Abstract
Cancerous cells display metabolic engineering through enhanced utilization of nutrients to support their increased requirements for proliferation, bioenergetics, biosynthesis, redox homeostasis, and cell signaling. To investigate the extent to which malignant cells rely on glycolysis and glutaminolysis, the effects of differential deprivation of nutrients such as d-glucose, l-glutamine, and pyruvate on proliferation, morphology, cell cycle, oxidative stress, mitochondrial function, autophagic vacuole formation, and migration in MDA-MB-231, HepG2, and HeLa cells were investigated in this study. Cell viability assay, cell morphology, and ATP assay showed higher dependence of MDA-MB-231 and HepG2 cells on glucose and glutamine, respectively, for cell survival, growth, ATP production, and proliferation, while HeLa cells were equally dependent on both. However, the combination of all three nutrients displayed maximum proliferation. Differential deprivation of glucose in the absence of glutamine resulted in G0/G1 plus G2/M arrest in MDA-MB-231, whereas G0/G1 arrest in HepG2 and S-phase arrest in HeLa cells occurred at 48 h. Although the differential withdrawal of nutrients revealed a varying degree of effect dependent on cell type, nutrient type, nutrient concentrations, and deprivation time, a general trend of increased oxidative stress, loss of mitochondrial membrane potential, and ATP and antioxidant (GSH) depletion led to mitochondrial dysfunction in all three cell lines and inhibition of cell migration in MDA-MB-231 and HeLa cells at 48 h. Extreme deprivation of nutrients formed autophagic vacuoles. Importantly, normal cells (HEK293) remained unaffected under most of the nutrient-deprived conditions examined. This study enhances our understanding of the impact of differential nutrient deprivation on critical characteristics of cancer cells, contributing to the development of metabolism-based effective anticancer strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite continuous evolution in cancer treatment methods, there is a rapid growth in cancer incidence and mortality worldwide. It is still the second leading cause of mortality worldwide, accounting for 9.96 million fatalities in 2020 (WHO fact sheets). Female breast cancer, liver cancer and cervical cancer rank first, seventh and ninth among 36 most frequently diagnosed cancer with an incidence rate of 11.7%, 4.7% and 3.1%, respectively and are the fourth, second and ninth leading cause of cancer death with a mortality rate of 6.9%, 8.3% and 3.4%, respectively (Sung et al. 2021).
Cancer cells exhibit metabolic reprogramming to maintain increased bioenergetic and biosynthetic demands, redox homeostasis and cell signaling to sustain uncontrolled proliferation and cell survival (Cluntun et al. 2017). These altered metabolic requirements of tumors are met by enhanced uptake and catabolism of two major substrates, D-glucose (G) and L-glutamine (Q), though it is also met by sodium pyruvate (Py) to some extent. The significance of cellular metabolism in tumorigenesis was first observed by Otto Warburg (Warburg effect). Although the Warburg effect is a far less efficient process of ATP production than oxidative phosphorylation (OxPhos), many human cancers show the Warburg effect (Cantor and Sabatini 2012). In the Warburg effect, even in high oxygen concentration, glycolysis occurring at a high rate preferentially catabolize G to lactate via pyruvate (aerobic glycolysis). Lactate then undergoes lactic acid fermentation in the cytosol. In contrast, mitochondrial respiration driven glucose catabolism occurs at a low rate (Akins et al. 2018). Thus, the Warburg effect permits the shunting of different intermediates of glycolysis into branching anabolic pathways supporting additional metabolic requirements of a proliferating cell. Q act as a reduced nitrogen source for the biosynthesis of nucleotides, a variety of nonessential amino acids, proteins, hexosamine and activation of cell signalling (Son et al. 2013; Hosios et al. 2016; Altman et al. 2016). Additionally, Q, as an important carbon source, replenishes TCA cycle intermediates, produces glutathione and synthesize lipids via reductive carboxylation and ATP under various conditions in cancer cells (Daye and Wellen 2012; Michalak et al. 2015; Yang et al. 2017). Replenishment of TCA cycle intermediates allows them to get diverted to various anabolic pathways during proliferation (Cantor and Sabatini 2012). The dependence of cancer cells on Q for survival is referred to as “glutamine addiction”. However, some cancer cell lines do not exhibit Q addiction (Cetinbas et al. 2016). The reasons behind this disparity are not well understood.
Although altered tumor metabolism has long been utilized as the basis for the development of new cancer therapeutics and strategies, most of the studies have been centered around changes in either G or Q metabolism alone. However, the complex cellular metabolism minimizes the possibility of the need for a single metabolite for multiple purposes in a tumor cell. Thus, a more comprehensive understanding of the major metabolic requirements of a cancer cell is vital. Differences in addiction to G and Q or tolerance to hypoglycemia, aglycemia, and Q starvation among various cancer types may be imperative while optimizing more specific strategies for treating a particular cancer type. Therefore, this study explores the in vitro effect and extent of interdependence of three major nutrient substrates—G, Q, and Py—on proliferation and survival across cancer cell lines from three different sites—breast (MDA-MB-231), liver (HepG2), and Cervix uteri (HeLa)—that will add to our current understanding of the development of effective cancer therapy approaches. Here, we have analyzed variation in cellular events and characteristics such as cell cycle, intracellular reactive oxygen species (ROS), mitochondrial superoxide (mtROS), mitochondrial membrane potential (MMP), intracellular ATP, antioxidant (GSH) and autophagic vacuole formation, which play critical roles in determining cells’ proliferation, migration potential, and survival.
Materials and methods
Materials
High glucose DMEM, DMEM without d-glucose, l-glutamine and sodium pyruvate, dimethyl sulfoxide (DMSO), 2′,7′-Dichlorodihydrofluorescin diacetate (H2DCFDA), propidium iodide (PI), ATP bioluminescent assay kit, glutathione (GSH), and acridine orange (AO) were purchased from Sigma-Aldrich, USA. Fetal bovine serum (FBS), trypsin, rhodamine 123 (R123) and MitoSOX red were procured from ThermoFisher Scientific. 3-(4,5-dimethylthiazol-2-yl)-2,5diphenyltetrazolium bromide (MTT) and all other chemicals used were of molecular grade.
Cell culture and treatments
Cancerous cell lines MDA-MB-231, HepG2, HeLa and normal cell line HEK293 were procured from a recognized national cell repository, National Centre for Cell Science, Department of Biotechnology, Pune, India. Cancer and normal cells were maintained in high glucose (HG, 4.5 g/L) and low glucose (1 g/L) DMEM, respectively, containing 10% FBS, penicillin (100U/mL)–streptomycin (100 μg/mL) and amphotericin B (100 μg/mL) at 37 °C in 5% CO2 atmosphere in a humidified incubator. At 70–80% confluency cells were subcultured using trypsin–EDTA. Subsequent to the first cycle of growth in HG DMEM containing 10% FBS, MDA-MB-231, HepG2, HeLa and HEK293 cell lines were seeded in experimental plates according to the required cell density in fresh HG-DMEM containing 5%, 2.5%, 2.5% and 2.5% FBS, respectively and maintained overnight or 24 h to adapt them to the changed serum concentrations. The end of this overnight cell growth period post-seeding denoted the start of the treatment period and marked as zero hours (0 h). Supplementation of HG-DMEM with 5%, 2.5%, and 2.5% FBS for MDA-MB-231, HepG2, and HeLa cell lines, respectively, was determined based on the cell viability assay, that showed a non-significant variation in cell proliferation of these cell lines when grown in HG-DMEM with altered serum concentrations for up to 72 h as compared to HG-DMEM with 10% FBS (Fig. S1). A comparative study of cell viability of these cells grown up to 72 h in low serum media and those cultured in media containing 10% dialyzed serum was also performed, that resulted in a non-significant change in cell viability (data not shown).
Treatments included replacement of HG-DMEM used for growing cells till 0 h with DMEM containing varying concentrations of the three energy substrates, G (25.5, 5.5, 2.8 and 0 mM), Q (4 mM) and Py (1 mM) either alone or in combinations and supplemented with 5%, 2.5%, 2.5% and 2. 5% FBS for MDA-MB-231, HepG2, HeLa, and HEK293 cell lines, respectively for 24, 48 and 72 h. Serum concentration was lowered to minimize its effect as a source of nutrient and to conduct a more targeted study of variation of major energy substrates on these cell lines. The nine varied metabolic conditions used as treatments in this study consisted of the DMEM containing 5% FBS (for MDA-MB-231) and 2.5% FBS (for HeLa, HepG2 and HEK293) and the following concentration of G, Q and Py:
-
(a)
Control: G 5.5 mM + Q 4 mM + Py 1 mM [PG]
-
(b)
Condition 1: G 25.5 mM + Q 4 mM + Py 1 mM [HG]
-
(c)
Condition 2: G 2.8 mM + Q 4 mM + Py 1 mM [Hypo G]
-
(d)
Condition 3: G 0 mM + Q 4 mM + Py 1 mM [GS]
-
(e)
Condition 4: only G 25.5 mM [HG’]
-
(f)
Condition 5: only G 5.5 mM [PG’]
-
(g)
Condition 6: only G 2.8 mM [Hypo G’]
-
(h)
Condition 7: without G, Q and Py [GS’]
-
(i)
Condition 8: only Py 1 mM
-
(j)
Condition 9: only Q 4 mM
DMEM containing 25.5 mM, 5.5 mM, 2.8 mM, and 0 mM glucose along with standard concentrations of Q and Py (4 mM and 1 mM, respectively) have been designated as high glucose (HG), normal or physiological glucose (PG), hypoglycemic glucose (Hypo G) (Heller 2017), and glucose starved (GS) media, respectively. DMEM lacking standard concentrations of Q and Py and containing only G as nutrient substrate in decreasing concentrations as mentioned above have been designated as HG’, PG’ and Hypo G’ media, respectively, whereas DMEM, also starved of G along with Q and Py, was termed as GS’ media.
Working solutions of dyes and fluorescent probes were prepared in the ten respective media (one used as control and nine used as treatments) containing 1% FBS to avoid post-treatment revival of cells during incubation with dyes and probes and interference with the activity of probes when a higher concentration of serum is used.
Cell viability/proliferation assay
Effects of different metabolic states on the proliferation of MDA-MB-231, HepG2, HeLa and HEK293 cell lines were studied by MTT assay as previously described (Khatua et al. 2022). Briefly, 3 × 103 MDA-MB-231 cells and 4 × 103 cells of HepG2, HeLa and HEK293 were seeded in 96-well plates and treated as explained in “Cell culture and treatments”. Media were removed 3 h before the completion of time points, and 4 µL MTT (5 mg/mL)/well/200 µL low-serum media were added and incubated for 3 h. Then formazan crystals were solubilised by adding 100 μl DMSO/well, and their absorbance was recorded at 570 nm in a multi-well plate reader (spectraMax M5-Molecular Devices, USA). Percentage change in cell viability was calculated relative to PG.
Morphological analysis
For the morphological study, approximately 0.6 × 105 MDA-MB-231 cells and 0.8 × 105 HepG2, HeLa and HEK293 cells were seeded in a 6-well plate and grown overnight. After that, they were treated with media comprising different metabolic states and allowed to grow for 24, 48 and 72 h. Morphological changes were captured using an inverted phase-contrast microscope (EA-Prime Inverted, Lmi microscopes, UK) at 20X magnification. For each treatment group, images of size 2.27 mm × 1.74 mm were captured, and all the cells with different morphologies on those images were quantified using the image analysis software- NIH Image J 1.52a; Java 1.8.0_112 (64-bit) and shown as bar graphs. Each cell of the images captured for each treatment and control group was classified in one of the three broad categories of cell morphologies, named ‘normal’, ‘spiked’ and ‘rounded’. ‘Normal’ represented standard and characteristic morphology, ‘spiked’ denoted echnoid spikes and membrane blebbing which represented morphology of the stressed/ less viable cells, and ‘rounded’ depicted cell shrinkage, cells with condensed nuclei, and apoptotic bodies which represented dead or very less viable cells or debris. For each cancer cell line and at a given time point, the percentage change in the number of cells in each of the three different morphological categories of a treatment group was shown relative to the percentage of cells in that particular morphological category in the control group (PG) of that time point of that particular cell line.
Cell cycle analysis
Cell cycle analysis was conducted as per (Roy et al. 2023a) with some modifications by flow cytometry utilizing propidium iodide (PI). PI is a red fluorescent DNA binding stain that helps determine phases of the cell cycle based on DNA quantification. Briefly, 1.5 × 105 MDA-MB-231cells and 2 × 105 HepG2 and HeLa cells, after overnight culture, were treated as mentioned in “Cell culture and treatments” for 24 and 48 h. Then cells were fixed with chilled ethanol (70% v/v) and stained with staining buffer containing PI and RNase (both at 50 μg/mL). Then 10,000 events per sample was acquired in a flow cytometer (BD FACS Verse), and percentage distribution of cells in G0/G1, S and G2/M phases was analyzed by ModFit LT 5.0 software (Verity Software House Inc., USA). During analysis, aggregates (doublets) removal was carried out by applying the appropriate gate on PI-area versus PI-width or PI-area versus PI-height dot plots.
Determination of intracellular and mitochondrial ROS generation
Effects of altered metabolic environments on total cytosolic ROS generation in cancer cells were analyzed by a flow cytometric assay performed as described by (Prasad et al. 2022). Briefly, 1.5 × 105 MDA-MB-231 cells and 2 × 105 HepG2 and HeLa cells were cultured in 6-well plates and treated for 24, 48 and 72 h, as described in “Cell culture and treatments”. On completion of time points, cells were loaded with 10 μM of H2DCFDA whose conversion by intracellular ROS to highly green fluorescent and oxidized form- 2ʹ,7ʹ-dichlorofluorescein (DCF) act as a ROS indicator. Mean DCF fluorescence from 10,000 cells of each sample was captured in FITC channel in BD FACS Verse flow cytometer using BD FACSuite software.
Similarly, effect of treatments on mitochondrial ROS generation in these cells was determined using mitochondrial superoxide indicator dye MitoSOX Red. 10 × 103 cells were seeded, grown for 24 h and then treated for 48 h in a 96-well plate. After that cells were washed and incubated with 2.5 μM of dye for 30 min in dark. Cells were again washed, lysed with cell lysis buffer and fluorescence was measured in a fluorimeter (SpectraMax iD5-Molecular Devices, CA, USA) at excitation/emission 396/610 nm. Fluorescence in the sample was normalized to total amount of protein and represented as fluorescence/μg protein. Data were represented as a fold change in fluorescence intensities of ROS indicators in the treated sample relative to control.
Flow cytometric determination of variation in MMP (Δѱ m)
Alterations in MMP due to the treatment of cancer cells under this study, with media differentially deprived of nutrients, were measured using R123 (10 μg/mL) following a method as explained earlier (Khatua et al. 2020). Loss of Δѱm or mitochondrial depolarization is correlated with leakage, thus loss of green fluorescence of the cationic fluorochrome R123, which is otherwise retained inside mitochondria. Cells were cultured, treated and R123 fluorescence from cells were captured and analyzed, similar to the previous assay for detecting intracellular ROS. Data were shown as the ratio of mean R123 fluorescence in a treated sample to control.
ATP assay
ATP content in MDA-MB-231, HepG2 and HeLa cells after 48 h exposure to modified metabolic conditions was assayed using ATP Bioluminescent Assay kit according to the manufacturer’s protocol and represented as the ratio of bioluminescence in the treated sample to control.
Measurement of reduced glutathione (GSH) level
GSH content was measured according to a standard protocol (Roy et al. 2023c), with minor modifications. Briefly, control and treated cells in a 6-well plate were harvested by trypsinization, followed by washing and cell lysate preparation using 100 µl RIPA lysis buffer, pH 8.0 (50 mM Tris, 150 mM NaCl, 0.05% deoxycholate, 1% NP-40, 0.1% SDS, 1 mM EDTA) and a brief ultrasonication. Thereafter, samples were centrifuged (10 K rpm, 10 min, 4 °C) and 25 µl of samples (supernatant) were added to 475 µl of the reaction mixture containing 100 mM NaH2PO4 (pH 8.0), EGTA (5 mM) and 25 µl of fluorescent substrate O-phthalaldehyde (1 mg/mL). After 15 min incubation, samples were analyzed with a spectrofluorophotometer (Shimadzu RF5301PC) at excitation 350 nm and emission 420 nm. Further, GSH content in samples was quantified from fluorescence intensity readings using a GSH standard curve and normalized with their per µg of protein (Fig. S2). Data were represented as a fold change in GSH content in treated groups compared to control. Then fold increase and fold decrease were calculated as fold change (treatment) ÷ fold change (control) and fold change (control) ÷ fold change (treatment) (Roy et al. 2023b).
Detection of acidic vesicular organelles (AVOs)
Fluorescence microscopy was employed to detect AVOs as the marker of autophagy using fluorescent dye acridine orange (AO) (Izdebska et al. 2019). Experimentally, cells were cultured on coverslips in 6-well plates (2 × 105 cells/well), and after 24 h, treated for 48 h. Then cells were washed with PBS and incubated with staining solution (1 µg/mL AO mixed in SFM) for 5 min in the dark. Then cells were washed two times with PBS and images were captured using a Zeiss fluorescence microscope equipped with an AxioCamMR3 camera and AxioVision software at 10X magnification in the green and red channels.
In vitro cell migration/wound healing assay
A cell migration assay was carried out following a method described by Liang et al. (2007) with some changes. In brief, cell monolayers with 80–90% confluency were formed in each well of six-well plates by growing cells seeded at 3 × 105 cells/ well. Subsequently, a 200 μl pipette tip was used to create a wound in each well and treated as described in “Culture and Treatments”. An inverted microscope for phase-contrast imaging (EA-Prime Inverted, Lmi microscopes, UK) was used to capture photographs of the wound at 0, 24 and 48 h. Differences in the rate of wound closure or percentage decrease in wound area (thickness) owing to treatment dependent variation in two-dimensional migration from the edges of the created wound were quantified using the image analysis software- NIH Image J 1.52a; Java 1.8.0_112 (64-bit).
Biplot analysis and classical hierarchical cluster analysis (HCA)
Biplot analysis and classical HCA were performed using past4.06b software on the results of 48 h (mean values) for each cancer cell line. Correlation matrix was applied for biplot analysis, whereas unweighted pair group method with arithmetic mean (UPGMA) algorithm and Euclidean similarity index were applied for classical HCA. Appropriate bootstrapping was also performed for both the analyses.
Statistical analysis
Each experiment was carried out three times (mostly in triplicates) and independently. Changes in treated conditions were represented as mean ± standard deviation (SD) relative to control (PG) of respective time points of the individual cell lines. To analyze the significance of these changes, one-way ANOVA combined with Tukey’s post hoc test for comparison among multiple groups was applied using the software GraphPad Prism 5 (San Diego CA, USA). Changes were considered significant at p < 0.05 versus control (PG), or between the analyzed groups.
Results
Differential depletion of glucose, glutamine and pyruvate differentially inhibits cancer cell proliferation
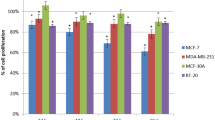
The three major nutrient sources used as a substrate by cancer cells for biosynthesis, survival, and proliferation are G, Q and Py (Elf and Chen 2014). The effect of these nutrient substrates on cell proliferation in MDA-MB-231, HepG2 and HeLa cells was investigated by MTT assay. Change in cell viability was dependent on time, substrate-type and substrate concentration. At 24 h, HG and Hypo G conditions did not show any significant change in the viability of all three cell lines relative to PG (Fig. 1A, B and C). GS and Q 4 mM media in MDA-MB-231 cells (Fig. 1A) and HG’ media in HepG2 cells (Fig. 1B) showed a significant reduction in cell viability relative to control by 18 ± 1.2%, 20 ± 1.6% and 36 ± 1.7%, respectively, whereas the other two cell lines remain unaffected in these conditions (Fig. 1A, B and C). A significant decrease (p < 0.001 or p < 0.01) in cell viability was observed when the cells were exposed to PG’, Hypo G’, GS’ and Py 1 mM media in all three cell lines except in MDA-MB-231 cells exposed to PG’, which resulted in less significant (p < 0.05) decrease in cell viability by 10 ± 0.6% (Fig. 1A). In these four conditions, HepG2 and HeLa cells showed the maximum and the minimum effects, respectively, with a reduction in viability by 41 ± 1.1% to 43 ± 1.3% in HepG2 (Fig. 1B) and 24 ± 2.1% to 24 ± 2.3% in HeLa cells (Fig. 1C) against 16 ± 1.3% to 41 ± 3.6% in MDA-MB-231 cells (Fig. 1A). Contrary to MDA-MB-231 cells, Q 4 mM media could maintain cell viability in HepG2 and HeLa cells for up to 24 h. Similar trends of cell viability continued at 48 and 72 h. However, for most of the metabolic conditions more pronounced decrease in viability was observed at 72 h and 48 h as compared to 24 h. GS condition decreased viability of HepG2 and HeLa cells less significantly (p < 0.05) at 48 h but failed to maintain the viability of MDA-MB-231 cells at 48 h and all three cell lines at 72 h. In GS medium, viability of MDA-MB-231, HepG2 and HeLa cells at 48 and 72 h decreased by 57 ± 4.3% and 72 ± 4.6% (Fig. 1A), 14 ± 0.9% and 27 ± 0.8% (Fig. 1B) and 13 ± 0.3% and 20 ± 0.8% (Fig. 1C) respectively, as compared to respective time point controls. At 48 h, viability of MDA-MB-231 and HeLa cells was less significantly compromised (p < 0.05, viability decreased by 10 ± 0.7% and 15 ± 0.4%, respectively) in HG’condition compared to HepG2 cells (p < 0.001, viability decreased by 60 ± 3%). A less significant (p < 0.05) decline in the viability of HepG2 and HeLa cells treated with Q 4 mM media was observed at 48 h, whereas the viability of MDA-MB-231 continues to decrease with high significance (p < 0.001). Complete deprivation of all three nutrient substrate (GS’) led to the maximum cell death in all three cell lines with a continuous increase from 24 to 72 h. For each time point of all three cell lines, the highest and the lowest cell viability was observed in HG and GS conditions, respectively. Thus, although G, Q and Py, all three affect the proliferation of these three cell lines, viability and survival of MDA-MB-231 cells are more dependent on G than Q. In contrast, the viability of HepG2 is more dependent on Q than G, and that of the HeLa cells is equally dependent on Q and G. The effect of conditions with differential nutritional deprivation explored in this study was also studied in the normal cell line HEK293. Cell viability of HEK293 remained unaffected under most conditions and was found to be only decreased in GSʼ and Py conditions by 67 ± 1.6% and 45 ± 2.9% at 24 h. At 48 h, together with these two conditions, which caused a reduction in cell viability of 88 ± 1.1% and 55 ± 2.7%, a less significant decrease in viability of 15 ± 2.8 and 13 ± 2.7% was observed in the Hypo G’ and Q 4 mM groups compared to the control group (Fig. 1D).
Effect of differential deprivation of nutrient substrates on proliferation in cancer and normal cells. MTT assay was used to determine the change in viability/proliferation in cancer cell lines A MDA-MB-231 B HepG2 C HeLa cell and D non-cancer cell line HEK293 upon treatment of cells with media differentially deprived of G, Q and Py for 24, 48 and 72 h, as discussed in “Cell culture and treatments”. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Significance of change in cell viability represented as, mean ± S.D of three independently performed experiments with n = 3, was tested by one-way ANOVA followed by Tukey’s multiple comparison test. *p < 0.05, **p < 0.01, ***p < 0.001 vs Control (PG) of each time point
To support cell viability and proliferation data from MTT assay, we morphologically examined the effect of differentially depleted nutrient substrates in MDA-MB-231, HepG2 and HeLa cells using phase-contrast light microscopy. In both PG (control) and HG conditions, ‘normal’ cell morphology was observed in all three cell lines with an increase in ‘normal’ cell density from 24 to 72 h (Figs. 2A and B, 3A and B and 4A and B). In Hypo G condition, MDA-MB-231 could maintain its morphology up to 48 h (Fig. 2C), whereas in GS condition, noticeable disruption in cell shape can be seen even at 48 h (Fig. 2D). In these two conditions, morphology was retained up to 72 and 48 h, respectively, with some elongated, ‘spiked’ and ‘rounded’ morphology of cells in HepG2 (Fig. 3C and D) and HeLa cell lines (Fig. 4C and D). Out of three cell lines, HepG2 was the least affected (Fig. 3C and D). MDA-MB-231 exhibited a time-dependent continuous decrease in ‘normal’ cell density (Fig. 2C and D), whereas it either increased or was not significantly affected till 48 h and then slightly decreased at 72 h in HepG2 (Fig. 3C and D) and HeLa (Fig. 4C and D). Contrary to the MDA-MB-231 morphology, which was not much affected by HG’ medium until 48 h or PG’ medium till 24 h (Fig. 2E and F), as also shown by (Gwangwa et al. 2019), when they propagated MDA-MB-231 in similar conditions, more number of shrunken, ‘spiked’ as well as ‘rounded’ cells were found at all time points in HepG2 (Fig. 3E and F) and at 48 and 72 h in HeLa cells (Fig. 4E and F). HG’ but not PG’ condition could maintain morphology of HeLa cells till 24 h. These two conditions also did not support the increase in ‘normal’ cell density with time. A more pronounced alteration in cellular morphology was observed when the three cell lines were propagated in Hypo G’, GS’ and Py 1 mM conditions and led to decreased ‘normal’ cell density. Increased number of shape-distorted cells, visualized as shrunken, ‘rounded’ and condensed MDA-MB-231 cells, were found at 48 and72 h (Fig. 2G–I) and shrunken, ‘spiked’, ‘rounded’ and highly condensed HepG2 (Fig. 3G–I) and HeLa cells (Fig. 4G–I) were found at all three-time points. In all cell lines, Q 4 mM medium had an effect similar to GS medium but had slightly less capacity to maintain the cellular morphology, and it could maintain the shape of MDA-MB-231 cells till 24 h (Fig. 2J) and HepG2 and HeLa cells till 48 h (Figs. 3J and 4J) In all conditions except PG and HG, a continuous increase in shrunken, ‘spiked’ and ‘rounded’ cells from 24 to 72 h was observed in all three cell lines (Figs. 2C–J, 3C–J and 4C–J). Morphological analysis of the non-cancer cell line HEK293 showed normal morphology in most treatments (Fig. 5B–F and J), and major alterations in morphology were observed only in GS’ and Py conditions at 24 and 48 h (Fig. 5H and I) and less distinct changes in Hypo G’ at 48 h (Fig. 5G).
MDA-MB-231 cells differentially deprived of the nutrients have distinct and altered morphology at a later time of growth. MDA-MB-231 cells were exposed to differential depletion of G, Q and Py for 24 h (left panel), 48 h (middle panel) and 72 h (right panel), as described in “Cell culture and treatments”. Morphological changes were captured by an inverted phase-contrast microscope at 20 × magnification. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Scale bar = 50 μm
Effect of differential depletion of nutrient substrate on morphology in HepG2 cells. HepG2 cells were treated with growth media containing varying concentrations G, Q and Py for 24 h (left panel) , 48 h (middle panel) and 72 h (right panel), as discussed in “Cell culture and treatments”. Morphological differences were captured by an inverted phase-contrast microscope at 20X magnification. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Scale bar = 50 μm
Morphological variations in HeLa cells under nutritional stress. HeLa cells were exposed to DMEMs supplemented with changed concentrations of G, Q and Py for 24 h (left panel), 48 h (middle panel) and 72 h (right panel), as discussed in “Cell culture and treatments”. Alterations in cellular morphology were captured by an inverted phase-contrast microscope at 20X magnification. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Scale bar = 50 μm
Differential depletion of nutrients caused no, or lessened, effects on morphological variations in HEK293 cells. Cells were treated with growth media containing varying concentrations of G, Q, and Py for 24 h (left panel) and 48 h (right panel), as mentioned in “Cell culture and treatments” Changes in cell morphology were captured by an inverted phase-contrast microscope at 20X magnification. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Scale bar = 50 μm
Furthermore, alteration in cell morphology at 24 and 48 h of exposure to modified nutritional states was quantified relative to control (PG). At 24 h and in HG and Hypo G conditions, variation in the number of ‘normal’, ‘spiked’ and ‘rounded’ cells was non-significant in all cell lines (Fig. 6A, B and C), whereas GS condition less significantly (p < 0.05) increased ‘spiked’ morphology by 4.9% and 5.3% in MDA-MB-231 (Fig. 6A) and HeLa cells (Fig. 6C), respectively, while HepG2 morphology remains unaffected (Fig. 6B). In HG’ medium, a less significant (p < 0.05) increase in only ‘spiked’ morphology by 5.0% and 8.3% was found in MDA-MB-231 (Fig. 6A) and HeLa cells (Fig. 6C), respectively, whereas in HepG2 cells, a highly significant (p < 0.001) decrease in ‘normal’ morphology by 32.9% and an increase in ‘rounded’ morphology by 21.7% was also observed along with a significant (p < 0.01) increase in ‘spiked’ morphology (Fig. 6B). Exposure to PG’ and Q 4 mM media for 24 h led to less significant (p < 0.05) decrease in ‘normal’ morphology by 11.6% and 8.3%, respectively, in MDA-MB-231 and by 15.3% and 10.9%, respectively, in HeLa cells. A significant (p < 0.01) increase in ‘spiked’ morphology by 7.2% and 5.2%, respectively, in MDA-MB-231 (Fig. 6A) and by 10.3% and 12.7%, respectively, in HeLa cells (Fig. 6C) was also seen in PG’ and Q 4 mM media. Compared to MDA-MB-231 and HeLa cells, a highly significant (p < 0.001) decrease in ‘normal’ morphology by 41.4%, increase in ‘rounded’ morphology by 28.1% and significant (p < 0.01) increase in ‘spiked’ morphology by 13.3% was observed in PG’ media, whereas less significant (p < 0.05) increase in only ‘spiked’ morphology by 4.7% was detected in Q 4 mM media in HepG2 cells (Fig. 6B). In Hypo G’, GS’ and Py 1 mM conditions, decrease in ‘normal’ morphology by 15.5%, 27.3% and 17.7%, respectively, in MDA-MB-231 (Fig. 5A), by 49.3%, 61.3% and 52.7%, respectively, in HepG2 cells (Fig. 6B) and by 18.8%, 35.70% and 21.1%, respectively, in HeLa cells (Fig. 6C); increase in ‘rounded’ morphology by 12.70%, 22.9 and 10.7%, respectively, in MDA-MB-231, by 36.5%, 41.1% and 33.3%, respectively, in HepG2 cells (Fig. 6B) and by 7.6%, 14.2% and 8.4% respectively in HeLa cells (Fig. 6C) was found. At 24 h, both HepG2 and HeLa cells displayed a highly significant (p < 0.001) increase in ‘spiked’ morphology in GS’and Py 1 mM conditions and significant (p < 0.01) increase in ‘spiked’ morphology in Hypo G’ condition, whereas MDA-MB-231 cells showed significant (p < 0.01) increase in ‘spiked’ morphology in Py 1 mM condition only. Compared to 24 h, cell quantification data at 48 h did not vary significantly in PG, HG and Hypo G condition except less significant (p < 0.05) increase in ‘normal’ morphology and decrease in ‘spiked’ morphology in HG condition in MDA-MB-231 cells (Fig. 6D) and less significant (p < 0.05) increase in ‘spiked’ morphology in Hypo G condition in HeLa cells (Fig. 6F). However, a highly significant (p < 0.001) decrease in ‘normal’ morphology and increase in ‘rounded’ morphology in GS condition was found in MDA-MB-231 cells (Fig. 6D). GS medium also led to a less significant (p < 0.05) increase in ‘spiked’ morphology in HepG2 cells (Fig. 6E) and decrease in ‘normal’ morphology HeLa cells (Fig. 6F). At 48 h and in HG’condition, maximum percentage variation in morphologies occurred in HepG2 cells, followed by HeLa cells and HepG2 cells. In Q 4 mM media, there was a highly significant (p < 0.001) decrease in ‘normal’ morphology and an increase in ‘spiked’ and ‘rounded’ morphology in MDA-MB-231 cells (Fig. 6D), whereas HepG2 cells showed a less significant (p < 0.05) decrease in ‘normal’ morphology and a significant increase (p < 0.001) in ‘spiked’ cells (Fig. 6E) and HeLa cells displayed a less significant (p < 0.05) decrease in ‘normal’ morphology and increase in ‘spiked’ and ‘rounded’ morphology (Fig. 6F). A more pronounced change in percentage of ‘normal’, ‘spiked’ and ‘rounded’ cells was observed in PG’, Hypo G’, GS’and Py 1 mM conditions in all three cell lines, but the overall trend of these changes was similar to 24 h.
Quantification of morphological changes in MDA-MB-231, HepG2 and HeLa cells under varied nutritional stress. Photographs captured for each treatment and control group (PG) were used to count different types of morphological variations. Each cell was classified in one of the three broad categories of cell morphologies, named as ‘normal’, ‘spiked’ and ‘rounded. Change in the number of ‘normal’, ‘spiked’ and ‘rounded’ cells in each of the treatment groups at 24 h (A–C) and 48 h (D–F) was represented in percentage and statistical significance (*p < 0.05, **p < 0.01 and ***p < 0.001) of the change was determined relative to control group of respective time point and cell line. ‘Normal’ represented standard and characteristic morphology, ‘spiked’ denoted morphology of the stressed/ less viable cells and ‘rounded’ represented dead or very less viable cells or debris
Cell cycle profiling during differential nutritional stress in MDA-MB-231, HepG2 and HeLa cells
To establish the effect of differential deprivation of nutrients (G, Q and Py) on cell proliferation in MDA-MB-231, HepG2 and HeLa cells, cell cycle analysis was carried out at 24 and 48 h post-treatment. At 24 h, the percentage of MDA-MB-231 cells in G0/G1 and S-phase remain unchanged when exposed to metabolic conditions used in this study, except GS’, Py 1 mM and Q 4 mM conditions which showed a significant (p < 0.01) decline in G0/G1 (Fig. 7A). Besides, a highly significant (p < 0.001) increase in S-phase cells was also found in GS’. G2/M arrest of MDA-MB-231 cells was limited to only Hypo G’, GS’, Py 1 mM and Q 4 mM media (Fig. 7A). In HepG2 cells, G2/M arrest was detected in GS and Q 4 mM media, whereas G0/G1 arrest together with a marked decrease in S-phase was observed in HG’, PG’, Hypo G’, GS’, Py 1 mM and Q 4 mM media (Fig. 7B). Treatment of HeLa cells with GS, HG’, PG’ and Hypo G’ media for 24 h resulted in S-phase arrest and a highly significant (p < 0.001) decline in G2/M cells (Fig. 7C). GS’, Py 1 mM and Q 4 mM conditions also exhibited a highly significant (p < 0.001) reduction in the G2/M cells, whereas GS’ condition also led to a less significant (p < 0.05) G0/G1 arrest (Fig. 7C).
Analysis of cell cycle profile in MDA-MB-231, HepG2 and HeLa cells under metabolic stress. Cells were fixed and stained with propidium iodide after treatment for 24 h (A–C) and 48 h (D–F) with media containing differentially depleted nutrient substrates. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Cell cycle phase distribution profile was recorded by flow cytometry and analyzed by ModFit LT 5.0 software. Data were represented as mean ± SD. *p < 0.05,**p < 0.01,***p < 0.001 versus control (PG) of each time point, after analysis with one-way ANOVA combined with Tukey’s test
At 48 h, out of nine varied metabolic conditions, MDA-MB-231 and HepG2 cell lines in HG medium showed significant (p < 0.01) decrease in G0/G1 and increase (p < 0.05 and p < 0.001, respectively) in S-phase cells. These two cell lines in GS, HG’, PG’and Hypo G’, Py 1 mM and Q 4 mM media exhibited highly significant (p < 0.001, except Py 1 mM medium in MDA-MB-231 where p < 0.05) increase in G0/G1 and decrease in S-phase cells (Fig. 7D and E). GS’ medium exhibited contrasting effects, i.e., highly significant (p < 0.001) decrease, no change and increase in G0/G1, S and G2/M-phase cells, respectively in MDA-MB-231 (Fig. 7D) and highly significant (p < 0.001) increase, decrease and no change in G0/G1, S and G2/M-phase cells, respectively in HepG2 cell line (Fig. 7E). Depletion of nutrients in HG’, PG’, Hypo G’, GS’ Py 1 mM and Q 4 mM media also led to G2/M arrest in MDA-MB-231 (Fig. 7D), whereas less significant (p < 0.05) decrease in G2/M-phase cells was observed in HepG2 cell line grown in Hypo G and GS media (Fig. 7E). HeLa cells cultured in HG and GS conditions showed a cell cycle profile similar to, but more prominent than MDA-MB-231 (Fig. 7D and F). In the Hypo G condition of HeLa cells, highly significant (p < 0.001) increase in G0/G1 and significant (p < 0.01) decrease in S-phase cells, an effect distinct from the other two cell lines was observed (Fig. 7F). Contrary to MDA-MB-231 and HepG2 cell lines, HeLa displayed a highly appreciable (p < 0.001) decrease in G0/G1 and increase in S-phase cells in HG’, PG’and Hypo G’ media, whereas cell cycle phase distribution pattern showed by HeLa cells in GS’, Py 1 mM and Q 4 mM conditions was similar to that of HepG2 cells apart from G2/M arrest in HeLa cells in GS’ condition (Fig. 7D, E and F). Thus, the studied nutrient derivation demonstrated cell-type dependent heterogeneity in cell cycle regulation.
Alteration in nutritional balance perturbs ROS homeostasis in cancer cells
MDA-MB-231 and HeLa cells displayed similar, significant (p < 0.05 or 0.01 or 0.001) and continuous rise in cytosolic ROS with increasing time points in all conditions from GS to GS’ relative to PG (control). In these four conditions, ROS generation varied between 3.2–6.9, 8–11.3, and 6.2–20.5-fold in MDA-MB-231, and 2.3–4.7, 3.6–9.4 and 5.4–10.7-fold in HeLa cells at 24, 48 and 72 h, respectively (Fig. 8A and C). Py 1 mM and Q 4 mM media also showed significant and continuously increasing ROS from 24 to 72 h, which rose by 5.7 and 3.4-fold, 11.2 and 8.7-fold, and 14.8 and 10.8-fold at 24, 48, and 72 h, respectively, in MDA-MB-231, and by 4.3 and 2.3-fold, 8.8 and 4.2-fold, and 9.3 and 6.6-fold at 24, 48, and 72 h, respectively, in HeLa cells. A significant (p < 0.01) and less significant (p < 0.05) increase in cytosolic ROS was also observed in Hypo G medium in MDA-MB-231 and HeLa cells, respectively at 72 h. In HepG2 cells, a significant increase in ROS by 2.1, 2.6, 3.5, 4.5, and 3.8-fold was examined in HG’, PG’, HypoG’, GS’ and Py 1 mM media at 24 h (Fig. 8B). ROS production also significantly increased in these metabolic conditions by 2.5, 2.9, 3.4, 3.7 and 3.6-fold, respectively at 48 h, and by 3.2, 3.6, 4.2, 6.4, and 5.2-fold respectively at 72 h. A slight decrease in ROS generation was observed in HypoG’, GS’ and Py 1 mM media at 48 h relative to 24 h, whereas a relatively higher and significant (p < 0.01 or 0.001) fold increase in ROS was detected in all conditions from HG’ to Py 1 mM at 72 h compared to 24 h and 48 h in HepG2 cells (Fig. 8B). GS media led to less significant (p < 0.05) rise in ROS production by 2.3-fold at 72 h, while Q 4 mM media resulted in less significant (p < 0.05) and significant (p < 0.01) increase in ROS production by 1.8-fold and 2.9-fold at 48 and 72 h, respectively (Fig. 8B). Thus, among the studied cell lines, MDA-MB-231cells showed the greatest variation in cytosolic ROS, whereas cytosolic ROS production in HepG2 cells was the least affected by changed metabolic states.
Nutrient-type, concentration and deprivation time-dependent induction of oxidative stress in MDA-MB-231, HepG2 and HeLa cells. As a marker of generation of oxidative stress condition, the presence of intracellular ROS was detected in A MDA-MB-231, B HepG2 and C HeLa cells using a probe, H2DCFDA by flow cytometry after growing cells in media with the differentially withdrawn nutrient substrates for 24, 48 and 72 h. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. ROS production was represented as mean ± SD and analyzed by one-way ANOVA followed by a Tukey’s test. MFI, mean fluorescence intensity and H2O2 (0.5 mM, 2 h) = positive control. *p < 0.05, **p < 0.01, ***p < 0.001 versus control (PG) of each time point
To examine the involvement of mitochondrial ROS in perturbation of overall ROS (intracellular/cytosolic and organelles-associated) homeostasis by differential nutrient depletion in MDA-MB-231, HepG2 and HeLa cells, we evaluated change in mtROS levels using mitochondrial superoxide indicator MitoSOX Red. Both, MDA-MB-231, and HepG2 cells showed similar increasing pattern of variation in mtROS production as that of intracellular ROS in HG’, PG’, HypoG’, GS’ and Py 1 mM conditions, where mitochondrial superoxide increased significantly between 2.0–2.7-fold and 3.0–4.7-fold, relative to control, in MDA-MB-231 and in HepG2 cells, respectively, at 48 h (Fig. 9A and B). GS and Q 4 mM induced excess mtROS in only MDA-MB-231 and HeLa cells (Fig. 9A and C). HeLa cells also exhibited significant increment in mtROS by 3.4 & 3.5-fold in GS’ and Py 1 mM group, respectively (Fig. 9C).
Nutrient deprivation-induced oxidative stress in MDA-MB-231, HepG2 and HeLa cells involved generation of mitochondrial ROS (mtROS). mtROS was detected in A MDA-MB-231, B HepG2 and C HeLa cells using a mitochondrial superoxide indicator, MitoSOX Red by fluorimetry after exposing cells to growth conditions differentially deprived of nutrients for 48 h. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. MFI, mean fluorescence intensity, and Rotenone (1 μM, 12 h) = positive control. *p < 0.05, **p < 0.01, ***p < 0.001 versus control (PG) after analysis with one-way ANOVA followed by a Tukey’s test
Effect of nutrient deprivation on mitochondrial function in cancer cells
Trends for the shift in MMP due to variation of nutrient substrates in cell lines under study were correlated with trends for ROS. At 24 h, a significant loss of MMP was observed when MDA-MB-231, HepG2 and HeLa cells were grown in Hypo G’, GS’ and Py 1 mM media where MMP was lost from the ratio of 1 (control) to the ratios 0.66, 0.43 and 0.67, respectively in MDA-MB-231 (Fig. 10A), 0.59, 0.56 and 0.55, respectively in HepG2 (Fig. 10B) and 0.59, 0.51 and 0.71, respectively in HeLa (Fig. 10C). At this time point, among the three cell lines, only MDA-MB-231 exhibited a significant (p < 0.05 and 0.01, respectively) fall in MMP in GS and Q 4 mM media to ratios 0.75 and 0.69, respectively (Fig. 10A), whereas a decrease in MMP brought about by PG’ medium was significant (p < 0.01) in HepG2 (Fig. 10B) and less significant (p < 0.05) in HeLa cells (Fig. 10C) and a less significant MMP loss to the ratio 0.69 by HG’medium was selective to only HepG2 (Fig. 10B). Compared to 24 h, a more pronounced mitochondrial membrane depolarization upon exposure to modified nutritional conditions was observed in all three cell lines at 48 and 72 h, following a trend of variations similar to 24 h of respective cell line (Fig. 10A, B and C). In contrast to 24 h, a significant loss of MMP was also caused by Hypo G and PG’ condition in MDA-MB-231 (Fig. 10A), Q 4 mM condition in HepG2 (Fig. 10B) and Hypo G, GS, HG’ and Q 4 mM conditions in HeLa cells (Fig. 10C) at both 48 and 72 h, whereas HG’ in MDA-MB-231 cells and Hypo G and GS conditions in HepG2 cells led to significant disruption of MMP only at 72 h.
Effect of differential depletion of energy substrates on mitochondrial function in cancer cells. Substrate-type, concentration and time-dependent variation in mitochondrial membrane potential was observed in A MDA-MB-231, B HepG2 and C HeLa cells using a fluorescent probe R123 by flow cytometry after differentially starving cells to the energy substrates by growing them in media containing limiting concentration of G, Q and Py for 24, 48 and 72 h. As an indicator of mitochondrial health and function, change in intracellular ATP content in response to the altered metabolic state of the growth media was also observed in D MDA-MB-231, E HepG2 and F HeLa. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py. Data represented as mean ± S.D of three independent experiments (n = 3) and analyzed by one-way ANOVA plus Tukey’s test. MFI mean fluorescence intensity, Rotenone (1 μM) = positive control. *p < 0.05, **p < 0.01, ***p < 0.001 versus control (PG) of each time point
Mitochondrial health status and functions are very well correlated with their ability to synthesize ATP. On observation of cellular ATP content after 48 h of differential withdrawal of energy substrates, MDA-MB-231 cells exhibited a 1.21-fold increase in ATP level in HG medium and decrease in ATP to the ratios 0.77, 0.91, 0.79, 0.66, 0.41and 0.69 in Hypo G, HG’, PG’, Hypo G’, Py 1 mM and Q 4 mM media, respectively, compared to control medium (PG). Complete depletion of G in the presence (GS) and absence (GS’) of Q and Py resulted in a reduction in ATP content to the ratios 0.73 and 0.35, respectively (Fig. 10D). Both HepG2 and HeLa cells showed a similar pattern of alterations in cellular ATP content. Exposure to HG’, PG’, HypoG’, GS’, Py 1 mM and Q 4 mM media led to decline in ATP level to the ratios 0.71, 0.66, 0.62, 0.45, 0.57 and 0.81, respectively, in HepG2 cells (Fig. 10E) and to the ratios of 0.79, 0.71, 0.54, 0.43, 0.49 and 0.78, respectively, in HeLa cells (Fig. 10F) relative to PG medium. A less significant (p < 0.05) change in ATP amount was found to be brought about by treatment of HeLa cells with HG and GS media (increase and decrease to the ratios 1.23 and 0.80, respectively), whereas variation due to HypoG medium was non-significant in both, HepG2 and HeLa cells (decrease to the ratios 0.93 and 0.88, respectively) (Fig. 10E and F).
Determination of glutathione (GSH) levels
We also evaluated the antioxidant GSH levels in the three cancer cell lines under different nutrient conditions. Except for the HG condition in MDA-MB-231 and the HG and Hypo G conditions in HeLa cells, all other conditions showed significantly decreased levels of GSH in both cell lines at 48 h. GS, HG’, PG’, Hypo G’, GS’, Py 1 mM, and Q 4 mM groups caused a decrease in GSH content by 2.2, 1.4, 1.5, 1.7, 3.3, 3.1, and 2.4 folds in MDA-MB-231 cells and 1.8, 1.6, 1.7, 2.5, 3.9, 1.9, and 1.8 folds in HeLa cells, respectively, compared to the control group (Fig. 11A and C). HG’, PG’, Hypo G’, GS’, and Py 1 mM conditions also reduced GSH levels by 1.3, 1.5, 1.4, 2.9, and 2. 8 folds in HepG2 cells. Interestingly, HepG2 cells also showed increased levels of GSH in the Q 4 mM (1.2-fold) condition (Fig. 11B).
Determination of the status of glutathione in cancer cells under varying nutritional deprivation. A fluorometric assay based on the detection of reduced glutathione (GSH) in cell lysate by conversion of a substrate, O-phthalaldehyde, to a fluorescent product measured the reduced glutathione (GSH) levels in A MDA-MB-231, B HepG2, and C HeLa cells after treating the cells with media differentially depleted in nutrients for 48 h. G, glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q, glutamine. Prime (ʹ) represents media without Q and Py. MFI, mean fluorescence intensity; Curcumin (20 μM, 12 h) = positive control. Data were represented as a fold change (mean ± SD) in GSH content in treated groups compared to the control group. *p < 0.05, **p < 0.01, ***p < 0.001 versus control (PG) after analysis with one-way ANOVA followed by a Tukey’s test
Detection of autophagy by acidic vesicular organelles (AVOs)
The presence of AVOs in the cells has been widely used as an autophagy marker. These AVOs are detected by fluorescence microscopy using acridine orange (AO) dye that stains differently on the nucleus and cytoplasm (green) and AVOs (red). In a merged fluorescence microscopy image (red + green channels), AVOs are visualized as red or orange-colored vacuolar structures. AVOs were not detected in HG, PG, and HypoG conditions in all three cancer cell lines, i.e., MDA-MB-231, HepG2, and HeLa (Fig. 12A, B, and C), whereas the GS condition caused AVO formation primarily in MDA-MB-231 cells, and very few AVOs were found in HeLa cells (Fig. 12D). Nutrient substrate deprivation in HG’, PG’, Hypo G’, GS, and Py 1 mM treatment groups displayed the presence of AVOs, thus autophagy, in all three cell lines (Fig. 12E–I). Only a fewer number of cells with AVOs or cells with a smaller number of AVOs were identified in growth conditions containing glutamine but lacking glucose and pyruvate in HepG2 cells, followed by HeLa and MDA-MB-231 cells (Fig. 12J). The negative control group (pre-treatment with 100 nM Bafilomycin A1 for 6 h in autophagy-inducing condition GS’) inhibited AVO formation (Fig. 12K).
Acridine orange (AO) staining-based preliminary detection of autophagy induced by nutrient deprivation in cancer cells. A–K Merged fluorescence microscopy images (red + green channels) showing red or orange-colored acidic vesicular organelles (AVOs) as well as the cytoplasm and nucleus (green fluorescence) in MDA-MB-231 (left panel), HepG2 (middle panel), and HeLa (right panel) cells after 48 h of treatment (B–J), followed by staining with dye AO. AVOs are detected by their red fluorescence in red channel and visualized as red-colored (in the absence of green fluorescence due to damage to the cytoplasm and nucleus) or orange-colored (in the presence of cytoplasm and nucleus that fluoresce green by AO staining) vacuolar structures in the merged image. K Negative control group included 6 h pre-treatment with 100 nM Bafilomycin A1 (Baf A1), followed by exposure to media completely deprived of glucose, glutamine and pyruvate (GS’). Scale bar: 20 µm
Comparable inhibition of cell migration due to differential glucose deprivation in MDA-MB-231 and HeLa cells is distinct from HepG2 cells
The initial steps of metastasis, namely cell migration and tissue invasion are clinically significant features of cancer (Efferth 2009). Tumor metastasis is prominently driven by diverse homotypic and heterotypic cell–cell interactions (Mishra et al. 2019). Therefore, determining migration and invasion capability of a tumor cell becomes highly relevant for developing new anticancer strategies. In this respect, cell culture two-dimensional cell migration or scratch assay or wound closure assay was performed to evaluate the difference in combined proliferation and migration potential of MDA-MB-231, HepG2 and HeLa in response to differential depletion of G and to probe its effectiveness as a probable strategy for the treatment of these cancer types. Both, PG (Fig. 13A.1, B.1 and C.1) and HG media (Fig. 13A.2, B.2 and C.2) resulted in significant (p < 0.01 or p < 0.001) migration of cells in all three cell lines at 24 and 48 h (Fig. 13A.5, B.5 and C.5). However higher rate of wound closure was found in the HG medium as compared to PG. Among the three cell lines, at 24 h, MDA-MB-231 exhibited the maximal potential for migration in PG medium (Fig. 13A.1 and A.5) followed by HeLa (Fig. 13C.1 and C.5) and HepG2 (Fig. 13B.1 and B.5), whereas at 48 h, the migration capacity decreased in the order of HepG2, MDA-MB-231 and HeLa. In HG media, the highest migration at both time points was shown by MDA-MB-231 (Fig. 13A.2 and A.5) followed by HepG2 (Fig. 13B.2 and B.5) and HeLa (Fig. 13C.2 and C.5). In Hypo G condition also, HepG2 cell line showed significant (p < 0.01), and highly significant (p < 0.001) migration of cells at 24 and 48 h, respectively (Fig. 13B.3 and B.5) comparable to its PG and HG conditions (Fig. 13B.1 and B.2). But MDA-MB-231 could exhibit only less significant (p < 0.05) and significant (p < 0.01) wound healing properties at 24 and 48 h, respectively, due to slowed migration resulting in prolonged wound closure time (Fig. 13A.3 and A.5). In HeLa cells, a less significant (p < 0.05) wound healing potential was found at 24 h compared to significant (p < 0.05) wound healing in HepG2 cells. Though highly significant (p < 0.001) migration was also observed in HeLa cells at 48 h, less migration (by 29.48%) was detected in it (Fig. 13C.3 and C.5) compared to HepG2 cells, which migrated by 53.79%. In contrast to MDA-MB-231 cells, which led to halting of cell migration due to inefficiency of cells to proliferate in GS condition at 24 and 48 h (Fig. 13A.4 and A.5), HepG2 cells showed significant and highly significant (p < 0.001) migration at 24 and 48 h respectively (Fig. 13B.4 and B.5), whereas this condition could only allow reduced migration (p < 0.05) in HeLa cells at both time points (Fig. 13C.4 and C.5).
Migration potential of MDA-MB-231, HepG2 and HeLa cells in glucose deprived condition. Cells were cultured for 24 h, and 48 h in PG, HG, Hypo G and GS media. Images were captured at the indicated time points, and a decrease in the wound area was calculated. Representative images of change in migration area in MDA-MB-231, HepG2 and HeLa cells in HG (A.2, B.2 and C.2), Hypo G (A.3, B.3 and C.3) and GS (A.4, B.4 and C.4) relative to control (A.1, B.1 and C.1) have been shown. Migratory capacity of cells was represented as (mean ± S.D) area of the three images for each group obtained from three separate experiments and analyzed by one-way ANOVA–Tukey’s multiple comparison test. *p < 0.05, **p < 0.01, ***p < 0.001 versus 0 h of each treatment or between indicated groups
Biplot analysis and classical hierarchical clustering
To see the overview of the effects of differential deprivation of nutrients, biplot analysis using correlation matrix and classical HCA were performed. Biplot is a combined representation of principal component analysis (PCA) plot and loading plot that helps in finding out the variables (phenomena or processes or characteristics) responsible for producing differences among clusters of samples. Based on the PCA scores, samples (cell growth conditions: PG, HG, Hypo G, GS, HG’, PG’, Hypo G’, GS’, Py 1 mM, and Q 4 mM) occupied different locations on the PCA plot and samples exhibiting similar effects formed clusters in MDA-MB-231 (Fig. 14A), HepG2 (Fig. 14C), and HeLa (Fig. 14E) cells. A loading plot displayed the influence or loading of each variable (phenomenon/ process/ characteristic: Cell viability, Cell morphology—normal, spiked, rounded, Cell cycle phase- G0, S, G2/M, ROS generation, MMP, ATP production and Cell migration) on a principal component (PC) by measuring the projections of these variables on each PC. In MDA-MB-231 (Fig. 14A), HepG2 (Fig. 14C), and HeLa (Fig. 14E) cell lines, the two variables were positively and negatively correlated when the angles formed by their vectors at the origin of the two PCs (PC1 = 0, PC2 = 0) were an acute and an obtuse angle, respectively. Smaller the angle (near to 0°), stronger is the positive correlation and larger the angle (near to 180°), stronger is the negative correlation between the two variables. HCA of effects observed in MDA-MB-231 (Fig. 14B), HepG2 (Fig. 14D), and HeLa (Fig. 14F) cell lines, further corroborated the cluster pattern of samples observed in PCA.
Biplot analysis and Hierarchical cluster analysis of effects observed at 48 h. Biplots for A MDA-MB-231 C HepG2 and E HeLa. Samples (cell growth conditions) shown as colored dots on the PCA plot and increase in variables (cellular phenomena or processes or characteristics) shown as green colored vectors on the loading plot. Classical hierarchical cluster dendrograms show arrangement of clusters of different cell growth conditions in B MDA-MB-231 D HepG2 and F HeLa. Distance on the vertical axis represents the relatedness or similarity in effects among cell growth conditions. G glucose; HG high glucose; PG physiological glucose; Hypo G hypoglycemic glucose; GS glucose starvation; Py pyruvate; Q glutamine. Prime (ʹ) represents media without Q and Py
Discussion
To study the metabolic adaptation in adenocarcinoma of the breast, hepatocellular carcinoma (HCC) and adenocarcinoma of the uterine cervix, we developed in vitro the nutrient-deprived conditions of tumor microenvironment in MDA-MB-231 (a highly aggressive triple-negative breast cancer), HepG2 and HeLa cells, respectively, by differential depletion of G, Q and Py in growth media. Then, we determined the effects of metabolic stress on cell viability, morphology, cell cycle profile, redox homeostasis, mitochondrial function, bioenergetics, antioxidant (GSH) status, autophagy induction and cell migration in these cells.
MDA-MB-231 is a basal-type malignant cancer cell that harbour mutation in the oncogene K-Ras (Balestrieri et al. 2012), and exhibit greater activity and expression of G13D, an activated oncogenic K-Ras protein (Kim et al. 2015). Similar to other fast-growing tumor cells (Rodríguez-Enríquez et al. 2010; Sandulache et al. 2011), MDA-MB-231 cells are highly glycolytic in nature which exhibit much-elevated consumption of G compared to Q with limited TCA cycle activity including, decreased pyruvate oxidation due to reduced activity of pyruvate dehydrogenase. MDA-MB-231 cells rely on carbon from glutamine for anaplerotic synthesis of nucleotides, proteins and fatty acids (Gaglio et al. 2011). HepG2 and HeLa cells express Myc protein which enhances Q metabolism by upregulating glutaminase (Liu et al. 2012). As reported by Yuneva et al. (2007), human fibroblast overexpressing Myc proto-oncogene have been shown to get killed by Myc dependent apoptosis under Q depletion but not by G deprivation. However, cell death due to Q withdrawal of K-Ras transformed NIH3T3 fibroblast cells containing an activated form of the K-Ras oncogene was marginal relative to that observed in cells overexpressing Myc (Gaglio et al. 2009), but these K-Ras transformed fibroblasts stop their growth and eventually die in G shortage (Chiaradonna et al. 2006).
We first determined the effect on proliferation due to differential deprivation of G, Q and Py by MTT assay, morphological examination and cell cycle analysis. Differential decrease in cell proliferation of cancer cells in modified nutrient conditions used in this study may be attributed to the activation of the specific pathway in these cancer cells upon differential depletion of nutrient substrates. The ability of MDA-MB-231 cells to support its proliferation and survive till 24 h in HG’and PG’ can be correlated with their enhanced dependency on G as nutrient substrate and availability of sufficiently high concentration of G for maintenance of viability. Differential change in viability in MDA-MB-231 and HeLa upon G depletion is speculated to be due to the differential expression of glutaminase (GLS1) and c-MYC (Wu et al. 2015). Proliferation and survival of HepG2 cells in PG, Hypo G GS and Q 4 mM conditions can be attributed to sestrin2 mediated and Q-dependent activation of PGC-1α, which is a mitochondrial biogenesis marker (Kumar et al. 2018). At later time points, HepG2 cells could sustain proliferation or could remain viable only when Q was present, which provided the essential macromolecules by anaplerosis, while HeLa cells supported their growth when together with Q, sufficient G was also present. Our qualitative and quantitative assessment of morphological differences was affirmative to induction of stress and cell death associated morphological characteristics like presence of echnoid spikes, membrane blebbing, cell shrinkage, condensed nuclei, and apoptotic bodies in growth conditions where nutrients were deprived to a greater degree and at later time points which corroborated our cell viability data. Cell viability and morphology results from normal cell line HEK293 further confirmed that normal cells remain unaffected in most of the nutrient-deprived conditions investigated in this study.
Reduction in viability and inhibition of tumor cell proliferation by many cytotoxic drugs and conditions of stress have been reported to involve cell cycle arrest in G0/G1 or S or G2/M-phase or their combinations (Shapiro and Harper 1999; Chua et al. 2009; ZHU et al. 2015; Wang et al. 2016; Han and Zhou 2016; Jingwen et al. 2017; Hubackova et al. 2020). Arrest in G0/G1 indicates growth and nutritional stress and the presence of inherited DNA lesions induced by replication stress of the previous cell cycle (Lezaja and Altmeyer 2018), whereas S and G2/M-phase arrests specify replication stress induced inhibition of replication and improper replication of DNA or excessive damage to DNA, respectively (Yi et al. 2015; Wenzel and Singh 2018; Pan et al. 2019). These cell cycle arrests are seen as a mechanism adopted by the damaged cells undergoing cell division to prevent their proliferation till the damage persist and to ensure transfer of error-free genetic material to the daughter cells once the damage is repaired. Otherwise, the cells harbouring irreparable DNA damage are eliminated by cell death processes. Display of cell cycle arrest at 48 h by MDA-MB-231 cells, in G0/G1 and G2/M phases due to GS and GS’ conditions, respectively, and in both, G0/G1 plus G2/M phases after exposure to HG’, PG’, Hypo G’, Py 1 mM and Q 4 mM conditions can be seen as an attempt to cope with nutritional stress and survive. Colombo et al. (2011), highlighted the role of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase, isoform 3 (PFKFB3) and GLS1 in causing S and G0/G1 arrest in conditions similar to GS and PG’, respectively, at 18 h working with synchronized HeLa cells. However, our study revealed S-phase arrest in both conditions at 24 h and G0/G1 and S-phase arrest in GS and PG’ conditions, respectively, at 48 h. Our finding of S-phase arrest in HG’ and PG’ is corroborated by the findings of Sun et al. (2019), who also reported S-phase arrest upon a similar condition of Q-deprivation for 48 h in T24 bladder cancer cell. Scarcity of G for 48 h in all three cell lines of the present study showed G0/G1 arrest similar to prostate, endometrial and ovarian cancer cells (Singh et al. 1999; Han et al. 2015; Sun et al. 2017). Oncogene modulated apoptotic sensitivity of transformed cells by arrest at energy-dependent G1 and G2/M checkpoints upon glucose deprivation has also been reported (Kansara and Berridge 2004). Recently stimulatory or inhibitory roles of different proteins resulting in cell cycle arrest of cancer cells in different phases under nutrient stress have been reported leading to their growth/survival or death. For example, induction of G2/M arrest and cell death by (β-N-acetylglucosamine)2-modified proteins due to increased p53 phosphorylation and extended upregulated expression of activation transcription factor 3 (ATF3) and G1/S arrest leading to cell survival under G depletion in renal carcinoma cells (Isono et al. 2014), and unchanged cell cycle profile leading to cell survival, by reduced expression of chk1 protein upon glucose deprivation for 18 h in lung (A549) and kidney (HEK293) cancer cells (Kim et al. 2011).
Cancer cells contain high ROS levels compared to normal cells due to increased metabolism, which could be pro-tumorigenic but also make them susceptible to oxidative stress-mediated cell death. Utilization of this ROS imbalance has remained one of the major strategies to kill cancer cells. Therefore, we also investigated cytosolic and mitochondrial ROS levels upon differential nutrient deprivation in growth media in MDA-MB-231, HepG2, and HeLa cells. Activation of adenosine monophosphate-activated kinase (AMPK), a pivotal sensor to nutritional stress, is linked with both, the survival of cancer cells by combating oxidative stress (Nogueira and Hay 2013), and inhibition of tumor cell proliferation (Shackelford and Shaw 2009). Non-induction of ROS in HG, PG and HypoG conditions till 72 h in all three cell lines (except at 72 h in HypoG condition in MDA-MB-231 and HeLa cells) of the present study can be correlated with the survival role of AMPK. In contrast, a continuous surge in intracellular ROS in these cells under other metabolic stress conditions (except GS condition at 24 and 48 h and Q 4 mM condition at 24 h in HepG2 cells) can be linked with its proliferation inhibition activity. Greater increase in oxidative stress in these cells grown in media lacking Q (HG’, PG’, Hypo G’, GS’ and Py 1 mM) is expected to be an effect of inhibition of the antioxidant activity of glutathione and other antioxidants due to Q depletion. These findings suggest that deprivation of essential nutrients for shorter durations and not-so-essential nutrients even for longer durations primes these cancer cells to adapt to the induced environment of early oxidative stress and activate survival mechanisms in these cells, whereas type, concentration, and time-dependent extreme depletion of nutrients create hostile oxidative environments that render these cancer cells susceptible to cell death pathways.
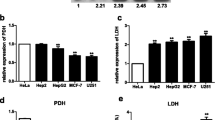
Then, we were interested in knowing the mitochondrial functional status by determining MMP and intracellular ATP upon differential withdrawal of nutrients. In cancer cells, the presence of mitochondria with more hyperpolarized MMP is seen as a pro-survival mechanism. The observed nutrient deprivation led to differential mitochondrial membrane depolarization was dependent on the nutrient and cell type and was probably a consequence of increased ROS generation. Reitzer et al. (1979), showed that Q, not G, was the main source of energy in HeLa cells. In a recent investigation, deviation from aerobic glycolysis was observed in HeLa cells where OxPhos was the major contributor (more than70%) in overall cellular ATP generation in normoxia condition (Rodríguez-Enríquez et al. 2010; Hernández-Reséndiz et al. 2015). The capability of HeLa and HepG2 cells to modulate their mitochondrial structure and function according to the availability of energy substrate facilitates the generation of energy exclusively from OxPhos by increased branching, constriction of the mitochondrial network and upregulated respiratory chain proteins (Rossignol et al. 2004; Plecitá-Hlavatá et al. 2008). On the contrary to HepG2 and HeLa cells, which showed ATP depletion primarily in media lacking Q, significant lowering of intracellular ATP content in Hypo G and GS conditions in MDA-MB-231 cells was possibly due to their inability to maximize utilization of Q aided OxPhos and greater dependence on glycolysis for energy, as also shown by Palorini et al. (2013).
Increasing ROS generation to destroy cancer cells in a targeted manner is probably challenging. As an alternative, since cancer cells contain high level of antioxidants to maintain homeostasis, inhibiting antioxidants to expose cancer cells to endogenously produced ROS is a viable treatment strategy (Hayes et al. 2020). A number of small molecule screens have identified drugs that selectively prevent the proliferation of transformed cells by increasing ROS due to decreased glutathione levels and inhibition of antioxidant pathways (Sullivan and Chanel 2014). Our results also showed significantly low levels of antioxidant glutathione (GSH) in all three cancer cell lines under conditions of severe nutrient starvation for 48 h. Higher GSH levels in HepG2 cells under Q 4 mM conditions can be attributed to the induction of compensatory antioxidant defense mechanisms in response to prolonged nutrient stress and the capacity of glutamine to synthesize GSH. Based on the measurement of GSH content, the trend of change in the NADH/NAD+ ratio (increase or decrease) can also be approximately estimated since a high NADH/NAD+ ratio means NADH is not getting converted to NAD+ , leading to low NADPH and consequently low GSH (Xie et al. 2020). The measurement of the NADH/NAD+ ratio helps in determining the dependency of cancer cells towards glycolysis (low NADH/NAD+ ratio) or OxPhos (high NADH/NAD+ ratio, which is linked to increased ROS generation) (Yang and Sauve 2016). Our data on GSH content analysis suggested that most of the nutrient-deprived conditions in the three cancer cell lines negatively impacted glycolysis, except the HG condition in MDA-MB-231, Hypo G, GS, and Q 4 mM conditions in HepG2, and HG and Hypo G conditions in HeLa cells. Despite the high NADH/NAD+ ratio expected in these unfavourable conditions for glycolysis, OxPhos might also be found to be reduced due to decreased GSH, excess ROS and lost membrane potential-mediated mitochondrial dysfunction. However, these speculations need to be validated with more direct estimates of NADH/NAD+ ratios and other targeted studies in the future.
Next, by a preliminary study based on detecting acidic vesicular organelles (autophagic vacuoles), we examined whether the differential nutrient deprivation also induced autophagy. Autophagy is mostly associated with cell survival and has been looked at as a process for recycling macromolecular precursors to supply nutrient sources and building blocks, maintaining cellular homeostasis. In cancer cells, the role of autophagy is rather controversial, as it exhibits tumor suppressing and tumor-promoting effects during the early and late stages of cancer, respectively (Lim et al. 2021). Inactivation of autophagy has been implicated in several cancers, including breast and liver (Lorin et al. 2013). Evidence of autophagy playing a ROS scavenger role in cervical cancer and immortalized mouse kidney cells (Scherz-Shouval et al. 2007; Mathew et al. 2009), when the ROS production is up to the level that promotes cancer (not excess ROS that can cause oxidative stress-mediated cell death), signifies its contribution to the inhibition of tumorigenesis by preventing DNA damage and maintaining genome integrity. Also, cells can either survive or die depending on the relative strength of the cell survival signal generated by autophagy (Noguchi et al. 2020). We found AVOs in the condition of excess nutrient depletion (PG’, Hypo G’, G’ and Py 1 mM) for 48 h, in all three cancer cell lines utilized in this study. This happened probably in response to the uncontrolled and prolonged activation of autophagy by oxidative stress, which may lead to cellular disintegration by excessive self-digestion or activation of apoptosis and ultimately cell death (Azad et al. 2009; Lim et al. 2021). Oxidative stress due to excess ROS and nutrient withdrawal has been shown to induce autophagic cell death (Azad et al. 2009). Also, there are many reports of cross-talk between autophagy and apoptosis and activation of apoptosis by autophagy (Noguchi et al. 2020). Furthermore, autophagy has also been reported to hinder tumor cell growth by reducing inflammation and inducing senescence (Lorin et al. 2013). Whether the autophagy induction observed in certain conditions of this study followed apoptosis, necrosis, and senescence, or whether autophagic cell death was independent of the other modes of cell death, needs to be explored in future studies.
Finally, to test the inhibition of the metastatic property due to differential glucose deprivation, a cell migration assay was conducted. The maximum speed and inhibition of migration of MDA-MB-231 cells detected in HG and GS conditions, respectively, were in line with a recent study (Santos and Hussain 2020), which showed higher concentrations of glucose lowered the expression of actin, elevated the expressions of kinesin, a motor protein, NFkB, reduced the nuclear stiffness, induced the hybrid epithelial-mesenchymal transition and increased the expression of Ki-67, a proliferation marker resulting in a considerable increase in the migration speed. Inhibition of migratory properties of MDA-MB-231 cells in GS conditions can be related to the decreased activity of K-Ras protein. Similar to the migration promoting effect of glucose shortage by liver kinase B1-AMPK-mediated MMP-9 activation (Endo et al. 2018), HepG2 cells displayed substantial migratory abilities in GS probably because of AMPK activation, as also shown. in other HCC cells (Ferretti et al. 2019).
Conclusion
Taken together, we presented the effect of graded deprivation of major nutrients and their extent of interdependence in affecting growth, proliferation, redox homeostasis, mitochondrial function, autophagy and cell migration characteristics of MDA-MB-231, HepG2, and HeLa cancer cells (Scheme 1). Differential nutritional stress-associated changes in these cells probably contributed to modulation of multiple metabolic pathways and cellular signaling related to cell survival, proliferation and death. This led to metabolic adaptations or cell death, depending on the nature of the deprivation and capacity to meet the cells’ energy, biosynthetic and redox demands. Importantly, normal cells remained largely unaffected by nutrient deprivation used in the present study, emphasizing the significance of this study in designing cancer cell targeted strategies. A thorough understanding of these metabolic changes in cancer cells in response to nutrient substrate deprivation will aid in the development of more appropriate and specific metabolism-based antitumor approaches for the prevention, diagnosis, control, and treatment of cancer by leveraging cancer cells' type-specific metabolic differences. Furthermore, our findings provide rationale for further research into the effects of existing or potential anticancer drugs in combination with conditions having differentially depleted nutrient to gain detailed insights into the mechanism of their additive, synergistic, or antagonistic action, as well as to assess their toxicity towards target and normal tissue in an animal model in vivo.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information file.
References
Akins NS, Nielson TC, Le HV (2018) Inhibition of glycolysis and glutaminolysis: an emerging drug discovery approach to combat cancer. Curr Top Med Chem 18:494–504. https://doi.org/10.2174/1568026618666180523111351
Altman BJ, Stine ZE, Dang CV (2016) From Krebs to clinic: glutamine metabolism to cancer therapy. Nat Rev Cancer 16:619–634. https://doi.org/10.1038/nrc.2016.71
Azad MB, Chen Y, Gibson SB (2009) Regulation of autophagy by reactive oxygen species (ROS): implications for cancer progression and treatment. Antioxid Redox Signal 11:777–790. https://doi.org/10.1089/ars.2008.2270
Balestrieri C, Vanoni M, Hautaniemi S et al (2012) Integrative transcriptional analysis between human and mouse cancer cells provides a common set of transformation associated genes. Biotechnol Adv 30:16–29. https://doi.org/10.1016/j.biotechadv.2011.06.013
Cantor JR, Sabatini DM (2012) Cancer cell metabolism: one hallmark, many faces. Cancer Discov 2:881–898. https://doi.org/10.1158/2159-8290.CD-12-0345
Cetinbas NM, Sudderth J, Harris RC et al (2016) Glucose-dependent anaplerosis in cancer cells is required for cellular redox balance in the absence of glutamine. Sci Rep 6:32606. https://doi.org/10.1038/srep32606
Chiaradonna F, Sacco E, Manzoni R et al (2006) Ras-dependent carbon metabolism and transformation in mouse fibroblasts. Oncogene 25:5391–5404. https://doi.org/10.1038/sj.onc.1209528
Chua P-J, Yip GW-C, Bay B-H (2009) Cell cycle arrest induced by hydrogen peroxide is associated with modulation of oxidative stress related genes in breast cancer cells. Exp Biol Med 234:1086–1094. https://doi.org/10.3181/0903-RM-98
Cluntun AA, Lukey MJ, Cerione RA, Locasale JW (2017) Glutamine metabolism in cancer: understanding the heterogeneity. Trends Cancer 3:169–180. https://doi.org/10.1016/j.trecan.2017.01.005
Colombo SL, Palacios-Callender M, Frakich N et al (2011) Molecular basis for the differential use of glucose and glutamine in cell proliferation as revealed by synchronized HeLa cells. Proc Natl Acad Sci 108:21069–21074. https://doi.org/10.1073/pnas.1117500108
Daye D, Wellen KE (2012) Metabolic reprogramming in cancer: unraveling the role of glutamine in tumorigenesis. Semin Cell Dev Biol 23:362–369. https://doi.org/10.1016/j.semcdb.2012.02.002
Efferth (2009) Molecular principles of cancer invasion and metastasis (Review). Int J Oncol. https://doi.org/10.3892/ijo_00000214
Elf SE, Chen J (2014) Targeting glucose metabolism in patients with cancer. Cancer 120:774–780. https://doi.org/10.1002/cncr.28501
Endo H, Owada S, Inagaki Y et al (2018) Glucose starvation induces LKB1-AMPK-mediated MMP-9 expression in cancer cells. Sci Rep 8:10122. https://doi.org/10.1038/s41598-018-28074-w
Ferretti AC, Hidalgo F, Tonucci FM et al (2019) Metformin and glucose starvation decrease the migratory ability of hepatocellular carcinoma cells: targeting AMPK activation to control migration. Sci Rep 9:1–14. https://doi.org/10.1038/s41598-019-39556-w
Gaglio D, Soldati C, Vanoni M et al (2009) Glutamine deprivation induces abortive S-phase rescued by deoxyribonucleotides in K-ras transformed fibroblasts. PLoS ONE 4:e4715. https://doi.org/10.1371/journal.pone.0004715
Gaglio D, Metallo CM, Gameiro PA et al (2011) Oncogenic K-Ras decouples glucose and glutamine metabolism to support cancer cell growth. Mol Syst Biol 7:523. https://doi.org/10.1038/msb.2011.56
Gwangwa MV, Joubert AM, Visagie MH (2019) Effects of glutamine deprivation on oxidative stress and cell survival in breast cell lines. Biol Res 52:15. https://doi.org/10.1186/s40659-019-0224-9
Han G, Zhou Q (2016) Dimethylfumarate induces cell cycle arrest and apoptosis via regulating intracellular redox systems in HeLa cells. Vitr Cell Dev Biol Anim 52:1034–1041. https://doi.org/10.1007/s11626-016-0069-2
Han J, Zhang L, Guo H et al (2015) Glucose promotes cell proliferation, glucose uptake and invasion in endometrial cancer cells via AMPK/mTOR/S6 and MAPK signaling. Gynecol Oncol 138:668–675. https://doi.org/10.1016/j.ygyno.2015.06.036
Hayes JD, At D-K, Tew KD (2020) Oxidative stress in cancer. Cancer Cell 38:167–197. https://doi.org/10.1016/j.ccell.2020.06.001
Heller SR (2017) Glucose concentrations of less than 3.0 mmol/L (54 mg/dL) should be reported in clinical trials: a joint position statement of the American diabetes association and the European association for the study of diabetes. Diabetes Care 40:155–157. https://doi.org/10.2337/dc16-2215
Hernández-Reséndiz I, Román-Rosales A, García-Villa E et al (2015) Dual regulation of energy metabolism by p53 in human cervix and breast cancer cells. Biochim Biophys Acta Mol Cell Res 1853:3266–3278. https://doi.org/10.1016/j.bbamcr.2015.09.033
Hosios AM, Hecht VC, Danai LV et al (2016) Amino acids rather than glucose account for the majority of cell mass in proliferating mammalian cells. Dev Cell 36:540–549. https://doi.org/10.1016/j.devcel.2016.02.012
Hubackova S, Davidova E, Boukalova S et al (2020) Replication and ribosomal stress induced by targeting pyrimidine synthesis and cellular checkpoints suppress p53-deficient tumors. Cell Death Dis 11:110. https://doi.org/10.1038/s41419-020-2224-7
Isono T, Chano T, Kitamura A, Yuasa T (2014) Glucose deprivation induces G2/M transition-arrest and cell death in N-GlcNAc2-modified protein-producing renal carcinoma cells. PLoS ONE 9:e96168. https://doi.org/10.1371/journal.pone.0096168
Izdebska M, Hałas-Wiśniewska M, Zielińska W et al (2019) Lidocaine induces protective autophagy in rat C6 glioma cell line. Int J Oncol 54(3):1099–1111. https://doi.org/10.3892/ijo.2018.4668
Jingwen B, Yaochen L, Guojun Z (2017) Cell cycle regulation and anticancer drug discovery. Cancer Biol Med 14:348. https://doi.org/10.20892/j.issn.2095-3941.2017.0033
Kansara M, Berridge MV (2004) Oncogenes modulate cell sensitivity to apoptosis induced by glucose deprivation. Anticancer Res 24:2503–2510
Khatua A, Prasad A, Priyadarshini E et al (2020) Emerging antineoplastic plant-based gold nanoparticle synthesis: a mechanistic exploration of their anticancer activity toward cervical cancer cells. J Clust Sci 31:1329–1340. https://doi.org/10.1007/s10876-019-01742-1
Khatua A, Prasad A, Behuria HG et al (2022) Evaluation of antimicrobial, anticancer potential and Flippase induced leakage in model membrane of Centella asiatica fabricated MgONPs. Biomater Adv 138:212855. https://doi.org/10.1016/j.bioadv.2022.212855
Kim AJ, Kim H-J, Jee HJ et al (2011) Glucose deprivation is associated with Chk1 degradation through the ubiquitin–proteasome pathway and effective checkpoint response to replication blocks. Biochim Biophys Acta Mol Cell Res 1813:1230–1238. https://doi.org/10.1016/j.bbamcr.2011.03.012
Kim R-K, Suh Y, Yoo K-C et al (2015) Activation of KRAS promotes the mesenchymal features of basal-type breast cancer. Exp Mol Med 47:e137–e137. https://doi.org/10.1038/emm.2014.99
Kumar A, Giri S, Shaha C (2018) Sestrin2 facilitates glutamine-dependent transcription of PGC-1α and survival of liver cancer cells under glucose limitation. FEBS J 285:1326–1345. https://doi.org/10.1111/febs.14406
Lezaja A, Altmeyer M (2018) Inherited DNA lesions determine G1 duration in the next cell cycle. Cell Cycle 17:24–32. https://doi.org/10.1080/15384101.2017.1383578
Liang CC, Park AY, Guan JL (2007) In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc 2:329–333. https://doi.org/10.1038/nprot.2007.30
Lim SM, Mohamad Hanif EA, Chin SF (2021) Is targeting autophagy mechanism in cancer a good approach? The possible double-edge sword effect. Cell Biosci 11:56. https://doi.org/10.1186/s13578-021-00570-z
Liu W, Le A, Hancock C et al (2012) Reprogramming of proline and glutamine metabolism contributes to the proliferative and metabolic responses regulated by oncogenic transcription factor c-MYC. Proc Natl Acad Sci 109:8983–8988. https://doi.org/10.1073/pnas.1203244109
Lorin S, Hamaï A, Mehrpour M et al (2013) Autophagy regulation and its role in cancer. Semin Cancer Biol 23:361–379. https://doi.org/10.1016/j.semcancer.2013.06.007
Mathew R, Karp CM, Beaudoin B et al (2009) Autophagy suppresses tumorigenesis through elimination of p62. Cell 137:1062–1075. https://doi.org/10.1016/j.cell.2009.03.048
Michalak KP, Maćkowska-Kędziora A, Sobolewski B, Woźniak P (2015) Key roles of glutamine pathways in reprogramming the cancer metabolism. Oxid Med Cell Longev 2015:1–14. https://doi.org/10.1155/2015/964321
Mishra AK, Campanale JP, Mondo JA, Montell DJ (2019) Cell interactions in collective cell migration. Development 146:dev172056. https://doi.org/10.1242/dev.172056
Noguchi M, Hirata N, Tanaka T et al (2020) Autophagy as a modulator of cell death machinery. Cell Death Dis 11:517. https://doi.org/10.1038/s41419-020-2724-5
Nogueira V, Hay N (2013) Molecular pathways: reactive oxygen species homeostasis in cancer cells and implications for cancer therapy. Clin Cancer Res 19:4309–4314. https://doi.org/10.1158/1078-0432.CCR-12-1424
Palorini R, De Rasmo D, Gaviraghi M et al (2013) Oncogenic K-ras expression is associated with derangement of the cAMP/PKA pathway and forskolin-reversible alterations of mitochondrial dynamics and respiration. Oncogene 32:352–362. https://doi.org/10.1038/onc.2012.50
Pan Z, Zhang X, Yu P et al (2019) Cinobufagin induces cell cycle arrest at the G2/M phase and promotes apoptosis in malignant melanoma cells. Front Oncol. https://doi.org/10.3389/fonc.2019.00853
Plecitá-Hlavatá L, Lessard M, Šantorová J et al (2008) Mitochondrial oxidative phosphorylation and energetic status are reflected by morphology of mitochondrial network in INS-1E and HEP-G2 cells viewed by 4Pi microscopy. Biochim Biophys Acta Bioenerg 1777:834–846. https://doi.org/10.1016/j.bbabio.2008.04.002
Prasad A, Khatua A, Mohanta YK et al (2022) Low-dose exposure to phytosynthesized gold nanoparticles combined with glutamine deprivation enhances cell death in the cancer cell line HeLa via oxidative stress-mediated mitochondrial dysfunction and G0/G1 cell cycle arrest. Nanoscale 14:10399–10417. https://doi.org/10.1039/d2nr02150a
Reitzer LJ, Wice BM, Kennell D (1979) Evidence that glutamine, not sugar, is the major energy source for cultured HeLa cells. J Biol Chem 254:2669–2676. https://doi.org/10.1016/S0021-9258(17)30124-2
Rodríguez-Enríquez S, Carreño-Fuentes L, Gallardo-Pérez JC et al (2010) Oxidative phosphorylation is impaired by prolonged hypoxia in breast and possibly in cervix carcinoma. Int J Biochem Cell Biol 42:1744–1751. https://doi.org/10.1016/j.biocel.2010.07.010
Rossignol R, Gilkerson R, Aggeler R et al (2004) Energy substrate modulates mitochondrial structure and oxidative capacity in cancer cells. Cancer Res 64:985–993. https://doi.org/10.1158/0008-5472.CAN-03-1101
Roy AC, Prasad A, Ghosh I (2023a) Phytochemical profiling of Tupistra nutans Wall. ex Lindl. inflorescence extract and evaluation of its antioxidant activity and toxicity in hepatocarcinoma (HepG2) and fibroblast (F111) cells. Appl Biochem Biotechnol 195:172–195. https://doi.org/10.1007/s12010-022-04145-7
Roy AC, Prasad A, Priya K et al (2023b) Anticancer effect of antioxidant-rich methanolic extract of Rauvolfia serpentina (L.) Benth ex. Kurz leaves in HepG2 and HeLa cells: a mechanistic insight. Biocatal Agric Biotechnol 50:102674. https://doi.org/10.1016/j.bcab.2023.102674
Roy AC, Prasad A, Singh S et al (2023c) Comparative bioactivity analysis of hydro-methanolic extracts of root leaf and inflorescence of Tupistra nutans Wall. ex Lindl: An in vitro study on antioxidant and anti-hepatocellular carcinoma activity. Biocatal Agric Biotechnol 52:102819. https://doi.org/10.1016/j.bcab.2023.102819
Sandulache VC, Ow TJ, Pickering CR et al (2011) Glucose, not glutamine, is the dominant energy source required for proliferation and survival of head and neck squamous carcinoma cells. Cancer 117:2926–2938. https://doi.org/10.1002/cncr.25868
Santos JM, Hussain F (2020) Higher glucose enhances breast cancer cell aggressiveness. Nutr Cancer 72:734–746. https://doi.org/10.1080/01635581.2019.1654527
Scherz-Shouval R, Shvets E, Fass E et al (2007) Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J 26:1749–1760. https://doi.org/10.1038/sj.emboj.7601623
Shackelford DB, Shaw RJ (2009) The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nat Rev Cancer 9:563–575. https://doi.org/10.1038/nrc2676
Shapiro GI, Harper JW (1999) Anticancer drug targets: cell cycle and checkpoint control. J Clin Invest 104:1645–1653. https://doi.org/10.1172/JCI9054
Singh G, Lakkis CL, Laucirica R, Epner DE (1999) Regulation of prostate cancer cell division by glucose. J Cell Physiol 180:431–438. https://doi.org/10.1002/(SICI)1097-4652(199909)180:3%3c431::AID-JCP14%3e3.0.CO;2-O
Son J, Lyssiotis CA, Ying H et al (2013) Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496:101–105. https://doi.org/10.1038/nature12040
Sullivan LB, Chanel NS (2014) Mitochondrial reactive oxygen species and cancer. Cancer Metab 2:17. https://doi.org/10.1186/2049-3002-2-17
Sun L, Yin Y, Clark LH et al (2017) Dual inhibition of glycolysis and glutaminolysis as a therapeutic strategy in the treatment of ovarian cancer. Oncotarget 8:63551–63561. https://doi.org/10.18632/oncotarget.18854
Sun N, Liang Y, Chen Y et al (2019) Glutamine affects T24 bladder cancer cell proliferation by activating STAT3 through ROS and glutaminolysis. Int J Mol Med. https://doi.org/10.3892/ijmm.2019.4385
Sung H, Ferlay J, Siegel RL et al (2021) Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Wang H, Zhang T, Sun W et al (2016) Erianin induces G2/M-phase arrest, apoptosis, and autophagy via the ROS/JNK signaling pathway in human osteosarcoma cells in vitro and in vivo. Cell Death Dis 7:e2247–e2247. https://doi.org/10.1038/cddis.2016.138
Wenzel ES, Singh ATK (2018) Cell-cycle checkpoints and aneuploidy on the path to cancer. In Vivo (brooklyn) 32:1–5. https://doi.org/10.21873/invivo.11197
Wu S, Yin X, Fang X et al (2015) c-MYC responds to glucose deprivation in a cell-type-dependent manner. Cell Death Discov 1:15057. https://doi.org/10.1038/cddiscovery.2015.57
Xie N, Zhang L, Gao W et al (2020) NAD+ metabolism: pathophysiologic mechanisms and therapeutic potential. Signal Transduct Target Ther 5(1):227. https://doi.org/10.1038/s41392-020-00311-7
Yang Y, Sauve AA (2016) NAD(+) metabolism: bioenergetics, signaling and manipulation for therapy. Biochim Biophys Acta 1864:1787–1800. https://doi.org/10.1016/j.bbapap.2016.06.014
Yang L, Venneti S, Nagrath D (2017) Glutaminolysis: a hallmark of cancer metabolism. Annu Rev Biomed Eng 19:163–194. https://doi.org/10.1146/annurev-bioeng-071516-044546
Yi X, Zhang X, Jeong H et al (2015) A novel bispidinone analog induces S-phase cell cycle arrest and apoptosis in HeLa human cervical carcinoma cells. Oncol Rep 33:1526–1532. https://doi.org/10.3892/or.2015.3722
Yuneva M, Zamboni N, Oefner P et al (2007) Deficiency in glutamine but not glucose induces MYC-dependent apoptosis in human cells. J Cell Biol 178:93–105. https://doi.org/10.1083/jcb.200703099
Zhu J, Chen M, Chen N et al (2015) Glycyrrhetinic acid induces G1-phase cell cycle arrest in human non-small cell lung cancer cells through endoplasmic reticulum stress pathway. Int J Oncol 46:981–988. https://doi.org/10.3892/ijo.2015.2819
Acknowledgements
A.P acknowledges the Indian Council of Medical Research (ICMR), Government of India for providing Senior Research Fellowship. A.C.R. received Junior Research Fellowship from the Department of Biotechnology (DBT), Government of India. We are thankful to CIF, SES, Jawaharlal Nehru University for providing flow cytometry facilities.
Funding
I.G received funding from DST-PURSE (Promotion of University Research and Scientific Excellence program of Department of Science and Technology, Government of India) and UGC UPE-II (University with Potential for Excellence, Phase II scheme of University Grant Commission, Government of India) via Jawaharlal Nehru University, New Delhi, India. The funding agencies had no role in study design, data acquisition, analysis, the decision to publish, and preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.P: Conceptualization, Methodology, Investigation, Writing–original draft, Writing–Review and Editing, Validation, Formal analysis, Visualization, Software. A.C.R: Methodology, Investigation, Writing–Review and Editing, Validation, Formal analysis. K. P: Methodology, Investigation, Validation. R.M: Investigation, Writing–Review and Editing, Formal analysis. I.G: Conceptualization, Writing–review and editing, Supervision, Project administration, Funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Financial interests
The authors declare they have no financial interests.
Non-financial interests
None.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Prasad, A., Roy, A.C., Priya, K. et al. Effect of differential deprivation of nutrients on cellular proliferation, oxidative stress, mitochondrial function, and cell migration in MDA-MB-231, HepG2, and HeLa cells. 3 Biotech 13, 339 (2023). https://doi.org/10.1007/s13205-023-03759-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-023-03759-w