Abstract
In plants, the HAK/KUP/KT family is the largest group of potassium transporters, and it plays an important role in mineral element absorption, plant growth, environmental stress adaptation, and symbiosis. Although these important genes have been investigated in many plant species, limited information is currently available on the HAK/KUP/KT genes for Pepper (Capsicum annuum L.). In the present study, a total of 20 CaHAK genes were identified from the pepper genome and the CaHAK genes were numbered 1 – 20 based on phylogenetic analysis. For the genes and their corresponding proteins, the physicochemical properties, phylogenetic relationship, chromosomal distribution, gene structure, conserved motifs, gene duplication events, and expression patterns were analyzed. Phylogenetic analysis divided CaHAK genes into four cluster (I–IV) based on their structural features and the topology of the phylogenetic tree. Purifying selection played a crucial role in the evolution of CaHAK genes, while whole-genome triplication contributed to the expansion of the CaHAK gene family. The expression patterns showed that CaHAK proteins exhibited functional divergence in terms of plant K+ uptake and salt stress response. In particular, transcript abundance of CaHAK3 and CaHAK7 was strongly and specifically up-regulated in pepper roots under low K+ or high salinity conditions, suggesting that these genes are candidates for high-affinity K+ uptake transporters and may play crucial roles in the maintenance of the Na+/K+ balance during salt stress in pepper. In summary, the results not only provided the important information on the characteristics and evolutionary relationships of CaHAKs, but also provided potential genes responding to potassium deficiency and salt stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potassium (K+) is an essential macronutrient, required for plant growth and development, and constituting 2% to 10% of plant dry matter (Leigh and Wyn Jones, 1984; Amrutha et al. 2007). As the most abundant cation in plant cells, K+ plays crucial roles in plant physiological processes, such as photosynthesis, transmembrane transport, enzyme activation, and response to stress (Leigh and Wyn Jones, 1984; Wang and Wu 2013; Luan et al. 2017). Due to heavy leaching losses, chemical fixation, and the low diffusion rate in the soil, the availability of K+ for plant roots is limited over large areas of agricultural land worldwide (Maathuis 2009; Rengel and Damon 2008).
As the K+ concentration in the rhizosphere rarely exceeds the range 0.01 – 1 mM, plants depend on a variety of K+ absorption systems to mediate K+ uptake under these conditions (Luan et al. 2009; Maathuis 2009; White 2013). It has been well recognized that there are two K+ uptake systems operating in roots, namely a low-affinity transport system (LATS), operating via K+ channels under high external K+ concentrations (> 0.5 mM), and a high-affinity transport system (HATS), operating via K+ transporters, which function at low external K+ concentrations (< 0.2 mM) (Epstein et al. 1963; Ashley et al. 2006; Chérel et al. 2014; Li et al. 2018). In plants, genes encoding K+ transporters are grouped into four major families: KT (K+ Transporter)/HAK (high-affinity K+ transporter)/KUP (K+ uptake permease), Trk (transport of K+)/HKT (high-affinity K+/Na+ transporter), KEA (K+ exchange antiporters), and CHX (cation/H+ exchanger) (Uozumi et al. 2000; Cellier et al. 2004; Aranda-Sicilia et al. 2016; Wang and Wu 2017). The HAK/KUP/KT group is the largest family and plays crucial roles in K+ uptake under different potassium conditions (Ahn et al. 2004; Li et al. 2018). In plants, the HAK/KUP/KT family comprises the majority of the plant K+ transporters identified so far and has been divided into four major clusters (I–IV) (Grabov 2007; Wang and Wu 2013). Multiple transporters in cluster I, such as HvHAK1, the first plant HAK/KUP/KT transporter member cloned from barley, AtHAK5 in Arabidopsis, and OsHAK1/OsHAK5 in rice, have high-affinity K+-uptake capacity and are up-regulated in response to low K+ conditions (Schachtman and Schroeder 1994; Santa-María et al. 1997; Gierth et al. 2005; Yang et al. 2014; Feng et al. 2021). OsHAK1 played a key role in the maintenance of K+ homeostasis and positively regulates tolerance to salt and drought stress (Chen et al. 2015). It was recently reported that OsHAK5 could impact on tiller number and development of roots by regulating an ATP-dependent auxin transporter in rice (Yang et al. 2020a, b). Members of cluster II of HAK/KUP/KT seem to be involved in plant development processes (Elumalai et al. 2002; Yang et al. 2009; Osakabe et al. 2013). In Arabidopsis, AtKUP4/TRH1 affects the growth of root hair tips by establishing auxin gradients in the root apex (Templalexis et al. 2021). The expression of AtKUP6 was induced by drought stress and involved in plant cell osmoregulation. Interestingly, AtKUP2, AtKUP6, and AtKUP8 function as efflux K+ transporters, and a triple knockout mutation of atkup2/6/8 significantly increased root cell size and decreased the sensitivity of guard cells and lateral root cells to abscisic acid (ABA) (Elumalai et al. 2002; Osakabe et al. 2013). In addition to the abiotic stress-responsive HAK genes, an increasing number of mutualistic symbiosis-induced HAK genes belonging to the cluster II HAK/KUP/KT family have been identified, and their functions have been shown to be associated with the mycorrhizal K+-uptake pathway (Desbrosses et al. 2004; Guether et al. 2009; Liu et al. 2019). Members of cluster III may maintain the balance of K+/Na+ (Okada et al. 2008). The potential functions and physiological roles of members of cluster IV had less reports than those in the other clusters (Grabov 2007; Han et al. 2016). Besides, the transcriptional regulation of HAK/KUP/KT transporters had also been reported. The OsAKT1 channel is modulated by the Calcineurin B-Like protein 1 (CBL1)–CBL-Interacting Protein Kinase 23 (CIPK23) complex (Li et al. 2014). The R2R3-type MYB transcription factor gene (AtMYB77) positively regulates the expression of AtHAK5 and enhanced K+ acquisition under low K+ stress (Feng et al. 2021) in Arabidopsis. Recently, Song et al. demonstrate that an endoplasmic reticulum-localized OsCYB5-2 protein can binding to OsHAK21, and then promoting OsHAK21-mediated K+ uptake and enhance salt tolerance in rice (Song et al. 2021).
Over the past two decades, the functional characterization of the HAK/KUP/KT family focused mainly on rice and Arabidopsis, in particular the members from the cluster I subfamily, such as AtHAK5, OsHAK1, OsHAK5 and OsHAK21 (Gierth et al. 2005; Yang et al. 2014; Chen et al. 2015; Shen et al. 2015). With the availability of whole complete genome sequences, a series of plant HAK/KUP/KT genes have been identified and characterized in a number of plant species, including 13, 27, 21, and 56 HAK/KUP/KT genes in Arabidopsis thaliana, rice (Oryza sativa), tomato (Solanum lycopersicum L.) and wheat (Triticum aestivum L.), respectively (Ahn et al. 2004; Yang et al. 2009; Hyun et al. 2014; Cheng et al. 2018).
Pepper (Capsicum annuum L.) is an economically important vegetable and spice plant of the Solanaceae family, is a rich source of vitamins, nutrients, and capsaicin, and is widely cultivated throughout the world (Lee et al. 2004; Moscone et al. 2007). In addition to promoting the growth of pepper, supplementary potassium can enhance the dry matter and chlorophyll concentrations under high salt treatments (Wamser et al. 2017; Kaya et al. 2020). As pepper is a salt-sensitive vegetable crop, maintaining high K+/Na+ ratios in the tissues is important in achieving salt tolerance. Whereas, HAK/KUP/KT transporter gene of pepper was poorly studied. CaHAK1 was cloned and its expression was inhibited by NH4+ when the exogenous K+ supply was interrupted was demonstrated, which indicated that CaHAK1 might play a crucial role in K+ uptake and tolerance to Na+ (Martınez-Cordero 2004; Martinez-Cordero et al. 2005). CcHAK1 gene, which is a high-affinity K+ transporter gene from habanero pepper, was induced by K+ starvation in roots and was not inhibited in the NaCl stress. Furthermore, K+ transporter activity of CcHAK1 in yeast is inhibited by external millimolar concentrations of NH4+ and Cs+, but not inhibited by Na+ (Ruiz-Lau et al. 2016).
Here, we identified putative HAK/KUP/KT transporter gene family members were not identified in pepper genome. In this study, we will identify 20 CaHAKs in genome-wide, then took the phylogenetic analysis of HAK/KUP/KT relationships among different plant species, provide conserved motifs in the corresponding proteins, as well as the features of gene structures, chromosomal locations, and characteristics of cis-regulatory elements in promoter regions. Then investigated the evolution and domestication of the CaHAK family members. In addition, the expression profiles of CaHAK family members in response to K+ deficiency or salt stress will catch to select the candidate genes involving in responding to K+ deficiency or salt stress. As a result, this study can provide a valuable information for further functional characterization of CaHAK genes and show insight to genetic improvement of potassium-utilization efficiency in the pepper breeding.
Methods
Plant materials and stress treatments
Pepper cultivar Zunla-1 was selected in this study, for its wide cultivation in Southern China and high-quality genome sequence. Seeds of pepper were surface-sterilized and germinated in a growth chamber with a photoperiod and temperature regime of 16 h light at 28 °C and 8 h dark at 20 °C. The seedlings were then transplanted to sterilized quartz sand for a 3-week culture period, meantime they were irrigated with full strength nutrient solution containing the following: 1 mM NH4+, 4 mM NO3−, 2 mM K+, 1 mM phosphate (Pi), 0.75 mM Ca2+, 0.5 mM Mg2+, 0.25 mM Cl−, 0.5 mM SO42−, 20 μM Fe2+, 9 μM Mn2+, 46 μM BO33+, 8 μM Zn2+, 3 μM Cu2+, and 0.03 μM MoO42+. After 3 weeks, the seedlings were then transferred to a hydroponic culture system in nutrient media to achieve different stress treatments. For the K+-deficiency treatment, 21-day-old pepper plants were cultured in the full strength nutrient medium solution containing 0.2 mM K+, with the pH adjusted to 5.5 with NaOH (Liu et al. 2019). Under the salt-stress treatment, 21-day-old pepper plants were cultured with full-strength nutrient medium containing 2 mM K+ and 200 mM NaCl (Liu et al. 2016a, b). The salt-stress treatment sample were harvested at 24 h after the treatments. Control plants for both treatments were cultured in the full-strength medium containing 2 mM K+. After each treatment was completed, the roots and leaves of the pepper plants (three replicates per treatment, three seedlings per replicate) were harvested, rinsed, and patted dry, then immediately snap-frozen in liquid nitrogen and stored at − 80 °C for subsequent RNA isolation.
Identification of the CaHAK gene family in pepper
To obtain all the HAK/KUP/KT family genes from the pepper genome, the Arabidopsis genes from this family were each queried in the pepper genome sequence downloaded from the NCBI genome database (https://www.ncbi.nlm.nih.gov/genome/?term=Capsicum+annuum+L). Each sequence was selected as a candidate HAK/KUP/KT protein with an e-value < 10–10 and a query of greater than 50%. All the candidate gene sequences obtained were submitted to the Pfam (http://pfam.xfam.org/) and SMART (http://smart.embl-heidelberg.de/) databases for further confirmative analysis (Finn et al. 2014; Letunic et al. 2020). Ultimately, a total of 20 HAK/KUP/KT genes were identified and named as CaHAK1-20 based on the corresponding orthologous genes from tomato. Physicochemical properties of the pepper HAK transporter genes, including gene name, ID, protein length, isoelectric point, predicted protein molecular weight, grand average of hydropathicity (GRAVY), and predicted protein subcellular localization were searched for using the ProtParam tool (http://web.expasy.org/protparam/) (Artimo et al. 2012). The subcellular locations of the CaHAK proteins were predicted by SOFTBERRY (http://linux1.softberry.com/) and Cell-PLoc 2.0 (http://www.csbio.sjtu.edu.cn/bioinf/Cell-PLoc-2/).
The full-length protein sequences encoded by HAK/KUP/KT from C. annuum (CaHAKs), O. sativa (OsHAKs) (Gupta et al. 2008), A. thaliana (AtHAK/KUP/KTs) (Ahn et al. 2004), C. reinhardtii (CrHAKs), P. patens (PpHAKs), V. vinifera L. (VvHAKs) (Nieves-Cordones et al. 2016), S. melongena (SmHAKs) and S. lycopersicum (SlHAKs) (Hyun et al. 2014; Liu et al. 2019) were determined using the program Clustal X (v1.8) with default parameters (gap opening penalty = 10, gap extension penalty = 0.2, protein weight matrix = Gonnet), and a phylogenetic tree was constructed by the Neighbor-Joining (NJ) method model with the MEGA7 software (Saito et al. 2013). The bootstrap parameter was set at 1,000 replicates (Kumar et al. 2016).
To further investigate the evolution of the HAK/KUP/KT family genes from algae to angiosperm like dicotyledons C. annuum, we identified HAK/KUP/KT genes in one algae (C. reinhardtii); one lycopodiophyta (S. moellendorffii); one early angiosperm (A. trichopoda); two bryophytes (M. polymorpha, P. patens); three monocotyledons (O. sativa, Z. mays, S. bicolor); five dicotyledons (A. thaliana, C. annuum, S. lycopersicum, S. tuberosum, V. vinifera), and count the numbers of HAK genes in four major clusters, then constructed the phylogenetic tree with MEGA7.0 using the NJ method (Saito et al. 2013) and bootstrap parameter with 1,000 replicates (Kumar et al. 2016).
Conserved motif recognition and gene structure analysis
The cDNAs and their corresponding DNA sequences from the CaHAK gene family were obtained from the genome database. Exon/intron structure analysis was carried out by comparing each cDNA sequence with the corresponding genomic DNA sequence. The 2,000-bp genomic DNA sequence upstream of each CaHAK gene was selected for promoter analysis from the pepper genome database. Then, the PlantCARE database was used to analyze the cis-regulatory elements in the promoters (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/search_CARE.html). Eventually, the cis-regulatory elements associated with stress response were selected to draw the distribution map in the promoter via TBtools software (Chen et al. 2020). The NCBICD (Conserved Domain) search was used for protein domain analysis (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi). The conserved motifs of the pepper CaHAK family proteins were identified using the MEME tool (http://meme-suite.org/tools/meme) (Bailey et al. 2009). The TMHHM Serve 2.0 tool (http://www.cbs.dtu.dk/services/TMHMM/) was used to predict the transmembrane (TM) domains of the CaHAK proteins.
Chromosome location and syntenic analysis
The localization information of the CaHAK gene family on the pepper chromosomes was obtained from the pepper GFF (general feature format) genome files. Then, TBtools was used to construct the chromosome localization map of the CaHAK gene. The orthologous genetic relationship among HAK genes from pepper, Arabidopsis, rice, grape, potato, eggplant and tomato was identified using the MCScanX program. The advanced Circos tool was used to visualize the syntenic relationship between different species. Non-synonymous (Ks), synonymous (Ka), and Ka/Ks values for each CaHAK gene pairs were calculated using the TBtools V 1.802 simple Ka/Ks calculator. The Ks value was used to estimate the divergence time of CaHAK duplicated gene pairs (T = Ks/2R Mya, Millions of years), where T refers to divergence time and R is the rate of synonymous substitutions (R = 1.5 × 10–8) (Edlund et al. 2004).
Transcriptome profiling analysis
To further explore the potential functions of the 20 CaHAK genes, the expression patterns of these members were analyzed in different tissues, namely roots, stems, leaves, flowers, bud tissues, and nine developmental stages of fruit, using public RNA-sequence data from pepper cultivar Zunla-1(Qin et al. 2014). The relative expression was expressed with fragments per kilobase of transcript per million mapped reads (FPKM) (Table S4) and showed with heat map. The heat map was drawn using TBtools v1.086.
Total RNA extraction and quantitative real-time PCR analysis
Total RNA was extracted from 100 mg fresh weight of pepper leaves and roots. The total RNA (approximately 1 μg) from the root sample was used to synthesize cDNA using a reverse transcription kit (Takara, Dalian, China). Using the synthesized cDNAs as templates, quantitative real-time PCR (qPCR) was performed in three technical replicates for each sample, using the SYBR Premix Ex Taq kit (Takara, Japan). The primer pairs used for CaHAK genes are listed in Table S1. All primers for RT-PCR were designed using Primer 5.0 software, and a melting curve was performed to check for gene-specific amplification (Fig. S1). The relative transcript abundance of each target gene was log2-normalized against β-Actin transcript level (Ma et al. 2019). Any changes in expression levels of CaHAK genes were quantified using the 2−ΔΔCt method (Li et al. 2009).
The RNA sequence data for tissue-specific expression profiling of CaHAKs (including roots, stems, leaves, buds, flowers, and the nine fruit developmental stages), were downloaded from the GEO Datasets (https://www.ncbi.nlm.nih.gov/gds/?term=GSE45037) (Qin et al. 2014). The relative expression levels of CaHAKs were analyzed by the fragments per kilobase per million reads (FPKM) values. Heatmaps of these genes were generated by TBtools v1.086 (Chen et al. 2020).
Statistical analysis
The experiment data were analyzed by ANONA (IBM SPSS Statistics 25.0), followed by Tukey’s test (*P < 0.05, **P < 0.01). The obtained data were processed by GraphPad Prism 8.0.1.
Results
Genome-wide identification of CaHAK gene family members in pepper
We identified 20 putative HAK/KUP/KT genes in pepper genome basing on the BLAST search results (Table S2). These putative genes were named CaHAK1 to 20, and all of them encoded proteins with, in most cases, similar physicochemical properties. Information about all the putative genes was listed in Table S2, including gene name, ID, protein length, isoelectric point, predicted protein molecular weight, grand average of hydropathicity (GRAVY), and predicted protein subcellular localization. The lengths of the 20 HAK/KUP/KT putative proteins ranged from 713 to 848 amino acids, the molecular weights and isoelectric points of these proteins varied from 79.2 to 94.4 kDa and from 6.04 to 9.34, respectively (Table S2). GRAVY analysis showed that all the proteins were hydrophobic, while the predicted subcellular locations of the CaHAK proteins were all at the plasma membrane (PM), the results of both properties suggested that the protein functions were to maintain K+ homeostasis in pepper.
Phylogenetic analyses of HAK/KUP/KT family genes
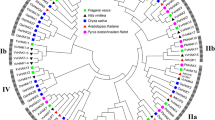
To reveal the genetic relationship among the CaHAK genes, a total of 144 plant HAK protein sequences from eight species, namely Capsicum annuum L, Solanum lycopersicum, Solanum tuberosum, Vitis vinifera, Oryza sativa, Arabidopsis, and Physcomitrella patens, Chlamydomonas reinhardtii, were aligned using the Clustal X program, and an unrooted tree was constructed using the Neighbor-Joining method using MEGA7 software. According to the phylogenetic tree, CaHAK family genes could be divided into four major clusters (I–IV) (Fig. 1). In cluster I, the six CaHAK members, CaHAK1, CaHAK5, CaHAK9, CaHAK13, CaHAK15, and CaHAK16 belong to cluster IA, and it is worth noting that cluster IB contained only members from rice. In cluster II, the eight CaHAK members, namely CaHAK2, CaHAK3, CaHAK4, CaHAK6, CaHAK7, CaHAK8, CaHAK10, and CaHAK17, were classified into three sub-clusters (cluster IIA, IIB, and IIC). In cluster III, CaHAK family contained five members and was composed of sub-cluster IIIA (CaHAK18 and CaHAK19) and III B (CaHAK11, CaHAK12, and CaHAK20). Cluster IV contained only one pepper gene, CaHAK14. Notably, several orthologous CaHAK members were found in tomato genome, indicated that a whole-genome triplication contributed to the expansion of the CaHAK gene family.
Phylogenetic analysis of CaHAK proteins from Capsicum annuum L. with orthologs from Arabidopsis thaliana, Oryza sativa, Physcomitrella patens, Chlamydomonas reinhardtii, Solanum lycopersicum, Solanum tuberosum, and Vitis vinifera. Different colors indicate different clusters. The phylogenetic tree was constructed by the Neighbor-Joining method, using MEGA7 software and 1,000 bootstrap replicates
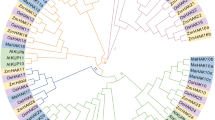
To further investigate the evolution of the HAK/KUP/KT family genes, we identified 223 HAK/KUP/KT genes in 14 species from algae, lycopodiophyta, early angiosperm, bryophytes, monocotyledons to dicotyledons. The counts of different cluster of HAK/KUP/KT genes are shown in Fig. 2. The number of HAK/KT/KUP family gene of algae and bryophyta was less than that in higher plants. Interestingly, the cluster IV was the largest subfamily both in algae and bryophyta species, indicated that the cluster IV began to expand during the plant terrestrialization. In addition, the numbers of cluster II and III family gene were highly conserved in magnoliophyte and mesangiospermae, respectively.
Phylogeny and diversity of HAK/KT/KUP genes in 14 species. The species tree of 14 selected species was constructed using the online software TIMETREE (http://timetree.org/). The number of HAK/KT/KUP family gene was visualized by TBtools v1.086. The different color stars and circles represent plant species
Chromosomal position and duplication analysis of CaHAK genes
The chromosomal locations of the CaHAK genes were mapped from the pepper genome database, using TBtools. As shown in Fig. S2, apart from CaHAK7, which was located on a scaffold chromosome (NW_015960857.1), all the CaHAK genes were mapped onto 9 of the 12 pepper chromosomes. Chromosome 3 carried the most CaHAK genes (five). Chromosome 2 carried four CaHAK genes, followed by chromosomes 5, 9, and 11 (two CaHAK genes on each chromosome), and chromosomes 1, 6, 8, and 12, each carrying a single CaHAK gene. A similar random distribution model of HAK/KUP/KT gene families was also observed in Arabidopsis and pears (Ahn et al. 2004; Wang et al. 2018).
In addition, the duplication modes of the CaHAK genes were analyzed in pepper, using TBtools. The results identified three segmentally duplicated gene pairs (CaHAK6/CaHAK8, CaHAK15/CaHAK2, and CaHAK4/CaHAK18), and one tandem duplication type gene pair (CaHAK13/CaHAK15) (Table 1). The frequencies of non-synonymous (Ka) and synonymous (Ks) substitutions and the corresponding ratio (Ka/Ks) can be used to estimate the evolutionary date and the direction of the subsequent selection pressure on these genes (Qiao et al. 2015; Wang et al. 2018). In the present study, the Ka/Ks ratios of these duplicated CaHAKs gene pairs were calculated. The results showed that all of the Ka/Ks ratios were < 0.5, indicating that purifying (negative) selection played the main force driving for the evolution of HAK genes. The time of divergence of segmentally/tandem CaHAK gene pairs was also estimated. According to the results, the divergence times of duplicated genes occurred between 2.81 and 8.29 million years ago (mya).
To further understand the phylogenetic relationship among pepper CaHAK genes, the collinear relationships with five other species were constructed, including four dicotyledonous plants (Arabidopsis, grape, eggplant and tomato) and one monocotyledonous plant (rice) (Fig. 3). In total, 27 syntenic gene pairs of CaHAKs with orthologs of the other four species were found in the pepper genome. The CaHAKs genes had eight syntenic gene pairs with tomato (Table S3), nine syntenic gene pairs with eggplant, six syntenic gene pairs with grape, and two syntenic gene pairs with each of A. thaliana and rice. Additionally, the numbers and chromosomal positions of the HAK/KUP/KT family genes in pepper were closely aligned with those of tomato and eggplant, indicating that HAK/KUP/KT genes of solanaceous species were highly conserved during evolutionary divergence.
Syntenic analysis of the HAK/KUP/KT family genes. The lines in different colors inside the circle indicate collinearity relationships among the genes from Capsicum annuum and Arabidopsis thaliana, Oryza sativa, Solanum lycopersicum, and Vitis vinifera. The duplications between HAKs were identified using TBtools software
Gene structure of HAK/KUP/KT family genes
To gain insights into the structural features of CaHAKs, the exon/intron structures of 20 CaHAKs were analyzed by comparing the corresponding coding sequences (CDS) and genomic sequences (Hu et al. 2015). As shown in Fig. 4, most HAK/KUP/KT genes in pepper contain 8 – 12 exons, and highly conserved exon/intron structures could be observed in the CaHAKs belonging to the same cluster. Notably, most of the exons common to all of the CaHAKs were 225 bp, 248 bp, 50 bp (or 53 bp), and 260 bp in length, a finding which is consistent with previous results from tomato and pear, indicating that this gene structure was evolutionarily conserved among these species (Liu et al. 2019; Wang et al. 2018). Interestingly, some of the CaHAK genes either lacked one intron (CaHAK6 and CaHAK8) or had gained one intron (CaHAK3, CaHAK10, and CaHAK16).
Gene structure characters of CaHAK genes from pepper and the architecture of the conserved motifs in CaHAK proteins. A Gene structures of CaHAK genes were based on phylogenetic relationship. Black boxes represent exons and lines represent introns. B Conserved protein motifs. Conserved motifs of CaHAK proteins were identified by the MEME tool and are shown in different-colored boxes
In general, the function of individual CaHAK proteins was reflected in their protein structure. All of the CaHAK proteins contained the K+ transporter domain (PF02705), which indicated that all 20 CaHAK genes could perform similar K+ acquisition functions. The distribution of 10 conserved motifs in the CaHAK proteins was analyzed, and all 20 CaHAK proteins contained at least eight typical motifs associated with K+ potassium acquisition, including motifs 2, 3, 4, 6, 7, 8, 9, and 10 (Fig. 4). Although most of the motifs were present and conserved in all 20 CaHAK proteins, some of the motifs were lost from individual CaHAK proteins, such as motif 16, 12 was absent from CaHAK15 and CaHAK10, respectively. This phenomenon had also been reported in HAK/KUP/KT family genes from other species, such as pear and wheat (Cheng et al. 2018; Wang et al. 2018).
Motif analysis of HAK/KUP/KT proteins
To further investigate the protein domains and conserved motifs of CaHAKs, the protein sequences were submitted to the online program MEME and NCBI conserved domain database (Bailey et al. 2009). A subsequent conserved domain analysis also confirmed all of the predicted CaHAK genes. As shown in Fig. S3, there are four types of members for “K_trans superfamily” (cl15781). The conserved domains named as “K_trans”, “PLN00149”, “PLN00151” were identified in CaHAK14, CaHAK8, and three members of cluster I (CaHAK11, CaHAK12, and CaHAK20), respectively. The other 15 CaHAK members harbor conserved domain of “K_trans superfamily”. This result is consistent with that encoded by the reported pear and barley HAK gene family (Wang et al. 2018; Cai et al. 2021). In total, 20 conserved motifs were identified in putative CaHAK proteins (Fig. S4). Generally, the 20 motifs were relatively highly conserved among CaHAK family members, a finding which was consistent with the exon/intron structures. It was worth noting that a highly conserved K-transport domain (GVVYGDLGTSPLY) (named motif 9) existed in all CaHAK proteins. In addition, motif 19 only existed in CaHAKs from clusters I and III, whereas motif 17 could only be found in members of cluster II. Based on the TMME2.0 program, it was predicted that the HAK/KUP/KT proteins in pepper had 10–14 transmembrane (TM) domains (Fig. S5), a finding which was similar to previous results from other higher plants (Li et al. 2017; Yang et al. 2020a, b).
Analysis of putative cis-regulatory elements in the CaHAK promoters
To better understand the expression differences and transcriptional regulation of CaHAKs, PlantCARE database was used to identify cis-regulatory elements (CREs) in the promoter regions. A total of seventeen types of CREs were identified (Fig. 4). Among the CREs responding to phytohormones, the numbers of abscisic acid- (35) and methyl jasmonate- (30) responsive elements were the highest. The auxin-responsive element was observed in the promoters of only three CaHAKs. A total of 13 CaHAKs contained the MYB-binding site, with eight CaHAK genes containing the MYBHv1-binding site (Fig. 4). Many CREs responding to different stresses, such as defense, anaerobiosis and low temperature, were identified among the CaHAKs. Of these, the anaerobic induction response element was the most common. There were also CREs involved in plant growth, including endosperm expression and “zein” metabolism regulatory elements, which were identified in the promoters of three and six CaHAK members, respectively (Fig. 5). Interestingly, as the most important environmental factor, light responsiveness was observed in the promoters of almost all CaHAKs, except for CaHAK16.
Tissue-specific expression patterns of CaHAKs
Each of the CaHAK genes was expressed in at least one pepper tissue, with 12 CaHAK genes exhibiting expression in all the tissues tested in this study (Fig. 6). The heat map of the CaHAKs showed that CaHAK4, CaHAK6, CaHAK18, and CaHAK19 had high levels of constitutive expression in all the tissues analyzed, whereas the expression levels of CaHAK1 and CaHAK16 showed much lower levels of expression than those shown by other members, suggesting functional divergence of CaHAK genes in different tissues of pepper. Notably, five CaHAKs (CaHAK3, 7, 9, 13, and 15) exhibited higher expression levels in flowers and flower buds than in other organs. As an indispensable mineral for higher plant, K+ plays a key role in pollen germination and tube growth via maintaining osmotic pressure. Similarly, OsHAK1, 19 and 20 clustered with CaHAK9, 13 and 15, also showed high expression levels in anthers and played an important role in receptor-like kinase mediates K+ homeostasis in pollen tube growth and integrity. This result indicating that these genes with high levels in flower might be involved in flower and bud development via K-mediated turgor pressure. As with the orthologs AtHAK5 and OsHAK5, CaHAK5 showed high expression levels in roots, implying that CaHAK5 might function particularly with respect to K+ acquisition by roots from the soil. In addition, CaHAK8 and CaHAK10 showed their highest expression levels in stem tissues. Based on the high expression level of CaHAK1 and CaHAK5 genes in pepper roots, we speculated that these genes might potentially play a crucial role in uptake or transport of K+/Na+ nutrition.
Expression analysis of pepper HAK/KUP/KT genes in response to K+-deficiency and salt stresses
To investigate the transcriptional response of pepper plant to low potassium conditions, the expression profiles of CaHAKs in leaves and roots at 7 days after exposing to low K+ stress were studied. Among the 20 CaHAK genes, nine genes (CaHAK9, CaHAK10, CaHAK12, CaHAK13, CaHAK14, CaHAK15, CaHAK16, CaHAK17, and CaHAK18) were not expressed under the experimental conditions. The expressions of CaHAK1, CaHAK3, CaHAK5, and CaHAK7 were upregulated in response to low K+ conditions (Fig. 7). The transcript abundance of CaHAK5 was significantly higher than that exhibited by the other CaHAKs. It is noteworthy that the expression levels of CaHAK3 and CaHAK7 were significantly upregulated in pepper roots in response to K+ deficiency. Hence, CaHAK3 and CaHAK7 were selected as candidates of high-affinity K+ uptake transporters in pepper plants.
Expression levels of CaHAK genes in response to K+ deficiency (− K) or salt stress (+ Na). The relative transcript levels of CaHAK genes in pepper leaves and roots were determined by quantitative real-time PCR (RT-PCR). Error bars represent standard error (SE) (n = 3). The asterisks indicate significant differences from the corresponding control (*P < 0.05, **P < 0.01), using analysis of variance (ANOVA)
To further determine how the expressions of CaHAKs were affected by salt stress, the expression profile of CaHAKs was determined in pepper plants after 24-h salt treatment (200 mM Na+). The results showed that most of the CaHAKs examined exhibited appreciably up-regulated expression levels in pepper roots, such as CaHAK3, CaHAK5, CaHAK6, CaHAK7, and CaHAK19, in responding to slat stress (Fig. 7). While, CaHAK2 and CaHAK11 showed down-regulated expression levels in pepper leaves under salt stress conditions.
Discussion
As one of the three essential plant macronutrients, K+ plays crucial roles in multiple physiological and biochemical processes (Anschutz et al. 2014). The HAK/KUP/KT family of genes has been reported to be involved in K+ uptake under different K+ supply conditions and in the presence of salt or drought stress, in species, such as Arabidopsis, rice, maize, wheat, and tea plants (Cheng et al. 2018; Yang et al. 2009; Yang et al. 2020a, b; Zhang et al. 2012). Still, there is limited information of the molecular mechanisms of potassium uptake in pepper. The completion of whole-genome sequencing in pepper allowed us to comprehensively investigate the regulation and structure of HAK/KUP/KT family genes in pepper (Qin et al. 2014).
In the present study, a total of 20 HAK/KUP/KT family genes were identified from the pepper genome, using BLASTN searches. The HAK/KUP/KT genes were distributed among four major clusters (I ~ IV) (Fig. 1). Additionally, CaHAK numbers from clusters II, III, and IV were distributed randomly among the subgroupings, indicating that the monocot/dicot split occurred before the significant duplication events that gave rise to clusters II, III and IV. In the current work, the number of exons in pepper HAK genes ranged from 8 to 12 (Fig. 4), a number which is highly conserved with respect to the orthologous HAK genes from rice and tomato (Hyun et al. 2014; Yang et al. 2009). Furthermore, the position and length of exons were also highly conserved in pepper and other plant species, such as tomato and pear plants (Hyun et al. 2014; Wang et al. 2018). Interestingly, CaHAK10 contains 12 exons, which is the largest exon number among the pepper HAK/KUP/KT genes, showing marked similarities with the tomato gene ortholog SlHAK10, which is the arbuscular mycorrhiza-induced K+ transporter gene (Liu et al. 2019). The conservation of gene structure allowed us to predict the potential functions of the pepper HAK/KUP/KT family genes.
Gene duplication contributes significantly to the expansion of gene numbers in plant species and leads to gene diversification, driving the evolution of paralogous genes (Maher et al. 2006). The duplication mechanisms include whole-genome duplication (WGD), segmental and tandem duplication, and rearrangements at the gene and chromosomal levels (Edger and Pires 2009; Jiao et al. 2011). In Chinese white pear, segmental duplication events played a more important role than did tandem duplication events in the expansion of the PbrHAK gene family (Wang et al. 2018). Likewise, the expansion of the HAK/KUP/KT family genes was driven primarily by segmental duplication events in poplar (He et al. 2012). This result showed that segmental and tandem duplication events both played vital roles in the evolution of CaHAK genes, with the segmental duplication type (representing 80% of duplication events) being more important than the tandem duplication type (20%), suggesting that the evolution of potassium absorption mainly evolved as a result of segmental duplication events, the genes involved showing high levels of conservation in pepper (Table 1). The selection pressure associated with duplicated gene pairs can be divided into three types, namely purifying (Ka/Ks < 1), positive (Ka/Ks > 1), and neutral selection (Ka/Ks = 1) (Yang 2007). In the present study, the Ka/Ks ratios of all the duplicated CaHAK gene pairs were < 0.5, suggesting that purifying selection may play the crucial role in adaptation to environmental changes during CaHAK gene evolutionary history. It should be noted that the time of divergence of tandem duplicated gene pairs (CaHAK1/CaHAK9) was less than for segmental duplication events. This finding is similar to previous observations, such as with the fatty acid desaturase (FAD) enzyme family genes in wheat (Hajiahmadi et al. 2020).
Syntenic analysis of HAK/KUP/KT family genes were performed to assess the evolutionary relationship of the CaHAK genes in four plant families, namely the Gramineae (O. sativa), the Brassicaceae (A. thaliana), the Vitaceae (V. vinifera), and the Solanaceae (tomato and eggplant). The results showed that the CaHAK genes had eight and nine syntenic gene pairs with tomato and eggplant, respectively, which was more than with rice (two pairs) grape (six pairs) or Arabidopsis (two pairs) (Fig. 3, Table S4). Qin et al. (2014) showed that the pepper diverged from tomato ~ 36 mya, and that approximately 38-Mb of genomic sequences of pepper was aligned to tomato, with 14% nucleotide divergence (Qin et al. 2014). There were more collinear gene pairs between pepper and tomato than between pepper and the other species, consistent with the fact that both pepper and tomato are members of the Solanaceae family.
The transcriptional and post-transcriptional regulation of genes encoding K+ transporters or channels are two common mechanisms in plants for achieving increased fitness to K+-deficiency conditions (Li et al. 2018; Wang and Wu 2013, 2017). Previous studies revealed that the HAK genes in cluster I and cluster V were mainly induced by potassium deficiency. For example, the expression of AtHAK5, AtKUP7, OsHAK1, OsHAK5, and HvHAK1 was significantly upregulated in roots under low-K+ conditions, maintaining K+ uptake and translocation from root to shoot (Chen et al. 2015; Gierth et al. 2005; Han et al. 2016; Santa-María et al. 1997; Yang et al. 2014). The AtHAK5 T-NDA insertion mutant plants showed a lower K+ accumulation in roots compared to wild-type plants under K+-deficient conditions. The K+ content in shoots but not in roots of atkup7 mutant plants exhibited a significantly decreased compared with those in the wild-type plants (Gierth et al. 2005; Han et al. 2016). Recent study revealed that the atkup9 mutant exhibited a short-root phenotype growth under low-K conditions. In addition, the Cs+ accumulation was significantly increased in the atkup9 mutant plants, indicated that AtKUP9 played an important role in root growth and Cs+ homeostasis in Arabidopsis. In this study, the expression of CaHAK1/CaHAK5 (cluster I) and CaHAK3/CaHAK7 (cluster II) was upregulated following exposure to low-K+ stress, implying that these genes are involved in mediating K+ uptake and transport in pepper under K+-deficiency conditions. Interestingly, the CaHAK3 and CaHAK7 genes showed similar expression patterns in response to either K+ deficiency or salt stress in pepper roots. As the two pepper HAK/KUP/KT paralogs, CaHAK3 and CaHAK7, showed a high degree of overlapping expression, this finding strongly implied genetic redundancy with the two genes.
In higher plant species, maintaining K+ uptake and the cellular K+/Na+ ratio is essential for salt tolerance (Cuin et al. 2003; Maathuis 2006; Shabala & Cuin 2008). It has been repeatedly documented that HAK/KUP/KT members are upregulated in response to salt stress. In Arabidopsis, transcript abundances of AtKUP6 and AtKUP11 were significantly upregulated in response to salt stress (Maathuis 2006). The OsHAK5 and OsHAK21 transcript levels were also upregulated under high Na+ conditions, playing crucial roles in salt stress tolerance by maintaining the K+/Na+ homeostasis in plant cells (Shen et al. 2015; Yang et al. 2014). In pepper, the expression level of CaHAK5 and CaHAK11 was significantly upregulated by salt stress, whereas CaHAK1 expression was rapidly downregulated under high-Na+ condition. These results implied that the CaHAK genes played a role in maintaining the K+/Na+ homeostasis in pepper under salt stress.
Conclusion
In the present study, a total of 20 CaHAK genes were identified in pepper (Capsicum annuum L). These genes were classified into four clusters (I – IV), based on the topology of the phylogenetic tree and on structural feature analysis. Exon/intron distribution and comparisons of domains/motifs were conserved in each cluster. Promoter analysis revealed associated with plant growth and development, stress-responsive and phytohormone elements in the CaHAKs promoter regions, indicating functional roles of CaHAKs in response to different stresses and developmental stages. CaHAK genes, except CaHAK7, were distributed on 9 of the 12 pepper chromosomes. According to the analysis of collinearity, the whole-genome triplication might contribute to the expansion of the CaHAK gene family. Moreover, the results of the Ka/Ks ratio revealed that tandem/segmental duplications have contributed to the expansion of the CaHAK family genes, and purifying selection played the key role in the divergence of the CaHAK genes in pepper. High expression of CaHAKs showed that CaHAK3, CaHAK7, CaHAK9, CaHAK13, and CaHAK15 in flower and buds, implied their potential functions in flower and buds development, even fruit setting. The analysis of expression patterns under low K+ and high Na+ conditions suggested that CaHAKs exhibited functional divergence in the processes of plant K+ uptake and salt stress tolerance. Taken together, this study provided insights into the HAK-mediated low K+ response of pepper at the transcriptional level and identified candidate genes that may be exploited to improve crop tolerance to abiotic stresses.
References
Ahn SJ, Shin R, Schachtman DP (2004) Expression of KT/KUP genes in Arabidopsis and the role of root hairs in K+ uptake. Plant Physiol 134:1135–1145. https://doi.org/10.1104/pp.103.034660
Amrutha RN, Sekhar PN, Varshney RK, Kishor PB (2007) Genome-wide analysis and identification of genes related to potassium transporter families in rice (Oryza sativa L.). Plant Sci 172:708–721. https://doi.org/10.1016/j.plantsci.2006.11.019
Anschutz U, Becker D, Shabala S (2014) Going beyond nutrition: regulation of potassium homoeostasis as a common denominator of plant adaptive responses to environment. J Plant Physiol 171:670–687. https://doi.org/10.1016/j.jplph.2014.01.009
Aranda-Sicilia MN, Aboukila A, Armbruster U, Cagnac O, Schumann T, Kunz HH, Jahns P, Rodriguez-Rosales MP, Sze H, Venema K (2016) Envelope K+/H+ antiporters AtKEA1 and AtKEA2 function in plastid development. Plant Physiol 172:441–449. https://doi.org/10.1104/pp.16.00995
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, Duvaud S, Flegel V, Fortier A, Gasteiger E, Grosdidier A, Hernandez C, Ioannidis V, Kuznetsov D, Liechti R, Moretti S, Mostaguir K, Redaschi N, Rossier G, Xenarios I, Stockinger H (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40:W597-603. https://doi.org/10.1093/nar/gks400
Ashley MK, Grant M, Grabov A (2006) Plant responses to potassium deficiencies: a role for potassium transport proteins. J Exp Bot 57(2):425–436. https://doi.org/10.1093/jxb/erj034
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202-208. https://doi.org/10.1093/nar/gkp335
Cai K, Zeng F, Wang J, Zhang G (2021) Identification and characterization of HAK/KUP/KT potassium transporter gene family in barley and their expression under abiotic stress. BMC Genom 22:317. https://doi.org/10.1186/s12864-021-07633-y
Cellier F, Conejero G, Ricaud L, Luu DT, Lepetit M, Gosti F, Casse F (2004) Characterization of AtCHX17, a member of the cation/H+ exchangers, CHX family, from Arabidopsis thaliana suggests a role in K+ homeostasis. Plant J 39:834–846. https://doi.org/10.1111/j.1365-313X.2004.02177.x
Chen G, Hu Q, Luo L, Yang T, Zhang S, Hu Y, Yu L, Xu G (2015) Rice potassium transporter OsHAK1 is essential for maintaining potassium-mediated growth and functions in salt tolerance over low and high potassium concentration ranges. Plant Cell Environ 38:2747–2765. https://doi.org/10.1111/pce.12585
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Cheng X, Liu X, Mao W, Zhang X, Chen S, Zhan K, Bi H, Xu H (2018) Genome-Wide identification and analysis of HAK/KUP/KT potassium transporters gene family in wheat (Triticum aestivum L.). Int J Mol Sci. https://doi.org/10.3390/ijms19123969
Chérel I, Lefoulon C, Boeglin M, Sentenac H (2014) Molecular mechanisms involved in plant adaptation to low K(+) availability. J Exp Bot 65(3):833–848. https://doi.org/10.1093/jxb/ert402
Cuin TA, Miller AJ, Laurie SA, Leigh RA (2003) Potassium activities in cell compartments of salt-grown barley leaves. J Exp Bot 54:657–661. https://doi.org/10.1093/jxb/erg072
Desbrosses G, Kopka C, Ott T, Udvardi MK (2004) Lotus japonicus LjKUP is induced late during nodule development and encodes a potassium transporter of the plasma membrane. Mol Plant Microbe Interact 17:789–797. https://doi.org/10.1094/mpmi.2004.17.7.789
Edger PP, Pires JC (2009) Gene and genome duplications: the impact of dosage-sensitivity on the fate of nuclear genes. Chromosome Res 17:699–717. https://doi.org/10.1007/s10577-009-9055-9
Edlund AF, Swanson R, Preuss D (2004) Pollen and stigma structure and function: the role of diversity in pollination. Plant Cell 16(Suppl):S84-97. https://doi.org/10.1105/tpc.015800
Elumalai RP, Nagpal P, Reed JW (2002) A mutation in the Arabidopsis KT2/KUP2 potassium transporter gene affects shoot cell expansion. Plant Cell 14:119–131. https://doi.org/10.1105/tpc.010322
Epstein E, Rains DW, Elzam OE (1963) Resolution of dual mechanisms of potassium absorption by barley roots. Proc Natl Acad Sci USA 5:684–692
Feng CZ, Luo YX, Wang PD, Gilliham M, Long Y (2021) MYB77 regulates high-affinity potassium uptake by promoting expression of HAK5. New Phytol. https://doi.org/10.1111/nph.17589
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222-230. https://doi.org/10.1093/nar/gkt1223
Genies L, Martin L, Kanno S, Chiarenza S, Carasco L, Camilleri V, Vavasseur A, Henner P, Leonhardt N (2021) Disruption of AtHAK/KT/KUP9 enhances plant cesium accumulation under low potassium supply. Physiol Plant 173(3):1230–1243
Gierth M, Maser P, Schroeder JI (2005) The potassium transporter AtHAK5 functions in K(+) deprivation-induced high-affinity K(+) uptake and AKT1 K(+) channel contribution to K(+) uptake kinetics in Arabidopsis roots. Plant Physiol 137:1105–1114. https://doi.org/10.1104/pp.104.057216
Grabov A (2007) Plant KT/KUP/HAK potassium transporters: single family - multiple functions. Ann Bot 99:1035–1041. https://doi.org/10.1093/aob/mcm066
Guether M, Balestrini R, Hannah M, He J, Udvardi MK, Bonfante P (2009) Genome-wide reprogramming of regulatory networks, transport, cell wall and membrane biogenesis during arbuscular mycorrhizal symbiosis in Lotus japonicus. New Phytol 182:200–212. https://doi.org/10.1111/j.1469-8137.2008.02725.x
Gupta M, Qiu X, Wang L, Xie W, Zhang C, Xiong L, Lian X, Zhang Q (2008) KT/HAK/KUP potassium transporters gene family and their wholelife cycle expression profile in rice (Oryza sativa). Mol Genet Genom 280:437–452
Hajiahmadi Z, Abedi A, Wei H, Sun W, Ruan H, Zhuge Q, Movahedi A (2020) Identification, evolution, expression, and docking studies of fatty acid desaturase genes in wheat (Triticum aestivum L.). BMC Genom 21:778. https://doi.org/10.1186/s12864-020-07199-1
Han M, Wu W, Wu WH, Wang Y (2016) Potassium transporter KUP7 is involved in K+ acquisition and translocation in Arabidopsis root under K+-limited conditions. Mol Plant 9:437–446. https://doi.org/10.1016/j.molp.2016.01.012
He C, Cui K, Duan A, Zeng Y, Zhang J (2012) Genome-wide and molecular evolution analysis of the Poplar KT/HAK/KUP potassium transporter gene family. Ecol Evol 2:1996–2004. https://doi.org/10.1002/ece3.299
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Hyun TK, Rim Y, Kim E, Kim J-S (2014) Genome-wide and molecular evolution analyses of the KT/HAK/KUP family in tomato (Solanum lycopersicum L.). Genes Genom 36:365–374. https://doi.org/10.1007/s13258-014-0174-0
Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang H, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, dePamphilis CW (2011) Ancestral polyploidy in seed plants and angiosperms. Nature 473:97–100. https://doi.org/10.1038/nature09916
Kaya C, Ashraf M, Alyemeni MN, Ahmad P (2020) The role of endogenous nitric oxide in salicylic acid-induced up-regulation of ascorbate-glutathione cycle involved in salinity tolerance of pepper (Capsicum annuum L.) plants. Plant Physiol Biochem 147:10–20. https://doi.org/10.1016/j.plaphy.2019.11.040
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lee YH, Kim HS, Kim JY, Jung M, Park YS, Lee JS, Choi SH, Her NH, Lee JH, Hyung NI, Lee CH, Yang SG, Harn CH (2004) A new selection method for pepper transformation: callus-mediated shoot formation. Plant Cell Rep 23:50–58. https://doi.org/10.1007/s00299-004-0791-1
Leigh RA, Wyn Jones RG (1984) A hypothesis relating critical potassium concentrations for growth to the distribution and functions of this ion in the plant cell. New Phytol 97:1–13
Letunic I, Khedkar S, Bork P (2020) SMART: recent updates, new developments and status in 2020. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaa937
Li XB, Fan XP, Wang XL, Cai. Yang WC. (2009) The cotton ACTIN1 gene is functionally expressed in fibers and participates in fiber elongation. Plant Cell 17:859–875
Li J, Long Y, Qi GN, Li J, Xu ZJ, Wu WH, Wang Y (2014) The Os-AKT1 channel is critical for K+ uptake in rice roots and is modulated by the rice CBL1-CIPK23 complex. Plant Cell 26:3387–3402. https://doi.org/10.1105/tpc.114.123455
Li W, Xu G, Alli A, Yu L (2018) Plant HAK/KUP/KT K(+) transporters: function and regulation. Semin Cell Dev Biol 74:133–141. https://doi.org/10.1016/j.semcdb.2017.07.009
Liu JL, Liu JJ, Chen AQ, Ji MJ, Chen JD, Yang XF, Gu M, Qu HY, Xu GH (2016a) Analysis of tomato plasma membrane H(+)-ATPase gene family suggests a mycorrhiza-mediated regulatory mechanism conserved in diverse plant species. Mycorrhiza 26(7):645–656. https://doi.org/10.1007/s00572-016-0700-9
Liu L, Zheng C, Kuang B, Wei L, Yan L, Wang T (2016b) Receptor-like kinase RUPO interacts with potassium transporters to regulate pollen tube growth and integrity in rice. PLoS Genet 12(7):e1006085. https://doi.org/10.1371/journal.pgen.1006085
Liu J, Liu J, Liu J, Cui M, Huang Y, Tian Y, Chen A, Xu G (2019) The potassium transporter SlHAK10 is involved in mycorrhizal potassium uptake. Plant Physiol 180:465–479. https://doi.org/10.1104/pp.18.01533
Lu Y, Chanroj S, Zulkifli L et al (2011) Pollen tubes lacking a pair of K+ transporters fail to target ovules in Arabidopsis. Plant Cell 23(1):81–93. https://doi.org/10.1105/tpc.110.080499
Luan S, Lan W, Chul LS (2009) Potassium nutrition, sodium toxicity, and calcium signaling: connections through the CBL-CIPK network. Curr Opin Plant Biol 12(3):339–346. https://doi.org/10.1016/j.pbi.2009.05.003
Luan M, Tang RJ, Tang Y, Tian W, Hou C, Zhao F, Lan W, Luan S (2017) Transport and homeostasis of potassium and phosphate: limiting factors for sustainable crop production. J Exp Bot 68(12):3091–3105. https://doi.org/10.1093/jxb/erw444
Ma X, Gai WX, Qiao YM, Ali M, Wei AM, Luo DX, Li QH, Gong ZH (2019) Identification of CBL and CIPK gene families and functional characterization of CaCIPK1 under Phytophthora capsici in pepper (Capsicum annuum L.). BMC Genom 20(1):775. https://doi.org/10.1186/s12864-019-6125-z
Maathuis FJ (2006) The role of monovalent cation transporters in plant responses to salinity. J Exp Bot 57:1137–1147. https://doi.org/10.1093/jxb/erj001
Maathuis FJ (2009) Physiological functions of mineral macronutrients. Curr Opin Plant Biol 12:250–258. https://doi.org/10.1016/j.pbi.2009.04.003
Maher C, Stein L, Ware D (2006) Evolution of Arabidopsis microRNA families through duplication events. Genome Res 16:510–519. https://doi.org/10.1101/gr.4680506
Martínez-Cordero MA, Martínez V, Rubio F (2004) Cloning and functional characterization of the high-affinity K+ transporter HAK1 of pepper. Plant Mol Biol 56(3):413–421. https://doi.org/10.1007/s11103-004-3845-4
Martinez-Cordero MA, Martinez V, Rubio F (2005) High-affinity K+ uptake in pepper plants. J Exp Bot 56:1553–1562. https://doi.org/10.1093/jxb/eri150
Moscone EA, Scaldaferro MA, Grabiele M, Cecchini NM, Sánchez García Y, Jarret R, Daviña JR, Ducasse DA, Barboza GE, Ehrendorfer F (2007) The evolution of chili peppers (Capsicum Solanaceae): a cytogenetic perspectives, 745 edn. International Society for Horticultural Science (ISHS), Leuven
Mouline K, Véry AA, Gaymard F et al (2002) Pollen tube development and competitive ability are impaired by disruption of a Shaker K+ channel in Arabidopsis. Genes Dev 16(3):339–350. https://doi.org/10.1101/gad.213902
Nieves-Cordones M, Ródenas R, Chavanieu A, Rivero RM, Martinez V, Gaillard I, Rubio F (2016) Uneven HAK/KUP/KT protein diversity among angiosperms: species distribution and perspectives. Front Plant Sci 7:127
Okada T, Nakayama H, Shinmyo A, Yoshida K (2008) Expression of OsHAK genes encoding potassium ion transporters in rice. Plant Biotechnol 25:241–245. https://doi.org/10.5511/plantbiotechnology.25.241
Osakabe Y, Arinaga N, Umezawa T, Katsura S, Nagamachi K, Tanaka H, Ohiraki H, Yamada K, Seo SU, Abo M, Yoshimura E, Shinozaki K, Yamaguchi-Shinozaki K (2013) Osmotic stress responses and plant growth controlled by potassium transporters in Arabidopsis. Plant Cell 25:609–624. https://doi.org/10.1105/tpc.112.105700
Qiao X, Li M, Li L, Yin H, Wu J, Zhang S (2015) Genome-wide identification and comparative analysis of the heat shock transcription factor family in Chinese white pear (Pyrus bretschneideri) and five other Rosaceae species. BMC Plant Biol 15:12. https://doi.org/10.1186/s12870-014-0401-5
Qin C, Yu C, Shen Y, Fang X, Chen L, Min J, Cheng J, Zhao S, Xu M, Luo Y, Yang Y, Wu Z, Mao L, Wu H, Ling-Hu C, Zhou H, Lin H, Gonzalez-Morales S, Trejo-Saavedra DL, Tian H, Tang X, Zhao M, Huang Z, Zhou A, Yao X, Cui J, Li W, Chen Z, Feng Y, Niu Y, Bi S, Yang X, Li W, Cai H, Luo X, Montes-Hernandez S, Leyva-Gonzalez MA, Xiong Z, He X, Bai L, Tan S, Tang X, Liu D, Liu J, Zhang S, Chen M, Zhang L, Zhang L, Zhang Y, Liao W, Zhang Y, Wang M, Lv X, Wen B, Liu H, Luan H, Zhang Y, Yang S, Wang X, Xu J, Li X, Li S, Wang J, Palloix A, Bosland PW, Li Y, Krogh A, Rivera-Bustamante RF, Herrera-Estrella L, Yin Y, Yu J, Hu K, Zhang Z (2014) Whole-genome sequencing of cultivated and wild peppers provides insights into Capsicum domestication and specialization. Proc Natl Acad Sci USA 111:5135–5140. https://doi.org/10.1073/pnas.1400975111
Rengel Z, Damon PM (2008) Crops and genotypes differ in efficiency of potassium uptake and use. Physiol Plant 133:624–636. https://doi.org/10.1111/j.1399-3054.2008.01079.x
Ruiz-Lau N, Bojorquez-Quintal E, Benito B, Echevarria-Machado I, Sanchez-Cach LA, Medina-Lara MF, Martinez-Estevez M (2016) Molecular cloning and functional analysis of a Na+-insensitive K+ transporter of Capsicum chinense Jacq. Front Plant Sci 7:1980. https://doi.org/10.3389/fpls.2016.01980
Saito T, Kobayashi NI, Tanoi K, Iwata N, Suzuki H, Iwata NTM (2013) Expression and functional analysis of the CorA-MRS2-ALR-type magnesium transporter family in rice. Plant Cell Physiol 54(10):1673–1683. https://doi.org/10.1093/pcp/pct112
Santa-María GE, Rubio F, Dubcovsky J, Rodríguez-Navarro A (1997) The HAK1 gene of barley is a member of a large gene family and encodes a high-affinity potassium transporter. Plant Cell 9:2281. https://doi.org/10.1105/tpc.9.12.2281
Schachtman DP, Schroeder JI (1994) Structure and transport mechanism of a high-affinity potassium uptake transporter from higher plants. Nature 370:655–658. https://doi.org/10.1038/370655a0
Shabala S, Cuin TA (2008) Potassium transport and plant salt tolerance. Physiol Plant 133:651–669. https://doi.org/10.1111/j.1399-3054.2007.01008.x
Shen Y, Shen L, Shen Z, Jing W, Ge H, Zhao J, Zhang W (2015) The potassium transporter OsHAK21 functions in the maintenance of ion homeostasis and tolerance to salt stress in rice. Plant Cell Environ 38:2766–2779. https://doi.org/10.1111/pce.12586
Song T, Shi Y, Shen L, Cao CJ, Shen Y, Jing W, Tian QX, Lin F, Li WY, Zhang WH (2021) An endoplasmic reticulum-localized cytochrome b 5 regulates high-affinity K(+) transport in response to salt stress in rice. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.2114347118
Templalexis D, Tsitsekian D, Liu C et al (2021) Potassium transporter TRH1/KUP4 contributes to distinct auxin-mediated root system architecture responses. Plant Physiol. https://doi.org/10.1093/phys/kiab472
Uozumi N, Kim EJ, Rubio F, Yamaguchi T, Muto S, Tsuboi A, Bakker EP, Nakamura T, Schroeder JI (2000) The Arabidopsis HKT1 gene homolog mediates inward Na(+) currents in xenopus laevis oocytes and Na(+) uptake in Saccharomyces cerevisiae. Plant Physiol 122:1249–1259. https://doi.org/10.1104/pp.122.4.1249
Wamser AF, Cecilio Filho AB, Nowaki RHD, Mendoza-Cortez JW, Urrestarazu M (2017) Influence of drainage and nutrient-solution nitrogen and potassium concentrations on the agronomic behavior of bell-pepper plants cultivated in a substrate. PLoS ONE 12:e0180529. https://doi.org/10.1371/journal.pone.0180529
Wang Y, Wu WH (2013) Potassium transport and signaling in higher plants. Annu Rev Plant Biol 64:451–476. https://doi.org/10.1146/annurev-arplant-050312-120153
Wang Y, Wu WH (2017) Regulation of potassium transport and signaling in plants. Curr Opin Plant Biol 39:123–128. https://doi.org/10.1016/j.pbi.2017.06.006
Wang Y, Lu J, Chen D, Zhang J, Qi K, Cheng R, Zhang H, Zhang S (2018) Genome-wide identification, evolution, and expression analysis of the KT/HAK/KUP family in pear. Genome 61:755–765. https://doi.org/10.1139/gen-2017-0254
White PJ (2013) Improving potassium acquisition and utilisation by crop plants. J Plant Nutr Soil Sci 176:305–316. https://doi.org/10.1002/jpln.201200121
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591. https://doi.org/10.1093/molbev/msm088
Yang Z, Gao Q, Sun C, Li W, Gu S, Xu C (2009) Molecular evolution and functional divergence of HAK potassium transporter gene family in rice (Oryza sativa L.). J Genet Genomics 36:161–172. https://doi.org/10.1016/S1673-8527(08)60103-4
Yang T, Zhang S, Hu Y, Wu F, Hu Q, Chen G, Cai J, Wu T, Moran N, Yu L, Xu G (2014) The role of a potassium transporter OsHAK5 in potassium acquisition and transport from roots to shoots in rice at low potassium supply levels. Plant Physiol 166:945–959. https://doi.org/10.1104/pp.114.246520
Yang T, Feng H, Zhang S, Xiao H, Hu Q, Chen G, Xuan W, Moran N, Murphy A, Yu L, Xu G (2020a) The potassium transporter OsHAK5 alters rice architecture via ATP-dependent transmembrane auxin fluxes. Plant Commun 1:100052. https://doi.org/10.1016/j.xplc.2020.100052
Yang T, Lu X, Wang Y, Xie Y, Ma J, Cheng X, Xia E, Wan X, Zhang Z (2020b) HAK/KUP/KT family potassium transporter genes are involved in potassium deficiency and stress responses in tea plants (Camellia sinensis L.): expression and functional analysis. BMC Genom 21:556. https://doi.org/10.1186/s12864-020-06948-6
Zhang Z, Zhang J, Chen Y, Li R, Wang H, Wei J (2012) Genome-wide analysis and identification of HAK potassium transporter gene family in maize (Zea mays L.). Mol Biol Rep 39:8465–8473. https://doi.org/10.1007/s11033-012-1700-2
Zonia L, Munnik T (2007) Life under pressure: hydrostatic pressure in cell growth and function. Trends Plant Sci 12(3):90–97. https://doi.org/10.1016/j.tplants.2007.01.006
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (32002123), Special project on Science and Technology of Anhui Province, China (201903b06020017), Major Science and Technology Projects in Anhui Province (Grant No. 18030701214), and funded by the Biotechnology in Plant Protection of Ministry of Agriculture and Rural Affairs and Zhejiang Province.
Author information
Authors and Affiliations
Contributions
JRZ, GHQ, and JJL planned and designed the research; JRZ, JJL, CYL and JYL performed the experiments; XLL, JRZ and JZ analyzed the data; GHQ, and JJL wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, J., Qin, G., Liu, X. et al. Genome-wide identification and expression analysis of HAK/KUP/KT potassium transporter provides insights into genes involved in responding to potassium deficiency and salt stress in pepper (Capsicum annuum L.). 3 Biotech 12, 77 (2022). https://doi.org/10.1007/s13205-022-03136-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03136-z