We conducted a pilot study to sequence the immune repertoire of peripheral blood mononuclear cells (PBMCs) from patients with ACS, including acute myocardial infarction (AMI) and unstable angina (UA), and quantitatively characterized BCR repertoires by bioinformatics analysis.
Results
We found that patients with AMI and UA had lower BCR repertoire diversity compared with controls with normal coronary arteries (NCA). Lower percentages of productive unique BCR nt sequences and higher percentages of top 200 unique BCR sequences were identified in AMI and UA patients than NCA controls. Patients had various preferential usage of V and J genes from B cell clones in accordance with the disease severity of coronary arteries. AMI patients had distinct CDR3 amino acids, and their frequency differed among patients with ACS.
Conclusions
Our results indicate that differential BCR signatures represent an imprint of distinct repertoires among ACS patients. This study thereby opens up the prospect of studying disease-relevant B cells to better understand and treat ACS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiovascular disease (CVD) is the main cause of death worldwide, leading to 17.3 million deaths per year, and CVD mortality is expected to rise to 23.6 million by 2030 (Laslett et al. 2012; Smith et al. 2012). Acute coronary syndrome (ACS) is the most common and severe type of coronary heart disease (CHD), mainly affecting elderly and postmenopausal women, and manifests as acute myocardial infarction (AMI) and unstable angina (UA) (Allan et al. 2013; Saeed et al. 2017). It is associated with smoking, hypertension, diabetes, hyperlipidemia, abdominal obesity, and a family history of early-onset CHD. Hemostatic abnormalities such as changes in procoagulant activity also play a role as a pivotal event in the pathophysiology of CHD (Słomka et al. 2017, 2018).
Adaptive immunity dysregulation has an important function in ACS development (Crea and Liuzzo 2013). The role of T cells in ACS is well characterized (Liuzzo et al. 2013). In our previous study, we had revealed the distinct T cell receptor (TCR) repertoires in patients with ACS and demonstrated the presence of disease associated T-cell clonotypes (Liu et al. 2020). But the role of B cells remains unclear. B cells produces antibody variety of antibodies role in responses to stress, injury, and infection. Through cell–cell contact, antigen presentation, and cytokine production, and B cells participate in systemic and local immune responses in atherosclerotic arteries (Sage et al. 2019). B cell-associated protective immunity was reported to reduce disease progression of atherosclerosis in animal models (Caligiuri et al. 2002), suggesting that targeting the B cell activating factor pathway or the anti-CD20 antibody could reduce atherosclerosis; the balance of B cell subtypes was also proposed to be involved in the development of atherosclerosis (Ait-Oufella et al. 2010).
Immune repertoire sequencing (IR-SEQ) can be used to identify the complementarity-determining region (CDR) diversity of B cell receptors (BCRs), so could help us assess the diversity of the immune system and analyze the relationship between immune functions and disease (Georgiou et al. 2014; Jackson et al. 2013). In the present study, we analyzed the BCR repertoire in peripheral blood mononuclear cells (PBMCs) from patients with ACS using IR-SEQ. Our work focused on producing a comprehensive, unrestricted BCR immunogenetic characterization from ACS patients.
Materials and methods
Ethics statement
This study was performed in accordance with ethical standards specified by the Declaration of Helsinki and its amendments, and was approved by the Ethics Committee of Human Clinical Research at Meizhou People’s Hospital, Meizhou Hospital Affiliated to Sun Yat-sen University, Guangdong Province, China (No.: MPH-HEC 2016-A-43). Written consent was obtained from each patient.
Patients
Patients were diagnosed with ACS in the Cardiology Department of Meizhou People’s Hospital according to the published guidelines (Damman et al. 2017; Müller 2012). Briefly, UA patients should meet the following conditions: (1) angina-like chest pain or ischemic equivalent; (2) electrocardiographic abnormalities compatible on at least two contiguous leads; (3) at least one major pericardial vessel with > 70 % stenosis. AMI patients should meet the above conditions plus: (4) abnormalities above the upper normal limit for myocardial necrosis biomarkers (troponin and/or CKMB). Normal coronary artery (NCA) patients were defined as no stenosis in coronary arteries by quantitative coronary angiography and served as controls in this study. Patients were excluded if having following conditions: impaired left ventricular ejection fraction ≤ 45 %, congestive heart failure, chronic kidney or hepatic disease, malignant disease.
Baseline data including smoking status, the presence of diabetes mellitus and dyslipidemia, and blood lipid levels. To assess drug usage, we collected the usage of angiotensin-converting enzyme inhibitors or angiotensin antibodies, nitrate esters, clopidogrel, statins, Ca2+ antagonists and β-blockers.
Sample collection
Venous blood samples were collected and placed in EDTA anticoagulant tubes. Peripheral blood mononuclear cells (PBMCs) were separated immediately using density gradient centrifugation (Ficoll-Paque, GE Healthcare, Boston, USA), and stored in − 80 °C for use.
RNA extraction and library preparation
Total RNA was extracted from PBMCs using TRIzol™ reagent (Invitrogen, California, USA). RNA concentration and integrity was determined on a NanoDrop ND-2000 spectrophotometer (Thermo Fisher Scientific, Massachusetts, USA) and agarose gel electrophoresis. For cDNA synthesis, a total of 2 µg RNA was used for reverse transcription with SuperScript II reverse transcriptase (Invitrogen, California, USA) according to the manufacturer’s protocol. The cDNA was used as a template for library construction. Briefly, a pool of primers containing 12 V-area specific forward primers and 4 J-area reverse primers was applied for amplification of 55 functional V genes and 6 functional J genes using a commercial QIAGEN Multiplex PCR Kit (Qiagen, Dusseldorf, Germany). The sequences of primers are shown in Table 1. Amplification cycling was performed as follows: pre-denaturation at 95 °C for 15 min, then 30 cycles of 94 °C for 30 s, 60 °C for 90 s, and 72 °C for 30 s, with a final extension at 72 °C for 5 min. The PCR products were purified using the QIAquick PCR purification kit (Qiagen, Dusseldorf, Germany) and the sequence adaptors were ligated. The qualified library was used for sequencing.
High throughput sequencing
Cluster generation, template hybridization, isothermal amplification, linearization, blocking, denaturation, and hybridization of the sequencing primers were performed on IMonitor as previously described (Zhang et al. 2015). Paired-end sequencing of samples was carried out with a read length of 150 bp on an Illumina HiSeq™ Xten platform (Illumina, California, USA) in ShenZhen Realomics Inc.
Bioinformatics analysis
First, a quality control check was performed to filter data suitable for subsequent analysis with following parameters: average ratio of Q30 > 80 % and average error rate < 0.1 %. Trimmomatic and FLASH software was used to filter out the sequences of joints, barcode sequences, and read overlaps (Magoč and Salzberg 2011). Next, paired reads were merged, then paired-reads and single end-reads were aligned with mismatches and indels. Then, MiXCR was used for BCR sequence alignment over the entire length of the reference sequence to extract valid clone information (Bolotin et al. 2015). The aligned sequences were mapped to V, D, and J gene reference sequences in the IMGT database (http://www.imgt.org/) as previously described (Giudicelli and Lefranc 2011). Finally, the correct sequences were grouped into clonal types, such as CDR3, and reads of poor quality were collected for further analysis. Subsequent cloning was performed based on the information distribution, gene recombination, diversity analysis, and other in-depth mining; the complete process is shown in Fig. 1.
Statistical analysis
Data was represented as the mean ± SEM. Comparisons between groups were performed using one-way analysis of variance using GraphPad Prism software (Version 6.01). BCR diversity index distribution was shown using the Shannon–Weiner coefficient. The BCR overlap was calculated as described (Liaskou et al. 2016), based on the number of common amino acid clonotypes in two samples: (number of common amino acid clonotypes in the two samples × 2) / (total number of amino acid clonotypes in sample 1 + total number of amino acid clonotypes in sample 2) ×100. The average of all samples in each group was reported. The CDR3 aa grand average of hydropathy (GRAVY) was calculated based on the Kyte and Doolittle method. *P < 0.05 was considered statistically significant.
Results
Data description and BCR repertoire distribution in ACS patients
A total of 30 individuals were enrolled in three groups (NCA, AMI, and UA) with 10 patients per group (Table 2). There were no differences in serum lipid profiles or medical histories among the groups.
Total sequence numbers for BCR repertoire distributions were: 3.8 × 106 to 5.3 × 106 for NCA, 3.9 × 106 to 7.0 × 106 for AMI, and 3.9 × 106 to 5.5 × 106 for UA. After bioinformatics analysis, high-quality merged sequence numbers were: 3.6 × 106 to 5.0 × 106 for NCA, 3.6 × 106 to 6.6 × 106 for AMI, and 3.6 × 106 to 5.1 × 106 for UA. Detailed sequence information is shown in Table S1.
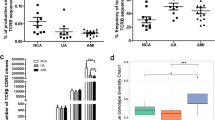
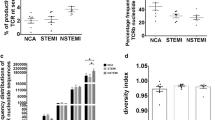
The clone distribution of BCR repertoires in NCA, AMI, and UA groups is shown in Fig. 1. The NCA group had the highest amino acid (aa) diversity (14.14 ± 0.53), which was significantly higher than that in AMI (11.75 ± 0.49, P < 0.01) and UA groups (12.02 ± 0.57, P < 0.05) (Fig. 2a), representing a higher diversity of BCR repertoires. The NCA group also had a higher percentage of productive unique BCR nucleotide (nt) sequences (Fig. 2b), but the lowest percentage of top200 B cell receptor repertoire nt sequences (27.38 ± 3.08) compared with AMI (45.07 ± 3.79, P < 0.01) and UA groups (42.69 ± 4.26, P < 0.01) (Fig. 2c). The number of BCR repertoires was similar among the three groups, but AMI patients had significantly higher copies of sequences present at > 1 % compared with the NCA group (18 and 50, respectively), and UA patients had significantly higher numbers present at 1–0.1 % compared with the NCA group (832 and 1,320, respectively, P < 0.05) (Fig. 2d). These results suggest that ACS patients have different clone distributions of BCR repertoires compared with other groups, while AMI and UA patients had more productive unique BCR nt sequences than individuals in the NCA group.
Clonal distribution of B cells in NCA, AMI, and UA patients. a The distribution of B cell clone frequencies through the measurement of amino acid diversity. b Percentage of productive unique BCR nt sequences in NCA, AMI, and UA patients. c The frequency of the top 200 BCR repertoire nt sequences in NCA, AMI, and UA patients. d The frequency distribution of BCR nt sequences from NCA, AMI, and UA patients. Data are represented as mean ± SEM of the group. Each dot represents a patient’s information. Differences between groups were compared using the t-test. **P < 0.01 and *P < 0.05. nt nucleotide
B cells from individuals in different groups show varying V and J gene usage
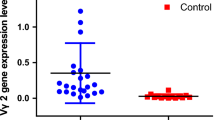
BCR repertoires were shown to differ among individuals, depending on genetic background, induced antigens, and viral infection. Because V-D-J segments form the oligoclonal expansion of B cells, we analyzed the distribution of V and J genes among individuals. Compared with the NCA group, UA patients had a significantly lower percentage usage of IGHV2 (5.89 ± 0.50 vs. 3.89 ± 0.56, P < 0.05), IGHV4 (17.72 ± 0.54 vs. 17.03 ± 0.90, P < 0.05), IGHV5 (13.64 ± 1.23 vs. 8.73 ± 1.21, P < 0.05), and IGHJ2P (1.00E-03 ± 1.92E-04, P < 0.05) segments, but a higher percentage usage of IGHV3 (24.14 ± 1.14, P < 0.05) (Fig. 3). Compared with the NCA group, AMI patients had a higher percentage usage of IGHV3 (24.14 ± 1.14 vs.31.69 ± 2.52, P < 0.05), but a lower usage of IGHV2 (5.89 ± 0.50 vs. 4.31 ± 0.54, P < 0.05) (Fig. 3). Regarding J gene usage, we only found a difference in IGHJ2P segment usage in AMI and UA patients compared with the NCA group (UA: 4.44E-04 ± 1.16 E-04, P < 0.05; AMI: 4.72E-04 ± 1.12 E-04, P < 0.05; and NCA: 1.00E-03 ± 1.92E-04). Other segments of V and J gene usage showed no difference among NCA, AMI, and UA patients (Figures S1 and S2). These data demonstrate a preferential usage of V and J genes from B cell clones in ACS patients.
V gene and J gene usage of clonotypes in the NCA, AMI, UA patients. Data show the percentage frequency of V gene (A: IGHV2, B: IGHV3, C: IGHV4, D: IGHV5) and J gene (E: IGHJ2P) usage by clonotypes in NCA, AMI, UA patients. Data show mean ± SEM frequency. Each dot represents each individual patient’s information. Data were compared to NCA group using paired t tests. **P < 0.01 and *P < 0.05
Overlap and distinct CDR3aa in BCR repertoires within rather than between groups
First, the degree of overlap in BCR repertoires both within groups and between groups was further analyzed. In the NCA group, an average overlap of 0.44 % was detected between any two patients, which was significantly higher than that in AMI patients (0.24 %, P < 0.05) and UA patients (0.27 %, P > 0.05) (Fig. 4a). For the unique clonotype overlap rate, 0.31 % was identified between NCA and AMI patients, and 0.28 % between NCA and UA patients (Fig. 4b). No difference was observed between groups. The hierarchical clustering of CDR3 aa found a total of 67 shared CDR3 aa between ACS and NCA patients (Fig. 5). Collectively, NCA patients had a higher average overlap than AMI and UA patients, and there was a sharing of BCR repertoires within AMI and UA groups.
The CDR3 region consists of V, D and J segments with no-template nucleotides inserted or deleted at junctional sites. To investigate immunology characteristics specific to each group, we analyzed the frequencies of CDR3 aa (Table 3). A total of 9 distinct CDR3 aa were found in the present study. Five CDR3 aa clonotypes, namely CASSIGRNTGELFF, CATSRDSSGANVLTF, CASSLVGANVLTF, CASSVGGGTYEQYF, CASSLVLDTQYF, which were prevalent in NCA (9/10) were absent in AMI patients, and were found in fewer UA patients (< 4/10). Two CDR3 aa clonotypes, namely CASSRAQETQYF and CASSLDRPYEQYF were less prevalent in AMI and UA patients compared to NCA. Another two CDR3 aa clonotypes, namely CASSLRYQETQYF and CSVEGQGYEQYF were prevalent in NCA and AMI patients, while only found in one UA patient. These data reveal frequency differences among distinct CDR3 aa clonotypes in ACS patients.
Discussion
High-throughput sequencing of BCR repertoire provides a way to advance our understanding of immune system by analyzing BCR properties in a large number of individuals. In this study, we characterized the BCR repertoire of ACS patients and found that: (1) AMI and UA patients have a lower CDR3 aa diversity compared with NCA patients;(2) AMI and UA patients have a lower percentage of productive unique BCR nt sequences, but a higher percentage of top200 BCR sequences compared with NCA patients; (3) a preferential usage of V and J genes exists among different ACS patients; (4) AMI patients have distinct CDR3 aa with different frequencies to ACS patients. Here, we found that five CDR3 aa (CASSIGRNTGELFF, CATSRDSSGANVLTF, CASSLVGANVLTF, CASSVGGGTYEQYF, and CASSLVLDTQYF) were absent in AMI patients but were present in nine of 10 NCA patients. The four CDR3 aa identified, CASSLRYQETQYF, CSVEGQGYEQYF, CASSRAQETQYF, and CASSLDRPYEQYF, were found in just one UA patient but in all NCA patients. Moreover, CASSRAQETQYF and CASSLDRPYEQYF were found in two AMI patients. Though we have identified different CDR3 aa among different groups of ACS patients, whether they affect the disease process or function to modulate inflammation in plaque formation remains unclear and requires further study.
ACS has been shown to correlate with inflammation. Moreover, inflammatory pathways not only regulate the formation of plaque in ACS patients, but also modulate the clinical consequences of thrombotic complications of atherosclerosis (Libby 2013). A balance exists between pro-inflammatory mechanisms and endogenous pathways that can promote the resolution of inflammation (Libby et al. 2014). T cells play an important role in the formation of plaque, similar to T-helper (Th) 1 cells. These secrete interferon-γ and activate macrophages to produce plaque-forming molecules such as Toll-like receptors, while Th2 and regulatory T cells secrete anti-inflammatory cytokines, like interleukin-10 and transforming growth factor-β, to delay inflammation (Ley et al. 2011; Libby et al. 2013). Studies have confirmed the role of macrophages and T cells in the formation and development of atherosclerotic plaque. It reveals the oxidized LDL and its oxidized specific antigen surface can recruit and activate macrophages and helper T cells, which can enlarge local inflammation and lead to atherosclerotic plaque progression.
Mature B cells express BCRs on the surface which specifically target exogenous antigens. The diversity of the BCR makes it possible for B cells to recognize thousands of millions self or non-self-invaders and protect against microbial infections. During the development of atherosclerosis, different B cell subsets such as B1 and B2 cell, play different roles. Natural IgM antibodies, mainly deriving from B1 cells, have been proven to mediate atheroprotective effects. The oxidized low density lipoprotein (oxLDL)–specific natural IgM antibodies produced by B1 cell, which can be enhanced by interleukin (IL)-5 stimulation, that block oxLDL uptake and foam cell formation(Fiskesund et al. 2010; Grönwall et al. 2012; Tsiantoulas et al. 2014; Tsimikas et al. 2012), but the role of other immunoglobulins, like IgG, in atherosclerosis is still unclear. B2 cells have been shown to promote inflammation and plaque complication via the B cell-activating factor (BAFF) (Kyaw et al. 2012, 2013), while the use of an anti-BAFFR antibody in ApoE−/− mice selectively deleted B2 cells and decreased plaque formation. A similar result was achieved using adoptive CD19+ B cell transfer or CD20 antibody treatment of ApoE−/− and Ldlr−/− mice (Ait-Oufella et al. 2010; Caligiuri et al. 2002; Kyaw et al. 2010).
Though the number of B cells in atherosclerotic plaques is small, a richness of B cells gather in the adventitia of atherosclerotic blood vessels to form local tertiary lymphoid tissue and regulate inflammatory reaction by antibodies and cytokines. The diversity of BCR repertoire has been found to be associated with many inflammatory diseases, such as multiple sclerosis (MS), systemic lupus erythematosus (SLE) and inflammatory bowel disease (IBD). Studies showed that variations in IGHV3 and IGHV 4, increased the risk of systemic lupus erythematosus (SLE), and SLE patients with such variations were more susceptible to nephritis, as well as higher titers of anti-DNA antibodies (Bashford-Rogers et al. 2018; Tipton et al. 2015). Previous study showed that B cell clonal expansion was observed in patients with MS, as well as the increased usage of the IGHV1 and IGHV4 genes (Beltrán et al. 2014). However, there are very limited studies on the relationship between BCR diversity and cardiovascular disease. A latest study by Zhang et al. found a skewed BCR repertoire in atherosclerosis plaques and revealed an association between BCR diversity and atherosclerosis (Zhang et al. 2021). In the present study, we found that ACS patients were more frequent in usage of IGVH3 gene, and the CDR3 aa diversity was significantly different in peripheral blood. We identified 9 distinct CDR3 aa clonotypes that were differentially distributed between NCA and ACS, which may have potential for clinical use as novel biomarkers.
There are some limitations in this study. First, the small sample size for IR-SEQ might compromise the conclusions. Second, the frequencies of CDR3 aa have not been further confirmed by clinical samples in a large scale. Third, the relationship between distinct CDR3 aa and ACS remains unclear, and more future work are needed to clarify it.
In summary, we report a comprehensive characterization of the immune profile of BCRs in ACS patients by IR-SEQ, and identified candidate distinct CDR3 aa. Our future work will investigate the role of CDR3 aa in the ACS disease process, characteristics of disease-associated clonotype aa, and antigenic triggers with the aim of developing target therapy for ACS patients.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
References
Ait-Oufella H, Herbin O, Bouaziz JD, Binder CJ, Uyttenhove C, Laurans L, Taleb S, Van Vré E, Esposito B, Vilar J et al (2010) B cell depletion reduces the development of atherosclerosis in mice. J Exp Med 207:1579–1587
Allan GM, Nouri F, Korownyk C, Kolber MR, Vandermeer B, McCormack J (2013) Agreement among cardiovascular disease risk calculators. Circulation 127:1948–1956
Bashford-Rogers RJM, Smith KGC, Thomas DC (2018) Antibody repertoire analysis in polygenic autoimmune diseases. Immunology 155:3–17
Beltrán E, Obermeier B, Moser M, Coret F, Simó-Castelló M, Boscá I, Pérez-Miralles F, Villar LM, Senel M, Tumani H et al (2014) Intrathecal somatic hypermutation of IgM in multiple sclerosis and neuroinflammation. Brain 137:2703–2714
Bolotin DA, Poslavsky S, Mitrophanov I, Mamedov IZ, Putintseva EV, Chudakov DM, Shugay M (2015) MiXCR: software for comprehensive adaptive immunity profiling. Nat Methods 12:380–381
Caligiuri G, Nicoletti A, Poirier B, Hansson GK (2002) Protective immunity against atherosclerosis carried by B cells of hypercholesterolemic mice. J Clin Invest 109:745–753
Crea F, Liuzzo G (2013) Pathogenesis of acute coronary syndromes. J Am Coll Cardiol 61:1–11
Damman P, van ‘t Hof AW, Ten Berg JM, Jukema JW, Appelman Y, Liem AH, de Winter RJ (2017) 2015 ESC guidelines for the management of acute coronary syndromes in patients presenting without persistent ST-segment elevation: comments from the Dutch ACS working group. Neth Heart J 25:181–185
Fiskesund R, Stegmayr B, Hallmans G, Vikström M, Weinehall L, de Faire U, Frostegård J (2010) Low levels of antibodies against phosphorylcholine predict development of stroke in a population-based study from northern Sweden. Stroke 41:607–612
Georgiou G, Ippolito GC, Beausang J, Busse CE, Wardemann H, Quake SR (2014) The promise and challenge of high-throughput sequencing of the antibody repertoire. Nat Biotechnol 32:158–168
Giudicelli V, Lefranc MP (2011) IMGT/junctionanalysis: IMGT standardized analysis of the V-J and V-D-J junctions of the rearranged immunoglobulins (IG) and T cell receptors (TR). Cold Spring Harb Protoc 2011:716–725
Grönwall C, Akhter E, Oh C, Burlingame RW, Petri M, Silverman GJ (2012) IgM autoantibodies to distinct apoptosis-associated antigens correlate with protection from cardiovascular events and renal disease in patients with SLE. Clin Immunol 142:390–398
Jackson KJ, Kidd MJ, Wang Y, Collins AM (2013) The shape of the lymphocyte receptor repertoire: lessons from the B cell receptor. Front Immunol 4:263
Kyaw T, Tay C, Khan A, Dumouchel V, Cao A, To K, Kehry M, Dunn R, Agrotis A, Tipping P et al (2010) Conventional B2 B cell depletion ameliorates whereas its adoptive transfer aggravates atherosclerosis. J Immunol 185:4410–4419
Kyaw T, Tay C, Hosseini H, Kanellakis P, Gadowski T, MacKay F, Tipping P, Bobik A, Toh BH (2012) Depletion of B2 but not B1a B cells in BAFF receptor-deficient ApoE mice attenuates atherosclerosis by potently ameliorating arterial inflammation. PLoS One 7:e29371
Kyaw T, Cui P, Tay C, Kanellakis P, Hosseini H, Liu E, Rolink AG, Tipping P, Bobik A, Toh BH (2013) BAFF receptor mAb treatment ameliorates development and progression of atherosclerosis in hyperlipidemic ApoE(-/-) mice. PLoS One 8:e60430
Laslett LJ, Alagona P Jr, Clark BA 3rd, Drozda JP Jr, Saldivar F, Wilson SR, Poe C, Hart M (2012) The worldwide environment of cardiovascular disease: prevalence, diagnosis, therapy, and policy issues: a report from the American College of Cardiology. J Am Coll Cardiol 60:S1–S49
Ley K, Miller YI, Hedrick CC (2011) Monocyte and macrophage dynamics during atherogenesis. Arterioscler Thromb Vasc Biol 31:1506–1516
Liaskou E, Klemsdal Henriksen EK, Holm K, Kaveh F, Hamm D, Fear J, Viken MK, Hov JR, Melum E, Robins H et al (2016) High-throughput T-cell receptor sequencing across chronic liver diseases reveals distinct disease-associated repertoires. Hepatology 63:1608–1619
Libby P (2013) Mechanisms of acute coronary syndromes and their implications for therapy. N Engl J Med 368:2004–2013
Libby P, Nahrendorf M, Swirski FK (2013) Monocyte heterogeneity in cardiovascular disease. Semin Immunopathol 35:553–562
Libby P, Tabas I, Fredman G, Fisher EA (2014) Inflammation and its resolution as determinants of acute coronary syndromes. Circ Res 114:1867–1879
Liu S, Zhong Z, Zhong W, Weng R, Liu J, Gu X, Chen Y (2020) Comprehensive analysis of T-cell receptor repertoire in patients with acute coronary syndrome by high-throughput sequencing. BMC Cardiovasc Disord 20:253
Liuzzo G, Montone RA, Gabriele M, Pedicino D, Giglio AF, Trotta F, Galiffa VA, Previtero M, Severino A, Biasucci LM et al (2013) Identification of unique adaptive immune system signature in acute coronary syndromes. Int J Cardiol 168:564–567
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963
Müller C (2012) New ESC guidelines for the management of acute coronary syndromes in patients presenting without persistent ST-segment elevation. Swiss Med Wkly 142:w13514
Saeed A, Kampangkaew J, Nambi V (2017) Prevention of Cardiovascular Disease in Women. Methodist Debakey Cardiovasc J 13:185–192
Sage AP, Tsiantoulas D, Binder CJ, Mallat Z (2019) The role of B cells in atherosclerosis. Nat Rev Cardiol 16:180–196
Słomka A, Korbal P, Piekuś A, Pawliszak W, Anisimowicz L, Żekanowska E (2017) Plasma levels of the A subunit of factor XIII in patients undergoing off-pump coronary artery bypass surgery. Pol Arch Intern Med 127:550–553
Słomka A, Piekuś A, Kowalewski M, Pawliszak W, Anisimowicz L, Żekanowska E (2018) Assessment of the Procoagulant Activity of Microparticles and the Protein Z System in Patients Undergoing Off-Pump Coronary Artery Bypass Surgery. Angiology 69:347–357
Smith SC Jr, Collins A, Ferrari R, Holmes DR Jr, Logstrup S, McGhie DV, Ralston J, Sacco RL, Stam H, Taubert K et al (2012) Our time: a call to save preventable death from cardiovascular disease (heart disease and stroke). Circulation 126:2769–2775
Tipton CM, Fucile CF, Darce J, Chida A, Ichikawa T, Gregoretti I, Schieferl S, Hom J, Jenks S, Feldman RJ et al (2015) Diversity, cellular origin and autoreactivity of antibody-secreting cell population expansions in acute systemic lupus erythematosus. Nat Immunol 16:755–765
Tsiantoulas D, Diehl CJ, Witztum JL, Binder CJ (2014) B cells and humoral immunity in atherosclerosis. Circ Res 114:1743–1756
Tsimikas S, Willeit P, Willeit J, Santer P, Mayr M, Xu Q, Mayr A, Witztum JL, Kiechl S (2012) Oxidation-specific biomarkers, prospective 15-year cardiovascular and stroke outcomes, and net reclassification of cardiovascular events. J Am Coll Cardiol 60:2218–2229
Zhang W, Du Y, Su Z, Wang C, Zeng X, Zhang R, Hong X, Nie C, Wu J, Cao H et al. (2015) IMonitor: A Robust Pipeline for TCR and BCR Repertoire Analysis. Genetics 201:459-472
Zhang S, Zhang S, Lin Z, Zhang X, Dou X, Zhou X, Wang X, Wang Z, Zhang Q (2021) Deep sequencing reveals 10 the skewed B-cell receptor repertoire in plaques and the association between pathogens and atherosclerosis. Cell Immunol 360:104256
Acknowledgements
The author would like to thank other colleagues whom were not listed in the authorship of Center for Cardiovascular Diseases and Center for Precision Medicine, Meizhou People’s Hospital (Huangtang Hospital), Meizhou Hospital Affiliated to Sun Yat-sen University for their helpful comments on the manuscript.
Funding
This study was supported by Key Scientific and Technological Project of Meizhou People’s Hospital, (Grant No.: MPHKSTP-20170101 and MPHKSTP-20180101 to MD Zhixiong Zhong), Science and Technology Program of Meizhou (Grant No.:2018B027 to Dr Sudong Liu), the Guangdong Provincial Key Laboratory of Precision Medicine and Clinical Translational Research of Hakka Population (Grant No.:2018B030322003).
Author information
Authors and Affiliations
Contributions
RQW and ZXZ conceived and designed the experiments; recruited subjects and collected clinical data were performed by SDL and XDG. Data analysis and manuscript writing were performed by RQW and SDL. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This study was conducted in accordance with the declaration of Helsinki. This study was conducted with approval from the Ethics Committee of the Meizhou People’s Hospital (Huangtang Hospital), Meizhou Hospital Affiliated to Sun Yat-sen University, Meizhou 514031, P. R. China (No.: MPH-HEC 2016-A-43). Written informed consent was obtained from each patient..
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Weng, R., Liu, S., Gu, X. et al. Characterization of the B cell receptor repertoire of patients with acute coronary syndrome. Genes Genom 44, 19–28 (2022). https://doi.org/10.1007/s13258-021-01110-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-021-01110-2