Abstract
Lipases, particularly microbial lipases, are important industrial biocatalysts. As a result, lipase enzyme screening, synthesis, and purification from microbial strains are constantly evolving to meet the needs of the pharmaceutical and food industries. Thus, the goal of this study was to identify the most potential lipase-producing bacterial strains from Aavin dairy industry effluent contaminated soil. Furthermore, growth parameters, such as pH, temperature, carbon and nitrogen sources, were optimized for lipase enzyme production from selected bacterial strains. According to the findings, 9 strains (V1–V9) of 15 bacterial isolates were found to be lipase producers. However, three strains (V1, V7, and V8) predominated and demonstrated significant lipase-producing activity. These V1, V7, and V8 bacterial strains were identified as Bacillus pumilus V1, Bacillus pumilus V7, and Bacillus subtilis V8 through 16S rRNA sequencing. About 16.6 to 27.8 µg mL−1 of lipase production was recorded under the optimal growth conditions: pH 8, temperature 37 °C, fructose and yeast extract as suitable carbon and nitrogen source. Among these 3 strains B. pumilus V1 showed excellent lipase productivity than others. The molecular weight of this lipase produced by bacteria was determined to be 35 kDa using sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In several biotechnology sectors, lipolytic enzymes (EC 3.1.1) can potentially catalyze a wide range of biological reaction molecules. Their usage ranges from food applications to biochemical applications in medicine, detergents production, pesticide synthesis, leather tanning, wastewater treatment (Whangchai et al. 2021), and cosmetics production (Sarmah et al. 2018, Anusha et al. 2021). Lipolytic enzymes are also used in alcoholics, acidolysis, esterification and aminolysis as biocatalysts (Mehta et al. 2021). Below is how lipases work as a biocatalyst: one of the most important benefits of rigorous biochemical research since the 1940s has been enzymes in illness diagnosis, providing a basis for clinical chemistry (Narayanan et al. 2021a). Lipases are found in every kingdom of life, including bacteria, archea and eukaryotes, including plants, livestock and fungi (Kandasamy et al. 2021a,b). Microbial lipase is more valuable than plant and animal enzymes because it is easy to control the genetically engineered and capable of rapid growth in a wide range of catalytic activity (Ponniah et al. 2021, Narayanan et al. 2021b). In addition, no seasonal oscillations impact microorganisms to ensure they are constantly supplied and high levels of fatty lipases from microbial cells are obtained. More stable than plants and animal derivatives are also microbial lipases, and their synthesis is easier and safer for industrial and research purposes (Filho et al. 2019; Egbuna et al. 2021). Due to their increased activity, neutral or alkaline pH optimum, bacterial enzymes are chosen over fungal enzymes (Kandasamy et al. 2021a,b). Bacterial cells are more easily genetically and environmentally modified due to their short generation times, simple dietary requirements, and the production of readily desirable qualities to increase cell yield and enzyme activity or secrete altered enzymes (Hunter et al. 2019, Saranyadevi et al. 2021). Bacterial lipases are mainly glycoproteins, but lipoproteins are some extracellular lipases (Kumarasamy et al. 2020). Bacterial extracellular lipases are often produced by nitrogen and carbon sources, inorganic salts, lipid presence, temperature, and oxygen availability (Soman et al. 2020a, Vijayan et al. 2020). The gram-positive and gram-negative bacteria create different lipases. Main lipases derive from Gram-positive, such as Bacillus sp., Staphylococcus sp., Streptococcus sp., and Enterococcus sp., are the most important bacterial species (Nguyen et al. 2020; Narayanan et al. 2021c). Few Gram-negative species, such as Achromobacter sp., Alcaligene sp., Burkholderi sp., and Chromo bacterium sp. Strains, were reported as lipase producer (Matias et al. 2021; Soman et al. 2020b; Narayanan et al. 2021d). The lipase enzyme produced from Bacillus strains is the most familiar and preferably involved in various industry sectors. However, the demand for lipase enzymes is increasing in industries; hence, the researchers focus on enumerate most potential lipase enzyme producing bacterial isolates from various sources. In this connection, this research was designed to enumerate the predominant and most efficient lipase-producing bacterial isolates from dairy (Aavin) effluent contaminated soil. Furthermore, the optimal growth conditions required for evaluating the maximum lipase enzyme producing competence of bacterial test isolates were studied.
Materials and methods
Sample collection
A muddy soil sample (10 g) contaminated with dairy effluent was collected at a 15–30 cm depth from the Aavin Dairy Industry in the Dharmapuri District of Tamil Nadu. The sample was contained in a sterile ziplock bag and immediately transported to the laboratory, where it was kept in a refrigerator until further processing (Uchida et al. 2018).
Enumeration of bacterial population and preliminary screening
The cultivable bacteria populations present in the collected dairy effluent contaminated muddy soil sample were enumerated by following standard serial dilution technique (Sirisha et al. 2010). In brief, about 1 g of collected muddy soil sample was dissolved in 10 mL of sterilized distilled water and used as stock solution. The standard serial dilution process was followed from this stock solution as 10–1–10–9. Then from the 10−6th dilution, 0.1 mL was taken and inoculated on sterilized Rhodamine olive oil agar (contains: peptone 5 g L−1, yeast extract 3 g L−1, NaCl 4 g L−1, agar 15 g L−1, rhodamine dye (1 mg mL−1) 10 mL L−1) Petri plates by spread plate method. The inoculated plates were incubated at 37 °C for 48 h. after 48 h of incubation, the colonies on the plates were observed, and 15 colonies were suspected as lipase producers based on colonies covered by orange fluorescent color under UV exposure. Then, as a preliminary screening step, these 15 probable colonies were inoculated individually on fresh Rhodamine olive oil agar media. The results were seen, and chosen strains were submitted to secondary screening.
Secondary screening
From the preliminary screening, 9 bacterial isolates (named as V1–V9) were subjected to secondary screening using Tween 20 agar plates (containing: peptone (667 g L−1), NaCl (5 g L−1), CaCl2.2H2O (0.1 g L−1), agar–agar (20 g L−1), and 10 mL of (156 g L−1) Tween20). A Loop full of these pure bacterial isolates was individually inoculated on sterilized Tween20 agar plates and incubated at 37 ºC for 24 h. After incubation, the colonies were observed and noted the colonies’ zone of clearance (Elhussiny et al. 2020).
Quantitative analysis by lipase activity assay
The quantitative analysis was performed to determine which strain (V1–V9) produced a significant volume of lipase enzyme through lipase activity assay (Sharma et al. 2017). Then 1 mL of each selected strain (3.0 × 107 CFU mL−1) was inoculated on freshly prepared production (50 mL) media containing 5 g L−1 (each) peptone and beef extract. After sterilization, olive oil was added as 10 mL L−1 (pH 7.0) in a 100 mL beaker. The culture (each) inoculated flasks were incubated at 37 ºC for 24 h. After incubation, the culture-grown medium was subjected to a centrifugation process at 5 K rpm for 15 min. Then the supernatant was subjected to lipase activity assay utilizing p-nitrophenyl laureate (p-NPL) as substrate and followed the standard estimation method. Then the absorbance of the sample was recorded at 380 nm using a nanodrop spectrophotometer (NanoDrop™ 2000, Thermo Scientific™, USA) and calculated the unit value using standard calculating formula.
Molecular characterization of bacterial test isolates
The lipase activity (quantitative) assay revealed that only three bacterial isolates (V1, V7, and V8) showed a significant volume of lipase production than others (Ilesanmi et al. 2020). Hence, these three strains were subjected to molecular characterization study by 16S rRNA sequencing for genus and species level identifications. The standard DNA extraction and 16S rRNA sequencing method were followed. Then obtained sequences of these test isolates were subjected to phylogenetic tree analysis to detect the percentage of genetic similarity using the NCBI-BLAST website.
Growth parameters optimization for lipase enzyme production
The microbes will express their maximum metabolic and growth activity under optimal conditions only. Hence, the most suitable (optimal) growth conditions for the lipase enzyme production efficiency test bacterial isolates, such as Bacillus pumilus V1, Bacillus pumilus V7, and Bacillus subtilis V8, were studied. The growth parameters, such as pH (6.0, 7.0, 8.0 and 9.0), temperature (25, 30, 35, 37 and 40 ºC), 300 mg L−1 (each) of carbon sources (glucose, sucrose, fructose, and maltose), and 300 mg L−1 (each) of nitrogen sources (beef extract, yeast extract, soybean, and peptone), on production medium are as per the requirement. Triplicates were performed for each growth parameter, and lipase yield was estimated by the nanodrop spectrophotometer method and a standard formula calculated yield (Sahoo et al. 2020).
Production of lipase
The extended lipase-producing activities of B. pumilus V1, B. pumilus V7, and B. subtilis V8, were studied under optimized conditions, such as pH 8, temperature 37 ºC, fructose (carbon source), and yeast extract (nitrogen source) in production medium and incubated for 48 h, in a shaking incubator. After incubation, the quantification (yield) of lipase enzyme was determined by spectrophotometer analysis and yield was calculated by a standard formula (Sharma et al. 2017).
Molecular weight determination of lipase by SDS-PAGE
The ammonium sulphate fractionation method was used to extract the lipase (protein) from bacterial test isolates (Sharma et al. 2017). In brief, 24 h-old bacterial culture was centrifuged at 10 K rpm for 10 min. The equal volume of ammonium sulphate (39 g 100 mL−1) solution was added to the supernatant and incubated at 4 ºC overnight. Then this reaction mix was centrifuged at 10 K rpm for 15 min in a cooling centrifuge. Then obtained pellet was dissolved in 0.1 M phosphate buffer (pH 7.0) and subjected to standard SDS-PAGE analysis.
Statistical analysis
The majority of the tests were done in triplicates to ensure accuracy and reproducibility, and the values mentioned in the results are the mean and standard error (SE) of triplicates.
Results and discussion
Isolation of lipase-producing bacteria
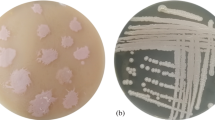
Lipases, particularly microbial lipases, are important industrial enzymes. As a result, lipase enzyme screening, synthesis, and purification from microbial strains are constantly evolving to meet the needs of the pharmaceutical and food industries. Interestingly, the results obtained in this study are most likely matched with the lipases found in bacteria that have been reported earlier. About 15 numbers bacterial isolates were enumerated from the Aavin dairy industry effluent contaminated muddy soil sample. In the preliminary screening, out of 15 bacterial isolates, 9 cultures were identified as lipase producers, and these 9 cultures were initially named V1–V9. The preliminary screening was confirmed by developing orange fluorescent color colonies on Rhodamine olive oil agar under UV exposure. It declared that the V1–V9 bacterial isolates possess lipase-producing potential. The secondary screening on Tween 20 agar plate analysis also confirmed that these V1–V9 bacterial isolates have lipase-producing competence and it was confirmed by zone of clearance around the colonies.
Similarly, around 9 bacterial strains from forty bacterial isolates enumerated from the oil contaminated soil have been reported as excellent lipase producers confirmed by fluorescent colonies on Rhodamine olive oil agar media (Lomthaisong et al. 2012). According to another report, the 8 bacterial strains screened from oil contaminated soil can produce lipase, which was confirmed by observing the opaque and pink colour fluorescent (halos around) colonies on olive oil and rhodamine B containing agar media (Alhamdani and Alkabbi 2016). Creating a unique interaction between uranyl fatty acid ion and cationic rhodamine B causes the orange colour fluorescence around the lipase-producing bacterial colonies when observed under UV light (Mohammed 2013). The generation of excited dimers of rhodamine B, which fluoresce at longer wavelengths than the exited monomer, may be related to the mechanism of this color formation (Arbeloa et al. 2007).
Lipase production by quantitative method
The extended lipase generating activity of these V1 to V9 bacterial isolates was examined by quantitative technique in olive oil as a substrate. The results showed that out of 9 isolates 3 strains, V1, V7, and V8 produced a significant volume of lipase as 31.2 µg mL−1, 22.9 µg mL−1, and 18.7 µg mL−1, respectively, confirmed by titration method. Furthermore, these strains were classified as a “very active strain” for lipase production than other strains. Several microbial species have been reported in the scientific community. For example, in bacteria, the species belongs to Bacillus, Achromobacter, Alcaligenes, Arthrobacter, and Pseudomonos, in fungi, Penicillium, Fusarium, Aspergillus, and so on have been recognized as a potential agent for lipase enzyme production (Chandra et al. 2020). The bacterial strain isolated from muddy soil from dairy effluents was suspected as belonging to the Bacillus genus and possessing substantial lipase generating capacity in this study. Willerding et al. (2011) identified around 75 bacterial strains out of 440 were identified as lipase producers through the qualitative method. Statistically, these strains were counted as 41% of the total bacterial diversity in Amazonia soil (Willerding et al. 2011). Another bacterial strain, namely Streptomyces exfoliates LP10 screened from petroleum contaminated soil, was recognized as an excellent lipase producer through the quantitative method and found that it yielded the lipase ranged from 1.5 to 6.9 IU mL−1 (Aly et al. 2012). Similarly, the B. cereus isolated from the contaminated soil produced around 225 IU mL−1, and it was quantified by titration method (Hassan et al. 2018).
Molecular characterization of bacterial test isolates
The 16S rRNA sequencing analysis results (1500 bp-sized bacterial sequences) were subjected to NCBI – BLAST analysis and performed similarity search. Obtained results revealed that the bacterial test isolates were B. pumilus V1, B. pumilus V7, and B. subtilis V8. Three strains had a computed value of 16S rRNA gene similarity of ~ 99% considering the reference data. It was revealed in Fig. 1 that the obtained isolates were phylogenetically connected. It could be inferred that B. pumilus V1, B. pumilus V7, and B. subtilis V8 were all relatives of the Bacillus genus. Strains V1, V7, and V8 were placed in their correct phylogenetic lineage using the neighbor-joining approach, and their comparative study was conducted. The tree with V1 = 0.11211510, V7 = 0.03374284 and V8 = 0.06522920 is presented here. While the tree branch lengths match those of the phylogenetic distances used to derive the phylogenetic tree, the tree itself is displayed to scale (Fig. 1). Similarly, the P. gessardi isolated from oil-contaminated soil has been reported as the considerable potential to produce industrially important lipase enzymes. This strain was characterized through a molecular approach (Yadav et al. 2021). The bacterial species reported from coconut oil mill contaminated soil also possess a significant lipase level producing bacterial diversity. Most of them were identified as Bacillus species, which was validated through a genetic characterization investigation (Bharathi and Rajalakshmi 2019). Another bacterial species, namely the Pseudomonas xinjiangensis strain CFS14 identified by 16S rRNA sequencing analysis and culture, was enumerated from oil contaminated soil with excellent lipase enzyme producing potential (Lomthaisong et al. 2012). Another species of Pseudomonas enumerated from the soil sample was also reported as an excellent lipase producer, and it was identified as Pseudomonas aeuriginosa through 16S rRNA sequencing (Ilesanmi et al. 2020).
Optimization of lipase production
The optimal growth conditions, such as pH, temperature, carbon, and nitrogen sources, are the essential factors determining bacteria lipase-producing competence (Abol-Fotouh et al. 2021). Because the optimal conditions could enhance and balance the metabolic activity of bacteria, facilitating the active lipase production (Hwang et al. 2014).
Effect of pH on lipase production
The pH is one of the most significant factors and that determine enzyme productivity. These B. pumilus V1, B. pumilus V7, and B. subtilis V8 showed maximum lipase productivity at pH 8 as 27.8 µg mL−1, 21.5 µg mL−1 20.8 µg mL−1, respectively (Fig. 2a). Furthermore, the lowest lipase production activity was found at pH 6 (2.08 µg mL−1) and pH 9 (8.33 µg mL−1). As a result of these findings, pH 8 was determined to be a suitable pH for lipase production by these bacterial strains (B. pumilus V1, B. pumilus V7, and B. subtilis V8). The pH of the culture strongly influences cell growth and enzyme production. Some key aspects to consider for the commercial application of enzymes included an enzyme’s maximum activity and stability under a mildly alkaline pH 8 (Ibrahim et al. 2021). The elevated lipase activity was observed in the pH range of 7–11, with pH 11 exhibiting the highest activity. Enzyme activity was found to be reduced in the acidic environment. According to the current report, the organism is an alkalophilic bacteria (Ilesanmi et al. 2020). Lipase-producing bacteria have been found to prefer pH levels around 8. Similarly, the suitable pH for B. stearothermophilus and S. rimosus was pH 7.0 and 8.0, respectively (Dikshit and Kim 2020). The suitable pH of lipase producers, such as Staphylococcus sp. (Sirisha et al. 2010) and Pseudomonas monteilli (Rasmey et al. 2017), was found as pH 7.0 pH 6.0, respectively, and produced around 35 IU mL−1.
Effect of temperature on lipase production
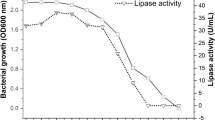
The influence of temperature on lipase synthesis by B. pumilus V1, B. pumilus V7, and B. subtilis V8 was examined with various temperatures, such as 25, 30, 35, 37, and 40 °C, for 48 h. The results revealed that the optimal temperature for these strains for lipase production was found as 37 °C. At this optimal temperature (37 °C), B. pumilus V1, B. pumilus V7, and B. subtilis V8 produced 27.3, 25.0, and 20.8 µg mL−1 lipase enzyme, respectively (Fig. 2b). Temperature influences lipase synthesis primarily through cell development, as demonstrated by a cell growth curve that increased correspondingly with increasing lipase activity (Rajendran et al. 2008). Several researchers investigated the effect of temperature on lipase fabrication over a temperature range of 30–80 °C while keeping all other conditions constant and noted that certain microbial strains are showed excellent lipase activity at increased temperature (Rodrigues et al. 2008). Temperature control enzyme synthesis at the mRNA transcription and, most likely, translation levels of proteins, increasing enzyme stability and production. Another reason for increased synthesis at higher temperatures could be because temperature affects their secretion, possibly by altering the physical features of the cell membrane. The thermostable Anoxybacillus sp. ARS-1 isolated from Taptapani Hotsprin showed the maximum lipase productivity of 29.4 IU g−1 at an optimal temperature of 57.5° (Sahoo et al. 2020).
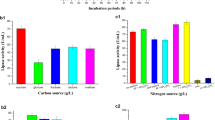
Effect of carbon and nitrogen sources on lipase production
Carbon supply is the primary contributor to the induction of lipase productivity. Both B. pumilus V1 and B. subtilis V8 prefer fructose as a carbon source and yielded 16.6 µg mL−1 and 14.5 µg mL−1 (Fig. 2c), respectively. The B. pumilus V7 has effectively utilized the maltose as a suitable carbon source and yielded 14.8 µg mL−1 of lipase (Fig. 2c). The isolated strains have not efficiently used glucose and sucrose as a carbon source for the lipase production metabolisms. Similarly, among various nitrogen sources, the yeast extract, followed by soybean and peptone, showed a significant level of influence on lipase production in B. pumilus V1 (yeast extract), B. pumilus V7 (soybean and peptone), and B. subtilis V8 (yeast extract) as 14.8, 13.8, and 8.3 µg mL−1 (Fig. 2d). While the beef extract has had no significant impact on lipase production by all the bacterial isolates. Carbon sources have proven essential to the production of lipase activity because this enzyme is an inducible one. A higher lipase synthesis in titration is caused by nitrogen-rich nutrients (Magdouli et al. 2017). In our study, the highest level of lipase synthesis was found in yeast extract at a concentration of 12.4 mg/mL. The study reported that minimum lipase activity was calculated to be 0.15 mg/mL in glucose and yeast extract.
The maximum lipase activity in glucose plus peptone was calculated to be 0.24 mg/mL. However, compared to our results, the study results demonstrate that the maximum lipase activity was obtained by providing olive oil to yeast extract for two days. On the other hand, increased lipase levels were produced using C. cylindracea to grow in a complex medium containing either 10 or 33% olive oil (Ciafardini et al. 2006). According to the latest study, olive oil is an excellent carbon source for lipase production by bacteria (Ilesanmi et al. 2020). For this study, we found that yeast extract, olive oil, and fructose all had a greater impact on lipase production in Bacillus sp. This enzyme can act as the oil–water interface, which makes it a great choice for industrial processes. The bacterial lipases are thermo-stable while plant and fungal lipases are non-specific substrates (Chandra et al. 2020).
Production of lipase
Under optimized conditions, such as pH 8, temperature 37 ºC, fructose (carbon source), yeast extract (nitrogen source), and bacterial test culture B. pumilus V1, B. pumilus V7 and B. subtilis V8 produced 27.8, 25.2, and 16.6 µg mL−1 of lipase, respectively. Because, under optimal conditions, cell proliferation, cell metabolism, and similar activities may be more effective, resulting in active production of primary and secondary metabolites (Murthy et al. 2014). Similarly, the growth conditions for Bacillus strain KS4 isolated from oil-contaminated soil samples were optimized as 1.16% olive oil, 0.12% tween 80, 5.99 mM MgCl2, and 7.16% inoculum size for highest lipase production as increased from 0.612 IU mL−1 to 2.17 IU mL−1. It was calculated as 3.54-fold higher than un-optimized conditions (Sharma et al. 2014). Another report stated that the optimized growth conditions for thermostable Staphylococcus warneri isolated from oil-contaminated soil were 55 °C, pH 8.0, 120 rpm, and 2% inoculums yielded 17.21 IU mL−1 (Yele and Desai 2015).
SDS-PAGE analysis
The molecular weight of the lipase enzyme produced from B. pumilus V1, B. pumilus V7, and B. subtilis V8 were determined by SDS-PAGE analysis. The obtained SDS-PAGE image (Fig. 3) analysis revealed that the lipases extracted from B. pumilus V1, B. pumilus V7, and B. subtilis V8 had a similar appearance on an SDS-PAGE gel. They were supposed to have a molecular weight of roughly 35 kDa based on the position of the protein marker (Fig. 3). This result was partially correlated with the findings of Balaji and Jayaraman (2014), who extracted and reported that the molecular weight of lipase enzyme derived from Bacillus sp. was found as 31.40 kDa to 50.0 (Balaji and Jayaraman 2014, Sharma et al. 2017). The Bacillus sp.-based lipase enzyme possesses excellent temperature stability and alkaline-intolerant detergent resistance, making them an industrial-applicable bacterium (Kroll et al. 2010). Pseudomonas aeruginosa HFE733, isolated from residential waste soil samples, generated alkaline lipase with a molecular weight of 51.0 kDa (Hu et al. 2018). Interestingly, another report states that the Bacillus sp. isolated from an oil contaminated soil, produced lipase molecular mass was found as 24 kDa (Sivaramakrishnan and Incharoensakdi 2016), and Pseudomonas aeruginosa LX1 yielded 56 kDa of lipase (Ji et al. 2010).
Conclusion
Extracellular lipases derived from bacteria have received far more attention than those derived from other microbial kingdoms. In this study, 9 bacterial isolates with lipase-producing activity were identified from Aavin dairy effluent-contaminated muddy soil. Nonetheless, three strains (V1, V7, and V8) out of nine demonstrated significant lipase production, which was confirmed by a quantitative lipase activity assay. These 3 strains were identified as B. pumilus V1, B. pumilus V7, and B. subtilis V8 through 16S rRNA sequencing. The optimal growth conditions for these test bacterial isolates for lipase production were found as pH 8, temperature 37 ºC, fructose (carbon source), and yeast extract (nitrogen source). Under these optimized conditions, the test bacterial culture B. pumilus V1, B. pumilus V7, and B. subtilis V8 yielded 27.8, 25.2, and 16.6 µg mL−1 of lipase correspondingly. Furthermore, the molecular size of this study was found as 35 kDa through SDS-PAGE analysis. These findings suggest that the bacterial strains identified from the dairy effluent-contaminated muddy soil (B. pumilus V1, B. pumilus V7, and B. subtilis V8) may be suitable for large-scale production after a fermentation and other growth parameter optimization study.
References
Abol-Fotouh D, AlHagar OE, Hassan MA (2021) Optimization, purification, and biochemical characterization of thermoalkaliphilic lipase from a novel Geobacillus stearothermophilus FMR12 for detergent formulations. Int J Biol Macromol 181:125–135. https://doi.org/10.1016/j.ijbiomac.2021.03.111
Alhamdani MA, Alkabbi HJJ (2016) Isolation and identification of lipase producing bacteria from oil-contaminant soil. J Biol Agric Healthc 6(2):1–7
Aly MM, Tork S, Al-Garni SM et al (2012) Production of lipase from genetically improved Streptomyces exfoliates LP10 isolated from oil-contaminated soil. Afr J Microbiol Res 6(6):1125–1137. https://doi.org/10.5897/AJMR11.1123
Anusha P, Natarajan D et al (2021) Heavy metal removal competence of individual and bacterial consortium, evolved from metal contaminated soil. Mater Today. https://doi.org/10.1016/j.matpr.2020.11.214
Arbeloa FL, Martínez VM, Arbeloa T et al (2007) Photoresponse and anisotropy of rhodamine dye intercalated in ordered clay layered films. J Photochem Photobiol c: Photochemy Rev 8(2):85–108. https://doi.org/10.1016/j.jphotochemrev.2007.03.003
Balaji L, Jayaraman G (2014) Metal ion activated lipase from halotolerant Bacillus sp. VITL8 displays broader operational range. Int J Biol Macromol 67:380–386. https://doi.org/10.1016/j.ijbiomac.2014.03.050
Bharathi D, Rajalakshmi G (2019) Microbial lipases: An overview of screening, production and purification. Biocatald Agric Biotechnol 22:101368. https://doi.org/10.1016/j.bcab.2019.101368
Chandra P, Enespa Singh R, Arora PK (2020) Microbial lipases and their industrial applications: a comprehensive review. Microb Cell Fact 19(1):169. https://doi.org/10.1186/s12934-020-01428-8
Ciafardini G, Zullo BA, Iride A (2006) Lipase production by yeasts from extra virgin olive oil. Food Microbiol 23(1):60–67. https://doi.org/10.1016/j.fm.2005.01.009
Dikshit PK, Kim BS (2020) Bacterial cellulose production from biodiesel–derived crude glycerol, magnetic functionalization, and its application as carrier for lipase immobilization. Int J Biol Macromol 153:902–911. https://doi.org/10.1016/j.ijbiomac.2020.03.047
Egbuna C, Awuchi CG, Kushwaha G et al (2021) Bioactive compounds effective against type 2 diabetes mellitus: A Systematic Review”. Cur Topics Med Chem 21:1. https://doi.org/10.2174/1568026621666210509161059
Elhussiny NI, Khattab AE-NA, El-Refai HA et al (2020) Biotransesterification capabilities of Mucorales whole-cell lipase isolates and mutants. Biocatald Agric Biotechnol. https://doi.org/10.1016/j.bcab.2020.101722
Filho DG, Silva AG, Guidini CZ (2019) Lipases: sources, immobilization methods, and industrial applications. Appl Microbiol Biotechnol 103(18):7399–7423. https://doi.org/10.1007/s00253-019-10027-6
Hassan SWM, Abd El Latif HH, Ali SM (2018) Production of cold-active lipase by free and immobilized marine Bacillus cereus HSS: application in wastewater treatment. Front Microbiol. https://doi.org/10.3389/fmicb.2018.02377/full
Hu J, Cai W, Wang C et al (2018) Purification and characterization of alkaline lipase production by Pseudomonas aeruginosa HFE733 and application for biodegradation in food wastewater treatment. Biotechnol Biotechnol Equip 32(3):583–590. https://doi.org/10.1080/13102818.2018.1446764
Hunter M, Yuan P, Vavilala D et al (2019) Optimization of protein expression in mammalian cells. Curr Protoc Protein Sci 95(1):e77. https://doi.org/10.1002/cpps.77
Hwang HT, Qi F, Yuan C et al (2014) Lipase-catalyzed process for biodiesel production: protein engineering and lipase production. Biotechnol Bioengi 111(4):639–653. https://doi.org/10.1002/bit.25162
Ibrahim AM, Hamouda RA, El-Naggar NEA et al (2021) Bioprocess development for enhanced endoglucanase production by newly isolated bacteria, purification, characterization and in-vitro efficacy as anti-biofilm of Pseudomonas aeruginosa. Sci Rep 11(1):9754. https://doi.org/10.1038/s41598-021-87901-9
Ilesanmi OI, Adekunle AE, Omolaiye JA et al (2020) Isolation, optimization and molecular characterization of lipase producing bacteria from contaminated soil. Sci Afri. https://doi.org/10.1016/j.sciaf.2020.e00279
Ji Q, Xiao S, He B et al (2010) Purification and characterization of an organic solvent-tolerant lipase from Pseudomonas aeruginosa LX1 and its application for biodiesel production. J Mol Catal b: Enzy 66(3):264–269. https://doi.org/10.1016/j.molcatb.2010.06.001
Kandasamy S, Devarayan K, Bhuvanendran N et al (2021a) Accelerating the production of biooil from hydrothermal liquefaction of microalgae via recycled biochar-supported catalysts. J Environ Chem Engi 9(4):105321. https://doi.org/10.1016/j.jece.2021.105321
Kandasamy S, He Z, Liu G et al (2021b) Current strategies and prospects in algae for remediation and biofuels: An overview. Biocatal Agric Biotechnol. https://doi.org/10.1016/j.bcab.2021.102045
Kroll J, Klinter S, Schneider C et al (2010) Plasmid addiction systems: perspectives and applications in biotechnology. Microb Biotechnol 3(6):634–657. https://doi.org/10.1111/j.1751-7915.2010.00170.x
Kumarasamy S, Subramanian V, Ranganathan M et al (2020) Microbial Stereo Inversion of (R) 3 Chloro-1, 2-Propandiol by Wickerhamomyces anomalous MGR6-KY209903. Chem Biointerface Res Appl. https://doi.org/10.33263/BRIAC105.61576166
Lomthaisong K, Buranarom A, Niamsup H (2012) Investigation of isolated lipase producing bacteria from oil-contaminated soil with proteomic analysis of its proteins responsive to lipase inducer. J Biol Sci 12(3):161–167. https://doi.org/10.3923/jbs.2012.161.167
Magdouli S, Guedri T, Tarek R et al (2017) Valorization of raw glycerol and crustacean waste into value added products by Yarrowia lipolytica. Bioresour Technol 243:57–68. https://doi.org/10.1016/j.biortech.2017.06.074
Matias RR, Sepúlveda AMG, Batista BN et al (2021) Degradation of Staphylococcus aureus biofilm using hydrolytic enzymes produced by amazonian endophytic fungi. Appl Biochem Biotechnol 193(7):2145–2161. https://doi.org/10.1007/s12010-021-03542-8
Mehta A, Guleria S, Sharma R et al (2021) The lipases and their applications with emphasis on food industry. In: Ray RC (ed) Microbial Biotechnology in Food and Health. Academic Press
Mohammed HJ (2013) Physicochemical factors affected the partial purified lipase activity of Acinetobacter baumannii local isolates. Iraqi J Pharm Sci. https://doi.org/10.31351/vol22iss1pp82-89
Murthy HN, Lee EJ, Paek KY (2014) Production of secondary metabolites from cell and organ cultures: strategies and approaches for biomass improvement and metabolite accumulation. Plant Cell Tiss Organ Cult 118(1):1–16. https://doi.org/10.1007/s11240-014-0467-7
Narayanan M, Gokila Devi P, Natarajan D et al (2021a) Green synthesis and characterization of titanium dioxide nanoparticles using leaf extract of Pouteria campechiana and larvicidal and pupicidal activity on Aedes aegypt. Environ Res. https://doi.org/10.1016/j.envres.2021.111333
Narayanan M, Gopi A, Natarajan D et al (2021b) Hepato and nephroprotective activity of methanol extract of Hygrophila spinosa and its antibacterial potential against multidrug resistant Pandoraea sputorum. Environ Res. https://doi.org/10.1016/j.envres.2021.111594
Narayanan M, Jayashree T, Kandasamy S et al (2021c) Antidermatophytic, antioxidant, and nephroprotectiveactivity of Solanum surattense: an invitro study. Process Biochem 109:178–185. https://doi.org/10.1016/j.procbio.2021.07.008
Narayanan M, Vigneshwari P, Natarajan D et al (2021d) Synthesis and characterization of TiO2 NPs by aqueous leaf extract of Coleus aromaticus and assess their antibacterial, larvicidal, and anticancer potential. Environ Res. https://doi.org/10.1016/j.envres.2021.111335
Nguyen MT, Matsuo M, Niemann S et al (2020) Lipoproteins in gram-positive bacteria: abundance, function, fitness. Front Microbiol. https://doi.org/10.3389/fmicb.2020.582582/full
Ponniah A, Natarajan D, Chinnathambi A et al (2021) Evaluation of chromium biosorption competence of indigenous Aspergillus tubingensis AF3. Chemosphere. https://doi.org/10.1016/j.chemosphere.2021.131055
Rajendran A, Palanisamy A, Thangavelu V (2008) Evaluation of medium components by Plackett-Burman statistical design for lipase production by Candida rugosa and kinetic modeling. Chin J Biotechnol 24(3):436–444. https://doi.org/10.1016/s1872-2075(08)60024-2
Rasmey AHM, Aboseidah AA, Gaber S (2017) Characterization and optimization of lipase activity produced by Pseudomonas monteilli 2403-KY120354 isolated from ground beef. Afr J Biotechnol 16(2):96–105
Rodrigues RC, Volpato G, Wada K et al (2008) Enzymatic synthesis of biodiesel from transesterification reactions of vegetable oils and short chain alcohols. J Am Oil Chem Soc 85(10):925–930. https://doi.org/10.1007/s11746-008-1284-0
Sahoo RK, Das A, Gaur M et al (2020) Parameter optimization for thermostable lipase production and performance evaluation as prospective detergent additive. Prepar Biochem Biotechnol 50(6):578–584. https://doi.org/10.1080/10826068.2020.1719513
Saranyadevi S, Suresh K, Muthusamy R et al (2021) Silver nanoparticles synthesized using asafoetida resin, characterization of their broad spectrum and larvicidal activity. Ann Romanian Soc Cell Biol 25(4):15035–15049
Sarmah N, Revathi D, Sheelu G et al (2018) Recent advances on sources and industrial applications of lipases. Biotechnol Prog 34(1):5–28. https://doi.org/10.1002/btpr.2581
Sharma D, Kumbhar BK, Verma AK et al (2014) Optimization of critical growth parameters for enhancing extracellular lipase production by alkalophilic Bacillus sp. Biocatal Agric Biotechnol 3(4):205–211. https://doi.org/10.1016/j.bcab.2014.04.004
Sharma P, Sharma N, Pathania S et al (2017) Purification and characterization of lipase by Bacillus methylotrophicus PS3 under submerged fermentation and its application in detergent industry. J Gene Engin Biotechnol 15(2):369–377. https://doi.org/10.1016/j.jgeb.2017.06.007
Sirisha E, Rajaseka N, Narasu ML (2010) Isolation and optimization of lipase producing bacteria from oil contaminated soils. Adv Biol Res 4(5):249–252
Sivaramakrishnan R, Incharoensakdi A (2016) Purification and characterization of solvent tolerant lipase from Bacillus sp for methyl ester production from algal oil. J Biosci Bioengin. https://doi.org/10.1016/j.jbiosc.2015.09.005
Soman S, Kumarasamy S, Narayanan M (2020) Biocatalyst: Phytase production in solid state fermentation by OVAT strategy. Chem Biointerface Res Appl. https://doi.org/10.33263/BRIAC105.61196127
Soman S, Suresh K, Muthusamy R et al (2020) Chemically defined medium for the production of phytase by Hanseniaspora guilliermondii S1, Pichia fermentans S2 and its secondary structure prediction of 16S rRNA. Biointerface Res Appl Chem. https://doi.org/10.33263/BRIAC105.62626272
Uchida Y, Mogi H, Hamamoto T et al (2018) Changes in denitrification potentials and riverbank soil bacterial structures along Shibetsu River. Japan Appl EnvironSoil Sci. https://doi.org/10.1155/2018/2530946
Vijayan S, Umadevi G, Mariappan R et al (2020) High luminescence efficiency of Copper doped Zinc Sulfide (Cu: ZnS) nanoparticles towards LED applications. Procee Mater Today. https://doi.org/10.1016/j.matpr.2020.11.214
Whangchai K, Hung T, Al-Rashed S et al (2021) Biodegradation competence of Streptomyces toxytricini D2 isolated from leaves surface of the hybrid cotton crop against β cypermethrin. Chemosphere. https://doi.org/10.1016/j.chemosphere.2021.130152
Willerding AL, Oliveira LAD, Moreira FW et al (2011) Lipase activity among bacteria isolated from amazonian soils. Enz Res. https://doi.org/10.4061/2011/720194
Yadav AN, Kour D, Kaur T et al (2021) Biodiversity, and biotechnological contribution of beneficial soil microbiomes for nutrient cycling, plant growth improvement and nutrient uptake. Biocatal Agric Biotechnol 33:102009. https://doi.org/10.1016/j.bcab.2021.102009
Yele VU, Desai K (2015) A new thermostable and organic solvent-tolerant lipase from Staphylococcus warneri; optimization of media and production conditions using statistical methods. Appl Biochem Biotechnol 175(2):855–869. https://doi.org/10.1007/s12010-014-1331-2
Acknowledgements
The authors acknowledge the DST-FIST (SR/FIST/LSI-673/2016) for strengthening the instrumentation facility of the Biotechnology Department of Periyar University, Salem, Tamil Nadu. This project was supported by Researchers Supporting Project number (RSP-2021/385) King Saud University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kandasamy, S., Vijayalakshmi, V.S., Salmen, S.H. et al. Screening, characterization, and optimization of lipase enzyme producing bacteria isolated from dairy effluents contaminated muddy soil. Appl Nanosci 13, 1443–1451 (2023). https://doi.org/10.1007/s13204-021-02062-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13204-021-02062-5