Abstract
Xanthomonas oryzae pv. oryzae (Xoo) is a destructive pathogen that causes bacterial blight disease of rice worldwide. Xoo uses T3SS (type III secretion system) effectors to subvert rice innate immunity. However, the comprehensive knowledge of rice genes involved in T3SS effectors-mediated interaction remains unclear. In this study, the transcriptome profiles of rice infected with a virulent Xoo strain from North-eastern region of India relatives to its avirulent strain (that lacks functional T3SS) were analyzed at early (2–6 hpi) and late (16–24 hpi) hours of infection. Out of total 255 differentially expressed genes (DEGs), during early infection, 62 and 70 genes were upregulated and downregulated, respectively. At late infection, 70 and 53 genes were upregulated and downregulated, respectively. The transcriptomic data identified many differentially expressed resistant genes, transposons, transcription factors, serine/threonine protein kinase, cytochrome P450 and peroxidase genes that are involved in plant defense. Pathway analysis revealed that these DEGs are involved in hormone signaling, plant defense, cellular metabolism, growth and development processes. DEGs associated with plant defense were also validated through quantitative real-time PCR. Our study brings a comprehensive picture of the rice genes that are being differentially expressed during bacterial blight infection. Nevertheless, the DEG-associated pathways would provide sensible targets for developing resistance to bacterial blight.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Xanthomonas oryzae pv. oryzae (Xoo), the causal agent of bacterial blight (BB), limits rice production across the globe (Mondal 2016). Under favorable environmental conditions such as humidity, warm and wet climate, Xoo can produce crop loss up to 70–100%. Xoo translocates the immune-suppressive effectors via type III secretion system (T3SS) to suppress the host immunity and to draw nutrients for its growth (White and Yang 2009; White et al. 2009). T3SS is, thus, a crucial component of Xoo pathogenesis but is dispensable for Xoo survival (Mondal et al. 2014). T3SS-secreted effectors are of two types: transcription activator-like effectors (TALEs) and Xanthomonas outer proteins (Xops). The interaction between the effector and host cell finally determines the pathogenicity (Mondal et al. 2016; Verma et al. 2018). TALEs possess the nucleotide-binding activity that targets the promoters of susceptible genes to induce pathogenicity (Cox et al. 2017; Yang et al. 2006). However, during evolution, plants developed alternative defence strategies by acquiring the mutation in TALE-binding site that leads to inhibition of Xoo-induced pathogenicity (Hutin et al. 2015). Additionally, plants also acquired new R-genes or E genes (newly classified as executor gene) whose activation is dependent on the pathogen’s TALE and results in the onset of defense responses like HR-induced cell death (Tian et al. 2014).
Despite several attempts to curb the bacterial blight disease via bactericides, none of the chemicals was highly effective. Genetic screening of resistant varieties and breeding approaches so far remained an effective method of BB disease management. The expression of these T3SS effectors was shown to be regulated by hrpX, a key regulator of hrp (hypersensitive response and pathogenicity gene) cluster (Mondal et al. 2016). Exploiting the host plant resistance via R-genes remains an effective means to control BB of rice. Out of 44 Xa genes identified in rice, 8 were cloned and characterized (Cheema et al. 2008; Kim et al. 2015). Mostly the pyramided rice lines with 2–4 genes combinations like xa13 + Xa21, xa5 + xa13 + Xa21 and Xa4 + xa5 + xa13 + Xa21 were found effective in providing resistance over the monogenic lines (Ellur et al. 2016b). However, Xa38 alone gave comparable resistance as found in the combination of xa13 + Xa21 (Ellur et al. 2016a). Besides resistance genes, an insight on rice transcripts that are involved, particularly in T3SS effectors-mediated interaction, is pertinent to have a comprehensive understanding of the resistant phenomenon. Xoo strains from north-western regions of India were widely studied for type III effectors (Mondal et al. 2014). Race 4, a virulent member from north-western Indian contains 21 Xop and 18 TAL T3SS effectors and its profile is almost similar to that of other Asian Xoo strains from Korea, Japanese and Philippines (Mondal et al. 2014). The key T3SS effectors of Xoo race 4, namely XopF and XopR that contribute to the bacterial virulence as well as to the disease production ability were deciphered (Mondal et al. 2016; Verma et al. 2018, 2019). However, Xoo strains from North-eastern regions, wherein rice is grown and eaten extensively, are yet to be studied. In this context, the present study aims to decipher the rice transcripts upon infection with a virulent North-eastern (NE) strain of Xoo (Assam isolate) relative to its avirulent strain (a functionally impaired T3SS mutant of NE strain). More precisely, our interest is to identify the major transcripts associated with plant defense against Xoo as well as to understand the pathways regulated by those transcripts. This detailed understanding of the rice::Xoo interaction would lead to the scope of exploring new targets for disease management.
Materials and methods
Plant material and Xoo inoculation
The rice variety Pusa basmati 1 (a popular variety from North-west India) with susceptibility to Xoo was used in this study. Race 2 of Xoo from Assam was used for inoculation. A Xoo mutant lacking hrpX (Xoo ΔhrpX) was constructed using a PCR based homologous recombination strategy (Verma et al. 2018). For Xoo inoculation, rice seedlings were grown in a glasshouse at 28 °C/25 °C (light/dark), with a 14 h photoperiod and > 85% relative humidity. The Xoo growth on YGCA (yeast glucose calcium-carbonate agar) medium at 28 °C for 48–72 h was harvested in 10 mM MgSO4·7H2O and the final concentration of the inoculum was adjusted to 108 CFU ml−1 by adding sterile water. Leaves of five-week-old rice plants were inoculated with the bacterial suspension using the needleless syringe-infiltration technique. The inoculated plants were misted with water twice a day to maintain high humidity and prolong leaf wetness for disease development.
RNA isolation, RNA-seq library preparation and sequencing
For total RNA isolation, the leaf samples were collected at 2, 6, 16 and 24 h of post-inoculation (hpi) using the TRIzol reagent (Invitrogen, USA) following the manufacturer’s protocol. Later, the RNA samples of 2 and 6 hpi were pooled together to use as early response while the samples of 16 and 24 hpi were pooled for use as a late response. Two independent replications were kept for each sample. The RNA concentration was quantified by NanoDrop 2000 Spectrophotometer (Thermo Fisher Scientific, USA), and the RNA integrity value (RIN) was evaluated by RNA 6000 Pico LabChip of Agilent 2100 Bioanalyzer (Agilent, USA). Eight libraries (Xoo wild and Xoo ∆hrpX treatment at 2–6 hpi and 16–24 hpi with two sets of replications) were constructed for RNA-seq analysis. Total RNA (200 μg) was incubated with DNase I (10 units) at 37 °C for 1 h, and the final sample volume was adjusted to 250 μl by adding nuclease-free water. Illumina Hiseq2500 platform was used for library preparation and RNA sequencing.
Preprocessing and rRNA removal
Raw reads were cleaned by removing adaptor sequences and unreliable low-quality bases using AdapterRemoval-v2 (version 2.2.0). rRNA sequences were removed from processed reads by aligning them with the silva database using bowtie2 (version 0.6.5), BamUtil (version 1.0.13) tools and subsequent workflow using sam-tools (version 0.1.19), sambamba (version 0.6.5), BamUtil (version 1.0.13) tools and in-house scripts.
Identification of differentially expressed genes
The filtered clean (pre-processed and rRNA removed) reads were aligned to the rice reference genome sequences available at MSU rice genome annotation project (http://rice.uga.edu/). STAR (version 2.5.3a) program was used for sequence alignment. The aligned reads were used for estimating expression of the genes and transcripts using cufflinks program (version 2.2.1). Differentially expressed genes (DEGs) were identified by cuffdiff program of cufflinks package (Supplementary Tables 1–4).
Gene ontology and pathway analysis
The gene ontology (GO) analysis of DEGs based on biological process, molecular function and cellular component were analyzed using Rice Net DB (Liu et al. 2013). The DEGs associated with various metabolic pathways were identified by Plant Reactome (Naithani et al. 2020). The transcription factors, peroxidase and resistant genes were identified by RiceFREND (Sato et al. 2013).
Quantitative real-time PCR analysis
A total of 14 genes involved in plant stress response were selected based on transcriptome data. The relative expression of these genes was performed using quantitative real-time PCR (qRT-PCR). The cDNA was synthesized using the same RNA templates as used in transcriptome analysis. The specific primer pair for each gene was synthesized (Supplementary table 5) and the relative expression experiment was performed using SYBR premix Ex Taq (TAKARA) in CFX96 RT-qPCR instrument (Bio-Rad, USA). The specific amplification of each gene was analyzed using a melting curve and by loading the PCR product on an agarose gel. The rice actin was taken as a reference gene. Three biological replicates were used for each sample and the real-time data was analyzed using 2−ΔΔCt method (Livak and Schmittgen 2001).
Results
Analysis of RNA-seq libraries and reads alignment
We used high-throughput RNA-sequencing to identify the DEGs in rice at early (2–6 hpi) and late phases (16–24 hpi) of Xoo or Xoo ΔhrpX infected plants. A total of 415,152,266 reads were obtained with a GC content between 48.57 and 50.53% and an average of 90.33% reads passed > = 30 phred score. After rRNA removal, the reads counts were 413,639,700. The raw reads were ranged from 49,063,634 to 55,255,308 and the filtered reads were ranged from 48,621,744 to 55,101,930. The 95.21–97.07% of the processed reads were aligned to the rice reference genome (http://rice.uga.edu) that showed a read alignment count between 46,637,313 and 53,096,533. The raw sequencing data of Xoo and Xoo ΔhrpX inoculated leaves at early and late bacterial infection were deposited into NCBI Sequence Read Archive (SRA) (Accession no. PRJNA591420).
Rice DEGs upon infection with Xoo and Xoo ∆hrpX mutant strains
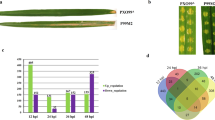
The transcriptome profiles of rice, infiltrated with wild Xoo (referred to as virulent having functional T3SS) and ∆hrpX mutant (referred to as avirulent having non-functional T3SS) were compared in early and late bacterial infection using Xoo ∆hrpX profile as a control. A total of 255 DEGs were identified in rice plants infected with NE Xoo strain relative to its Xoo ∆hrpX mutant strain. The number of DEGs in early bacterial infection (132 genes) was higher than in late infection (123 genes). In an overview, 62 genes were upregulated and 70 genes were downregulated during the early phase of infection. In late infection, 70 genes were upregulated and 53 genes were downregulated (Fig. 1a). Further, the Venn diagram exposed some common genes in both early and late bacterial infection (Fig. 1b). One overlapping upregulated gene (LOC_Os03g14654) encoding for LTPL108-protease inhibitor/seed storage/LTP family protein was identified in both early and late bacterial infection. Likewise, one overlapping downregulated gene (LOC_Os05g28830) was also identified in both early and late bacterial infection. It encodes PMR5 protein that was previously reported to provide resistance against Magnaporthe oryzae in Arabidopsis thaliana (Maeda et al. 2009). One gene (LOC_Os05g45380) is upregulated in early infection but downregulated in late infection. It encodes a C-terminally encoded peptide 9 (CEP9). These small peptides are important for cellular signaling, growth and defense responses. We also identified two genes that are downregulated in early infection but upregulate in late infection. These genes code for a hypothetical protein (LOC_Os02g17590) and an NB-ARC domain resistance protein (LOC_Os02g18070). Most resistance proteins have a central nucleotide-binding (NB-ARC) domain. These proteins are involved in pathogen recognition and subsequent activation of defense responses (Van Ooijen et al. 2008). The transcriptome analysis revealed that DEGs were upregulated 2–4 times in the early stages of infection. Non-symbiotic hemoglobin 2 (LOC_Os03g12510), aminotransferase (LOC_Os04g52440), and protease inhibitor/seed storage/LTP family protein precursors; LTPL108 (LOC_Os03g14654), LTPL153 (LOC_Os05g47730), and LTPL157 (LOC_Os10g36100) were the most significantly upregulated genes. The downregulated genes showed 2–3.7-fold drop in their expression. The significantly downregulated genes were terpene synthase (LOC_Os02g02930), NB-ARC domain protein (LOC_Os02g18070), SAM-dependent carboxyl methyltransferase (LOC_Os02g48770), lipoxygenase (LOC_Os03g52860), peptide transporter PTR2 (LOC_Os04g50940) and laccase precursor (LOC_Os11g42200). In late infection, DEGs showed 2–4.27-fold upregulation relative to Xoo ∆hrpX inoculation. The significantly upregulated genes were peroxidase precursor (LOC_Os01g19020), expansin precursor (LOC_Os01g60770), NB-ARC domain protein (LOC_Os02g18070), GDSL-like lipase/acylhydrolase (LOC_Os06g06250), beta-galactosidase precursor (LOC_Os06g37560), RGH1A (LOC_Os08g07330) and many uncharacterized genes (LOC_Os01g42520, LOC_Os08g28784, LOC_Os11g31770). The significantly downregulated genes include receptor-like protein kinase 5 precursor (LOC_Os02g40180), nicotianamine synthase (LOC_Os03g19427), NB-ARC domain protein (LOC_Os04g53120), NBS-LRR disease resistance protein (LOC_Os11g15670), MLA12 (LOC_Os12g17480), white-brown complex homolog protein 11 (LOC_Os12g22284) and many uncharacterized genes (LOC_Os01g32270, LOC_Os01g58670, LOC_Os02g33335, LOC_Os03g54240, LOC_Os06g05440, LOC_Os11g15624, LOC_Os12g20390). In addition, many genes showed unaltered expression in both early and late bacterial infection as shown by differential expression FPKM plots (Fig. 1c). Further, we have also identified many DEGs isoforms in both early and late bacterial infection (Table 1). An isoform of WRKY11 (LOC_Os01g43650.2) was upregulated 2.08529-fold in early infection. Ectopic expression of rice WRKY11 (OsWRKY11) was shown to enhance Xoo resistance against Xoo and tolerance to drought stress. It activates many defense-responsive genes to combat pathogen attack (Lee et al. 2018). In particular, two isoforms of an uncharacterized gene (LOC_Os12g18410.1 and LOC_Os12g18410.2) was identified in early upregulated genes suggesting the role of this gene in plant immunity against Xoo invasion. However, further work is needed to elaborate its function in plant defense.
Graph showing the total number of DEGs at early and late bacterial infection relative to its T3SS-defective avirulent strain (a). VENN diagram showing the overlapping DEGs in early and late Xoo infection relative to its T3SS-defective avirulent strain (b). FPKM plots showing upregulated, downregulated and non-differential expression of differentially expressed genes at early and late bacterial infections (c)
We also identified 37 genes (14 in early and 23 in late infection), which were specifically expressed in wild Xoo infection relative to Xoo ∆hrpX inoculation (Table 2). Most of these genes are uncharacterized. Likewise, we also got 23 genes (11 in early and 13 in late infection) that were not expressed in wild Xoo infection relative to Xoo ∆hrpX infection (Table 3). Most of these genes are also uncharacterized. These genes may play an important role during Xoo pathogenesis. However, further confirmation of their precise role during rice and Xoo interaction is required using wet-lab experimentations.
Identification of resistant genes, peroxidase and transcription factors
In both early and late bacterial infections, many resistant genes, transcription factors and peroxidase genes were identified. In early infection, cinnamoyl CoA reductase (LOC_Os01g18120) is upregulated 2.09-fold. It is involved in lignin deposition in plant cell wall. The expression of this gene is increased after both abiotic and biotic stresses involving infection by Xoo and Magnaporthe grisea (Park et al. 2017). Other upregulated genes are cytochrome P450 (LOC_Os08g39730) and non-symbiotic hemoglobin 2 (LOC_Os03g12510). The upregulated transcription factors belong to ethylene-responsive transcription factor TINY (LOC_Os02g13710) and bZIP (LOC_Os06g50600) families. The downregulated genes important for defense response were NB-ARC domain proteins (LOC_Os02g18070, LOC_Os02g18080), cytochrome p450 (LOC_Os03g12500, LOC_Os03g40600), chalcone and stilbene synthases (LOC_Os07g34260), peroxidase (LOC_Os07g48060), disease resistance RPP13-like protein 1 (LOC_Os08g43010) and beta-expansin (LOC_Os09g29710). Notably, we also identified many downregulated wall-associated kinase (OsWAK6, OsWAK81, OsWAK112d and OsWAK124) in early infection. These kinases are important for plant resistance against both fungal and bacterial pathogens. The rice line expressing OsWAK25 showed enhanced resistance against Xoo and Magnaporthe oryzae pathogens (Harkenrider et al. 2016). In addition, overexpression of CsWAK08 showed enhanced resistance against Xanthomonas citri subsp. citri in Citrus sinensis probably by ROS and jasmonic acid-mediated signaling (Li et al. 2020). In late infection, many upregulated defense-related genes including peroxidases (LOC_Os01g19020, LOC_Os01g28030, LOC_Os10g02040), NBS-LRR disease resistance proteins (LOC_Os02g02670, LOC_Os11g29520), NB-ARC domain-containing protein (LOC_Os02g18070), NB-ARC/LRR disease resistance protein (LOC_Os11g29090) were identified. In addition, two upregulated disease resistance gene analogs (RGAs), RGA1A (LOC_Os12g33160, LOC_Os08g07330) were also identified that may play important role in plant defense against Xoo invasion (Sekhwal et al. 2015). The upregulated TFs belong to MYB family (LOC_Os01g47370, LOC_Os12g33950). Many downregulated TFs were also identified in late bacterial infection. They belong to helix–loop–helix DNA-binding domain protein (LOC_Os01g09930, LOC_Os03g55550) and B3 DNA-binding domain protein (LOC_Os03g06850) families. The downregulated defense-related genes were receptor-like protein kinase 5 precursor (LOC_Os02g40180), NB-ARC domain-containing protein (LOC_Os04g53120) and NBS-LRR disease resistance protein (LOC_Os11g15670). The data suggest that Xoo effectors modulate many defense-related genes. However, further research is required to decipher the function of these genes in rice defense against BB disease.
Functional classification of rice DEGs
DEGs were functionally classified by GO analysis based on the biological process, cellular component and molecular function. In comparison to Xoo ∆hrpX mutant, the wild Xoo expressed a total of 72 DEGs which were classified into 17 functional groups during the early infection. In molecular function, the majority of DEGs belong to physiological processes, metabolism, and cellular processes. In the cellular component section, the cell was the largest group followed by an external encapsulating structure (7), cell wall (7) and extracellular region (6). An early phase of infection, induced several DEGs predominantly related to the catalytic activity (39) and binding (33) followed by transferase activity (18), transferring phosphorous-containing groups (10), kinase activity (10) and oxygen binding (5) (Fig. 2a). In the late phase of infection, majority of the DEGs corresponds to physiological process (44) and metabolism (34) in molecular function. Among the cellular component section, majority of the DEGs belong to cell (37 genes) and rest belongs to the cellular components like envelope, organelle envelope, endomembrane system and nuclear envelope (3 genes for each category). In biological processes, catalytic activity and binding constitute the largest group followed by hydrolase activity (Fig. 2b). DEGs were categorized into ten functional groups based on subcellular localisation. Genes related to chloroplast, cytosol, extracellular, mitochondria and nucleus were deferentially expressed in high numbers in both early and late phases of infection; whereas, genes belonging to peroxisome, vacuole and golgi body are expressed in less number (Fig. 3). In overview, Xoo type III effectors had a strong influence on the expression of rice genes that are localized in various organelles and are associated with biological processes leading to stress recognition/responses for pathogen-induced susceptibility or resistance.
Metabolic pathway analysis of DEGs
The tight regulation of metabolic processes and energy transactions is required for plant growth and defense against diverse pathogens (Huot et al. 2014). Xoo early infection caused differential expression of genes involved in carbon metabolism, lipid metabolism, ethylene biosynthesis and signaling, protein/amino acid metabolism, translation elongation and termination, seed development, transport and reproductive structure development. In early infection, 4 genes out of 132 were associated with different pathways. The gene coding for glucose-1-phosphate adenylyltransferase large subunit (LOC_Os05g50380) is upregulated (2.30551-fold) in early infection. It is linked to starch biosynthesis (Fig. 4a) and converts alpha-D-glucose 1-phosphate into ADP-D-glucose in the cell cytoplasm. The data suggest that T3SS effectors enhance starch biosynthesis for uninterrupted Xoo growth in rice tissues. The other upregulated gene is 60S acidic ribosomal protein P0 (LOC_Os12g03880). This protein is essential for translation initiation, elongation and termination. It is upregulated 2.00405-fold suggesting that during infection, T3SS effectors increase protein synthesis in rice cells. Two genes were downregulated in early infection. One gene, aminocyclopropane carboxylate synthetase (LOC_Os04g48850) belongs to the methionine salvage pathway. It is downregulated 2.06887-fold. Besides, it is also involved in the ethylene biosynthesis from methionine via catalyzing the formation of 1-aminocyclopropane-1-carboxylate which is an essential precursor for ethylene biosynthesis in higher plants (Fig. 4b). This enzyme uses pyridoxal phosphate (PPi) as a cofactor. Ethylene governs many cellular signaling and is also involved in the activation of many genes. This study demonstrates that Xoo T3SS effectors may suppress ethylene biosynthesis to induce blight disease. The other downregulated gene is phospholipase D (LOC_Os06g40170). It is downregulated 2.18836-fold and is linked to choline biosynthesis III (Fig. 4c). It catalyzes the hydrolysis of phosphatidylcholine (PC) to choline (Cho) and phosphatidic acid (PA) at the endoplasmic reticulum (ER) membrane. In late infection, 4 genes out of 123 were found associated with different pathways. One gene ent-kaurene synthase (LOC_Os04g10060) is present in the plastid. It is linked to momilactone biosynthesis present in stroma of plastids and catalyses the conversion of syn-copalyl diphosphate into 9 beta-pimara-7,15-diene in plastid stroma (Fig. 5a). It is upregulated 2.39106-fold relative to Xoo ΔhrpX. Momilactone A and B are plant defensive compounds and exhibit antibacterial and antifungal activities (Okada et al. 2007). Another gene (LOC_Os08g14760: AMP-binding domain-containing protein) is linked to suberin biosynthesis from L-Phenylalanine. It is upregulated 2.4593-fold in wild Xoo infection relative to Xoo ΔhrpX. In the pathway, it catalyzes the conversion of caffeate to caffeoyl-CoA and ferulate to feruloyl-CoA (Fig. 5b). Suberin deposition provides plant protection against pathogens and the data suggest that T3SS effectors enhance suberin deposition. Nicotianamine synthase2 (NAS2; LOC_Os03g19420) is downregulated to 2.31735-fold and is linked to mugineic acid biosynthesis (Fig. 5c). Mugineic acid is an amino acid that is secreted under iron deficiency for iron solubilization. Another gene (LOC_Os11g44600) coding for calmodulin-binding protein (CBP60g) is downregulated to 2.8662-fold. It is linked to salicylic acid signaling and is involved in the formation of activated CaM/CBP60g/SARD1 complex in the nucleoplasm. Salicylic acid is a phytohormone and has diverse roles. It is involved in plant growth, photosynthesis, transpiration and iron uptake. Besides, it is also involved in plant defense by inducing systemic acquired resistance and expression of pathogenesis-related proteins. The data suggest that Xoo T3SS effectors suppress salicylic acid signaling to overcome plant defense.
Validation of major rice transcripts through RT-qPCR
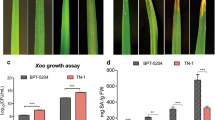
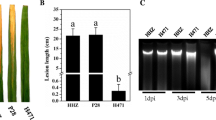
Validation of the rice transcripts as obtained from RNA-seq data was carried out through RT-qPCR of 14 selected DEGs. The sequences of these genes were retrieved from MSU rice genome annotation project (http://rice.uga.edu). We found that the genes like non-symbiotic hemoglobin 2 (LOC_Os03g12510) and cinnamoyl COA reductase (LOC_Os01g18120) were upregulated 2.7- and 3.12-fold, respectively, in early Xoo infection relative to Xoo ∆hrpX mutant. The downregulated genes were beta-expansin precursor (LOC_Os09g29710), chalcone and stilbene synthases (LOC_Os07g34260), peroxidase precursor (LOC_Os07g48060), SAM-dependent carboxyl methyltransferase (LOC_Os02g48770), decarboxylase (LOC_Os08g04540) and laccase precursor (LOC_Os01g62600) (Fig. 6a). At late hours of bacterial infection, few stresses responsive genes like NB-ARC domain-containing protein (LOC_Os02g18070) and RGH1A (LOC_Os08g07330 and LOC_Os12g33160) showed 2.87- to 4-fold upregulation. A significant fold of downregulation of BURP domain-containing protein (LOC_Os05g12410), white–brown complex homolog protein 11 (LOC_Os12g22284) and MLA12 (LOC_Os12g17480) were also noted upon challenged with Xoo over Xoo ∆hrpX mutant (Fig. 6b). The RT-qPCR results were also consistent with the data obtained from transcriptome analysis. Our data suggest that T3SS-dependent Xoo virulence led to the modulation of disease-resistant gene expression from the very beginning of the interaction. The resultant suppression of disease-resistant genes during interaction leads to the unrestricted spread of Xoo into rice plants.
Discussion
In this study, we compared the rice transcriptome upon infection with virulent Xoo compared to its avirulent counterpart (a functionally retarded T3SS mutant of wild Xoo race2) at both early and late bacterial infection. We have shown that out of a total of 255 DEGs, 62 and 70 genes were up-and downregulated at early hours of infection, respectively. At late hours of infection, 70 and 53 genes were up- and downregulated, respectively. Further, VENN diagram revealed some overlapping gene in both early and late bacterial infection. We also analyzed the transcriptome data using GO analysis that showed many DEGs in various activities like physiological processes, metabolism and stress responses. The subcellular localization indicated the DEGs are present in various cell parts including chloroplast, endoplasmic reticulum, plasma membrane etc. Our data showed that upon infection, Xoo targets diverse metabolic pathways of plant, mainly for the two broad motives. First, the bacterium targets those pathways that are associated with nutrient-/energy-supply like starch, choline, mugineic acid biosynthesis to get uninterrupted nutrient supply for its growth and multiplication. Concurrently, Xoo craves to destabilize the plant defense response by interfering with pathways associated with defense like suberin, momilactone, methionine salvage, ethylene and salicylic acid signaling. The analysis showed that in early infection, Xoo T3SS effectors enhance expression of glucose-1-phosphate adenylyltransferase large subunit which is linked to starch biosynthesis. Starch is an essential stored nutrient in plants and pathogen usually target to utilize this available food material for their carbon source. Xoo is a vascular pathogen that spreads systemically through the xylem tissue. Xylem is poor in essential nutrients that could support Xoo growth. To enrich nutrient abundance in the xylem vessels, Xoo targets plant food storage including starch biosynthesis. There are several reports wherein the Xoo stimulated rice gene associated with starch biosynthesis that possibly alters the source–sink relationships of the infected tissue, resulting in enhanced nutritional conditions (Yang et al. 2006). The continued growth of Xoo in the invaded plant tissues is ascertained by the defunct defense responses. The defense pathways that are the common targets by the rice pathogens include phytoalexin biosynthesis (like momilactone), suberin (cell-wall-associated defence polymers), ethylene, salicylic acid biosynthesis. In our study, we evident the modulation of genes associated with these pathways.
We evident several rice genes that contribute to the disease resistance are suppressed upon infection with wild Xoo (race 2 from Assam). Peroxidases are responsible for plant defences against various pathogens. Notably, the peroxidase proteins do participate in the thickening of the cell wall via deposition of lignin and suberin, metabolism of reactive oxygen and reactive nitrogen species, hypersensitive response and programmed cell death (Almagro et al. 2009). We found significant downregulation of peroxidase precursor, at early bacterial infection. This suggests that peroxidase genes are the main target by the Xoo and its suppression favors Xoo to multiply into the rice plant. The modulated expression of peroxidase genes was reported by several researchers in rice upon infection with Xoo as well as blast fungus (Sasaki et al. 2004; Velazhahan et al. 2006). Several classes of R-genes are known, while the most abundant are those having a centrally located nucleotide-binding site (NBS) domain and a carboxy-terminus leucine-rich repeat (LRR) domain (Li et al. 2019; DeYoung and Innes 2006). We witnessed significant downregulation of disease-resistant genes having NBS-LRR, NBS-ARC domains during rice::Xoo interaction. The NBS-ARC domain is involved in the induction of hypersensitive response in plant upon infection with biotic stresses. The transcriptome profile of Eucalyptus grandis demonstrated the pathogen's target for R-genes having NBS-ARC domain family when challenged with a Chrysoporthe austroafricana (a fungal pathogen) and Leptocybe invasa (insect pest) (Christie et al. 2016). Rice resistance proteins of NBS-LRR family, namely RGA4, RGA5 are demonstrated to interact with Magnaporthe oryzae effector protein AVR1-CO39 (Cesari et al. 2013). The other downregulated disease resistance genes during early hours of infection include RPP13 and SAM-dependent carboxyl methyltransferase. The molecular characterization of the RPP13 locus in different accessions indicated that the locus comprises a single gene predicted to encode an NBS-LRR protein with an amino‐terminal LZ motif. The amino acid sequence also suggests that more of the LRR structure is available for interaction with target molecules than has previously been reported for other disease‐resistance genes (Bittner-Eddy et al. 2000). The RPP13 locus, mapped to the bottom arm of chromosome 3, was shown to confer resistance to five Perosnospera parasitica isolates, including Maks9, in the Arabidopsis accession Nd‐1 (Bittner-Eddy et al. 2000). Our data suggest that Xoo effectors target several disease-resistant genes to induce rice susceptibility against BB.
Altogether, the present study depicts a clear picture of the rice transcripts that are differentially expressed upon infection with a virulent Xoo strain from Assam relative to its avirulent member. The transcriptome data suggest that diverse physiological and biochemical pathways are targeted by Xoo. Comparative analysis of expressed genes at two distinct periods indicated the major DEGs associated with early and late hours of infection. We further verified the DEGs associated with disease resistance using quantitative real-time PCR. In conclusion, our study not only informs about the rice targets but also unravels the associated pathways that are modulated upon Xoo infection. This insight would be of immense significance for sensible manipulation towards developing resistance to bacterial blight.
Accession number
All the RNA-Seq data have been submitted to NCBI Sequence Read Archive under the accession number PRJNA591420.
References
Almagro L, Gómez Ros LV, Belchi-Navarro S, Bru R, Ros Barceló A, Pedreño MA (2009) Class III peroxidases in plant defence reactions. J Exp Bot 60:377–390. https://doi.org/10.1093/jxb/ern277
Bittner-Eddy PD, Crute IR, Holub EB, Betnon JL (2000) RPP13 is a simple locus in Arabidopsis thaliana for alleles that specify downy mildew resistance to different avirulence determinants in Peronospora parasitica. Plant J 21:177–188. https://doi.org/10.1046/j.1365-313x.2000.00664.x
Cesari S, Thilliez G, Ribot C, Chalvon V, Michel C, Jauneau A, Rivas S, Alaux L, Kanzaki H, Okuyama Y, Morel JB (2013) The rice resistance protein pair RGA4/RGA5 recognizes the Magnaporthe oryzae effectors AVR-Pia and AVR1-CO39 by direct binding. Plant Cell 25:1463–1481. https://doi.org/10.1105/tpc.112.107201
Cheema KK, Grewal NK, Vikal Y, Sharma R, Lore JS, Das A, Bhatia D, Mahajan R, Gupta V, Bharaj TS, Singh K (2008) A novel bacterial blight resistance gene from Oryza nivara mapped to 38 kb region on chromosome 4L and transferred to Oryza sativa L. Genet Res (camb) 90:397–407. https://doi.org/10.1017/S0016672308009786
Christie N, Tobias PA, Naidoo S, Külheim C (2016) The Eucalyptus grandis NBS-LRR gene family: physical clustering and expression hotspots. Front Plant Sci 6:1238. https://doi.org/10.3389/fpls.2015.01238
Cox KL, Meng F, Wilkins KE, Li F, Wang P, Booher NJ, Carpenter SC, Chen LQ, Zheng H, Gao X, Zheng Y (2017) TAL effector driven induction of a SWEET gene confers susceptibility to bacterial blight of cotton. Nat Commu 8:15588. https://doi.org/10.1038/ncomms15588
DeYoung BJ, Innes RW (2006) Plant NBS-LRR proteins in pathogen sensing and host defense. Nat Immunol 7:1243–1249. https://doi.org/10.1038/ni1410
Ellur RK, Khanna A, Gopala Krishnan S, Bhowmick PK, Vinod KK, Nagarajan M, Mondal KK, Singh NK, Prabhu KV, Singh AK (2016a) Marker-aided incorporation of Xa38, a novel bacterial blight resistance gene, in PB1121 and comparison of its resistance spectrum with xa13 + Xa21. Sci Rep 6:29188. https://doi.org/10.1038/srep29188
Ellur RK, Khanna A, Yadav A, Pathania S, Rajashekara H, Singh VK, Krishnan SG, Bhowmick PK, Nagarajan M, Vinod KK, Prakash G (2016b) Improvement of Basmati rice varieties for resistance to blast and bacterial blight diseases using marker assisted backcross breeding. Plant Sci 242:330–341. https://doi.org/10.1016/j.plantsci.2015.08.020
Harkenrider M, Sharma R, De Vleesschauwer D, Tsao L, Zhang X, Chern M, Canlas P, Zuo S, Ronald PC (2016) Overexpression of rice wall-associated kinase 25 (OsWAK25) alters resistance to bacterial and fungal pathogens. PLoS ONE 11:e0147310. https://doi.org/10.1371/journal.pone.0147310
Huot B, Yao J, Montgomery BL, He SY (2014) Growth–defense tradeoffs in plants: a balancing act to optimize fitness. Mol Plant 7:1267–1287. https://doi.org/10.1093/mp/ssu049
Hutin M, Pérez-Quintero AL, Lopez C, Szurek B (2015) MorTAL Kombat: the story of defense against TAL effectors through loss-of-susceptibility. Front Plant Sci 6:535. https://doi.org/10.3389/fpls.2015.00535
Kim SM, Suh JP, Qin Y, Noh TH, Reinke RF, Jena KK (2015) Identification and fine-mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theor Appl Genet 128:1933–1943. https://doi.org/10.1007/s00122-015-2557-2
Lee H, Cha J, Choi C, Choi N, Ji HS, Park SR, Lee S, Hwang DJ (2018) Rice WRKY11 plays a role in pathogen defense and drought tolerance. Rice 11:5. https://doi.org/10.1186/s12284-018-0199-0
Li Z, Huang J, Wang Z, Meng F, Zhang S, Wu X, Zhang Z, Gao Z (2019) Overexpression of Arabidopsis nucleotide-binding and leucine-rich repeat genes RPS2 and RPM1 (D505V) confers broad-spectrum disease resistance in rice. Front Plant Sci 10:417. https://doi.org/10.3389/fpls.2019.00417
Li Q, Hu A, Qi J, Dou W, Qin X, Zou X, Xu L, Chen S, He Y (2020) CsWAKL08, a pathogen-induced wall-associated receptor-like kinase in sweet orange, confers resistance to citrus bacterial canker via ROS control and JA signaling. Hortic Res 7:42. https://doi.org/10.1038/s41438-020-0263-y
Liu L, Mei Q, Yu Z, Sun T, Zhang Z, Chen M (2013) An integrative bioinformatics framework for genome-scale multiple level network reconstruction of rice. J Integr Bioinform 10:94–102. https://doi.org/10.2390/biecoll-jib-2013-223
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Maeda K, Houjyou Y, Komatsu T, Hori H, Kodaira T, Ishikawa A (2009) AGB1 and PMR5 contribute to PEN2-mediated preinvasion resistance to Magnaporthe oryzae in Arabidopsis thaliana. Mol Plant Microbe Interact 22:1331–1340. https://doi.org/10.1094/MPMI-22-11-1331
Mondal KK (2016) Emerging phytobacterial diseases in india: research status and challenges. In: Perspectives of plant pathology in genomic era. India Phytopathological Society, New Delhi, pp 167–178
Mondal KK, Meena BR, Junaid A, Verma G, Mani C, Majumder D, Khicher M, Kumar S, Banik S (2014) Pathotyping and genetic screening of type III effectors in Indian strains of Xanthomonas oryzae pv. oryzae causing bacterial leaf blight of rice. Physiol Mol Plant Pathol 86:98–106. https://doi.org/10.1016/j.pmpp.2014.03.005
Mondal KK, Verma G, Manju JA, Mani C (2016) Rice pathogen Xanthomonas oryzae pv. oryzae employs inducible hrp-dependent XopF type III effector protein for its growth, pathogenicity and for suppression of PTI response to induce blight disease. Eur J Plant Pathol 14:311–323. https://doi.org/10.1007/s10658-015-0768-7
Naithani S, Gupta P, Preece J, D’Eustachio P, Elser JL, Garg P, Dikeman DA, Kiff J, Cook J, Olson A, Wei S (2020) Plant Reactome: a knowledgebase and resource for comparative pathway analysis. Nucleic Acids Res 48:D1093–D1103. https://doi.org/10.1093/nar/gkz996
Okada A, Shimizu T, Okada K, Kuzuyama T, Koga J, Shibuya N, Nojiri H, Yamane H (2007) Elicitor induced activation of the methylerythritol phosphate pathway toward phytoalexins biosynthesis in rice. Plant Mol Biol 65:177–187. https://doi.org/10.1007/s11103-Mol.Plant-MicrobeInteract.007-9207-2
Park HL, Bhoo SH, Kwon M, Lee SW, Cho MH (2017) Biochemical and expression analyses of the rice cinnamoyl-CoA reductase gene family. Front Plant Sci 8:2099. https://doi.org/10.3389/fpls.2017.02099
Sasaki K, Iwai T, Hiraga S, Kuroda K, Seo S, Mitsuhara I, Miyasaka A, Iwano M, Ito H, Matsui H, Ohashi Y (2004) Ten rice peroxidases redundantly respond to multiple stresses including infection with rice blast fungus. Plant Cell Physiol 45:1442–1452. https://doi.org/10.1093/pcp/pch165
Sato Y, Namiki N, Takehisa H, Kamatsuki K, Minami H, Ikawa H, Ohyanagi H, Sugimoto K, Itoh JI, Antonio BA, Nagamura Y (2013) RiceFREND: a platform for retrieving coexpressed gene networks in rice. Nucleic Acids Res 41:D1214–D1221. https://doi.org/10.1093/nar/gks1122
Sekhwal MK, Li P, Lam I, Wang X, Cloutier S, You FM (2015) Disease resistance gene analogs (RGAs) in plants. Int J Mol Sci 16:19248–19290. https://doi.org/10.3390/ijms160819248
Tian D, Wang J, Zeng X, Gu K, Qiu C, Yang X, Zhou Z, Goh M, Luo Y, Murata-Hori M, White FF (2014) The rice TAL effector-dependent resistance protein XA10 triggers cell death and calcium depletion in the endoplasmic reticulum. Plant Cell 26:497–515. https://doi.org/10.1105/tpc.113.119255
Van Ooijen G, Mayr G, Kasiem MM, Albrecht M, Cornelissen BJ, Takken FL (2008) Structure-function analysis of the NB-ARC domain of plant disease resistance proteins. J Exp Bot 59:1383–1397. https://doi.org/10.1093/jxb/ern045
Velazhahan R, Jayaraj J, Khabbaz S, Kumar RS, Muthukrishnan S (2006) Xanthomonas oryzae pv. oryzae infection triggers accumulation of phenolics, defense-related enzymes and thaumatin-like proteins in rice leaves. Arch Phytopathol Plant Prot 39:329–339. https://doi.org/10.1080/03235400500222172
Verma G, Sharma M, Mondal KK (2018) XopR TTSS-effector regulates in planta growth, virulence of Indian strain of Xanthomonas oryzae pv. oryzae via suppressing reactive oxygen species production and cell wall-associated rice immune responses during blight induction. Funct Plant Biol 45:561–574. https://doi.org/10.1071/FP17147
Verma G, Mondal KK, Kulshreshtha A, Sharma M (2019) XopR T3SS-effector of Xanthomonas oryzae pv. oryzae suppresses cell death-mediated plant defense response during bacterial blight development in rice. 3 Biotech 9:272. https://doi.org/10.1007/s13205-019-1802-9
White FF, Yang B (2009) Host and pathogen factors control-ling the rice-Xanthomonas oryzae interaction. Plant Physiol 150:1677–1686. https://doi.org/10.1104/pp.109.139360
White FF, Potnis N, Jones JB, Koebnik R (2009) The type III effectors of Xanthomonas. Mol Plant Pathol 10:749–766. https://doi.org/10.1111/j.1364-3703.2009.00590.x
Yang B, Sugio A, White FF (2006) Os8N3 is a host disease-susceptibility gene for bacterial blight of rice. Proc Natl Acad Sci USA 103:10503–10508. https://doi.org/10.1073/pnas.0604088103
Acknowledgements
The authors acknowledge the Director, Joint Director (Research), Head (Division of Plant Pathology), ICAR-Indian Agricultural Research Institute, New Delhi, India for their support whenever required during the execution of the project. The authors are also grateful to the scientific team of AgriGenome Labs Pvt Ltd, Smart City Kochi, Kerala, India for their support in generating the transcriptome data.
Funding
This research was funded by Department of Biotechnology, Government of India, grant number BT/PR16238/NER/95/103/2015.
Author information
Authors and Affiliations
Contributions
Conceptualization: KKM, PJH; methodology: AK, YR, GV, AB, AQ, AL, KR, MS, TG, R, ER, SM, K, NS, CM; Formal analysis and investigation: AK, KKM, DB; writing—original draft preparation: KKM, AK; writing—review and editing: KKM, AK; Funding acquisition: KKM, PJH; resources: KKM, PJH; supervision: KKM, PJH.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Research involving human participants and/or animals
This research does not include any human or animal participants.
Informed consent
This research does not include any human or animal participants.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mondal, K.K., Kulshreshtha, A., Handique, P.J. et al. Rice transcriptome upon infection with Xanthomonas oryzae pv. oryzae relative to its avirulent T3SS-defective strain exposed modulation of many stress responsive genes. 3 Biotech 12, 130 (2022). https://doi.org/10.1007/s13205-022-03193-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03193-4