Abstract
Background
Short hairpin RNAs (shRNAs) expressed from vectors have been used as an effective means of exploiting the RNA interference (RNAi) pathway in mammalian cells. Of several methods to express shRNA, a method of transcribing shRNAs embedded in microRNA precursors has been more widely used than the one that directly expresses shRNA from RNA polymerase III promoters because the microRNA precursor form of shRNA is known to cause lower levels of cytotoxicity and off-target effects than the overexpressed shRNAs from the RNA polymerase III promoters.
Objective
We study the primary sequence features of microRNA precursors, which enhance their processing into mature form, helps design more potent shRNA precursors embedded in microRNA precursors.
Methods
We measure the enhancement of gene knockdown efficiency by adding CNNC motifs in the 3′ flanking region of shRNA precursor embedded in the human miR-30a microRNA precursor.
Results
By systemically adding three CNNC motifs in the 3′ flanking region of shRNA precursor, we found that addition of two CNNC motifs saturates their enhanced knockdown ability of shRNA and that the CNNC motif in the + 17 to + 20 from the drosha cleavage site is most important for the shRNA-mediated target gene knock down. We also did see little knockdown of target gene expression by the shRNA precursor lacking CNNC motif.
Conclusion
Since genetic studies generally require techniques that could reduce gene expression at different degrees, the findings in this study will allow us to use RNAi for genetic studies of reducing gene expression at different degrees.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNA interference (RNAi) is a post-transcriptional mechanism that inhibits gene expression, mediated by double-stranded RNA and evolutionarily conserved in a variety of eukaryotes (Agrawal et al. 2003; Safe 2013). Based on the understanding of the RNAi mechanism, RNAi triggers, exogeneous materials that initiate RNAi, have been developed to inhibit gene expression in DNA or RNA sequence-specific manner (Elbashir et al. 2001; Agrawal et al. 2003; Tijsterman and Plasterk 2004; Buchman 2005).
The microRNAs, cellular and endogenous double-stranded RNAs that initiate RNAi response, are expressed as hairpin like structures in the primary RNA transcripts (pri-miRNA) (Gregory and Shiekhattar 2005; Kim 2005; Winter et al. 2009; Griffiths-Jones 2010). In the nucleus, a microprocessor, Drosha/DGCR8 complex, cleaves the basal stem of pri-miRNAs to release precursor miRNA (pre-miRNA) that is further processed by Dicer in the cytoplasm. The Dicer nuclease cleaves the hairpin region of pre-miRNA to produce a 19 ~ 21 bp (bp) RNA duplex with 2-nucleotide, 3′ overhang on each end of RNA duplex, of which one strand serves as guide to down-regulate protein translation or degrade complementary mRNA (Grishok et al. 2001; Lee et al. 2003; Denli et al. 2004; Liu et al. 2004; Song et al. 2004; Nakanishi et al. 2005).
DNA vectors that are introduced into cells by transfection or viral gene delivery system are often used to express shRNAs, single stranded RNAs with stem-loop structure, that inhibit the expression of target genes (Abbas-Terki et al. 2002; Song and Yang 2010). In the early version of DNA vectors, RNA polymerase III promoters such as H1 and U6 were used to express pre-form of shRNA because they have defined transcription start sites (Xia et al. 2003; Borchert et al. 2006; Mäkinen et al. 2006). Since these promoters highly express pre-shRNA, they often saturate the endogenous microRNA pathway, eventually causing cytotoxicity and increasing off-target effect due to the inaccurate cleavage of the stem-loop structure by Dicer (Grimm et al. 2006; Khan et al. 2009; Van Gestel et al. 2014). To circumvent these problems and to use shRNA technology in a tissue-specific manner, shRNA sequence was embedded in the microRNA precursor such as miR-30, miR-155 and miR17-92, which was then located in the intronic region of reporter gene whose transcription was carried out by RNA polymerase II (Zeng et al. 2002; Siolas et al. 2005; Stegmeier et al. 2005; Chung et al. 2006; Liu et al. 2008; Fellmann et al. 2013; Bofill-De Ros and Gu 2016).
Several studies have examined structural or sequence features that can affect the intracellular processing of pri-miRNAs. The structural features include the length of the hairpin with the stem (33 ~ 36 nt), the size of apical loop (10 ~ 23 nt), and the position as well as the size of bulge (GHG motif). Primary sequence motifs are UG motif at the basal stem junction, UGU motif at the apical loop, and CNNC motif at the 3′ flanking region (+ 16 or + 18 from the Drosha cleavage site) (Auyeung et al. 2013; Fellmann et al. 2013; Fang and Bartel 2015).
Among the primary sequence motifs, CNNC motif was analyzed in detail and shown to be essential for the processing of pri-miRNA to pre-miRNA. In fact, the processing efficiency of the microprocessor (pri- to pre-miRNA conversion) was increased when the CNNC motif was present in the 3′ flanking region of the pri-miRNA. This phenomenon was significantly reduced even when only one of the two “C” bases constituting the CNNC motif was mutated and the motif was highly conserved among the vertebrate species of the pri-miRNAs. About 59.4% of the human pri-miRNAs possess the CNNC motif. Typically, miR-30a and miR-16 pri-miRNAs showed significant differences in the processing efficiency of microprocessor depending on the presence of CNNC motif (Auyeung et al. 2013).
In this study, we investigated the significance of CNNC motif of the intronic shRNA precursors for their gene knockdown efficiency. We measured the effect of intronic miR-30a-based shRNA on the gene knockdown by the alteration of CNNC motifs in the 3′ flanking region of the shRNA precursor. First, to evaluate gene knockdown efficiency of various forms of shRNA precursor against murine p53, a stable cell line (CHO K1-EGFP-p53) were constructed in CHOK1 cell line in which EGFP fluorescent protein gene and murine p53 gene were expressed in the form of a fusion protein. Secondly, we constructed the expression vectors of shRNA precursors composed of different sequence motifs in the 3′ flanking region, transfected them into the CHO K1-EGFP-p53 stable cell line, and measured the reduction of EGFP fluorescence by FACS. As a result, the shRNA precursor containing both CNNC 1 and 2 motifs showed the highest degree of gene knockdown. The CNNC motif in the + 17 to + 20 from the drosha cleavage site is most important for the shRNA-mediated target gene knock down, and little knockdown of target gene expression was observed by the shRNA precursor lacking CNNC motif.
Materials and methods
Cell culture and transfection
CHO-K1 cells were grown in Dulbecco’s modified Eagle’s medium (DMEM) (CORNING, Corning, USA) supplemented with 10% FBS (Lonza, Basel, Switzerland), penicillin (100 U/ml) and streptomycin (100 µg/ml) at 37 °C and 5% CO2. To generate a stable cell line that expressed EGFP-murine p53 fusion protein, CHO K1 cells were transfected using the JetPRIME transfection reagent (PolyPlus, New York, USA) and selected in a medium containing 100 µg/ml hygromycin-B. Then, a homogeneous population expressing GFP was enriched by fluorescence-activated cell sorting. The CHO-K1 stable cell line expressing EGFP-p53 was maintained in DMEM containing 10% FBS, 1% penicillin/streptomycin, and 50 ug/ml hygromycin B.
For transient expression of the plasmids, 5 × 105 cells seeded into six-well plates 24 h before transfection were transfected with plasmids expressing various forms of shRNA (2 ug) using JetPRIME transfection reagent following the manufacturer’s instruction. Doxycycline was added 6 h after transfection to a final concentration of 1 ug/ml, and fresh medium with doxycycline (1 ug/ml) was added every 24 h.
Plasmids
The pCDNA3.1 plasmid was used to construct shRNA precursor sequence embedded in the miR-30a pri-miRNA in the intronic region of a reporter gene, TurboRFP. Its ORF was obtained by PCR amplification from PSD-95-pTagRFP (Addgene, Watertown, USA) and cloned right after CMV promoter in pCDNA3.1. The 5′ and 3′ regions of miR-30a precursor sequence were prepared by the synthesis of oligonucleotides, in which EGFP ORF was ligated between them using Xho I and EcoR I sites, eventually creating 5′-miR-30a region-Xho I-EGFP-EcoR I-3′-miR-30a region cassette. Then, the cassette was cloned in the intronic region after TurboRFP ORF. The final vector sequences were verified by sanger sequencing.
To assemble shRNA precursor sequences, shRNA sequences against target genes were cloned between 5′ and 3′ regions of the precursor using Xho I (NEB, Ipswich, USA) and EcoR I (NEB, Ipswich, USA) sites. In this cloning scheme, EGFP ORF was flanked by 5′ and 3′ region sequences of miR-30a precursor and released from the vector by treating Xho I and EcoR I.
The following sequence represents the intronic miR-30a shRNA precursor, in which sense and antisense strands were represented by underlined small letters, hairpin loop by italic bold capital letters: “TTGAATGAGGCTTCAGTACTTTACAGAATCGTTGCCTGCACATCTTGGAAACACTTGCTGGGATTACTTCTTCAGGTTAACCCAACAGAAGGCTCGAGAAGGTATATTGCTGTTGACAGTGAGCGAccctgtcatcttttgtcccttTAGTGAAGCCACAGATGTAaagggacaaaagatgacagggGTGCCTACTGCCTCGGAATTCAAGGGGCTACTTTAGGAGCAATTATCTTGTTTACTAAAACTGAATACCTTGCTATCTCTTTGATACATTTTTACAAAGCTGAATTAAAATGGTATAAATTAAATCACTTT”. This shRNA precursor was located in the intron after TurboRFP ORF.
For the experiments to reduce the expression of EGFP-p53 using p53-specific miR-30a, following shRNA sequences that have been known to knockdown the expression of murine p53 expression were replaced with the Xho I (CTCGAG)/EcoR I (GAATTC) fragment in the miR30a shRNA precursor.
p53-393 shRNA
CTCGAGAAGGTATATTGCTGTTGACAGTGAGCGAccctgtcatcttttgtcccttTAGTGAAGCCACAGATGTAaagggacaaaagatgacagggGTGCCTACTGCCTCGGAATTC.
p53-814 shRNA
CTCGAGAAGGTATATTGCTGTTGACAGTGAGCGCccactacaagtacatgtgtaaTAGTGAAGCCACAGATGTAttacacatgtacttgtagtggATGCCTACTGCCTCGGAATTC.
Negative control shRNA
CTCGAGAAGGTATATTGCTGTTGACAGTGAGCGAggatgtttcaccaaggaaataTAGTGAAGCCACAGATGTAtatttccttggtgaaacatccGTGCCTACTGCCTCGGAATTC.
The above oligonucleotide templates were PCR-amplified with the primers (Forward primer: 5′-GTGCAATGCTCGAGAAGGTATATTG-3′, Reverse primers: For CCTC motif, 5′-TAAGCGATGAATTCCGAGGCAGTAG-3′; for CATA motif, 5′-CTACTGCATAGGAATTCA TCGCTTA-3; for CATA–CTTC motif, 5′-CTACTGCATAGGACTTCAAGAATTCATCGCTTA-3; for CCTC–CTTC motif, 5′-CTACTGCCTCGGACTTCAAGAATTCATCGCTTA-3; for CCTC–CTTC–CTTC motif, 5′-CTACTGCCTCGG ACTTCGGCCTTCAAGAATTCATCGCTTA-3′).
Then Xho I/EcoR I digest of the PCR products was cloned into the same RE sites in the vector containing miR30a shRNA precursor.
Flow cytometric analysis
To quantitate the expression of fluorescence proteins by flow cytometer, CHO K1-EGFP-p53 stable cells transiently transfected with shRNA expression vector. After 48 h transfection, cells were harvested by the trypsin (0.25%) treatment, briefly washed with 1xPBS, and resuspended in the FACS buffer (1X PBS, 2% FBS).
A Moflo XDP flow cytometer (Beckman Coulter, Brea, USA) was used for the analysis. The instrument settings were as follows: log forward scatter (FSC) and log side scatter (SSC) at 440 V. The EGFP was excited at 488 nm and fluorescence was collected through 529/28-nm bandpass filter. The singlet cell population was gated on FSC and SSC. The typical sampling rate was 500 cells per second, and the typical sample size was 50,000 cells per measurement unless otherwise stated. The data were analyzed with Kaluza software (Beckman Coulter, Brea, USA).
Results and discussion
Flow cytometric assay system to quantitate the efficiency of gene knock-down by RNAi
The understanding about micro RNA biogenesis from pri-miRNA to mature microRNA is important to design effective shRNA with reduced cytotoxicity and off-target effects. Here we examined the significance of CNNC motif known to enhance the processing of pri-miRNA to pre-miRNA, for the processing of intronic shRNA precursor embedded in the human miR-30a pri-miRNA, in which the stem region of the pri-miRNA was replaced with the shRNA sequence complementary to a target gene, p53, without any miss-matched base pairs (Fig. 1). Thus, shRNA precursor becomes transcribed in an intron of a marker gene from an RNA polymerase II promoter, processed into siRNA following micro-RNA processing pathways of Drosha and Dicer complexes.
Sequence and 2° structure of the shRNA precursor embedded in the miR-30a miRNA. Stem region comprises of the 21 nucleotides long sense and antisense double-stranded RNA. Red arrows represent the canonical cleavage sites of Drosha and green arrows for the ones of Dicer. The regions highlighted by blue and red circles mean the cleavage sites of restriction enzymes of Xho I and EcoR I, which were used to clone shRNA precursors in the intronic shRNA expression vectors. In several vectors expressing intronic shRNA precursors, which have been widely used for expressing individual shRNA and genome-scale shRNA libraries, A CNNC motif located at the + 17 to + 20 nucleotides from the 3′ Drosha cleavage site in miR-30a miRNA (labeled as CNNC 2) was mutated to create an EcoR I site for the cloning of shRNA precursors (C residue highlighted in green hexagon was mutated into A residue highlighted in red circle). The miR-30a miRNA contains two CNNC motifs (labeled as CNNC 1 and CNNC 2) located at the + 10 to + 13 and + 17 to + 20 nucleotides from the 3′ Drosha cleavage site in miR-30a miRNA
Several studies showed that a CNNC motif at the 3′ flanking region (located + 17 ~ + 20 nucleotides from the Drosha cleavage site) of the pri-miR-30a is evolutionary conserved and the key factor determining the efficiency of its processing into its mature form. Since miR-30a pri-miRNA contains second CNNC motif at the 3′ flanking region (+ 10 ~ + 13 nucleotides from the Drosha cleavage site), we investigate whether the presence of multiple CNNC motifs in the 3′ flanking region of shRNA precursor embedded in the miR-30a pri-miRNA could additively enhance the processing of the precursor by measuring knockdown efficiency of a target gene, EGFP-p53 (Fig. 1).
For the quantitative measurement of a target gene knockdown by shRNA, we first established stable cell lines that uniformly express murine EGFP-p53 in CHO K1 cells. Out of several stable cell lines that showed different degrees of GFP signal, we chose the one expressing the highest level of GFP for subsequent study (Fig. 2), which is more suitable to measure the difference of knockdown efficiency by the DNA constructs having different numbers of CNNC motifs, in which the shRNA precursor against murine p53 located in the artificial intronic, 3′ UTR region of Turbo-RFP reporter gene (Fig. 3). Thus, the EGFP-p53 stable cells that express the shRNA precursor were identified by the expression of Turbo-RFP using flow cytometry or fluorescent microscopy, and the knockdown efficiency of EGFP-p53 was calculated as the decrease of GFP signal in a population of the stable cells expressing Turbo-RFP and thus intronic shRNA precursors in flow cytometric analyses.
A stable cell line expressing EGFP-murine p53. a A diagram of the vector used to make a stable cell line. The SRa promoter expresses a gene that confers hygromycin B-resistance. UBC promoter expresses EGFP-murine p53 fusion protein. b Microscopic analysis of the CHO K1 stable cell line. All cells in the field express EGFP-p53 protein. c FACS analysis of the stable cell line showing that almost all stable cells express equal level of GFP. Gray area: CHO K1 cells, Green area: CHO K1 stable cells expressing EGFP-p53. The GFP-positive cells were enriched once (used for the experiment in Fig. 4) or twice (used for the experiment in Fig. 5) by FACS
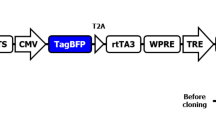
Vectors used to create fluorescent reporter for murine p53 knockdown and to express intronic shRNA precursors. a To create a CHO-K1 cell line that uniformly expressing murine p53-EGFP fusion protein, a plasmid vector was constructed in which a gene to confer hygromycin B-resistance was expressed from SRa promoter and EGFP-p53 from UBC promoter. After its transfection into CHO-K1 cell line, hygromycin B-resistant colonies from individual cells were independently grown to measure the expression levels and distribution of GFP by flow cytometry. The colony showing most uniform and highest level of GFP expression was used for the subsequent experiments. The p53-393 and p53-814 shRNA sequences are against the murine p53 ORF, known to efficiently knockdown its expression. b To express shRNA precursors from intron, a plasmid vector was constructed to express Turbo-RFP reporter and shRNA precursor in the intron region from CMV promoter. SD and SA represent spliced donor and spliced acceptor sites, respectively. To express shRNA precursor embedded in the miR-30a miRNA, 5′ and 3′ parts of the miRNA were cloned in the intronic region. For cloning different shRNA sequences, EcoR I (E)/Xho I (X) digest of PCR products prepared by using shRNA precursor oligonucleotides in Fig. 1 as template was replaced into the vector already containing a shRNA precursor
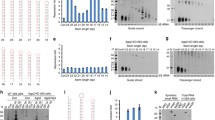
Effect of CNNC motifs on the shRNA-mediated gene knockdown. a Sequence information of the miR-30a 3′ flanking region containing one or two CNNC motifs. The first one (CNNC1) located at the + 10 ~ + 13 nucleotides from the Drosha cleavage site and the second one (CNNC2) at the + 17 ~ + 20 nucleotides. b Flow cytometric analysis of the CHO-K1 stable cell line expressing EGFP-p53 48 h after transient transfection of the constructs that have different combination of the CNNC motifs. NC represents negative control with scrambled shRNA, p53-814 shRNA is specific to murine p53. The value in percentage represent the fraction of cells containing reduced GFP signals compared to the control. Each FACS analysis was carried out with 1.0 × 104 cells. c The percentage of cells transiently transfected with the vectors containing p53-814 and reduced GFP signal. All four different constructs with scrambled shRNA (negative control), regardless of the presence of CNNC motifs, showed 8% of cells expressing basal level of GFP (dotted black line). The results represent the mean ± SD (n = 3). d The total amounts of GFP signal measured by transient transfection experiment with each p53-814 construct (without CNNC, with CNNC1, with CNNC2, with both CNNC1 and 2) were divided by the number of cells subjected to each flow cytometric analysis. The average EGFP intensity with the negative control was the same in all four constructs 94% (dotted black line) because a small fraction of stable cells used for the experiments express decreased amounts of GFP. The results represent the mean ± SD (n = 3)
Two CNNC motifs additively enhance the shRNA-mediated gene knockdown. a Sequence information of the miR-30a 3′ flanking region containing one to three CNNC motifs. The first one (CNNC1) located at the + 10 ~ + 13 nucleotides from the Drosha cleavage site, the second one (CNNC2) at the + 17 ~ + 20 nucleotides, and the third one (CNNC3) at the + 24 ~ + 27 nucleotides. b Flow cytometric analysis of the CHO-K1 stable cell line expressing EGFP-p53 48 h after transient transfection of the constructs that have different combination of the CNNC motifs. NC represents negative control with scrambled shRNA, p53-393 and p53-814 shRNAs are specific to murine p53. The value in percentage represent the fraction of cells containing reduced GFP signals compared to the control. c The percentage of cells transiently transfected with the vectors containing p53-393 or p53-814 and reduced GFP signal. The vectors also contain CNNC2 (yellow) CNNC1 and 2 (dark gray), and CNNC1, 2 and 3 (black). Flow cytometric data was analyzed to obtain percentage of cells separated from the main population of GFP-expressing cells after transient transfection of each construct. The results represent the mean ± SD (n = 2). d The total amounts of GFP signal measured by transient transfection experiment were divided by the number of cells subjected to each flow cytometric analysis as described in Fig. 4d. The average EGFP intensities with the negative controls were below 100% because a small fraction of stable cells used for the experiments express decreased amounts of GFP. The results represent the mean ± SD (n = 2)
CNNC motifs in the 3′ flanking region of the shRNA precursor enhance the knockdown efficiency of shRNA
Two CNNC motifs appear to additively enhance the knockdown efficiency of shRNA, although the one located at the + 17 ~ + 20 nucleotides from the Drosha cleavage site is the primary element that enhances the knockdown efficiency. Here, using a well-known shRNA sequence, p53-814, that specifically knockdowns the expression of murine p53, we measured whether the CNNC motifs could enhance the efficiency when the shRNA sequence was embedded in the intronic miR-30a pri-miRNA (Fig. 4). The first CNNC motif (CCTC) was located at the + 10 ~ + 13, and the second (CTTC) at the + 17 ~ + 20 nucleotides from the Drosha cleavage site. As shown in Fig. 3b, different degrees of GFP expression levels are observed by the flow cytometric analysis of GFP expression in the CHO K1-p53-GFP stable cells expressing Turbo-RFP after transiently transfecting the constructs having different combinations of CNNC motifs (Fig. 4a). The constructs expressing a scrambled, negative control shRNA did not cause any decreases of p53-GFP expression, regardless of the presence of CNNC motifs (Fig. 4). The p53-814 construct without any CNNC motifs or with the motif at the + 10 ~ + 13 nucleotides from the Drosha cleavage site did not cause any decrease of p53-GFP expression, but the construct with a CNNC motif at the + 17 ~ + 20 nucleotides from the Drosha cleavage site knockdown the expression of p53-GFP, which were measured by the decrease of GFP signal in Turbo-RFP-positive CHO-K1-p53-GFP stable cells and by the population of the cells showing reduced expression of p53-GFP (Fig. 4c, d). Interestingly, addition of the second CNNC motif at the + 10 ~ + 13 nucleotides to the construct containing the motif at the + 17 ~ + 20 nucleotides from the Drosha cleavage further enhanced the knockdown efficiency. Thus, two CNNC motifs in the 3′ flanking region of the shRNA precursor appear to additively enhance the knockdown efficiency of shRNA, which was expressed from the shRNA precursor in intron of a reporter gene whose transcription was derived from RNA polymerase II promoter.
Two CNNC motifs additively enhance the knockdown efficiency of shRNA
The success of increasing knockdown efficiency by introducing two CNNC motifs in the 3′ flanking region of shRNA precursor inspired us to examine the effect of adding a third CNNC motif on the region. As shown in the Fig. 5, the third CNNC motif was added at the + 24 ~ + 27 nucleotides from the Drosha cleavage. Thus, all three CNNC motifs were introduced 3 nucleotides after the previous one. In this experiment, two shRNA sequences against murine p53, p53-393 and p53-814, were used to measure the effect of adding multiple CNNC motifs in the constructs on the knockdown efficiency. The same kind of flow cytometric analysis was carried out as in the experiment described in Fig. 3. As explained above, both p53 shRNA sequences from the constructs having two CNNC motifs knockdown more efficiently than the constructs having one CNNC motif at the + 17 ~ + 20 nucleotides region, but addition of the third CNNC motif did not increase the knockdown efficiency compared to the constructs containing two CNNC motifs (Fig. 5).
Conclusions
Combined the experiments of adding multiple CNNC motifs to measure the knockdown efficiency of shRNA, we concluded that CNNC motif located at the + 17 ~ + 20 nucleotides from the Drosha cleavage site is the primary motif that could enhance the processing of shRNA precursor into shRNA, and the processing could be enhanced by the presence of the second CNNC motif, not by the subsequent, third motif. Since genetic studies generally require techniques that could reduce gene expression at different degrees, the findings in this study will allow us to use RNAi for genetic studies of reducing gene expression at different degrees.
References
Abbas-Terki T, Blanco-Bose W, Déglon N et al (2002) Lentiviral-mediated RNA interference. Hum Gene Ther 13:2197–2201
Agrawal N, Dasaradhi PVN, Mohmmed A et al (2003) RNA interference: biology, mechanism, and applications. Microbiol Mol Biol Rev 67:657–685
Auyeung VC, Ulitsky I, McGeary SE, Bartel DP (2013) Beyond secondary structure: primary-sequence determinants license Pri-miRNA hairpins for processing. Cell 152:844–858
Bofill-De Ros X, Gu S (2016) Guidelines for the optimal design of miRNA-based shRNAs. Methods 103:157–166
Borchert GM, Lanier W, Davidson BL (2006) RNA polymerase III transcribes human microRNAs. Nat Struct Mol Biol 13:1097–1101
Buchman TG (2005) RNAi. Crit Care Med 33:S441–S443
Chung KH, Hart CC, Al-Bassam S et al (2006) Polycistronic RNA polymerase II expression vectors for RNA interference based on BIC/miR-155. Nucleic Acids Res 34:e53
Denli AM, Tops BBJ, Plasterk RHA et al (2004) Processing of primary microRNAs by the microprocessor complex. Nature 432:231–235
Elbashir SM, Lendeckel W, Tuschl T (2001) RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev 15:188–200
Fang W, Bartel DP (2015) The menu of features that define primary microRNAs and enable de novo design of microRNA genes. Mol Cell 60:131–145
Fellmann C, Hoffmann T, Sridhar V et al (2013) An optimized microRNA backbone for effective single-copy RNAi. Cell Rep 5:1704–1713
Gregory RI, Shiekhattar R (2005) MicroRNA biogenesis and cancer. Cancer Res 65:3509–3512
Griffiths-Jones S (2010) MiRBase: microRNA sequences and annotation. Curr Protoc Bioinform 1291–12910
Grimm D, Streetz KL, Jopling CL et al (2006) Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441:537–541
Grishok A, Pasquinelli AE, Conte D et al (2001) Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106:23–34
Khan AA, Betel D, Miller ML et al (2009) Transfection of small RNAs globally perturbs gene regulation by endogenous microRNAs. Nat Biotechnol 27:549–555
Kim VN (2005) MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 6:376–385
Lee Y, Ahn C, Han J et al (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425:415–419
Liu J, Carmell MA, Rivas FV et al (2004) Argonaute2 is the catalytic engine of mammalian RNAi. Science 80-:305:1437–1441
Liu YP, Haasnoot J, ter Brake O et al (2008) Inhibition of HIV-1 by multiple siRNAs expressed from a single microRNA polycistron. Nucleic Acids Res 36:2811–2824
Mäkinen PI, Koponen JK, Kärkkäinen AM et al (2006) Stable RNA interference: comparison of U6 and H1 promoters in endothelial cells and in mouse brain. J Gene Med 8:433–441
Nakanishi Y, Oikawa M, Kumagai T et al (2005) Elastic property of Kondo semiconductor CeOs4Sb12. Phys B Condens Matter 359–361:907–909
Safe S (2013) RNA interference. Brenner’s Encycl Genet Second Ed 418:288–289
Siolas D, Lerner C, Burchard J et al (2005) Synthetic shRNAs as potent RNAi triggers. Nat Biotechnol 23:227–231
Song H, Yang P-C (2010) Construction of shRNA lentiviral vector. N Am J Med Sci 598–601
Song JJ, Smith SK, Hannon GJ, Joshua-Tor L (2004) Crystal structure of argonaute and its implications for RISC slicer activity. Science 80(305):1434–1437
Stegmeier F, Hu G, Rickles RJ et al (2005) A lentiviral microRNA-based system for single-copy polymerase II-regulated RNA interference in mammalian cells. Proc Natl Acad Sci 102:13212–13217
Tijsterman M, Plasterk RHA (2004) Dicers at RISC: the mechanism of RNAi. Cell 117:1–3
Van Gestel MA, Van Erp S, Sanders LE et al (2014) shRNA-induced saturation of the microRNA pathway in the rat brain. Gene Ther 21:205–211
Winter J, Jung S, Keller S et al (2009) Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 11:228–234
Xia XG, Zhou H, Ding H et al (2003) An enhanced U6 promoter for synthesis of short hairpin RNA. Nucleic Acids Res 31:e100
Zeng Y, Wagner EJ, Cullen BR (2002) Both natural and designed micro RNAs can inhibit the expression of cognate mRNAs when expressed in human cells. Mol Cell 9:1327–1333
Acknowledgements
This work was supported by the SGER Program through the NRF by the Ministry of Education to Y. Kee (Grant Number NRF-2015R1D1A1A02060346); Basic Science Research Program through the NRF by the Ministry of Education (Grant Number NRF-2015R1D1A3A01015641), the grant from the National R&D Program for Cancer Control, Ministry of Health & Welfare, Republic of Korea (Grant Number HA17C0035), and 2015 Research Grant from Kangwon National University to B.J. Hwang.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Park, S.K., Kee, Y. & Hwang, B.J. Enhancement of gene knockdown efficiency by CNNC motifs in the intronic shRNA precursor. Genes Genom 41, 491–498 (2019). https://doi.org/10.1007/s13258-019-00783-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-019-00783-0