Abstract
The agave weevil, Scyphophorus acupunctatus, is a pest of agave. Its larvae cause damage to agaves by boring holes in the plant. Boring requires that the insect consume the constituents of its host plant, which contains sugars and many recalcitrant polymers. It has been hypothesized for many years that the gut bacterial communities of S. acupunctatus play a role in its ability to metabolize agave components. However, studies exploring this insect's gut bacterial communities have yet to be performed. In this work, we used a 16S rRNA gene-based metabarcoding approach to characterize the gut bacterial communities of field-collected agave weevils from different localities in Mexico. We found that external factors, including host plants, have important effects on the structure of the gut bacterial communities of S. acupunctatus. Despite this variability, we found a discrete core bacterial community mainly composed of the genera Prevotella, Pectinatus, Liquorilactobacillus, Secundilactobacillus, Paucilactobacillus, and Pseudomonas. These genera may be necessary for S. acupunctatus as metabolic helpers and/or gatekeepers. Additional studies are needed to fully assess the functionality of the gut bacterial community of this species in terms of its metabolic contribution, which may help to decipher their potential ecological implications. The information we provided here is the first step for guiding further questions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The agave weevil, Scyphophorus acupunctatus Gyllenhal, 1838, is a native American beetle species belonging to the family Curculionidae. Currently, it is considered the major agave insect pest worldwide and is well-established in Central and South America, the Caribbean, Africa, and Asia (da Silva Brito et al. 2021). S. acupunctatus is present in all of Mexico's agave-producing regions and affects both wild and cultivated agaves, including economically significant species such as blue agave (Agave tequilana Weber) and mezcal magueys (Agave angustifolia Haw. and Agave cupreata Trel. & Berger) (Solís-Aguilar et al. 2001; Figueroa-Castro et al. 2016; Vega-Petlacalco et al. 2018; Cruz-Jardón et al. 2018).
S. acupunctatus larvae cause damage to agaves by boring holes in the plant's outer and inner layers while feeding. The insect spends its entire life cycle within plant structures, strongly limiting the effectiveness of chemical control methods based on the use of synthetic insecticides (Valdés-Rodríguez et al. 2004; Figueroa-Castro et al. 2016; da Silva Brito et al. 2021). Boring also allows opportunistic microorganisms to colonize damaged tissues, provoking a disease known as Bud Soft Rot (BSR), which results in reduced or stunted development or even plant death (Waring and Smith 1986; González-Hernández 2007; Cruz Faustino et al. 2019). The initial signs of this disease are necrotic lesions at the tips of the stalks. In most cases, lesions appear first on the apical spine or in the lateral spines then progress toward the center and base of the bud, causing a downward rotting with a soft consistency that ultimately reaches the pineapple. This damage disintegrates the tissue and leaves the pineapple hollow, leaving only the fibers and causing the death of the plant (Rubio-Cortés 2007; Palemón-Alberto et al. 2021).
To date, there has been limited research on the details of the gut bacterial communities of S. acupunctatus. However, for many years it has been hypothesized that S. acupunctatus may be involved in symbiotic relationships with microorganisms that help break down plant tissues (Waring and Smith 1986). This hypothesis seems plausible considering that the gut bacterial communities of many herbivorous insects can influence the insect’s ability to exploit resources by facilitating the digestion of recalcitrant plant materials, including lignocellulosic substrates (Flint et al. 2008; Xie et al. 2019). There are several examples of such relationships. For example, the gut microbiota of termites enzymatically convert lignocellulosic substrates into assimilable derivatives (Dumond et al. 2021), while the larvae of the scarabaeid Protaetia brevitarsis efficiently break down lignocellulose despite not having complete lignocellulose-breaking enzymes in their genome (Wang et al. 2022). The European corn borer larvae (Ostrinia nubilalis) has been found to degrade maize cellulose aided by its bacterial microbiota (Li et al. 2022), and the gut bacterial communities of the wood-feeding palm weevil, Rhynchophorus ferrugineus, also plays a major role in lignocellulose hydrolysis (Angzzas et al. 2016).
The gut bacterial communities of insects has also been linked to other important roles, such as development, immunity, and behavior, as well as digestion and nutrition, where it aids in the detoxification of toxic substances and the synthesis of essential nutrients that are beneficial to the host, such as nitrogen, vitamins, or cofactors (Ceja-Navarro et al. 2015; Suárez-Moo et al. 2020; Wang et al. 2018). For example, in herbivorous insects, the gut bacterial communities is frequently involved in maintaining nitrogen balance through the recycling or elimination of nitrogen compounds (Rozadilla et al. 2020). These functions are especially relevant in insects that rely on food sources that are low in nitrogen, as occurs with dung beetles (Suárez-Moo et al. 2020; Rozadilla et al. 2020; Huang et al. 2021). In the case of curculionid beetles, several studies have demonstrated deep functional interactions with fungal and bacterial symbionts (Chakraborty et al. 2020; Ibarra-Juárez et al. 2020).
Given this background, we consider that it is reasonable to suspect that the gut bacterial communities of S. acupunctatus contribute to its ability to utilize agave as a sole food source. Confirming this hypothesis requires specific studies that characterize the gut bacterial communities of S. acupunctatus and determine how it interacts with agave plants. In addition, if some members of the gut bacterial communities of S. acupunctatus are key for its physiology, then they can be considered a potential target for controlling this pest species, as manipulating the gut microbiota of insects has been proposed as a possible way to control insect pest species (He et al. 2021; Poveda 2021). For these reasons, in this work, we surveyed the gut bacterial communities of S. acupunctatus from different regions of Mexico using 16S rRNA gene-based metabarcoding to test a) whether the gut bacterial communities of S. acupunctatus vary among different hosts and localities, and b) whether there is a core bacterial community.
2 Materials and methods
2.1 Sampling sites
We collected adult specimens of S. acupuntactus between November 2021 to May 2022 from cultivated agave fields in four different localities (Fig. 1) that included three host plant species (A. cupreata, A. tequilana, and A. salmiana). Six individuals (three males and three females) were collected from each locality and transported to the laboratory in plastic containers with holes for ventilation.
Map showing the sampling sites within Mexico where adult specimens of Scyphophorus acupunctatus were collected. Three different host plant species were included: A. cupreata (Michoacan and Guerrero), A. salmiana (Tlaxcala), A. tequilana (Jalisco). The geographic coordinates and the climate of each sampling site are also shown according to the Köppen climate classification. Aw: tropical wet and dry climate; Cwa: Monsoon-influenced humid subtropical climate; Cwb: dry-winter subtropical highland climate. The geographic coordinates of each site were recorded and plotted in R using the ggmap, osmdata, and ggspatial packages (Kahle and Wickham 2013; Clemens 2015; Dunnington and Thorne 2020)
2.2 Gut dissection and gDNA isolation
Specimens were taken for gut dissection under aseptic conditions using sterilized tweezers and dissecting scissors. Prior to dissection, the specimens were thoroughly washed to decontaminate the surfaces of the insects using a three-step washing protocol, each step consisting of 70% ethanol, 4% chlorine, and sterile distilled water for 3 min each. Six individual guts (three males and three females) per site were used for genomic DNA (gDNA) extraction, which were placed in a 2.0 mL tube with 200 µL of NaCl2 0.9%, and milled in a tissue disruptor for 5 min. The gDNA was isolated using the Quick-DNA Fungal/Bacterial Miniprep Kit (Zymo Research, CA. USA) and DNA clean and concentrator kit (Zymo Research, CA. USA). The gDNA concentration and integrity were checked using a nanodrop and 1.2% agarose gel electrophoresis, respectively.
2.3 Library preparation and sequencing
The V3-V4 region of the bacterial 16S rRNA gene was amplified from gDNA derived from gut samples using the S-D-Bact-0341-b-S-17 = 5′ TCG TCG GCA GCG TCA GAT GTG TAT AAG AGA CAG CCT ACG GGN GGC WGC AG and the S-D-Bact-0785-a-A-21 = 5′ GTC TCG TGG GCT CGG AGA TGT GTA TAA GAG ACA GGA CTA CHV GGG TAT CTA ATC C primers (Klindworth et al. 2013) plus Illumina adapters (in bold). The 25 μL PCR reactions contained 12.5 μL of DreamTaq Green PCR Master Mix (2X; Thermo Scientific), 125 nM each of forward and reverse 16S rRNA gene primers, and 2 µL (30–60 ng) of gDNA. The amplification was performed with a thermal cycling program under the following conditions: initial denaturation for 3 min at 95 °C, followed by 30 cycles of denaturation for 15 s at 95 °C, annealing for 30 s at 53 °C, extension for 60 s at 72 °C, and a final extension for 7 min at 72 °C. PCR amplicons were resolved on a 1.8% agarose gel. PCR for the no-template control was included.
The libraries were prepared following the instructions of the 16S Metagenomic Sequencing Library Preparation for Illumina©. The concentration of the amplicons was assessed using a Qubit 2.0 Fluorometer with the dsDNA high sensitivity kit (Thermo Scientific©, USA). Amplicon clean-up was carried out using AMPure XP beads (Beckman Coulter). Amplicons were indexed with the Illumina sequencing adapters using the Nextera XT Index Kit, purified using AMPure XP beads, and quantified fluorometrically with the dsDNA high sensitivity kit (Thermo Scientific©, USA. The integrity and quality of the final library were verified using the Agilent High Sensitivity DNA kit on the Agilent Bioanalyzer 2100 system (Agilent Technologies, USA). Libraries were pooled in equimolar concentrations and then sequenced in a paired-end (2 × 300 bp) sequencing format with a MiSeq Reagent Kit V3 (600 cycles) using the MiSeq platform (Illumina, San Diego, CA, USA). Sequencing was performed in the Sequencing Unit at the Instituto de Ecología, A.C. (Inecol), Mexico.
2.4 Data analysis
2.4.1 Bioinformatic analysis
The raw data resulting from the sequencing experiments were processed with the DADA2 package to resolve amplicon sequence variants (ASVs) (Callahan et al. 2016), using the following filtering criteria: i) an error threshold of one base in sense and two in antisense reads, and ii) removal of sequences with ambiguous bases. From the filtered sequences, error modeling was performed (Callahan et al. 2016, 2017), and surviving sequences were merged and filtered to remove chimeric sequences using the “removeBimeraDenovo” algorithm with the “consensus” method (Callahan et al. 2016, 2017). The resulting sequences were used to obtain the consensus sequences. Taxonomic assignment of sequences was performed with the Bayesian classification method (Wang et al. 2007) using the SILVA database version 138 (Quast et al. 2013). The results were integrated with the metadata into a phyloseq object with the Phyloseq package (McMurdie and Holmes 2013). Sequences not identified at the Phylum level and those identified as "Mitochondrion" or "Chloroplast" at any taxonomic level were removed with the Phyloseq base functions (Callahan et al. 2016, 2017). The phyloseq object was transformed into an MPSE object with the MicrobiotaProccess package (Xu et al. 2023), and all samples were rarefied with the mp_cal_rarecurve function. With the microbiome package (Lahti and Shetty 2012), alpha diversity values (Observed and Shannon) were evaluated for each group of samples according to their origin and analyzed for statistical significance using a Wilcoxon test. To test whether the bacterial communities differed among sites, we statistical tested the Beta diversity of the bacterial communities using a PERMANOVA analysis (pairwise test with 999 permutations and including the Climate + Species + Sex interaction), based on Bray distances. A PCoA estimated with the Hellinger abundances and the Bray distances were plotted to visualize clustering patterns in the data according to the host agave species. These analyses were also done using MicrobiotaProcess package (Xu et al. 2023). We also performed a hierarchical cluster analysis using the Hellinger transformed abundances and Bray distances to find the clustering pattern of samples. The results were visualized with the ggtree (Yu 2020), ggtreeExtra (Xu et al. 2021), and ggplot2 (Wickham 2011) packages using the top 20 most abundant genera. To find enriched genera among the bacterial communities from the different host agave species, a linear discriminant analysis with effect size (LEfSe) was performed with the microbiomeMarker package (Cao et al. 2022) using a p-value cut-off of 0.05 for both the Wilcoxon and Kruskal-Wallis tests. The core bacterial community was determined using the 'core' function of the microbiome package (Lahti and Shetty 2012) using 0.001 and 0.5 for the detection and prevalence parameters, respectively. In all cases, plots were customized with the functions of the ggplot2 package in (Wickham 2011), and all analyses were done under the R environment (R Core Team 2022).
3 Results
Although we sampled six biological replicates per site in the field, we lost some samples during the DNA extraction/amplification, and/or sequencing steps and were unable to preserve the original number of samples. So, we finally retained four datasets for Guerrero, five for Jalisco and Tlaxcala, and six for Michoacán (Figs. 1 and 2, and Supplementary Table 1). The raw data are available in the NCBI SRA under the Bioproject PRJNA1004019.
Relative abundances at the family level. The relative abundance profile of the gut bacterial communities in the samples vary widely according to host plant. Composition profiles show Prevolletacea, Lactobacillaceae, and Enterobacteracea families as the dominant families. Gue: Guerrero; Mich: Michoacán; Tlax: Tlaxcala; Jal: Jalisco
The total raw reads obtained was ≈285 × 103 ± 150 × 103 (mean ± SD) while the reads after filtering was ≈64.4 × 103 ± 38.4 × 103 (mean ± SD) (Detailed data in Supplementary Fig. 1). The rarefaction curves constructed to estimate the extent of the analysis showed that the curves of all libraries reached or nearly reached the plateau phase, supporting the depth of the sequencing effort (Supplementary Fig. 1). Across all the samples, the ASVs identified were distributed among 23 bacterial phyla, 43 classes, 106 orders, 154 families, and 247 genera. In Fig. 2, we show the relative abundances by family, which varied widely depending on the location and the host plant. The dominant families were Prevolletacea, Lactobacillaceae, and Enterobacteracea. In general, within each host plant, gut bacterial communities exhibited similar alpha diversity with no statistically significant differences among sexes or hosts (Fig. 3A and B, Table 1).
Alpha diversity comparisons of the gut microbiota of S. acupunctatus from different host plants and between insect sexes. The gut microbiota exhibited similar alpha diversity values among comparison groups, with no statistically significant differences among A) hosts or B) sexes. The p-value of the Wilcoxon test is indicated above each comparison
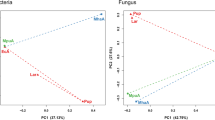
Hierarchical clustering analyses based on the relative abundance of the 20 most abundant taxa showed that the gut bacterial communities of adult S. acupunctatus were strongly influenced by host plant and climate (Fig. 4). The PCoA analysis showed clustering of the bacterial communities according to their origin (Fig. 5), which was supported by significant differences among host plants and climate in the PERMANOVA test (Table 1). However, we did not find differences among sexes (Table 1). In addition, using the LEfSe analysis, we identified some genera that are enriched in the gut bacterial communities of S. acupuntactus depending on the plant host (Fig. 6). Finally, the size and composition of the core bacterial community are shown in Fig. 7, which belonged to the genera Prevotella, Pectinatus, Liquorilactobacillus, Secundilactobacillus, Paucilactobacillus, and Pseudomonas.
4 Discussion
S. acupuntactus is a pestiferous insect species for agaves. The adults oviposit in the most tender parts of the buds, and the emerging larvae feed on the agave pineapples, sweeping towards the interior of the plants, forming galleries and perforating the leaves, stems, roots, and stalks. This boring causes rot, either by oxidation of the plant tissues or by the development of phytopathogens, leading to the decline and death of the affected plants (Solís-Aguilar et al. 2001; Palemón-Alberto et al. 2022).
To a large extent, S. acupunctatus could not cause this degree of damage to agave plants if it were not able to feed on them. As a result, the nutritional exploitation of agave by S. acupunctatus must be understood, and the metabolic contribution of S. acupunctatus' gut bacterial communities to its ability to digest agave tissues must be investigated. For this reason, we consider it important to understand the assemblage of the bacterial gut bacterial communities of S. acupunctus. By doing so, we attempted to clarify two key points: the plasticity of the bacterial communities in different environmental contexts, and the existence of a core bacterial community that remains stable across environments. In general, we found that the gut bacterial communities of S. acupunctatus vary according to its environment. However, there is a core bacterial community that we assume displays a closer association with its host. Nevertheless, it is important to conduct a thorough analysis of the bacterial communities found in both the host plant and the beetles in order to validate this inference. It is worth noting that this is the first survey of the gut bacterial communities of S. acupuntactus.
We found no differences in alpha diversity among samples based on their site of origin or the sex of the insect. However, there were significant differences in beta diversity, where we identified that climate and host plants are drivers of the gut bacterial assemblage in S. acupuntactus, which concurs with results in other species (Yun et al. 2014; Jones et al. 2019). Since alpha diversity is the diversity of local collection sites, while beta diversity is the spatial change in composition between localities, these results show that that the bacterial communities of S. acupuntactus exhibits a certain level of plasticity.
To understand the differences in the structure of the bacterial communities among sites, we consider it necessary to assume that the foraging environment exerts a strong pressure on S. acupuntactus, forcing it to make adjustments to its metabolism and physiology (Adams et al. 2010; Yun et al. 2014; Shukla et al. 2016; Chakraborty et al. 2023). In general, agave sap is rich in sugars (with approximately 75% of its dry matter being mainly sucrose, fructose, and glucose), as well as fructo-oligosaccharides, cellulose, non-cellulosic polysaccharides, lignin, acetate, protein, and minerals (Ortiz-Basurto et al. 2008; Munive et al. 2014; Corbin et al. 2015). Moreover, the specific composition of agave sap is strongly affected by the agave species, the ripening stage of the plant, and the climate (Rendón-Salcido et al. 2009; Corbin et al. 2015; Ortiz-Basurto et al. 2008).
Many polymers in the agave sap are well recognized as not easily digestible, requiring the assistance of several coordinated enzymatic steps to be assimilated. Here, it seems plausible that the core bacterial community we detected can play a central role in the digestion of these polymers and the sugar components. For example, many members of the genus Prevotella are found in the gut and rumen bacterial communities of several animal species, where they play major roles in carbohydrate and nitrogen metabolism (Kim et al. 2017). Members of this genus are reported to degrade hemicellulose, pectin, and soluble xylan (Ueki et al. 2007; Dodd et al. 2010; Kabel et al. 2011). They are active hemicellulolytic components of the depolymerization of soluble wheat arabinoxylan during ruminal digestion thanks to the diverse endoxylanases and carbohydrate esterases they produce (Miyazaki et al. 1997; Dodd et al. 2010). Prevotella genes involved in fixing nitrogen have also been found to be active in the gut bacterial communities of vegan-fed mice (Gálvez et al. 2020). In the same study, it was also found that eating complex plant polysaccharides, especially arabinoxylans, makes Prevotella more abundant (Gálvez et al. 2020). This suggests a potential role of Prevotella as a metabolic helper for the degradation of complex sugars and as a nitrogen supplier for S. acupuntactus. We consider that such metabolic capacities could facilitate the digestion of complex carbohydrate polymers and deal with the high carbon:nitrogen ratio found in agave biomass.
Another genus found as a component of the core bacterial community is Pectinatus, a bacterial genus that has been extensively studied due to its role in beer spoilage. Pectinatus can use a wide range of sugars as carbon sources, leading to the metabolic production of acetic acid, propionic acid, lactic acid, succinic acid, acetoin, sulfhidric acid, and other sulfur compounds (Lee et al. 1978; Juvonen and Suihko 2006; Paradh 2015). Although we did not find reports on the role of Pectinatus in insects, its potential metabolic capacities to metabolize sugars and produce a wide range of metabolites can be useful for S. acupunctatus. For example, propionate is required for the synthesis of the juvenile hormone (Brindle et al. 1987) while the acetoin produced by gut microbiota has been demonstrated that acts as a pheromone for many insects, including weevils (Robacker and Lauzon 2002; Saïd et al. 2005; Farine et al. 2017). Due to the limited knowledge on the function of Pectinatus in insects, we strongly recommend conducting a comprehensive analysis of this genus in S. acupunctatus to establish its functional role.
Prevotella and Pectinatus are strict anaerobes (Chelack and Ingledew 1987; Miyazaki et al. 1997), which suggests that S. acupunctatus has compartments in its gut where conditions are mostly or fully anoxygenic. In accordance, other members of the core bacterial community belong to the general group of Lactic Acid Bacteria (LAB), such as the genera Liquorilactobacillus, Secundilactobacillus, and Paucilactobacillus. LAB are gram-positive bacteria that tend to dominate in anaerobic, carbohydrate‐containing environments characterized by an acidic pH and that use carbohydrates as the only or main carbon source to produce lactic acid (Axelsson and Ahrné, 2000; George et al. 2018). Many LAB have been identified as probiotics in insects, as occurs with honeybees (Vásquez et al. 2012), and for their host immune protection (Daisley et al. 2020; Iorizzo et al. 2022). Therefore, it is likely that the function of these LAB in S. acupunctus includes the utilization of sugars and that they mediate gate-keeping activities against pathogens.
In the case of Pseudomonas, it is now recognized that most species within this genus are facultative anaerobes (Kampers et al. 2021), which can help explain the coexistence of this genus with the other core members described. Finding members of Pseudomonas among the core bacterial community of this insect is relevant because several species have been recognized as ligninolytic microorganisms (i.e., they participate in the lignin degradation; Grgas et al. 2023). Furthermore, members of Pseudomonas have been related to detoxification activities in other curculionide beetles and to insecticide resistance in other insects, such as S. frugiperda larvae, due to their high efficacy for the biodegradation of toxic compounds (Ceja-Navarro et al. 2015; Almeida et al. 2017). Many Pseudomonas species also are nitrogen fixers under microaerophilic conditions (Desnoues et al. 2003; Fox et al. 2016; Sanow et al. 2023), supporting the idea that S. acupunctatus' gut bacterial communities provides nitrogen to its host. However, all these ideas require functional testing.
The gut bacterial communities of S. acupunctus may be also shaped by its interaction with the bacterial communities associated with agaves. For example, it has been reported that the bacterial communities in the episphere of agaves vary among agave species, while those in the endosphere are affected by the season (Desgarennes et al. 2014). It is likely that some of the bacteria found in its gut have been recruited from the host plant and incorporated as metabolic helpers. However, to obtain a clearer idea of the contribution of the plant bacterial communities to the gut bacterial communities of S. acupunctus, a comparison of the gut bacterial communities of S. acupunctus with those of the episphere and endosphere of healthy and S. acupunctus-infested agaves must be done.
It is plausible that some components of the bacterial communities of the gut act as opportunistic pathogens that proliferate on the plant tissues affected by the borer activity of S. acupunctus, resulting in the Bud Soft Rot (BSR). In fact, the idea that S. acupunctus is the vector of the microorganisms causing the BSR has been largely discussed in the literature (Solís-Aguilar et al. 2001; Cuervo-Parra et al. 2019). For example, it has been proposed that this disease is caused by different pathogens, including bacteria and fungi such as Pectobacterium carotovorum, Erwinia cacticida, Bacillus pumilus, Pantoea agglomerans, Pseudomonas sp., Pantoea dispersa, and Fusarium oxysporum (Aquino-Bolaños et al. 2020; Reyes-Zambrano et al. 2020; Palemón-Alberto et al. 2021). So, given the multifactorial nature of the potential causative agents of the BSR, it has been postulated that BSR constitutes a syndrome rather than a disease (López-Bautista et al. 2020). If we observe the enriched genera in the gut of S. acupunctus in relation to the host species, we can determine that the majority of them are anaerobes and facultative anaerobes, which is consistent with the notion of the anoxygenicity of the gut of S. acupunctus, and that many of them have the potential to cause soft rot, as reported in other plant species (Sawada et al. 2021; Zhao et al. 2022; Osei et al. 2022). This may explain why it is difficult to associate BSR with a single microbial infection for all agave species, and it could instead be a pathobiome-caused syndrome. At this point, we encourage a metabarcoding survey to explore the microbial communities associated with BSR in different host species and interpret the information in light of the pathobiome concept and using culture-independent techniques. This idea can open the door to considering the BSR as a complex, multifactorial phenomenon that involves a complete pathobiome that is structured differently according to the plant host and the surrounding bacterial communities. Special mention should be made of the overrepresentation of Prevotella in the gut of S. acupunctus on A. tequilana, not only because it is the most abundant member of the core, but also because this genus has been positively correlated as a major contributor to the development and aggravation of Dickeya zeae, the causal agent of the rice foot rot (Bez et al. 2021). It is probable that Prevotella is involved in the development of the BSR, but the lack of previous mention of this genus could reflect the fact that most studies reporting cultured bacteria associated with the BSR use general, non-specialized culture media and aerobic conditions. For this reason, we encourage further research to extend these culture-based surveys to the analysis of anaerobes, where the use of culturomics approaches could be advantageous.
One of the limitations of this study is that we focused solely on the S. acupunctus gut bacterial communities. Bacteria are only one part of a larger story in which fungi and other groups can play additional roles. Thus, it remains to be determined how the fungal communities of this insect is structured and what role it may play for its host. Examining the bacterial communities associated with S. acupunctus at different stages of its life cycle could provide useful insights into the development effect on the establishment and assembly of microbial communities in this species.
5 Conclusion
In this work, we surveyed the bacterial community present in the gut of S. acupunctus for the first time and determined that its assemblage is shaped by the host plant and the climate. Despite this, we found a core bacterial community that we propose to be functionally relevant. These data are relevant because they provide the basis for proposing hypotheses to guide further work to understand the complex interactions between S. acupunctus and their host plants. We encourage testing of the functional role of the microorganisms associated with S. acupunctus here described.
References
Adams AS, Adams SM, Currie CR, Gillette NE, Raffa KF (2010) Geographic variation in bacterial communities associated with the red turpentine beetle (Coleoptera: Curculionidae). Environ Entomol 39(2):406–414
Almeida LGD, Moraes LABD, Trigo JR, Omoto C, Consoli FL (2017) The gut microbiota of insecticide-resistant insects houses insecticide-degrading bacteria: A potential source for biotechnological exploitation. PLoS ONE 12(3):e0174754. https://doi.org/10.1371/journal.pone.0174754
Angzzas SMK, Ashuvila MA, Dayang NFAZ (2016) Potential lignin degraders isolated from the gut of Rhynchophorus ferrugineus. In: International Conference on Mechanics, Materials and Structural Engineering (ICMMSE 2016). Atlantis Press, pp 124–130. https://doi.org/10.2991/icmmse-16.2016.22
Aquino-Bolaños T, Sánchez-García JA, Ortíz-Hernández YD, Hernández-Cruz J, Cortés-Martínez CI (2020) Carrier and vector of Pectobacterium carotovorum subsp. carotovorum and its handling through a base of entomopathogenic fungi in Agave sp. Fla Entomol 103(2):243–246
Axelsson L, Ahrné S (2000) Lactic acid bacteria. Applied microbial systematics. Springer Netherlands, Dordrecht, pp 367–388
Bez C, Esposito A, Thuy HD et al (2021) The rice foot rot pathogen dickeya zeae alters the in-field plant microbiome. Environ Microbiol 23(12):7671–7687
Brindle PA, Baker FC, Tsai LW, Reuter CC, Schooley DA (1987) Sources of propionate for the biogenesis of ethyl-branched insect juvenile hormones: role of isoleucine and valine. Proc Natl Acad Sci U S A 84(22):7906–7910. https://doi.org/10.1073/pnas.84.22.7906
Cao Y, Dong Q, Wang D, Zhang P, Liu Y, Niu C (2022) MicrobiomeMarker: an R/Bioconductor package for microbiome marker identification and visualization. Bioinformatics 38(16):4027–4029. https://doi.org/10.1093/bioinformatics/btac438
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583
Callahan BJ, McMurdie PJ, Holmes SP (2017) Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J 11(12):2639–2643
Ceja-Navarro JA, Vega FE, Karaoz U, Hao Z et al (2015) Gut microbiota mediate caffeine detoxification in the primary insect pest of coffee. Nat Commun 6:7618. https://doi.org/10.1038/ncomms8618
Chakraborty A, Ashraf MZ, Modlinger R, Synek J, Schlyter F, Roy A (2020) Unravelling the gut bacteriome of Ips (Coleoptera: Curculionidae: Scolytinae): identifying core bacterial assemblage and their ecological relevance. Sci Rep 10:18572
Chakraborty A, Purohit A, Khara A, Modlinger R, Roy A (2023) Life-stage and geographic location determine the microbial assemblage in Eurasian spruce bark beetle, Ips typographus L. (Coleoptera: Curculionidae). Front For Glob Change 6(1176160):10–3389
Chelack BJ, Ingledew WM (1987) Anaerobic Gram-negative bacteria in brewing - a review. J Am Soc Brew Chem 45(4):123–127
Clemens K (2015) Geocoding with openstreetmap data. GEOProcessing 10
Corbin K, Byrt C, Bauer S, DeBolt S, Chambers D et al (2015) Prospecting for energy-rich renewable raw materials: agave leaf case study. PLoS ONE 8(10):e0135382. https://doi.org/10.1371/journal.pone.0135382
Cruz Faustino JJ, Figueroa Castro P, Alcántara Jimenéz JA, López Martínez V, Silva García F (2019) Vegetal synergists for trapping the adult of Scyphophorus acupunctatus Gyllenhal, in pheromone baited traps, in Agave angustifolia Haw., in Morelos, Mexico. Acta Zool Mex 35. https://doi.org/10.21829/azm.2019.3502187
Cruz-Jardón LF, Figueroa-Castro P, López-Martínez V, Pérez-Figueroa M (2018) Semiochemicals-baited traps for detecting and estimating the population density of Scyphophorus acupunctatus Gyllenhal (Coleoptera: Dryopthoridae), in agaves, in Tlaquiltenango, Morelos. Acta Zool Mex 34. https://doi.org/10.21829/azm.2018.3412163
Cuervo-Parra JA, Pérez-España VH, Pérez PAL, Morales-Ovando MA, Arce-Cervantes O, Aparicio-Burgos JE, Romero-Cortes T (2019) Scyphophorus acupunctatus (Coleoptera: Dryophthoridae): a weevil threatening the production of agave in Mexico. Fla Entomol 102:1–9
Da Silva Brito SS, Villa M, Benhadi-Marín J, da Silva F, Pereira JA (2021) The temporal and spatial variation of arthropod associations inhabiting non-crop vegetation in a Sisal crop, Agave sisalana in the Caatinga biome. Appl Sci 11(14):6498. https://doi.org/10.3390/app11146498
Daisley BA, Pitek AP, Chmiel, et al (2020) Lactobacillus spp. attenuate antibiotic-induced immune and microbiota dysregulation in honey bees. Commun Biol 3(1):534. https://doi.org/10.1038/s42003-020-01259-8
Desgarennes D, Garrido E, Torres-Gomez MJ et al (2014) Diazotrophic potential among bacterial communities associated with wild and cultivated Agave species. FEMS Microbiol Ecol 90(3):844–857. https://doi.org/10.1111/1574-6941.12438
Desnoues N, Lin M, Guo X, Ma L, Carreño-Lopez R, Elmerich C (2003) Nitrogen fixation genetics and regulation in a Pseudomonas stutzeri strain associated with rice. Microbiology 149(8):2251–2262
Dodd D, Moon YH, Swaminathan K, Mackie RI, Cann IK (2010) Transcriptomic analyses of xylan degradation by Prevotella bryantii and insights into energy acquisition by xylanolytic bacteroidetes. J Biol Chem 285(39):30261–30273. https://doi.org/10.1074/jbc.M110.141788
Dumond L, Lam LPY et al (2021) Termite gut microbiota contribution to wheat straw delignification in anaerobic bioreactors. ACS Sustain Chem Eng 9(5):2191–2202. https://doi.org/10.1021/acssuschemeng.0c07817
Dunnington D, Thorne B (2020) ggspatial: Spatial Data Framework for ggplot2. R package version 1.1.4
Farine JP, Habbachi W, Cortot J, Roche S, Ferveur JF (2017) Maternally-transmitted microbiota affects odor emission and preference in Drosophila larvae. Sci Rep 7(1):6062. https://doi.org/10.1038/s41598-017-04922-z
Figueroa-Castro P, González-Hernández H, Carrillo-Sánchez JL, Solís-Aguilar JF, Real-Laborde JId, Rojas JC (2016) Effect of the height and distribution pattern of pheromone-baited traps on the capture of Scyphophorus acupunctatus (Coleoptera: Dryophthoridae) on blue agave (Asparagales: Asparagaceae). Fla Entomol 2(99):297–299. https://doi.org/10.1653/024.099.0222
Flint HJ, Bayer EA, Rincon MT, Lamed R, White BA (2008) Polysaccharide utilization by gut bacteria: potential for new insights from genomic analysis. Nat Rev Microbiol 6(2):121–131. https://doi.org/10.1038/nrmicro1817
Fox AR, Soto G, Valverde C et al (2016) Major cereal crops benefit from biological nitrogen fixation when inoculated with the nitrogen-fixing bacterium Pseudomonas protegens Pf-5 X940. Environ Microbiol 18(10):3522–3534
George AS et al (2018) Interactions of Salmonella enterica serovar Typhimurium and Pectobacterium carotovorum within a tomato soft rot. Appl Environ Microbiol 84(5):e01913-e1917
Gálvez EJ, Iljazovic A, Amend L, Lesker TR, Renault T, Thiemann S, Strowig T (2020) Distinct polysaccharide utilization determines interspecies competition between intestinal Prevotella spp. Cell Host & Microbe 28(6):838–852
González-Hernández H, Solís-Aguilar JF, Pacheco Sánchez C, Flores-Mendoza FJ, Rubio-Cortés R, Rojas-León JC (2007) Insectos barrenadores del agave tequilero. In: González-Hernandéz H, del Real Laborde JI, Solís-Aguilar JF (eds), Manejo de Plagas del Agave Tequilero. Colegio de Postgraduados and Tequila Sauza, S.A. de C.V., Zapopan, Jalisco, México, pp 39–67
Grgas D, Rukavina M, Bešlo D, Štefanac T, Crnek V, Šikić T, ... Landeka Dragičević T (2023) The bacterial degradation of lignin—a review. Water 15(7):1272
He B, Chen X, Yang H, Cernava T (2021) Microbiome structure of the aphid Myzus persicae (Sulzer) is shaped by different solanaceae plant diets. Front Microbiol 12:667257. https://doi.org/10.3389/fmicb.2021.667257
Huang K, Wang J, Huang J et al (2021) Host phylogeny and diet Shape gut microbial communities within bamboo-feeding insects. Front Microbiol 12:633075. https://doi.org/10.3389/fmicb.2021.633075
Ibarra-Juárez LA et al (2020) Evidence for succession and putative metabolic roles of fungi and bacteria in the farming mutualism of the ambrosia beetle Xyleborus affinis. Msystems 5(5):e00541-e620. https://doi.org/10.1128/mSystems.00541-20
Iorizzo M, Letizia F, Ganassi S et al (2022) Functional properties and antimicrobial activity from lactic acid bacteria as resources to improve the health and welfare of honey bees. Insects 13(3):308. https://doi.org/10.3390/insects13030308
Jones AG, Mason CJ, Felton GW, Hoover K (2019) Host plant and population source drive diversity of microbial gut communities in two polyphagous insects. Sci Rep 9:2792
Juvonen R, Suihko M (2006) Megasphaera paucivorans Sp. Nov, Megasphaera sueciensis sp. nov. and Pectinatus haikarae sp. nov., isolated from brewery samples, and emended description of the genus Pectinatus. Int J Syst Evol Microbiol 4(56):695–702. https://doi.org/10.1099/ijs.0.63699-0
Kabel MA, Yeoman CJ, Han Y et al (2011) Biochemical characterization and relative expression levels of multiple carbohydrate esterases of the xylanolytic rumen bacterium Prevotella ruminicola 23 grown on an ester-enriched substrate. Appl Environ Microbiol 77(16):5671–5681. https://doi.org/10.1128/AEM.05321-11
Kahle DJ, Wickham H (2013) ggmap: spatial visualization with ggplot2. R J 5(1):144
Kampers LF, Koehorst JJ, van Heck RJ, Suarez-Diez M, Stams AJ, Schaap PJ (2021) A metabolic and physiological design study of Pseudomonas putida KT2440 capable of anaerobic respiration. BMC Microbiol 21:1–15. https://doi.org/10.1186/s12866-020-02058-1
Kim JN, Méndez-García C, Geier RR et al (2017) Metabolic networks for nitrogen utilization in Prevotella ruminicola 23. Sci Rep 7:7851. https://doi.org/10.1038/s41598-017-08463-3
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41(1), e1-e1
Lahti L, Shetty S (2012) microbiome R package (2012–2019)
Lee SY, Mabee MS, Jangaard NO (1978) Pectinatus, a new genus of the family Bacteroidaceae. Int J Syst Bacteriol 4(28):582–594. https://doi.org/10.1099/00207713-28-4-582
Li J, Wang S, Zhao J, Dong Z, Shao T (2022) Gut microbiota of Ostrinia nubilalis larvae degrade maize cellulose. Front Microbiol 13:816954. https://doi.org/10.3389/fmicb.2022.816954
López-Bautista V, Mora-Aguilera G, Gutiérrez-Espinosa MA et al (2020) Morphological and molecular characterization of Fusarium spp. associated to the regional occurrence of wilt and dry bud rot in Agave tequilana. Rev Mex Fitopat 38:79–106. https://doi.org/10.18781/r.mex.fit.1911-4
Miyazaki K, Martin JC, Marinsek-Logar R, Flint HJ (1997) Degradation and utilization of xylans by the rumen anaerobe Prevotella bryantii (formerly P. ruminicola subsp. brevis) B14. Anaerobe 3(6):373–381. https://doi.org/10.1006/anae.1997.0125
McMurdie, P. J., & Holmes, S. (2013). phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PloS one 8(4):e61217
Munive F, Páez M, Romero-Granja C, Espín N, Casa-Villegas M (2014) Fermentation of Agave americana L. sap produced in Cayambe-Ecuador. Rev Bionatura 8:15
Ortiz-Basurto R, Pourcelly G, Doco T, Williams P, Dornier M, Belleville M (2008) Analysis of the main components of the aguamiel produced by the maguey-pulquero (Agave mapisaga) throughout the harvest period. J Agric Food Chem 10(56):3682–3687. https://doi.org/10.1021/jf072767h
Osei R, Yang C, Cui L, Ma T, Li Z, Boamah S (2022) Isolation, identification, and pathogenicity of Lelliottia amnigena causing soft rot of potato tuber in China. Microb Pathog 164:105441. https://doi.org/10.1016/j.micpath.2022.105441
Palemón-Alberto F, Ortega-Acosta SA, Domínguez-Monge S, Castañeda-Vildozola A, Reyes-García G, Cruz-Lagunas B, Flores-Simon OU (2021) First report of bud soft rot on Agave angustifolia caused by Pantoea dispersa in México. Plant Dis 105(10):3286. https://doi.org/10.1094/PDIS-02-21-0316-PDN
Palemón-Alberto F, Ortega-Acosta SA, Castañeda-Vildozola A, Reyes-García G et al (2022) Damage by Scyphophorus acupunctatus Gyllenhal in Species of Agave. Southwest Entomol 47(2):437–442. https://doi.org/10.3958/059.047.0219
Paradh AD (2015) Gram-negative spoilage bacteria in brewing. In: Brewing microbiology. Woodhead Publishing, pp 175–194
Poveda J (2021) Insect frass in the development of sustainable agriculture. A Review. Agron Sustain Dev 41:5. https://doi.org/10.1007/s13593-020-00656-x
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO. 2013. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41(Database issue):D590–6. https://doi.org/10.1093/nar/gks1219
R Core Team (2022) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/ Accessed 23 Mar 2023
Rendón-Salcido LA, Colunga-GarcíaMarín P, Barahona-Pérez LF et al (2009) Sugars and alcoholic byproducts from henequen (Agave fourcroydes) as influenced by plant age and climate. Rev Fitotec 32:39–44
Reyes-Zambrano SJ, Lecona-Guzmán CA, Gutiérrez-Miceli, et al (2020) Scanning electron microscopy and enzymatic analysis in Agave americana during Fusarium oxysporum infection. Rev Mex Fitopatol 38(3):408–419. https://doi.org/10.18781/r.mex.fit.2005-3
Robacker DC, Lauzon CR (2002) Purine metabolizing capability of Enterobacter agglomerans affects volatiles production and attractiveness to Mexican fruit fly. J Chem Ecol 28:1549–1563. https://doi.org/10.1023/a:1019920328062
Rozadilla G, Cabrera N, Virla E, Greco N, McCarthy C (2020) Gut Microbiota of Spodoptera frugiperda (J.E. Smith) larvae as revealed by metatranscriptomic analysis. J Appl Entomol 5(144):351–363. https://doi.org/10.1111/jen.12742
Rubio-Cortés R (2007) Enfermedades del cultivo de agave. In: Rulfo-Vilchis O, Pérez-Domínguez JF, del Real Laborde JI, Byerly-Murphy KF (eds), Conocimiento y prácticas agronómicas para la producción de Agave tequilana Weber en la zona de denominación de origen del tequila. Instituto Nacional de Investigaciones Forestales, Agrícolas y Pecuarias. Centro de Investigación Regional del Pacífico Centro. Libro técnico Núm. 4. México, pp. 169–195
Saïd I, Renou M, Morin JP, Ferreira JM, Rochat D (2005) Interactions between acetoin, a plant volatile, and pheromone in Rhynchophorus palmarum: Behavioral and olfactory neuron responses. J Chem Ecol 31:1789–1805. https://doi.org/10.1007/s10886-005-5927-4
Sanow S, Kuang W, Schaaf G et al (2023) Molecular mechanisms of Pseudomonas assisted plant nitrogen uptake-opportunities for modern agriculture. Mol Plant Microbe Interact. https://doi.org/10.1094/MPMI-10-22-0223-CR
Sawada H, Fujikawa T, Tsuji M, Satou M (2021) Pseudomonas allii sp. nov., a pathogen causing soft rot of onion in Japan. Int J Syst Evol Microbiol 71:004582. https://doi.org/10.1099/ijsem.0.004582
Shukla SP, Sanders JG, Byrne MJ, Pierce NE (2016) Gut microbiota of dung beetles correspond to dietary specializations of adults and larvae. Mol Ecol 25(24):6092–6106
Solís-Aguilar JF, González-Hernández H, Leyva-Vázquez JL, Equihua-Martínez A, Flores Mendoza FJ, Martínez-Garza A (2001) Scyphophorus acupunctatus Gyllenhal, plaga del agave tequilero en Jalisco, México. Agrociencia 35(6):663–670
Suárez-Moo P, Cruz-Rosales M, Ibarra-Laclette E et al (2020) Diversity and composition of the gut microbiota in the developmental stages of the dung beetle Copris incertus Say (Coleoptera, Scarabaeidae). Front Microbiol 11:1698. https://doi.org/10.3389/fmicb.2020.01698
Ueki A, Akasaka H, Satoh A, Suzuki D, Ueki K (2007) Prevotella paludivivens sp. nov., a novel strictly anaerobic, gram-negative, hemicellulose-decomposing bacterium isolated from plant residue and rice roots in irrigated rice-field soil. Int J Syst Evol Microbiol 57:1803–1809. https://doi.org/10.1099/ijs.0.64914-0
Valdés-Rodríguez S, Ramírez-Choza JL, Reyes-López J, Blanco-Labra A (2004) Respuestas del insecto Max (Scyphophorus acupunctatus Gyllenhal [Coleoptera: Curculionidae] hacia algunos compuestos atrayentes del henequén. Acta Zool Mex 20(3):157–166
Vásquez A, Forsgren E, Fries I et al (2012) Symbionts as major modulators of insect health: lactic acid bacteria and honeybees. PLoS ONE 7(3):e33188. https://doi.org/10.1371/journal.pone.0033188
Vega-Petlacalco M, Arzuffi R, Valdez J et al (2018) Food quality influences ovarian development Inscyphophorus acupunctatus (Coleoptera: Dryophthoridae). Fla Entomol 3(101):447–452. https://doi.org/10.1653/024.101.0301
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73(16):5261–5267
Wang L, Liu T, Wu Y et al (2018) Bacterial microbiota assemblage in Aedes albopictus mosquitoes and its impacts on larval development. Mol Ecol 14(27):2972–2985. https://doi.org/10.1111/mec.14732
Wang K, Gao P, Geng L, Liu C, Zhang J, Shu C (2022) Lignocellulose degradation in Protaetia brevitarsis larvae digestive tract: refining on a tightly designed microbial fermentation production line. Microbiome 10:1–16
Waring GL, Smith RL (1986) Natural history and ecology of Scyphophorus acupunctatus (Coleoptera: Curculionidae) and its associated microbes in cultivated and native agaves. Ann Entomol Soc Am 79(2):334–340. https://doi.org/10.1093/aesa/79.2.334
Wickham H (2011) ggplot2. Wiley interdisciplinary reviews: computational statistics. 3(2):180–185
Xie S, Lan Y, Sun C, & Shao, Y. (2019). Insect microbial symbionts as a novel source for biotechnology. World J Microbiol Biotechnol 35:1–7
Xu S et al (2021) ggtreeExtra: compact visualization of richly annotated phylogenetic data. Mol Biol Evol 38(9):4039–4042
Xu S, Zhan L, Tang W et al (2023) MicrobiotaProcess: a comprehensive R package for deep mining microbiome. The Innovation 4(2). https://doi.org/10.18129/B9.bioc.MicrobiotaProcess
Yu G (2020) Using ggtree to visualize data on tree-like structures. Curr Protocol Bioinforma 69:e96
Yun J et al (2014) Insect gut bacterial diversity determined by environmental habitat, diet, developmental stage, and phylogeny of host. Appl Environ Microbiol 80(17):5254–5264
Zhao X, Tian Y, Yue L et al (2022) Identification and characterization of pathogenicity of Lelliottia nimipressuralis causing soft rot of Codonopsis pilosula (dangshen) roots in China. Plant Pathol 71(8):1801–1811. https://doi.org/10.1111/ppa.13606
Acknowledgements
Salazar-Rivera G.I. thanks Mexico’s Consejo Nacional de Humanidades, Ciencias y Tecnología (CONAHCYT) for postdoctoral fellowship 239732 and the Centro de Investigación y Asistencia Tecnológica del Estado de Jalisco (CIATEJ). The anonymous reviewers deserve gratitude for their constructive feedback, which significantly enhanced this article. This research was supported by CONAHCYT-FORDECYT 292474 and 296369.
Funding
JNEV, EIL, JAZB.
Author information
Authors and Affiliations
Contributions
Conceived and designed the analysis: GSR, JNEV, APS, JAZB.
Collected the data: GSR, MOM, JNEV.
Contributed data or analysis tools: GSR, MOM, JNEV, JAZB.
Performed the analysis: GSR, AGMAC, APS, JAZB.
Review and edit the manuscript: AGMAC, EIL, IMHV.
Wrote the paper: GSR, APS, IMHV, JNEV, JAZB.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Salazar-Rivera, G.I., Pereira-Santana, A., Hernández-Velázquez, I.M. et al. Disentangling the gut bacterial communities of the agave weevil, Scyphophorus acupunctatus (Coleoptera: Curculionidae). Symbiosis 92, 381–392 (2024). https://doi.org/10.1007/s13199-024-00978-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-024-00978-4