Abstract
The giant landrace of maize Jala is a native crop cultured in Nayarit and Jalisco States in the occident of México. In this study, after screening 374 rhizospheric and endophytic bacteria isolated from rhizospheric soil, root, and seed tissues of maize Jala, a total of 16 bacterial strains were selected for their plant-growth-promoting potential and identified by 16S rRNA phylogenetic analysis. The isolates exhibited different combinations of phenotypic traits, including solubilisation of phosphate from hydroxyapatite, production of a broad spectrum of siderophores such as cobalt, iron, molybdenum, vanadium, or zinc (Co2+, Fe3+, Mo2 +, V5+, Zn2+), and nitrogen fixation capabilities, which were detected in both rhizospheric and endophytic strains. Additional traits such as production of 1-aminocyclopropane-1-carboxylate deaminase and a high-rate production of Indoleacetic Acid were exclusively detected on endophytic isolates. Among the selected strains, the rhizospheric Burkholderia sp., and Klebsiella variicola, and the endophytic Pseudomonas protegens significantly improved the growth of maize plants in greenhouse assays and controlled the infection against Fusarium sp. 50 on fresh maize cobs. These results present the first deep approach on handling autochthonous microorganisms from native maize with a potential biotechnological application in sustainable agriculture as biofertilizers or biopesticides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Jala maize (Zea mays) is a Mexican giant maize landrace cultured in the fertile volcanic valley of Jala in the State of Nayarit, Mexico (Kempton 1924). Its farming demands waterlogged soils, producing plants up to 5 m long and cobs between 45 and 60 cm long after a life cycle of 7–8 months (Rice 2007; Prasanna 2012). Outside the Jala Valley, the growth of Jala maize do not reach its normal size (Aguilar-Castillo et al. 2006). Extensive agriculture, economic pressures, and market demands have significantly reduced the number of landraces cultivated in the country and have displaced many native varieties, including Jala (Rice 2007). Moreover, intensive cropping homogenizes bacterial diversity and reduces the complexity of bacterial networks in soils (Figuerola et al. 2015; Karimi et al. 2019; Wang et al. 2020). Geographical, climatic, and edaphic differences between crop soils can be used to predict the high and specific microbial diversity associated with each native landrace, soil type, and agricultural practice (Aira et al. 2010; Bouffaud et al. 2012; Zhou et al. 2017). The conservation of the current diversity of maize landraces and their germplasm represents a valuable cultural heritage and a profitable biological resource for the development of improved varieties with suitable phenotypes. Research on maize ecology should include the study of its own germplasm and microbiome as a holobiont.
Plant-growth-promoting bacteria (PGPB) colonize plant tissues without causing any symptom of disease or competition for nutrients. PGPB facilitate nutrient uptake, participate in rhizoremediation, modulate plant stress control, reduce disease damage, antagonize soil-borne pathogens, and induce systemic resistance, among other activities (Saleem et al. 2007; Beneduzi et al. 2012; Weber et al. 2018). The culturable fraction of autochthonous PGPB associated with maize has an important biotechnological application in sustainable agriculture as biofertilizers and/or biopesticides.
Several rhizospheric and endophytic bacteria have been isolated from maize plants with potential plant-growth-promoting (PGP) characteristics such as biological nitrogen fixation, phosphate solubilization, siderophore secretion, heavy metal stress tolerance, and 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase production. Among these maize-associated bacteria, Azospirillum, Enterobacter, Brevundimonas, Pseudomonas, Pantoea, Arthrobacter, and Achromobacter are frequently described as PGPB; Bacillus and Pseudomonas are highlighted as antagonists of Fusarium species; and Klebsiella, Herbaspirillum, Burkholderia, and Rhanella are known nitrogen-fixing bacteria on grass-like plants including maize (Reis et al. 2004; Montañez et al. 2009; Pereira and Castro 2014; Douriet-Gámez et al. 2018). Due to the rhizospheric effect, the largest population of these bacteria has been found to be associated to the root, which is an environment of high metabolic activity between the plant and microorganisms (Bulgarelli et al. 2013; Gaiero et al. 2013).
A recent report on microbial endophytes from ten different varieties (landraces) of maize from Canada, United States of America, and Mexico, described an isolate of Pantoea ananatis from Jala seeds that produces auxins (Johnston-Monje and Raizada 2011). Other works have reported the biofertilization potential of a mixture of Sphingomonas sp., Azospirillum brasiliense, and Acinetobacter sp. when applied to an autochthonous blue maize from the state of Tlaxcala, Mexico (non-specified landrace), which improves plant growth under desiccation stress (Molina-Romero et al. 2017).
Although information about the endophytic bacteria of maize is extensive, few works address the bacterial diversity within native maize of Mexico, which is the origin centre, domestication, and diversification of this grass (Buckler and Stevens 2006). The purpose of this study was to evaluate the biotechnological potential of a collection of PGPB isolated from the rhizosphere and maize tissues of the native landrace Jala. This collection of culturable PGPB might harbour an important potential for species-specific biofertilization and biopesticide application on maize.
Materials and methods
Sample collection and soil characterization
Soil and maize plants were collected from the Jala municipality (14° 25.2′ 24′′ N; 21° 05.7′ 6′′ E) in the state of Nayarit, Mexico. Plants were carefully removed from soil, collected in sterile plastic bags, and stored at 4 °C for the isolation of PGPB. Soil was collected from five equidistant points within the field, using a soil profile sampler at 20–30 cm depth. Soil samples were mixed before storage at 4 °C to be later analysed. The soil had a neutral pH in water (6.97), a loamy texture, and was composed of 39.32% sand, 40.36% silt, 20.32% clay, 3.08% organic matter, 12 ppm inorganic N, 120 ppm Bray P, 478 ppm K, and 94.1 ppm Fe.
Bacterial isolation and growth conditions
Root and grain samples were disinfected for 5 min with a 2% sodium hypochlorite solution containing 100 μL of Tween-20 followed by a 3-min rinse with 70% ethanol. Samples were washed five times with sterile distilled water and aseptically macerated in a mortar. One gram of rhizospheric soil attached to the roots was washed out and processed. Direct isolation and enrichment cultivation were performed in solid and semi-solid nitrogen-free media, including Winogradsky, Bridges, Haahtela with 1% of glucose and Haahtela with 1% of mannitol and sucrose as carbon sources (Bridges 1981; Haahtela et al. 1983; Atlas 2010). Direct isolation of bacteria was performed using the serial dilution method. The enrichment cultures were obtained after inoculating 1 g of the samples into nitrogen free semi-solid agar. They were incubated at 28 °C and transferred to fresh media every 15 days for 60 days. Pure cultures were obtained from both direct isolation and enrichment cultivation and preserved at − 70 °C in a solution of 25% glycerol (v/v). All cultures were incubated under aerobic conditions at 28 °C.
DNA extraction and PCR amplification of 16S rRNA
DNA extraction was performed as described by Hoffman and Winston (1987) DNA was diluted in 20 µL of water and quantified by measuring the absorbance at 260 nm.
Bacterial isolates were identified by the analysis of the 16S rRNA sequence using the primers and the conditions reported by Relman (1993) using 100 ng of template DNA, 1 × Green GoTaq Flexi Buffer, 10 pmol L–1 of a dNTP mixture, 1 × GoTaq Flexi DNA, 10 pmol L–1 of each primer, and 3.5 mmol L–1 of MgCl2 in a final volume of 25 µL. PCR products (1.4 kb) were purified using the ZymocleanTM Gel DNA Recovery Kit (ZYMO RESEARCH, USA) and sequenced through a DNA sequencing service (Macrogen Inc., Korea).
Sequence analysis
The resultant nucleotide sequences were compared using the BLAST tool (Altschul et al. 1997) and aligned with the most similar sequences and with sequences from related species collected from the non-redundant GenBank database of 16S ribosomal RNA from the National Center for Biotechnology Information (NCBI; http://www.ncbi.nlm.nih.gov).
All the sequences were aligned and manually edited with SEAVIEW 3.2 software (Galtier et al. 1996). The most appropriate model of nucleotide substitution was selected using the software jModelTest 2.0 (Darriba et al. 2012). The similarity percentage between sequence pairs was calculated using MatGAT software (Campanella et al. 2003). All sequences were submitted to the GenBank database.
Characterization of isolates as PGPB
Phosphate solubilization
The phosphate solubilization capacities of the strains were tested on the National Botanical Research Institute Phosphate (NBRIP) agar medium, which uses Ca3(PO4)2 as a sole source of insoluble phosphate (Mehta and Nautiyal 2001). The agar plates were inoculated with 5 µL of bacterial solution adjusted to an optical density at 600 nm (OD600) of 0.8. After incubation at 28 °C for 3 days, the P-solubilizing halo zone was measured, and the phosphate solubilization index (PSI) was calculated as the ratio between the diameter of the halo and the diameter of the colony.
The quantitative phosphate solubilization test was performed by the modified molybdenum blue method (He and Honeycutt 2005; Bashan et al. 2013) with hydroxyapatite as the sole source of insoluble phosphorus.
Siderophore production
Siderophore production was detected following the universal chrome azurol S (CAS) agar plate assay (Schwyn and Neilands 1987) supplemented with five different ions from the inorganic salts: ferric sulfate Fe2(SO4)3, copper sulfate CuSO4, sodium molybdate Na2MoO4, sodium metavanadate NaVO3 and zinc sulfate ZnSO4. CAS-blue agar plates were inoculated with a spot of 5 µL of a bacterial suspension adjusted to 0.8 (OD600) and incubated at 28 °C for 48 h. The positive production of siderophore was detected by the production of a yellow halo around the bacterial growth. The ion capture index was calculated as the PSI.
Quantification of indoleacetic acid (IAA)
Bacterial indole-3-acetic acid production was estimated by a colorimetric assay using Salkowski’s reagent (35% HClO4, 0.01 mol L–1 FeCl3), which is a common method for indole-type molecule estimation, including auxins (Patten and Glick 1996). The isolates were grown in Tryptic Soy Broth (TSB) and TSB supplemented with L-tryptophan as a precursor of IAA (Gordon and Weber 1951; Leveau and Lindow 2005). The isolates were incubated at 28 °C for 4 days. The supernatant was obtained after 15 min of centrifugation at 18,516×g. The reaction was performed by mixing 0.5 mL of the supernatant with 1 mL of Salkowski’s reagent. The mixture was incubated in darkness for 30 min. The absorbance was estimated at 530 nm, and the IAA concentration was determined with an IAA standard curve (Szkop et al. 2012).
ACC deaminase activity
Isolates were tested to growth on a medium with 1-aminocyclopropane-1-carboxylic acid (ACC) as the sole source of nitrogen. Liquid Glucose Ivo medium (LGI) broth was supplemented with 1 µL of 2 mol L–1 ACC solution prepared in deionized water and sterilized by filtration (Johnston-Monje and Raizada 2011). Non-supplemented LGI broth was used as a control of nitrogen fixation activity. A total of 10 µL of the bacterial suspension (OD600 of 0.8) was added to 190 µL of the LGI broth in a sterile 96-well plate. The plate was incubated at 28 °C for 10 days, and the optical density (OD600) was measured every 24 h. Those isolates growing exclusively in supplemented LGI media were considered to display a deaminase activity for the substrate ACC.
In vitro nitrogen-fixation activity
The nitrogen-fixation activity of the isolates was measured by the acetylene reduction assay (ARA) using Haahtela’s medium. Isolates were grown in closed vessels (10 mL) containing 5 mL of semi-solid media with 50 µL of bacterial suspension (OD600 of 0.8). The tubes were sealed, and after 72 h of incubation at 28 °C, 1 mL (v/v) of the head atmosphere of the test tube was exchanged with an equal volume of acetylene gas, followed by another 24 h of incubation at 28 °C. Closed vessels without bacteria and vessels inoculated with Klebsiella variicola 6A3 were used as negative and positive controls, respectively.
The ethylene concentration was measured on a Varian 3300 (Walnut Creek, CA, USA) gas chromatograph fitted with a hydrogen flame ionization detector and a 3.2 mm × 2 m stainless steel column packed with Poropak N (80–100 mesh). The column was operated isothermally at 95 °C with nitrogen as the carrier gas at a flow rate of 30 mL min− 1. The detector temperature was 105 °C. Ethylene production was estimated by integration of the area under the curve and transformed into nmol of ethylene h−1 units.
PCR amplification of the nifD gene
Based on the identity of bacteria from this work, a pair of primers nifD-JalaF (5′-CGBGGCTGCGCCTAYGC-3′) and nifD-JalaR (5′-CGGGCRAAGATGGCGAAGC-3′) was designed to amplify nifD genes present among the isolates from the collection, using the following conditions: 1 min at 94 °C; 1 min at 55 °C; and 1 min at 72 °C for 35 cycles, using 100 ng of template DNA, 10 pmol L−1 of each primer and 3.0 mmol L–1 of MgCl2. The expected size of the amplicon was approximately 1.2 kb. PCR products were purified, sequenced, and analysed.
Plant growth promotion effect on maize
A total of 12 relevant PGPB strains belonging to five species were tested on “Conejo” maize plants. Bacterial suspensions (OD600 of 1.0) were applied as a seed drench to surface sterilized seeds at a ratio of 10 mL per 20 seeds. Sterile distilled water was used as the untreated negative control.
Seedlings were grown in a 1-kg plastic bag with vermiculite, under a 12.5 h light/11.5 dark cycle in greenhouse conditions at an average temperature of 20 °C. The seedlings were watered every 2 days with 150 mL per bag of Hoagland solution (Hoagland and Arnon 1950). After 50 days, growth parameters such as height and dry weight of stems and roots were measured. Each experiment included three plants per bacterial strain treatment with three replications.
Statistical analysis
Indoleacetic quantification and phosphate solubilisation were expressed as average calculation and its standard deviation calculations. Differences between treatment data from plant height and weight were determined using a two-way ANOVA test followed by a Dunnett’s multiple comparison test using GraphPad Prism software version 6.00 for Windows (GraphPad Software, La Jolla California USA; www.graphpad.com).
Detection of antagonistic isolates
Antagonistic effects against the maize fungal pathogen Fusarium sp. 50 was tested through fungi–bacteria dual culture. Forty-eight-hour bacterial cultures were harvested from Tryptic Soy Agar (TSA) and adjusted to an OD600 of 0.8 with sterile distilled water. Bacterial suspension spots of 5 µL were placed peripherally on a Yeast Peptone Dextrose (YPD) plate. A 5 mm cylinder taken from the mycelial fungal growth plate was placed at the centre of a YPD plate. Plates were incubated at 28 °C for 96 h. Fungal growth inhibition zones indicated an antagonistic effect of bacteria against fungi.
Maize protection against fungal infection
The potential antagonist capabilities of PGPB against Fusarium sp. 50 were evaluated in a maize cob model. Fresh maize cobs (Zea mays var. Conejo) were surface disinfected as described above. A total of 10 μL of bacterial suspensions adjusted to an OD600 of 1.0 (approximately 8 × 106 CFU mL–1) were injected in four equidistant points to each cob and incubated in plastic boxes for 24 h at room temperature. After incubation, conidial suspensions (105 per 10 μL) of Fusarium sp. 50 were inoculated at the centre of a cob and marked with a toothpick. Cobs were incubated at room temperature and monitored daily until the appearance of signs of fungal infection. Each experiment included three cobs per treatment.
Results
Soil characterization
Soil of Jala is a neutral substrate (pH of 6.6–7.3) with a high saturation point (52%), which is a desirable feature according to the maize crop demands. This soil is fertile and rich in organic matter, nitrogen, phosphorus, and other minerals. It presents a low level of salinity (pH = 7.05, CE = 0.74 dS m–1, RAS = 1.07, and PSI = 1.31). These characteristics explain why this soil is suitable to sustain the cultivation of maize.
Isolation of endophytic and rhizospheric PGPB from maize Jala
A total of 374 strains were isolated from the soil, grain, and roots of 10 Jala maize landrace plants (Table 1). Among the isolates, 2 rhizospheric: Klebsiella variicola R3J3HD7 (MK855126), Burkholderia sp. R3J3HD10 (MK855131); 5 grain endophytic: Herbaspirillum seropedicae E2WL3 (MK855130), Pantoea sp. E2K4 (MK855135), Pseudomonas protegens E1BL2 (MK855121), Pseudomonas protegens E2HL9 (MK855122), Rhizobium pusense E2K3 (MK855124); and 9 root endophytic strains: Achromobacter xylosoxidans Z2K8 (MK855127), Achyromobacter xylosoxidans Z3AD5a (MK855134), Burkholderia sp. Z2K9 (MK855132), Chryseobacterium gleum Z1HL3 (MK855133), Kosakonia sp. Z2WD1 (MK855128), Pantoea sp. Z2WD2 (MK855136), Phytobacter diazotrophicus Z2WL1 (MK855129), Pseudomonas alcaligenes Z1K6 (MK855125), Pseudomonas japonica Z2K3 (MK855123) were selected by their PGP capabilities and identified by 16S rRNA gene analysis. The identity percentages of the isolates are shown in Table 2.
Isolate characterization as potential PGPB
Phosphate solubilization
All 374 isolates were cultured in NBRIP, and the isolates that displayed a solubilization halo were cultured in broth media to confirm and quantify the solubilization rate. Consistent semi-qualitative and quantitative results of phosphate solubilization were found for the strains Pantoea sp. Z2WD2, P. protegens E2HL9, Pantoea sp. E2K4, P. japonica Z2K3, H. seropedicae E2WL3, P. protegens E1BL2, Kosakonia sp. Z2WD1, and P. diazotrophicus Z2WL1 (Table 3). The rhizospheric strains Kosakonia sp. Z2WD1 and Pantoea sp. Z2WD2 were the most efficient phosphate solubilizers (Fig. 1).
Phosphate-solubilizing activity of bacteria isolated from Jala maize landrace. Production of clear halo zones of solubilization on NBRIP agar plates by Pantoea sp. E2K4. Weak halos formed by Pseudomonas protegens E1BL2 and Phytobacter diazotrophicus Z2WL1 strains, and no halo was observed with Rhizobium pusense E2K3, which was used as the negative control
Siderophore production
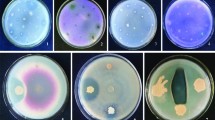
Almost all 374 strains were able to secrete some type of siderophore. However, A. xylosoxidans Z2K8 and Z3AD5a, P. japonica Z2K3, Chryseobacterium gleum Z1HL3, and P. protegens E2HL9 and E1BL2 strains exhibited higher ion-chelating capabilities (Table 3) (Fig. 2).
Detection of siderophore production by colour change on CAS-blue agar supplemented with five different ions (Co2+, Fe3+, Mo2+, V5+, Zn2+) after 48 h of incubation by the strains (clockwise sense) (1) Klebsiella variicola R3J3HD7, (2) Achromobacter xylosoxidans Z3AD5a, (3) Pseudomonas japonica Z2K3, (4) Achromobacter xylosoxidans Z2K8, (5) Pseudomonas protegens E2HL9, (6) Pseudomonas protegens E1BL2, (7) Pantoea sp. Z2WD2 and (8) Chryseobacterium gleum Z1HL3
Auxin production (IAA)
Auxin production was quantified at the stationary growth phase. More than 50 isolates demonstrated IAA production. Pantoea sp. Z2WD2, E2K4, and P. diazotrophicus Z2WL1 were the best producers of auxins, accumulating more than 20 µg mL–1 in the culture media. All the strains could produce IAA with and without the addition of tryptophan; among them, P. diazotrophicus Z2WL1 doubled its production with the addition of tryptophan, while H. seropedicae E2WL3 produced more auxin-like molecules in medium without tryptophan (Table 3).
A total of 16 strains with the best potential PGP phenotypes (IAA and siderophore production and phosphate solubilization) were selected for evaluation under a second stage of tests that included the ACC deaminase activity test, ARA assay, and amplification and sequencing of nifD genes.
Growth in ACC as a sole nitrogen source
Pantoea sp. Z2WD2, P. protegens E1BL2, R. pusense E2K3, Pantoea sp. E2K4, Burkholderia sp. Z2K9, and A. xylosoxidans Z3AD5a and Z2K8 grew on ACC as a sole nitrogen source. K. varicola R3J3HD7 is a diazotrophic bacteria that grew in the control medium without any nitrogen source. For this reason, its deamination activity cannot be recognized (Fig. 3).
Growth kinetics of the utilization of ACC as a sole nitrogen source by bacteria isolated from Jala maize landrace. Panels A1 to 4 LGI medium supplemented with ACC as the sole source of nitrogen. Panels B1 to 4 show bacteria growing on LGI medium without any source of nitrogen (nitrogen fixation control)
In vitro nitrogen-fixation activity
A total of four strains were able to fix nitrogen in vitro; however, only H. seropedicae and K. variicola R3J3HD7 exhibited acetylene reduction rates higher than 19 nmol of ethylene h–1 (Table 4).
Amplification, sequencing, and phylogeny of the nifD gene
The gene nifD was amplified on H. seropedicae E2WL3, P. diazotrophicus Z2WL1, and P. protegens E2HL9 strains from root or grain endophytes and K. variicola R3J3HD7 from the rhizosphere. All the ARA-positive isolates harboured nifD genes (Table 4) except for the P. protegens isolate E2HL9, which was unable to reduce acetylene under the assayed conditions.
Plant growth promotion of bacteria on Conejo landrace maize
Differences within the inoculated plants were measured and registered. The plant-growth-promoting (PGP) effect of the strains was evident after 50 days in a greenhouse. The Burkholderia sp. isolate Z2K9, P. protegens E2HL9 and E1BL2, K. variicola R3J3HD7, and H. seropedicae E2WL3 significantly increased the length of the roots, leaves, and dry weight of entire maize plants when compared to the uninoculated control (Fig. 4).
Growth promotion assay on maize plants from the landrace “Conejo” inoculated individually with either endophytic or rhizospheric PGPB isolated from Jala maize. A Maize plants of 50 days inoculated with PGPB under the following scheme (from left to right): non-inoculated plant, K. variicola R3J3HD7, P. protegens E2HL9, P. protegens E1BL2, Burkholderia sp. R3J3HD10, and H. seropedicae E2WL3. B Dry weight of the maize plants on 50th day. C Stem and root length of treated plants after 50 days. Data analysis was performed using GraphPad PRISM statistics program. Differences between treatments and control means were determined using two-way ANOVA with multiple comparison (*p < 0.05)
Antagonistic interaction tests
P. protegens E1BL2 and E2HL9, K. variicola R3J3HD7, and Burkholderia sp. R3J3HD10 demonstrated the broadest and highest growth control of Fusarium sp. 50 in vitro. In addition, P. protegens E1BL2 and Burkholderia sp. R3J3HD10 protected Conejo maize cobs from infection and rot by Fusarium sp. 50 (Fig. 5).
Antagonistic interaction of Fusarium sp. 50 with the bacterial isolates Burkholderia sp. R3J3HD10, K. variicola R3J3HD7, and two strains of P. protegens E1BL2 and E2HL9. A Growth of Fusarium sp. 50 in potato dextrose agar (PDA). B Inhibition of fungal growth by P. protegens E1BL2 and E2HL9. C Inhibition of fungal growth by Burkholderia sp. R3J3HD10 and K. variicola R3J3HD7. D Maize cob protection assay on “Conejo” maize cobs inoculated with PGPB, after 7 days of infection with Fusarium sp. 50. Toothpicks mark the inoculation point of Fusarium sp. 50
Discussion
In this study, the PGP potential of endophytic and rhizospheric bacteria associated with Jala landrace native Mexican maize was explored as an effort to highlight the importance and potential of the maize-bacteria symbiosis. The soil where the Jala maize grows has been described as poor with low N, K, and organic matter contents and low cation-exchange capacity (Aguilar-Castillo et al. 2006). Though, in this work, the samples of Jala maize soils showed silt, fertile, and well-structured soil with low salinity and high water-holding ability. These differences are possibly derived from the differences between the sites, the time of year, and the sampling strategies.
The isolation of bacteria performed in this work involved the use of three solid media without a nitrogen source to promote the biased isolation of nitrogen-fixing bacteria. However, many other non-nitrogen-fixing bacteria were isolated in these “nitrogen-free media,” probably, because agar contains between 1 and 178 mmol L–1 (0.1%) of total nitrogen (Scholten and Pierik 1998). The nitrogen content in aboveground and belowground tissues of grasses range be between 0.2 and 1.1% (Zhang et al. 2014), but this nitrogen source is not easily available for endophytic bacteria, which face a restrictive environment of assimilable inorganic or organic nitrogen sources. This could explain the high frequency of isolation of diverse endophytic/diazotrophic and non-diazotrophic bacteria from maize plant tissues (Baldani et al. 2014; Ji et al. 2014; Pérez-Montaño et al. 2014).
The nitrogen-fixing capability of the isolated bacteria was confirmed by PCR detection and sequencing of nifD genes and by the acetylene reduction test, which is an indirect test to demonstrate nitrogen fixation. Diazotrophic bacteria from both rhizosphere soil and plant tissues were isolated and identified to the genus or species level by phylogenetic and similitude analyses. Though there is no consensus on the threshold for differentiating bacterial species, in this work a nucleotide similitude in the 16S rRNA sequence above ≥ 98.65% was used to define the species (Kim et al. 2014). Klebsiella variicola R3J3HD7 and Herbaspirillum seropedicae E2WL3 had high acetylene reduction rates in vitro, and Pantoea sp. E2K4 and Phytobacter diazotrophicus Z2WL1 had more modest rates under the assayed conditions. These quantitative differences are not important and have no relevant biological meaning, because these in vitro estimations depend on the nitrogen-free media used and the environmental conditions assayed to find the best expression of diazotrophic phenotype (Rao 1978; Raven 1988; Morales-Jiménez et al. 2013).
Among the strains associated with Jala maize, rhizospheric K. variicola R3J3HD7 showed important in vitro PGP features, the growth promotion of “Conejo” maize landrace, and the highest rate of ethylene production. This diazotrophic enterobacteria of the Klebsiella pneumoniae complex has also been frequently found in the rhizosphere and as an endophyte of roots, stems, and leaves of sugarcane, banana, rice, and maize, among other plants (Wei et al. 2014; Lin et al. 2015; Reyna-Flores et al. 2018).
The endophytic isolates Pantoea sp. E2K4 and Z2WD2 are located in the Pantoea ananatis/Pantoea allii clade. Both PGPB species exhibited prominent features, including siderophore and IAA production, ACC utilization as a sole nitrogen source, phosphate solubilization, and growth promotion of “Conejo” maize landrace, additionally Pantoea sp. E2K4 was also able to reduce acetylene and harboured the nifD gene. P. ananatis has been reported mainly as a phytopathogen of a wide variety of hosts, such as maize, rice, onion, and eucalyptus, but as PGPB of maize, potato, and pepper (Coutinho and Venter 2009; Brady et al. 2011; Ikeda et al. 2013). The strains isolated in our study were not phytopathogenic for maize in the greenhouse assay. Instead, they exhibited a PGP phenotype. The P. ananatis pan-genome harbour a sizeable accessory genome of 1690 protein coding sequences, which reveals a wide intraspecific diversity and multiple niche occupation (Weller-Stuart et al. 2017).
The endophytic strain H. seropedicae E2WL3 was isolated from grains and displayed several in vitro and greenhouse PGP properties including nitrogen fixation, IAA and siderophore production, phosphate solubilization, and growth promotion of “Conejo” maize landrace. All these phenotypic traits have been commonly reported in H. seropedicae isolated from tissues or the rhizosphere of maize (Balsanelli et al. 2016; da Fonseca Breda et al. 2019; Dall'Asta et al. 2019).
Among the strains in this study, the highest yield of phosphate solubilization in the hydroxyapatite assay was detected in the root endophyte Kosakonia sp. Z2WD1 (142 mg mL–1). After nitrogen, phosphorus is the most important plant nutrient and the phosphate solubilization capacity is a common phenotype feature among rhizospheric isolates. The low similitude of the 16S RNA gene of the strain with other formally described species suggests that this strain is a new species within the genus. Other soil, endophytic, phyllospheric, and rhizospheric Kosakonia species stand out for their ability to promote plant growth through several mechanisms, including nitrogen fixation, inorganic phosphate solubilization, mineralization of organic phosphate, siderophore production, and auxin synthesis and efflux. Strains of Kosakonia radicincitans, Kosakonia arachidis, Kosakonia oryzae, Kosakonia sacchari, and Kosakonia pseudosacchari have been isolated from wheat, maize, groundnut, rice, sugar cane, banana, and yerba mate (Peng et al. 2009; Madhaiyan et al. 2010; Witzel et al. 2012; Zhu et al. 2013; Bergottini et al. 2015; Kämpfer et al. 2016; Cruz Barrera et al. 2019). To our knowledge, only one K. radicincitans strain was associated with bacterial wilt in banana plants (Suhaimi et al. 2014). Although most species of Kosakonia harbour functional nif genes and fix nitrogen (Madhaiyan et al. 2010; Witzel et al. 2012; Zhu et al. 2013), any nif gene nor acetylene reduction activity was detected for the strain of Kosakonia sp. Z2WD1.
Several strains of three species of Pseudomonas were isolated from grain and root maize tissues, but only P. protegens E2HL9 and E1BL2 expressed PGP features. Both P. protegens E2HL9 and E1BL2 strains isolated from grain tissue exhibited notable PGP characteristics and fungal antagonism against Fusarium sp. 50, but they were unable to fix nitrogen. P. protegens and many other Pseudomonas species have never been recognized as nitrogen-fixing bacteria, only Pseudomonas azotifigens and P. stutzeri A15 harbour an entire set of functional nitrogen-fixation genes that may have been horizontally acquired from other diazotrophs (Desnoues et al. 2003; Yan et al. 2008). The ability of both strains to produce siderophores has also been recognized in other Pseudomonas species (Sharma and Johri 2003). The main function of the siderophores pyoverdine and enantio-pyochelin of P. protegens is iron scavenging, but they also increase the survival of the strains in soil, confer resistance to fungal fusaric acid, and promote competence for iron uptake with other bacteria (Ruiz et al. 2015; Drehe et al. 2018). The strain P. protegens E1BL2 exhibited antifungal properties against Fusarium sp. 50, both in vitro and in a cob infection model, and produced HCN (data not shown), a volatile antifungal agent. Actual evidence suggests that P. protegens could be worth exploitation as an efficient biocontrol agent to reduce citrus canker caused by Xanthomonas citri subsp. citri. (Michavila et al. 2017).
Although root-associated Phytobacter diazotrophicus Z2WL1 showed some phenotypic traits usually associated with PGPB, such as being able to reduce acetylene and harbour the nifD gene, it did not promote the growth of maize plants in any of the greenhouse tests. There is not much information about this species but a report in wild rice and the sequenced genome of an isolate from the oil palm without further details (Salha et al. 2020; Zhang et al. 2008). For these reasons, more studies are needed to evaluate the importance of this species isolated from plants.
Rhizobium pusense E2K3 isolated from tissue grain exhibited modest PGP in vitro features, but the nifD gene and acetylene reduction were not detected. This species (syn. Agrobacterium pusense) has been isolated from bean nodules and chickpea rhizosphere (Panday et al. 2011; Aguilar et al. 2016) and was reported as the main human pathogen within the genus Agrobacterium/Rhizobium (Aujoulat, et al. 2015). For this reason, it was not considered for the PGP experiments in maize. On the other hand, neither Chryseobacterium gleum Z1HL3 nor Achromobacter xylosoxidans Z2K8 and Z3AD5a strains were included in the maize growth promotion experiments due to the frequent association of these species with human infections, and their PGP potential is scarce (Swenson and Sadikot 2015; Arouna et al. 2017).
Burkholderia sp. Z2K9 and Burkholderia sp. R3J3HD10 are two closely related strains. Both strains expressed some PGP features in vitro, but only Burkholderia sp. R3J3HD10 was able to promote maize growth in the greenhouse assay and to antagonize Fusarium sp. 50. There are many papers that report the fungal antagonism and/or PGP capacities of Burkholderia spp. in many plants, including maize and teosinte of non-identified landraces (Ikeda et al. 2013; Zhao et al. 2014; Pande et al. 2017; Esmaeel et al. 2018; Tagele et al. 2018, 2019; You et al. 2020). Burkholderia sp. R3J3HD10 was closely related to Burkholderia contaminans, a PGPB strain isolated from the rhizospheric soil of maize reported to control banded leaf and sheath blight of maize caused by Rhizoctonia solani in greenhouse assays (Tagele et al. 2018). The potential of Burkholderia spp. in agriculture is a debated subject due to the disadvantages reported by its pathogenic potential for humans despite its clear PGP and antifungal suppression phenotype (Eberl and Vandamme 2016; Rosenblueth et al. 2018).
This concern is not exclusive for Burkholderia spp. for practically all the species isolated in this work, have been associated to clinical cases from immunocompromised and elderly patients (Aujoulat et al. 2015; Bhatti et al. 2017; Pillonetto et al. 2018; Rodríguez-Medina et al. 2019; Mertschnigg et al. 2020). The potential pathogenic features, the molecular epidemiology registers, and the high resemblance among strains isolated from plants and human infections, support the idea of a cross-kingdom bacteria that might follow a route of transmission from plants to humans (Barrios-Camacho et al. 2019). Although clinical cases of these species vary from very rare to infrequent, it has been proposed that their use in agriculture should not be encouraged (Rosenblueth et al. 2018). In our opinion, since there are few cases of human infections related to PGPB species from agricultural origin, the potential use as a biofertilizer, bioinoculant, or antagonistic of each species must be evaluated on a case-by-case basis through a comparative whole-genome analysis to identify putative virulence factors that discern the plant beneficial strains from potential human pathogenic strains (Mertschnigg et al. 2020). In general, many potential virulence genes associated to genomic islands have been detected only in clinical strains (Faoro et al. 2019). In the future, pan-genomic and pathogenesis tests in animal models should be mandatory, to determine the strains innocuousness before incorporating them into a bioinoculants formulation. In this study, maize plants responded both positively and neutrally to individual bacterial inoculation, discarding the possibility of handling pathogenic bacteria. The main PGPB in the greenhouse assays were Burkholderia sp. R3J3HD10, H. seropedicae E2WL3, Pantoea sp. Z2WD2, P. protegens E2HL9, P. protegens E1BL2, and K. variicola R3J3HD7.
If the plant is considered a holobiont, it could be expected that its microbiome is also diverse and represents an important potential as a PGP resource, as demonstrated in this study. Face to the obstacles derived from the social opposition to the use of genetically modified organisms, culturable bacteria represents an important reserve of germplasm that can contribute to a more sustainable agriculture.
Conclusions
Jala landrace is colonized by bacteria that display PGP features, such as nitrogen fixation, phosphate solubilization, utilization of ACC as a sole nitrogen source, and siderophore and auxin production. These features were detected for both rhizospheric and endophytic bacteria except for the production of ACC deaminase, found exclusively in the endophytic bacteria from grain and root tissue. Most of the outstanding strains were isolated from root and grain tissues and gather more than one PGP ability. Maize plants responded both positively and neutrally to the individual bacterial inoculation, discarding the possibility of having isolated pathogenic bacteria. Under greenhouse conditions, maize plants growth is improved by Burkholderia sp. R3J3HD10, H. seropedicae E2WL3, Pantoea sp. Z2WD2, P. protegens E2HL9, P. protegens E1BL2, and K. variicola R3J3HD7.
Availability of data and material
Not applicable for this section.
Accession Numbers: Uploaded to the National Center for Biotechnology Information database (NCBI): K855127—Achromobacter xylosoxidans Z2K8; MK855134—Achyromobacter xylosoxidans Z3AD5a; MK855132—Burkholderia sp. Z2K9; MK855131—Burkholderia sp. R3J3HD10; MK855133—Chryseobacterium gleum Z1HL3; MK855130—Herbaspirillum seropedicae E2WL3; MK855126—Klebsiella variicola R3J3HD7; MK855128—Kosakonia sp. Z2WD1; MK855135—Pantoea sp. E2K4; MK855136—Pantoea sp. Z2WD2; MK855129—Phytobacter diazotrophicus Z2WL1; MK855121—Pseudomonas protegens E1BL2; MK855122—Pseudomonas protegens E2HL9; MK855125—Pseudomonas alcaligenes Z1K6; MK855123—Pseudomonas japonica Z2K3; MK855124—Rhizobium pusense E2K3.
Code availability
Not applicable for this section.
References
Aguilar A, Peralta H, Mora Y, Díaz R, Vargas-Lagunas C, Girard L, Mora J (2016) Genomic comparison of Agrobacterium pusense strains isolated from bean nodules. Front Microbiol 7:1720. https://doi.org/10.3389/fmicb.2016.01720
Aguilar-Castillo JA, Carballo-Carballo A, Castillo-González F, Santacruz-Varela A, Mejía-Contreras JA, Crossa-Hiriartte J, Baca-Castillo G (2006) Diversidad fenotípica y variantes distintivas de la raza Jala de maíz. Agric Técnica en México 32:57–66. http://www.scielo.org.mx/scielo.php?script=sci_arttext&pid=S0568-25172006000100006&lng=es&tlng=es
Aira M, Gómez-Brandón M, Lazcano C, Bååth E, Domínguez J (2010) Plant genotype strongly modifies the structure and growth of maize rhizosphere microbial communities. Soil Biol Biochem 42:2276–2281. https://doi.org/10.1016/j.soilbio.2010.08.029
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Arouna O, Deluca F, Camara M, Fall B, Fall B, Ba Diallo A, Docquier JD, Mboup S (2017) Chryseobacterium gleum in a man with prostatectomy in Senegal: a case report and review of the literature. J Med Case Rep 11:1–5. https://doi.org/10.1186/s13256-017-1269-4
Atlas RM (2010) Handbook of microbiological media, 4th edn. CRC Press, Boca Raton
Aujoulat F, Marchandin H, Zorgniotti I, Masnou A, Jumas-Bilak E (2015) Rhizobium pusense is the main human pathogen in the genus Agrobacterium/Rhizobium. Clin Microbiol Infect 21:472.e1–472.e5. https://doi.org/10.1016/j.cmi.2014.12.005
Baldani JI, Reis VM, Videira SS, Boddey LH, Divan Baldani VL (2014) The art of isolating nitrogen-fixing bacteria from non-leguminous plants using N-free semi-solid media: a practical guide for microbiologists. Plant Soil 384:413–431. https://doi.org/10.1007/s11104-014-2186-6
Balsanelli E, Tadra-Sfeir MZ, Faoro H, Pankievicz VCS, de Baura VA, Pedrosa FO, de Souza EM, Dixon R, Monteiro RA (2016) Molecular adaptations of Herbaspirillum seropedicae during colonization of the maize rhizosphere. Environ Microbiol 18:2343–2356. https://doi.org/10.1111/1462-2920.12887
Barrios-Camacho H, Aguilar-Vera A, Beltran-Rojel M, Aguilar-Vera E, Duran-Bedolla J, Rodriguez-Medina N, Lozano-Aguirre L, Perez-Carrascal OM, Rojas J, Garza-Ramos U (2019) Molecular epidemiology of Klebsiella variicola obtained from different sources. Sci Rep 9:1–10. https://doi.org/10.1038/s41598-019-46998-9
Bashan Y, Kamnev AA, De-Bashan LE (2013) A proposal for isolating and testing phosphate-solubilizing bacteria that enhance plant growth. Biol Fertil Soils 49:1. https://doi.org/10.1007/s00374-012-0756-4
Beneduzi A, Ambrosini A, Passaglia LMP (2012) Plant growth-promoting rhizobacteria (PGPR): their potential as antagonists and biocontrol agents. Genet Mol Biol 35:1044–1051. https://doi.org/10.1590/S1415-47572012000600020
Bergottini VM, Filippidou S, Junier T, Johnson S, Chain PS, Otegui MB, Zapata PD, Junier P (2015) Genome sequence of Kosakonia radicincitans strain YD4, a plant growth-promoting rhizobacterium isolated from yerba mate (Ilex paraguariensis St. Hill.). Genome Announc. https://doi.org/10.1128/genomeA.00239-15
Bhatti MD, Kalia A, Sahasrabhojane P, Kim J, Greenberg DE, Shelburne SA (2017) Identification and whole genome sequencing of the first case of Kosakonia radicincitans causing a human bloodstream infection. Front Microbiol 8:62. https://doi.org/10.3389/fmicb.2017.00062
Bouffaud ML, Kyselková M, Gouesnard B, Grundmann G, Muller D, Moënne-Loccoz Y (2012) Is diversification history of maize influencing selection of soil bacteria by roots? Mol Ecol 21:195–206. https://doi.org/10.1111/j.1365-294X.2011.05359.x
Brady CL, Goszczynska T, Venter SN, Cleenwerck I, de Vos P, Gitaitis RD, Coutinho TA (2011) Pantoea ananatis sp. nov., isolated from onion plants and seed. Int J Syst Evol Microbiol 61:932–937. https://doi.org/10.1099/ijs.0.022921-0
Bridges JR (1981) Nitrogen-fixing bacteria associated with bark beetles. Microb Ecol 137:131–137. https://doi.org/10.1007/BF02032495
Buckler IV, ES, Stevens NM (2006) Maize origins, domestication, and selection. In Motley TJ, Zerega N, Cross H (ed) Darwin's harvest: New approaches to the origins, evolution, and conservation of crops, Columbia University Press, pp 67–90
Bulgarelli D, Schlaeppi K, Spaepen S, van Themaat EVL, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838. https://doi.org/10.1146/annurev-arplant-050312-120106
Campanella JJ, Bitincka L, Smalley J (2003) MatGAT: an application that generates similarity/identity matrices using protein or DNA sequences. BioMed Cent 4:1–4. https://doi.org/10.1186/1471-2105-4-29
Coutinho TA, Venter SN (2009) Pantoea ananatis: an unconventional plant pathogen. Mol Plant Pathol 10:325–335. https://doi.org/10.1111/j.1364-3703.2009.00542.x
Cruz Barrera M, Jakobs-Schoenwandt D, Gómez MI, Becker M, Patel AV, Ruppel S (2019) Salt stress and hydroxyectoine enhance phosphate solubilisation and plant colonisation capacity of Kosakonia radicincitans. J Adv Res 27:91–97. https://doi.org/10.1016/j.jare.2019.03.012
da Fonseca Breda FA, da Silva TFR, Dos Santos SG, Alves GC, Reis VM (2019) Modulation of nitrogen metabolism of maize plants inoculated with Azospirillum brasilense and Herbaspirillum seropedicae. Arch Microbiol 201:547–558. https://doi.org/10.1007/s00203-018-1594-z
Dall’Asta P, Velho AC, Pereira TP, Stadnik MJ, Arisi ACM (2019) Herbaspirillum seropedicae promotes maize growth but fails to control the maize leaf anthracnose. Physiol Mol Biol Plants 25:167–176. https://doi.org/10.1007/s12298-018-0616-2
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772. https://doi.org/10.1038/nmeth.2109
Desnoues N, Lin M, Guo X, Ma L, Carreño-Lopez R, Elmerich C (2003) Nitrogen fixation genetics and regulation in a Pseudomonas stutzeri strain associated with rice. Microbiology 149:2251–2262. https://doi.org/10.1099/mic.0.26270-0
Douriet-Gámez NR, Maldonado-Mendoza IE, Ibarra-Laclette E, Blom J, Calderón-Vázquez CL (2018) Genomic analysis of Bacillus sp. strain B25, a biocontrol agent of maize pathogen Fusarium verticillioides. Curr Microbiol 75:247–255. https://doi.org/10.1007/s00284-017-1372-1
Drehe I, Simonetti E, Ruiz JA (2018) Contribution of the siderophores pyoverdine and enantio-pyochelin to fitness in soil of Pseudomonas protegens Pf-5. Curr Microbiol 75:1560–1565. https://doi.org/10.1007/s00284-018-1560-7
Eberl L, Vandamme P (2016) Members of the genus Burkholderia: good and bad guys. Research. https://doi.org/10.12688/f1000research.8221.1
Esmaeel Q, Sanchez L, Robineau M, Dorey S, Clément C, Jacquard C, Barka EA (2018) Draft genome sequence of plant growth-promoting Burkholderia sp. strain BE12, isolated from the rhizosphere of maize. Genome Announc 6:e00299-e318. https://doi.org/10.1128/AEM.67.11.5285-5293.2001
Faoro H, Oliveira WK, Weiss VA, Tadra-Sfeir MZ, Cardoso RL, Balsanelli E, Brusamarello-Santos LCC, Camilios-Neto D, Cruz LM, Raittz RT, Marques ACQ, LiPuma J, Fadel-Picheth CMT, Souza EM, Pedrosa FO (2019) Genome comparison between clinical and environmental strains of Herbaspirillum seropedicae reveals a potential new emerging bacterium adapted to human hosts. BMC Genomics 20:630. https://doi.org/10.1186/s12864-019-5982-9
Figuerola ELM, Guerrero LD, Türkowsky D, Wall LG, Erijman L (2015) Crop monoculture rather than agriculture reduces the spatial turnover of soil bacterial communities at a regional scale. Environ Microbiol 17:678–688. https://doi.org/10.1111/1462-2920.12497
Gaiero JR, McCall CA, Thompson KA, Day NJ, Best AS, Dunfield KE (2013) Inside the root microbiome: bacterial root endophytes and plant growth promotion. Am J Bot 100:1738–1750. https://doi.org/10.3732/ajb.1200572
Galtier N, Gouy M, Gautier C (1996) SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny. Bioinformatics 12:543–548. https://doi.org/10.1093/bioinformatics/12.6.543
Gordon SA, Weber RP (1951) Colorimetric estimation of indoleacetic acid. Plant Physiol 26:192–195. https://doi.org/10.1104/pp.26.1.192
Haahtela K, Kari K, Sundman V (1983) Nitrogenase activity (acetylene reduction) of root-associated, cold-climate Azospirillum, Enterobacter, Klebsiella, and Pseudomonas species during growth on various carbon sources and at various partial pressures of oxygen. Appl Environ Microbiol 45:563–570. https://doi.org/10.1128/AEM.45.2.563-570.1983
He Z, Honeycutt CW (2005) A modified molybdenum blue method for orthophosphate determination suitable for investigating enzymatic hydrolysis of organic phosphates. Commun Soil Sci Plant Anal 36:9–10. https://doi.org/10.1081/CSS-200056954
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil. College of Agriculture, University of California, Circular, p 347
Hoffman CS, Winston F (1987) A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene 57:267–272. https://doi.org/10.1016/0378-1119(87)90131-4
Ikeda AC, Bassani LL, Adamoski D, Stringari D, Cordeiro VK, Glienke C, Steffens MB, Hungria M, Galli-Terasawa LV (2013) Morphological and genetic characterization of endophytic bacteria isolated from roots of different maize genotypes. Microb Ecol 65:154–160. https://doi.org/10.1007/s00248-012-0104-0
Ji SH, Gururani MA, Chun S-C (2014) Isolation and characterization of plant growth promoting endophytic diazotrophic bacteria from Korean rice cultivars. Microbiol Res 169:83–98. https://doi.org/10.1016/j.micres.2013.06.003
Johnston-Monje D, Raizada MN (2011) Conservation and diversity of seed associated endophytes in Zea across boundaries of evolution, ethnography and ecology. PLoS ONE 6:e20396
Kämpfer P, McInroy JA, Doijad S, Chakraborty T, Glaeser SP (2016) Kosakonia pseudosacchari sp. nov., an endophyte of Zea mays. Syst Appl Microbiol 39:1–7. https://doi.org/10.1371/journal.pone.0020396
Karimi B, Dequiedt S, Terrat S, Jolivet C, Arrouays D, Wincker P, Cruaud C, Bispo A, Chemidlin N, Prévost-Bouré NC, Ranjard L (2019) Biogeography of soil bacterial networks along a gradient of cropping intensity. Sci Rep 9:3812. https://doi.org/10.1038/s41598-019-40422-y
Kempton JH (1924) Jala maize, a giant variety from Mexico. J Hered 15:337–344
Kim M, Oh H-S, Park S-C, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0
Leveau JHJ, Lindow SE (2005) Utilization of the plant hormone indole-3-acetic acid for growth by utilization of the plant hormone indole-3-acetic acid for growth by Pseudomonas putida strain 1290. Appl Environ Microbiol 71:2365–2371. https://doi.org/10.1128/AEM.71.5.2365-2371.2005
Lin L, Wei C, Chen M, Wang H, Li Y, Li Y, Yang L, An Q (2015) Complete genome sequence of endophytic nitrogen-fixing Klebsiella variicola strain DX120E. Stand Genomic Sci 10:22. https://doi.org/10.1186/s40793-015-0004-2
Madhaiyan M, Poongushali S, Lee JS, Saravanan VS, Lee KC, Santhanakrishnan P (2010) Enterobacter arachidis sp. nov., a plant-growth-promoting diazotrophic bacterium isolated from rhizosphere soil from groundnut. Int J Syst Evol Microbiol 60:1559–1564. https://doi.org/10.1099/ijs.0.013664-0
Mehta S, Nautiyal CS (2001) An efficient method for qualitative screening of phosphate-solubilizing bacteria. Curr Microbiol 43:51–56. https://doi.org/10.1007/s002840010259
Mertschnigg T, Patz S, Becker M, Feierl G, Ruppel S, Bunk B, Spröer C, Overmann J, Zarfel G (2020) First report of Kosakonia radicincitans bacteraemia from Europe (Austria)—Identification and whole-genome sequencing of strain DSM 107547. Sci Rep 10:1948. https://doi.org/10.1038/s41598-020-58689-x
Michavila G, Adler C, De Gregorio PR, Lami MJ, Caram Di Santo MC, Zenoff AM, de Cristobal RE, Vincent PA (2017) Pseudomonas protegens CS1 from the lemon phyllosphere as a candidate for citrus canker biocontrol agent. Plant Biol 19:608–617. https://doi.org/10.1111/plb.12556
Molina-Romero D, Baez A, Quintero-Hernández V, Castañeda-Lucio M, Fuentes-Ramírez LE, Bustillos-CristalesRodríguez-Andrade MRO, Morales-García YE, Munive A, Muñoz-Rojas J (2017) Compatible bacterial mixture, tolerant to desiccation, improves maize plant growth. PLoS One 12:e0187913. https://doi.org/10.1371/journal.pone.0187913
Montañez A, Abreu C, Gill PR, Hardarson G, Sicardi M (2009) Biological nitrogen fixation in maize (Zea mays L.) by 15N isotope-dilution and identification of associated culturable diazotrophs. Biol Fertil Soils 45:253–263. https://doi.org/10.1007/s00374-008-0322-2
Morales-Jiménez J, Vera-Ponce de León A, García-Domínguez A, Martínez-Romero E, Zúñiga G, Hernández-Rodríguez C (2013) Nitrogen-fixing and uricolytic bacteria associated with the gut of Dendroctonus rhizophagus and Dendroctonus valens (Curculionidae: Scolytinae). Microb Ecol 66:200–210. https://doi.org/10.1007/s00248-013-0206-3
Panday D, Schumann P, Das SK (2011) Rhizobium pusense sp. nov., isolated from the rhizosphere of chickpea (Cicer arietinum L.). Int J Syst Evol Microbiol 61:2632–2639. https://doi.org/10.1099/ijs.0.028407-0
Pande A, Pandey P, Mehra S, Singh M, Kaushik S (2017) Phenotypic and genotypic characterization of phosphate solubilizing bacteria and their efficiency on the growth of maize. J Genet Eng Biotechnol 15:379–391. https://doi.org/10.1016/j.jgeb.2017.06.005
Patten CL, Glick BR (1996) Bacterial biosynthesis of indole-3-acetic acid. Can J Microbiol 42:207–220. https://doi.org/10.1139/m96-032
Peng G, Zhang W, Luo H, Xie H, Lai W, Tan Z (2009) Enterobacter oryzae sp. nov., a nitrogen-fixing bacterium isolated from the wild rice species Oryza latifolia. Int J Syst Evol Microbiol 59:1650–1655. https://doi.org/10.1099/ijs.0.005967-0
Pereira SIA, Castro PML (2014) Diversity and characterization of culturable bacterial endophytes from Zea mays and their potential as plant growth-promoting agents in metal-degraded soils. Environ Sci Pollut Res 21:14110–14123. https://doi.org/10.1007/s11356-014-3309-6
Pérez-Montaño F, Alías-Villegas C, Bellogín RA, del Cerro P, Espuny MR, Jiménez-Guerrero I, López-Baena FJ, Ollero FJ, Cubo T (2014) Plant growth promotion in cereal and leguminous agricultural important plants: from microorganism capacities to crop production. Microbiol Res 169:325–336. https://doi.org/10.1016/j.micres.2013.09.011
Pillonetto M, Arend L, Gomes SMT, Oliveira MAA, Timm LN, Martins AF, Barth AL, Mazzetti A, Hersemann L, Smits THM, Mira MT, Rezzonico F (2018) Molecular investigation of isolates from a multistate polymicrobial outbreak associated with contaminated total parenteral nutrition in Brazil. BMC Infect Dis 18:397. https://doi.org/10.1186/s12879-018-3287-2
Prasanna BM (2012) Diversity in global maize germplasm: characterization and utilization. J Biosci 37:843–855. https://doi.org/10.1007/s12038-012-9227-1
Rao VR (1978) Effect of carbon sources on asymbiotic nitrogen fixation in a paddy soil. Soil Biol Biochem 10:319–321. https://doi.org/10.1016/0038-0717(78)90029-9
Raven JA (1988) The iron and molybdenum use efficiencies of plant growth with different energy, carbon and nitrogen sources. New Phytol 109:279–287. https://doi.org/10.1111/j.1469-8137.1988.tb04196.x
Reis VM, Estrada-de los Santos P, Tenorio-Salgado S, Vogel J, Stoffels M, Guyon S, Mavingui P, Baldani VLD, Schmid M, Baldani JI, Balandreau J, Hartmann A, Caballero-Mellado J (2004) Burkholderia tropica sp. nov., a novel nitrogen-fixing, plant-associated bacterium. Int J Syst Evol Microbiol 54:2155–2162. https://doi.org/10.1099/ijs.0.02879-0
Relman DA (1993) Universal bacterial 16S rRNA amplification and sequencing. In: Persing DH, Smith TF, Tenover FC, White TJ (eds) Diagnostic molecular microbiology: principles and applications. ASM Press, Washington, pp 489–495
Reyna-Flores F, Barrios-Camacho H, Dantan-Gonzalez E, Ramírez-Trujillo JA, Beltrán FLA, Rodríguez-Medina N, Garza-Ramos U, Suárez-Rodríguez R (2018) Draft genome sequences of endophytic isolates of Klebsiella variicola and Klebsiella pneumoniae obtained from the same sugarcane plant. Genome Announc. https://doi.org/10.1128/genomeA.00147-18
Rice E (2007) Conservation in a changing world: in situ conservation of the giant maize of Jala. Genet Resourc Crop Evol 54:701–713. https://doi.org/10.1007/s10722-006-0023-3
Rodríguez-Medina N, Barrios-Camacho H, Duran-Bedolla J, Garza-Ramos U (2019) Klebsiella variicola: an emerging pathogen in humans. Emerg Microbes Infect 8:973–988. https://doi.org/10.1080/22221751.2019.1634981
Rosenblueth M, Ormeño-Orrillo E, López-López A, Rogel MA, Reyes-Hernández BJ, Martínez-Romero JC, Reddy PM, Martínez-Romero E (2018) Nitrogen fixation in cereals. Front Microbiol 9:1794. https://doi.org/10.3389/fmicb.2018.01794
Ruiz JA, Bernar EM, Jung K (2015) Production of siderophores increases resistance to fusaric acid in Pseudomonas protegens Pf-5. PLoS One 10:e0117040. https://doi.org/10.1371/journal.pone.0117040
Saleem M, Arshad M, Hussain S, Bhatti AS (2007) Perspective of plant growth promoting rhizobacteria (PGPR) containing ACC deaminase in stress agriculture. J Ind Microbiol Biotechnol 34:635–648. https://doi.org/10.1007/s10295-007-0240-6
Salha Y, Sudalaimuthuasari N, Kundu B, AlMaskari RS, Alkaabi AS, Hazzouri KM, AbuQamar SF, El-Tarabily KA, Amiri KMA (2020) Complete genome sequence of Phytobacter diazotrophicus strain UAEU22, a plant growth-promoting bacterium isolated from the date palm rhizosphere. Microbiol Resour Announc. https://doi.org/10.1128/MRA.00499-20
Scholten H, Pierik R (1998) Agar as a gelling agent: chemical and physical analysis. Plant Cell Rep 17:230–235. https://doi.org/10.1007/s002990050384
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophore. Anal Biochem 160:47–56. https://doi.org/10.1016/0003-2697(87)90612-9
Sharma A, Johri BN (2003) Growth promoting influence of siderophore-producing Pseudomonas strains GRP3A and PRS9 in maize (Zea mays L.) under iron limiting conditions. Microbiol Res 158:243–248. https://doi.org/10.1078/0944-5013-00197
Suhaimi NS, Yap KP, Ajam N, Thong KL (2014) Genome sequence of Kosakonia radicincitans UMEnt01/12, a bacterium associated with bacterial wilt diseased banana plant. FEMS Microbiol Lett 358:11–13. https://doi.org/10.1111/1574-6968.12537
Swenson CE, Sadikot RT (2015) Achromobacter respiratory infections. Ann Am Thorac Soc 12:252–258. https://doi.org/10.1513/AnnalsATS.201406-288FR
Szkop M, Sikora P, Orzechowski S (2012) A novel, simple, and sensitive colorimetric method to determine aromatic amino acid aminotransferase activity using the Salkowski reagent. Folia Microbiol 57:1–4. https://doi.org/10.1007/s12223-011-0089-y
Tagele SB, Kim SW, Lee HG, Kim HS, Lee YS (2018) Effectiveness of multi-trait Burkholderia contaminans KNU17BI1 in growth promotion and management of banded leaf and sheath blight in maize seedling. Microbiol Res 214:8–18. https://doi.org/10.1016/j.micres.2018.05.004
Tagele SB, Kim SW, Lee HG, Lee YS (2019) Potential of novel sequence type of Burkholderia cenocepacia for biological control of root rot of maize (Zea mays L.) caused by Fusarium temperatum. Int J Mol Sci 20:1005. https://doi.org/10.3390/ijms20051005
Wang Y, Xu X, Liu T, Wang H, Yang Y, Chen X, Zhu S (2020) Analysis of bacterial and fungal communities in continuous-cropping ramie (Boehmeria nivea L. Gaud) fields in different areas in China. Sci Rep 10:3264. https://doi.org/10.1038/s41598-020-58608-0
Weber NF, Herrmann I, Hochholdinger F, Ludewig U, Neumann G (2018) PGPR-Induced growth stimulation and nutrient acquisition in maize: Do root hairs matter? Sci Agric Bohem 49:164–172. https://doi.org/10.2478/sab-2018-0022
Wei CY, Lin L, Luo LJ, Xing YX, Hu CJ, Yang LT, Li YR, An Q (2014) Endophytic nitrogen-fixing Klebsiella variicola strain DX120E promotes sugarcane growth. Biol Fertil Soils 50:657–666. https://doi.org/10.1007/s00374-013-0878-3
Weller-Stuart T, De Maayer P, Coutinho T (2017) Pantoea ananatis: genomic insights into a versatile pathogen. Mol Plant Pathol 18:1191–1198. https://doi.org/10.1111/mpp.12517
Witzel K, Gwinn-Giglio M, Nadendla S, Shefchek K, Ruppel S (2012) Genome sequence of Enterobacter radicincitans DSM16656(T), a plant growth-promoting endophyte. J Bacteriol 194:5469. https://doi.org/10.1128/JB.01193-12
Yan Y, Yang J, Dou Y, Chen M, Ping S, Peng J, Lu W, Zhang W, Yao Z, Li H, Liu W, He S, Geng L, Zhang X, Yang F, Yu H, Zhan Y, Li D, Lin Z, Wang Y, Elmerich C, Lin M, Jin Q (2008) Nitrogen fixation island and rhizosphere competence traits in the genome of root-associated Pseudomonas stutzeri A1501. Proc Natl Acad Sci USA 105:7564–7569. https://doi.org/10.1073/pnas.0801093105
You M, Fang S, MacDonald J, Xu J, Yuan ZC (2020) Isolation and characterization of Burkholderia cenocepacia CR318, a phosphate solubilizing bacterium promoting corn growth. Microbiol Res 233:126395. https://doi.org/10.1016/j.micres.2019.126395
Zhang GX, Peng GX, Wang ET, Yan H, Yuan QH, Zhang W, Lou X, Wu H, Tan ZY (2008) Diverse endophytic nitrogen-fixing bacteria isolated from wild rice Oryza rufipogon and description of Phytobacter diazotrophicus gen. nov. sp. nov. Arch Microbiol 189:431–439. https://doi.org/10.1007/s00203-007-0333-7
Zhang J, Wang X-J, Wang J-P, Wang W-X (2014) Carbon and nitrogen contents in typical plants and soil profiles in Yanqi Basin of Northwest China. J Integr Agric 13:648–656. https://doi.org/10.1016/S2095-3119(13)60723-6
Zhao K, Penttinen P, Zhang X, Ao X, Liu M, Yu X, Chen Q (2014) Maize rhizosphere in Sichuan, China, hosts plant growth promoting Burkholderia cepacia with phosphate solubilizing and antifungal abilities. Microbiol Res 169:76–82. https://doi.org/10.1016/j.micres.2013.07.003
Zhou WP, Shen WJ, Li YE, Hui DF (2017) Interactive effects of temperature and moisture on composition of the soil microbial community. Eur J Soil Sci 68:909–918. https://doi.org/10.1111/ejss.12488
Zhu B, Zhou Q, Lin L, Hu C, Shen P, Yang L, An Q, Xie G, Li Y (2013) Enterobacter sacchari sp. nov., a nitrogen-fixing bacterium associated with sugar cane (Saccharum officinarum L.). Int J Syst Evol Microbiol 63:2577–2582. https://doi.org/10.1099/ijs.0.045500-0
Acknowledgements
All the authors would like to thank Proof-Reading-Service.com Ltd. for review of the English version of the manuscript. B.R.G., E.D.U.C., and J.E.G.C. are grateful to the Consejo Nacional de Ciencia y Tecnología (CONACYT) and PIFI/BEIFI-IPN for the scholarships. L.V.T. and C.H.R. are fellows of EDI-IPN, COFAA-IPN, and SNI-CONACYT. This work was supported by the Secretaría de Investigación y Posgrado-IPN (SIP-20181779, 20195643, 20200782) and the project 16131431995 “Estudio de la diversidad microbiana asociada a suelo rizosférico de 10 variedades de maíz” by the Instituto Nacional de Investigaciones Forestales Agrícolas y Pecuarias (INIFAP).
Author information
Authors and Affiliations
Contributions
BRG, CVG and JEGC performed the isolation and bacterial characterization; BRG, NMJ and EVC performed greenhouse experiments; RIAG, LVT and CHHR provided the samples, designed, and coordinated the study; BRG wrote the first draft manuscript; EVC RIAG, NMJ and CHHR contributed to manuscript editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Ethics approval
Not applicable for this section.
Consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
Not applicable for this section.
Rights and permissions
About this article
Cite this article
Rios-Galicia, B., Villagómez-Garfias, C., De la Vega-Camarillo, E. et al. The Mexican giant maize of Jala landrace harbour plant-growth-promoting rhizospheric and endophytic bacteria. 3 Biotech 11, 447 (2021). https://doi.org/10.1007/s13205-021-02983-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-021-02983-6