Abstract
Buckwheat is a pseudocereal that belongs to family Polygonaceae. It is cultivated as a traditional crop in Asia, central Europe and eastern Europe and is used as food as well as medicine. It possesses high quality protein that lacks gluten and is rich in many nutraceuticals and antioxidants; due to which it is presently in great demand. In the present investigation, we collected the buckwheat germplasm from various regions of north western Himalayas of Jammu & Kashmir and Ladakh. The material was purified and then used for molecular variability studies. We used 15 Simple Sequence Repeat (SSRs) to study polymorphism among 52 genotypes of common buckwheat (Fagopyrum esculentum) and further, evaluated these SSRs for transferability in other buckwheat species. Among 15 SSRs isolated from common buckwheat; only seven SSRs could amplify the other buckwheat species. These SSRs were then used to study the genetic variability among 110 genotypes of buckwheat based on the allele frequency. These SSRs amplified a total number of 136 alleles, with 30 alleles amplified by Fem-1322. The major allele frequency ranged from 0.11 to 0.42 and the expected heterozygosity ranged from 0.04 to 0.46. Fem-1322 represents highest Polymorphic Information Content (PIC) of 0.93; followed by Fem-1303 and Fem-1407 with PIC 0.89. The gene diversity ranged from 0.72 to 0.93. Further Darwin based cluster analysis revealed the formation of two major groups by the 110 genotypes. Furthermore, model based STRUCTURE analysis generated two subpopulations which correspond to distance based groups. High level of genetic diversity was observed within the population. The results obtained provided insights about transferability of SSR markers among different species and these findings have further implications in buckwheat breeding as well as conservation strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Buckwheat (2x = 16), a pseudocereal is a dicotyledonous and broadleaved crop belonging Polygonaceae family and genus Fagopyrum. Buckwheat is called as ‘beech wheat’ due to its triangular seed shape. The genus Fagopyrum currently consists of 20 species including Fagopyrum esculentum (Common buckwheat), Fagopyrum tartaricum (Tartary buckwheat) and Fagopyrum dibotrys (perennial buckwheat) as three important cultivated buckwheat species [13]. Since decades, these species are well explored as these are in use both as edible form and for medicinal purpose [11]. It is rich in carbohydrates, fibers, lipids, vitamins, minerals and antioxidants [2, 22, 26]. Unlike other cereals, buckwheat is rich source of amino acids such as methionine, tryptophan, lysine and histidine [23]. Besides, its protein comprises of 18.2% albumin, 0.8% prolamin, 43.3% globulin and 22.7% glutelin [9]. The flour of buckwheat is used in a similar way as that of other cereals. It is deficient of gluten and thus is beneficial for the patients suffering from celiac disease [27]. Buckwheat is an indispensable crop for various medicinal purposes as it aids in tumor inhibition, lowering of cholesterol level, inflammation control, regulation of hypertension and diabetes, protection against childhood asthma and prevention against breast as well as colon cancer [23]. Moreover, common buckwheat sprouts are also known to contain several flavonoids, including vitexin, isovitexin, orientin, isoorientin, rutin and quercetrin [2]. Among all these flavanoids, rutin is most important flavanoid found is three important buckwheat species with the highest concentration (3–6% of dry weight) in Fagopyrum tartaricum [2]. It increases strength of blood vessels and also protects the vital organs such as heart and brain [2]. Tremendous genetic diversity based on morphological parameters have been observed in buckwheat species of Himalayan region, due to different pattern of rainfall, altitude and different soil pattern [24]. There is a general consensus among scientists that buckwheat is native to temperate East Asia, particularly to the South-West China and eastern side of the Himalayas [2]. It has also been reported that the buckwheat species originated from Yunnan province and the mountain area between Yunnan and Sichuan provinces [19]. Buckwheat is primarily grown in northern hemisphere, chiefly in Eastern and central Europe, Russia, East Asia, Nepal, Kazakhstan, Korea, China and Japan [3]. Since ages, farmers are employing cultivation practices of a particular crop for its acclimatization to that environment [12]. For the conservation and improvement of buckwheat crop, information on genetic diversity and population structure among inbreeds is necessary for essential derivation identification and legal protection of germplasm and selection of elite lines [8, 12, 25]. Earlier, studies using conventional methods such as allozyme analysis, utilization of morpho-agronomic traits in order to establish genetic relationship among commercial cultivars, wild cultivars and landraces have been conducted within and among a particular [1]. From breeding point of view self incompatibility (SI) is an important trait in common buckwheat for its improvement. Self incompatibility (SI) of a plant is referred to as inability of that plant to set seed after self pollination. This process is important for out breeding mechanism (cross pollination) for normal fruit set. SI promotes gene flow and associated genetic diversity on which selection can act [18]. SI can also help in development of hybrid plants. Self incompatibility trait is governed by single gene S allele. Self incompatibility could be changed to self compatibility by mutations including loss of S function allele in pollen or style, modifier genes mutation outside the S locus or recombination between a stylar incompatibility gene and a pollen incompatibility gene in the S locus [17, 18]. Common buckwheat (Fagopyrum esculentum) is self incompatible crop whereas tartary buckwheat (Fagopyrum tartaricum) is self compatible crop [29]. As such there is need of identifying a marker linked with S allele that may help in breeding of buckwheat. Advancement in molecular biology like use of DNA based markers for assessing genetic diversity have proved to be more efficient as compared to formally used morphological and biochemical markers [1]. Among the PCR-based techniques, certain molecular markers like RAPD, AFLP, SSR markers are mostly used for assessing diversity in various crops [12]. AFLP and RAPD markers suffer from a certain drawbacks over SSR markers such as dominant mode of inheritance and inference of homology from band co-migration [7]. However, SSR markers are ideal for estimating genetic diversity due to its co-dominant nature, hyper variability, random distribution within genome and high polymorphic information content [7, 28]. Thus, keeping in view the importance of buckwheat and the need for its improvement, this study was undertaken to evaluate genetic diversity among Fagopyrum esculentum genotypes with the help of SSR markers and checking their transferability from Fagopyrum esculentum to Fagopyrum tartaricum.
Material and methods
Plant material, DNA isolation and quantification
The seed material in the present study comprised of 110 genotypes of two Fagopyrum species mostly collected from different regions of Jammu and Kashmir and some procured from NBPGR (Supplementary Table-1). Genomic DNA was isolated from the leaves of three weeks old seedlings by CTAB method [4].
SSRs selection for characterization of buckwheat
A set of 15 common buckwheat SSRs selected from available literature detailed in Supplementary Table 2 were used for amplification of different genotypes in the present study. These 15 SSRs were firstly amplified on Fagopyrum esculentum germplasm and then were used for studying transferability in Fagopyrum tartaricum.
PCR amplification and gel electrophoresis of SSRs in common buckwheat and Fagopyrum tartaricum
Each 5 μL reaction mixture contained 50 ng of template DNA, 10 µM of each forward and reverse primer, and 3 µl of kappa master mix (reaction buffer, dNTP, Mgcl2 and Taq polymerase). DNA amplification was carried out in a thermal cycler (Applied biosystems, thermoscientific) with the given profile as step I (initial denaturation at 94 °C for 4 min), 35 cycles of step II (denaturation at 94 °C for 30 s, annealing at 50–60 °C for 30 s and extension at 72 °C for 30 s), following step III final extension at 72 °C for 7 min (Supplementary Table 2). The amplified product were resolved on agarose gel (3%) and SSRs which does not resolved on 3% agarose properly were resolved on silver staining PAGE (polyacrylamide gel electrophoresis) gels (Fig. 1). Allelic format was used for scoring the bands within the range of expected base pair (bp).
SSR data analysis
Fifteen SSRs were employed for studying various discriminatory parameters among 52 Fagopyrum esculentum germplasm by using Power Marker [14] and then these SSRs were used to check the transferability into Fagopyrum tartaricum. Cluster analysis was performed based on dissimilarity matrix using a shared allele index with the help of DARwin software version 5.0 [20]. Additionally, principal coordinate analysis (PCoA) was calculated using DARwin software [20]. 14 SSRs were used to study diversity among 52 Fagopyrum esculentum genotypes whereas 7 SSRs which amplified in both Fagopyrum esculentum and Fagopyrum tartaricum were used for studying genetic diversity among 110 genotypes of buckwheat. Population structure of germplasm was accessed based on bayesian approach in STRUCTURE ver 2.3.4 software [21]. K value was identified by using burn-ins of 50,000 followed by 50,000 MCMC (Markov Chain Monte Carlo). The data was run ten times for each K value of 1 to 10 with the possibility of admixture. The most probable value of K corresponds to the peak in an ad hoc statistic D(K) which is associated with L(K), L’(K) and L”(K) [5]. The D(K) perceives the rate of change in log probability of the data with respect to the number of groups inferred by STRUCTURE. The probability value of genotypes higher than standard threshold value of > 70% were assigned to a particular population and genotypes with probability value < 70% were considered as admixtures. STRUCTURE software was also used for the construction of structure bar plot [21] and for estimating genetic differentiation and Fst values [30].
Results and discussion
Discriminatory parameters of SSR markers in Fagopyrum esculentum
Analyzing diversity in buckwheat through SSR markers have proved to be an efficient way. 15 SSRs were utilized to study the polymorphism among 52 genotypes of Fagopyrum esculentum. Out of 15 SSRs, 14 were found polymorphic in Fagopyrum esculentum genotypes. The total number of alleles identified was 143 in which most of the alleles were polymorphic with the average number of 9 alleles per primer (Supplementary Table-3). The maximum number of alleles was identified in the primer GB-FE-035 with 36 alleles. The Polymorphic information content (PIC) of each primer in common buckwheat ranged from 0.29 (primer GB-FE-054) to 0.92 (primer GB-FE-035) with an average of 0.56. Similarly, gene diversity also ranged from 0.31 (primer GB-FE-054) to 0.92 (primer GB-FE-035) with an average of 0.58. The primer GB-FE-035 has been considered as the most efficient primer because of its highest PIC value (0.91, 0.93) as reported by earlier studies conducted by Ma et al. [15] and Song et al. [20] respectively. The heterozygosity of primers ranged from 0.17(GB-FE-054) to 0.61 (Fem 1303) with an average of 0.15. Major allele frequency (MAF) ranges from 0.17(GB-FE-1322) to 0.96 (GB-FE-014) with an average frequency of 0.53. The difference in the results from earlier studies might be due to the different germplasm used in this study.
Genetic diversity study in 52 accessions of buckwheat
Clustering using neighbor joining method (NJ) exhibited that 52 buckwheat genotypes are grouped into three major clusters. Among three major clusters (Clusters I, II and III) were represented by 52 genotypes (Fig. 2). Cluster I consists of 20 genotypes; Cluster II comprising of 22 genotypes whereas Cluster III consists of 10 genotypes procured from NBPGR. Based on dissimilarity matrix, the genotypes (BWZ-6, BWZ-29) were identified to be more dissimilar with the value 0.9 while as the genotypes (BWZ-28, BWZ-29) and (BWZ-73, BWZ- 74) were identified to be similar with the value 0.21 and 0.25 respectively. The highly diverse genotypes found in this study could be utilized efficiently in future buckwheat breeding programmes.
Transferability of Fagopyrum esculentum markers into Fagopyrum tartaricum
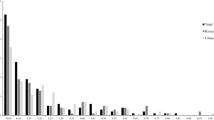
The primers of Fagopyrum esculentum were also employed in Fagopyrum tartaricum in order to check its transferability in other species including Fagopyrum tartaricum. The data in Fagopyrum tartaricum was analyzed in the similar way as in the common buckwheat. Out of 15 primers, seven primers exhibited good amplification in Fagopyrum tartaricum (Table 1). A total number of 136 alleles were obtained in which most the alleles were polymorphic with the average number of 19 alleles per primer (Table 1). Maximum number of alleles was identified in the primer Fem 1322 possessing 30 alleles while minimum number of alleles was identified in primer GB-FE-080 having nine alleles only. Similarly, in an earlier investigation, the primer Fem 1322 exhibited highest PIC with 68 alleles [10]. It can be concluded that high polymorphism exhibition of this primer is might be due to InDel (insertion-deletion) mutations at the flanking regions of SSR having longer repeat unit. Polymorphic information content (PIC) of each primer ranged from 0.69 (primer GB-FE- 080) to 0.93 (Fem 1322) with an average of 0.86. Heterozygosity of each primer ranged from 0.04 (GB-FE-080) to 0.46 (Fem 1303) with an average of 0.25. Gene diversity ranged from 0.72 (GB-FE-080) to 0.93(Fem-1322) with an average of 0.87. Further, Major allele frequency ranged from 0.11 (Fem 1322) to 0.42 (GB-FE-080) with an average major allelic frequency of 0.22. A significant correlation was observed between polymorphic information content (PIC), allele number, gene diversity. Taking into account various parameters mentioned above, Fem 1322 was considered to be the best primer for genetic diversity analysis among selected varieties and primer GB-FE-080 was found to be least efficient. Moreover, PIC (0.93), allele number (30), and gene diversity (0.93) and a significant correlation of these values, primer Fem 1322 depicts its potential for evaluation of diversity. The different parameter of buckwheat SSR which amplified in tartaricum species is given in Table 1.
Genetic diversity and population structure analysis of 110 buckwheat genotypes
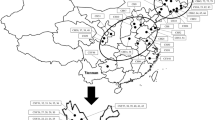
In the current study, the genetic diversity and population structure of common and Fagopyrum tartaricum accessions collected from diverse regions of Jammu & Kashmir was studied based on SSRs in order to provide useful information for conservation and utilization of buckwheat genetic resources in Jammu and Kashmir. Neighbor joining (NJ) method divided 110 buckwheat genotypes into two major clusters (Fig. 2). Cluster I contains 54 genotypes from different districts of Kashmir and from NBPGR. However, cluster II consists of 55 genotypes and there is formation of one cluster which consists of only one genotype BWZ-83 (Fig. 3). Further, principle component analysis (PCoA) revealed similar result as observed in cluster analysis (Fig. 4). The diversity of Fagopyrum esculentum and Fagopyrum tartaricum is attributed to the cross incompatibility and low gene flow between the two species which also suggests that there is conservation of the two species [6, 16]. Further, based on dissimilarity matrix, the genotype BWZ83 was identified to be most diverse genotype with the value 1.00 among 110 genotypes while as genotypes (BWZ77, BWZ 90) were identified to be the similar genotypes with the value 0.14. Thus, the genotype BWZ 83 can be utilized in future breeding programmes for buckwheat improvement.
The model based structure analysis revealed a clear D (K) at K = 2 in the plots of L (K) versus D (K) with maximum mean probability of likelihood value of L (K) = − 2671.49 confirms the assignment of germplasm to two subpopulations (Fig. 5a). Out of 110 genotypes, 57 (51.8%) genotypes are present in population I, 52 genotypes (47.2%) are present in population II and one admixture genotype BWZ63 (Fig. 5b and supplementary table 4). The two sub-populations had moderate differentiation, with Fst value of 0.1500 for population I and 0.0771 for population II and the expected heterozygosity at given locus is 0.6988 and 0.7392 respectively for population I and II (Table 2). The genetic differentiation based on Fst values between two buckwheat sub-populations identified by population structure analysis is 0.1987. Thus, it is clear from the dendrogram analysis that the two species of buckwheat are diverse and the seven markers employed in the study are sufficient to distinguish the two species of buckwheat. Similarly, in an investigation of Facho et al. [6] two buckwheat species (Fagopyrum esculentum and Fagopyrum tartaricum) were clearly divided into two populations based on microsatellite data analysis.
a Graphical representation of the optimal number of groups in the program STRUCTURE inferred using the criterion of Evano et al. [5]. The analysis was based on data obtained from 7 microsatellite loci in buckwheat collection evaluated for genetic diversity. b STRUCTURE plot of membership coefficients for all the accessions of buckwheat in the study sample sorted in the same order and classified according to successive selected preset K values ranging from 1 to 10. For K = 2 the groups are identified
Conclusion
In order to explore the diversity of large number of genotypes, the present study has assisted in identification of highly polymorphic SSRs such as GB-FE-035 in case of Fagopyrum esculentum and Fem 1322 in case of Fagopyrum tartaricum. These polymorphic markers will provide important genetic resources for studies of buckwheat species. All the statistical analysis including cluster analysis, structure analysis and principal component analysis (PCoA) exhibited similar results which provided clear insight about the divergence of two indispensable species of buckwheat i.e., Fagopyrum esculentum and Fagopyrum tartaricum. This study also revealed that the SSRs used for Fagopyrum esculentum could be used for assessing genetic diversity relatedness in Fagopyrum tartaricum. The diversity between the two species of buckwheat can be harnessed for conservation and utilization of germplasm for breeding programmes. The study is helpful to improve buckwheat crop genetically and to develop resistant varieties.
References
Blair MW, Hedetale V, McCouch SR. Fluorescent labeled microsatellite panels useful for detecting allelic diversity in cultivated rice (Oryza sativa L.). Theor Appl Genet. 2009;105:449–57.
Campbell GC. Buckwheat Fagopyrum esculentum Moench. Promoting the conservation and use of underutilized and neglected crops. Institute of Plant Genetics and Crop Plant Research, Gatersleben, Germany and the IPGRI, Rome, Italy; 1997.
Chrungoo NK, Dohtdong L, Chettry U. Genome plasticity in buckwheat. In V. R. Rajpal et al. (eds.). Gene pool diversity and crop improvement. . Sustainable development and biodiversity. Berlin: Springer; 2016, vol. 10, pp. 227–39.
Doyle JJ, Doyle JL. Isolation of plant DNA from fresh tissue. Focus. 1990;12(13):39–41.
Evano G, Regnaut S, Goudet J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol. 2005;14:2611–20.
Facho ZH, Farhatullah TW, Ali S. Species divergence and diversity in buckwheat landraces collected from the western Himalayan region of Pakistan. Pak J Bot. 2019;51(6):2215–24.
Goldstein DB, Schlötterer C. Microsatellites. Evolution and application. New York: Oxford University Press; 1999, pp. 449–57.
Hallauer AR, Russell WA, Lamkey KR. Corn breeding. Agronomy Publications; 1988, p. 259.
Ikeda K, Asami Y. Mechanical characteristics of buckwheat noodles. Fagopyrum. 2000;17:67–72.
Iwata H, Imon K, Tsumur Y, Ohsaw R. Genetic diversity among Japanese indigenous common buckwheat (Fagopyrum esculentum) cultivars as determined from amplified fragment length polymorphism and simple sequence repeat markers and quantitative agronomic traits. Genome. 2005;48(3):367–77.
Jing R, Li HQ, Hu CL, Jiang YP, Qin LP, Zheng CJ. Phytochemical and pharmacological profiles of three Fagopyrum buckwheats. Int J Mol Sci. 2016;17(4):589.
Kump B, Javornik B. Evaluation of genetic variability among common buckwheat (Fagopyrum esculentum Moench) populations by RAPD markers. Plant Sci. 1996;114(2):149–58.
Li AR. “Fagopyrum” flora of China. Beijing: Science Press and the Missouri Botanical Garden Press; 2003.
Liu K, Muse SV. Power marker: an integrated analysis environment for genetic marker analysis. Bioinformatics. 2005;21(9):2128–9.
Ma KH, Kim NS, Lee GA, Lee SY, Lee JK, Yi JY, Park YJ, Kim TS, Gwag JG, Kwon SJ. Development of SSR markers for studies of diversity in the genus Fagopyrum. Theor Appl Genet. 2009;119:1247–54.
Marshall HG. Isolation of self-fertile, homomorphic forms in buckwheat, Fagopyrum sagittatum Gilib. Crop Sci. 1969;9:651–3.
Mizuno N, Yasui Y. Gene flow signature in the S-allele region of cultivated buckwheat. BMC Plant Biol. 2019;19:125. https://doi.org/10.1186/s12870-019-1730-1.
de Nettancourt D. Incompatibility and incongruity in wild and cultivated plants. Berlin: Springer; 2001.
Ohnishi O. Search for the wild ancestor of buckwheat III. The wild ancestor of cultivated common buckwheat and of tatary buckwheat. Econ Bot. 1998;52(2):123.
Perrier X, Jacquemoud-Collet JP. DARwin software. https://darwin.cirad.fr/darwin. 2006.
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155:945–59.
Przybylski R, Gruczyńska E. A review of nutritional and nutraceutical components of buckwheat. Eur J Plant Sci Biotechnol. 2009;3(1):10–22.
Song JY, Lee GA, Yoon MS, Ma KH, Choi YM, Lee JR., Lee MC. Analysis of genetic diversity and population structure of buckwheat (Fagopyrum esculentum Moench.) landraces of Korea using SSR markers. In: Buckwheat germplasm in the world. New York: Academic Press; 2011, pp. 315–31.
Sood M, Chuahan S. Biodiversity conservation in Himalayan region. Int J Appl Res. 2015;2(2):610–5.
Smith JSC, Ertl DS, Orman BA. Identification of maize varieties. In: Wrigley CW, editor. Identification of food grain varieties. St. Paul: American Association for Cereal Chemists; 1995. p. 253–264.
Sytar O, Brestic M, Zivcak M, Phan Tran LS. The contribution of buckwheat genetic resources to health and dietary diversity. Curr Genomics. 2016;17(3):193–206.
Tang Y, Ding MQ, Tang YX, Wu YM, Shao JR, Zhou ML. Germplasm resources of buckwheat in China. In: Zhou M, Kreft I, Woo SH, Chrungoo NK, Wieslander G, editors. Molecular breeding and nutritional aspects of buckwheat. The Netherlands: Academic Press; 2016. p. 13–20.
Tautz D. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res. 1989;17(16):6463–71.
Woo SH, Kamal AHM, Tatsuro S, Campbell CG, Adachi T, Yun SH, Chung KY, Choi JS. Buckwheat (Fagopyrum esculentum Moench.): concepts, prospects and potential. Eur J Plant Sci Biotech. 2010;4:1–16.
Wright S. The genetical structure of populations. Ann Eugenics. 1951;15:323–54.
Acknowledgements
SMZ is grateful to NMHS GBPNIHESD, Almora, Uttrakhand, India for financial support of this work (Project Sanction Order No. GBPNI/NMHS17-18/SG24/622).
Author information
Authors and Affiliations
Contributions
EB have contributed in conducting the experiment and preparing first draft of manuscript. RM did the analysis and helped in preparing the manuscript. RAM helped in preparing the manuscript. WAD has helped in collection and field work. SMZ has designed the experiment, helped in analysis, prepared the final manuscript and got grant for undertaking this research work.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bashir, E., Mahajan, R., Mir, R.A. et al. Unravelling the genetic variability and population structure of buckwheat (Fagopyrum spp.): a collection of north western Himalayas. Nucleus 64, 93–101 (2021). https://doi.org/10.1007/s13237-020-00319-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13237-020-00319-y