Abstract
Background
Cancer cell growth, metastasis, and drug resistance are major challenges in treating liver hepatocellular carcinoma (LIHC). However, the lack of comprehensive and reliable models hamper the effectiveness of the predictive, preventive, and personalized medicine (PPPM/3PM) strategy in managing LIHC.
Methods
Leveraging seven distinct patterns of mitochondrial cell death (MCD), we conducted a multi-omic screening of MCD-related genes. A novel machine learning framework was developed, integrating 10 machine learning algorithms with 67 different combinations to establish a consensus mitochondrial cell death index (MCDI). This index underwent rigorous evaluation across training, validation, and in-house clinical cohorts. A comprehensive multi-omics analysis encompassing bulk, single-cell, and spatial transcriptomics was employed to achieve a deeper insight into the constructed signature. The response of risk subgroups to immunotherapy and targeted therapy was evaluated and validated. RT-qPCR, western blotting, and immunohistochemical staining were utilized for findings validation.
Results
Nine critical differentially expressed MCD-related genes were identified in LIHC. A consensus MCDI was constructed based on a 67-combination machine learning computational framework, demonstrating outstanding performance in predicting prognosis and clinical translation. MCDI correlated with immune infiltration, Tumor Immune Dysfunction and Exclusion (TIDE) score and sorafenib sensitivity. Findings were validated experimentally. Moreover, we identified PAK1IP1 as the most important gene for predicting LIHC prognosis and validated its potential as an indicator of prognosis and sorafenib response in our in-house clinical cohorts.
Conclusion
This study developed a novel predictive model for LIHC, namely MCDI. Incorporating MCDI into the PPPM framework will enhance clinical decision-making processes and optimize individualized treatment strategies for LIHC patients.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liver hepatocellular carcinoma (LIHC) is a highly prevalent malignancy worldwide and ranks as the third most common cause of cancer-related deaths globally [1]. Despite substantial advances in cancer diagnosis and therapy, most LIHC patients are diagnosed at an advanced stage, precluding the possibility of curative interventions and resulting in dismal clinical outcomes. Therefore, identifying more sensitive diagnostic and prognostic biomarkers, as well as effective predictors to stratify patients for personalized therapy, is of great scientific significance and clinical importance in the treatment of LIHC. Immunotherapies, such as immune checkpoint blockade (ICB) therapy, have exhibited clinical efficacy in a subset of LIHC patients. However, due to the intricate tumor immune microenvironment (TME) and the high heterogeneity of LIHC, the cellular mechanisms of the LIHC response to immunotherapy remain poorly understood, and only a minority of patients respond favorably to ICI treatment [2]. Sorafenib was the first approved first-line targeted therapy for advanced LIHC; nevertheless, the inevitable development of acquired resistance limits its efficacy [3]. Therefore, the exploration of the underlying mechanisms concerning the response to immunotherapy and targeted therapy in LIHC can provide potential targets and a theoretical basis for drug design and clinical treatment.
Mitochondria play a significant role in regulating cell death-related pathways, and dysregulation of mitochondrial function can activate alternative cell death signals, including those related to apoptosis, autophagy, necroptosis, ferroptosis, pyroptosis, cuproptosis, and disulfidptosis, which are implicated in the development and progression of various cancers, such as LIHC, lung cancer, and breast cancer [4,5,6,7,8,9]. Apoptosis is a well-studied form of programmed cell death (PCD) that prevents cancer development by eliminating damaged or abnormal cells through a series of morphological and biochemical changes [7, 9, 10]. Multiple experiments, both in vivo and in vitro, have demonstrated that defects in apoptosis are closely associated with the proliferation and differentiation of hepatocellular carcinoma cells [9, 10]. Autophagy is a cellular process in which damaged organelles and proteins are degraded, and this process plays a crucial role in maintaining cellular homeostasis. Dysregulation of autophagy is also connected with the development and progression of multiple cancers [11, 12]. Accumulating evidence demonstrates that autophagy can affect various biological behaviors of LIHC, including occurrence, development, recurrence, metastasis, and drug resistance [13]. Ferroptosis is another type of proinflammatory cell death characterized by the accumulation of lipid peroxides and iron-dependent reactive oxygen species [14], which correlates with the occurrence and development of various diseases, such as Parkinson’s disease, ischemia–reperfusion injury, and tumors. It has been reported that LIHC shows varying degrees of iron metabolism disorders and ferroptosis characteristics, such as lipid peroxidation accumulation leading to the progression of LIHC and resistance to sorafenib treatment [15]. Recently, Xiaoguang Liu et al. identified a new type of mitochondrial metabolism-related PCD named disulfidptosis [5]. According to the study, high expression of SLC7A11 in kidney cancer cells accelerates the depletion of NADPH in the cytoplasm under glucose-deprived conditions, which results in an inalterable accumulation of disulfides and induces disulfide stress, ultimately leading to disulfidptosis. However, the role of disulfidptosis-related genes in LIHC remains unclear. In addition, there are also various forms of PCD that are related to mitochondrial metabolism function, including necroptosis, pyroptosis, and cuproptosis, which also play a significant role in the carcinogenesis and development of LIHC [6, 11]. Although the role of the main mitochondrial cell death forms in cancers has been widely studied, the association between these death forms as a whole and antitumor immunity in LIHC requires further elucidation.
Working hypothesis
In the field of modern medicine, the concept of 3P medicine, encompassing predictive, preventive, and personalized strategies, has emerged as a pivotal approach to enhancing patient care. Predictive medicine aims to forecast disease risk and therapeutic outcomes, preventive medicine focuses on measures to avert disease onset or progression, and personalized medicine tailors treatments to an individual’s genetic and lifestyle factors. By integrating these principles, we can better address the challenges posed by LIHC, where early detection and effective, tailored interventions are crucial for improving patient outcomes. Therefore, exploring the role of mitochondrial cell death (MCD)-related genes in LIHC could provide a better understanding of the molecular mechanism of LIHC carcinogenesis and a novel perspective for personalized LIHC therapy in the context of 3P medicine.
Study design
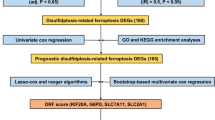
In the present study, we identified nine mitochondria-associated cell death signature genes (MCDS) in LIHC and constructed a mitochondria cell death index (MCDI) to explore the association of MCDS and MCDI with the carcinogenesis of LIHC at the bulk, single-cell, and spatial transcriptomic levels. The genetic and mutation landscape of these genes in LIHC was investigated, and a predictive model for LIHC was constructed. Moreover, the association of MCDS levels and the MCDI with the immune system features was examined. In addition, the immunotherapy and targeted therapy response of MCDS/MCDI in LIHC was also investigated and verified. The purpose of our study is to offer guidance for prognosis forecasting and therapeutic strategies, aligning with the principles of 3P medicine. The workflow of our study is visually represented in the graphic abstract, illustrating the methodological approach undertaken.
Materials and methods
Data acquisition and preprocessing
RNA-sequencing (RNA-seq) expression data (log2 converted) of 371 LIHC patients and 276 normal samples were obtained from The Cancer Genome Atlas (TCGA): https://portal.gdc.com and the GTEx databases: https://commonfund.nih.gov/GTEx, respectively. Transcriptomic expression profiling data of three LIHC cohorts were downloaded from the Gene Expression Omnibus (GEO): https://www.ncbi.nlm.nih.gov/geo/ (GSE14520), International Cancer Genome Consortium (ICGC): https://dcc.icgc.org/ (ICGC-LIRI), and Wang’s study: [16,17,18,19], respectively. In addition, the corresponding clinicopathological characteristics of the LIHC patients from the cohorts mentioned above were also obtained. Single-cell RNA sequencing (scRNA-seq) data of LIHC patients were collected from the GSE156625 dataset: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE156625.
MCD gene list
The MCD-related genes were collected from the KEGG, GSEA gene sets, and previous articles by combining seven mitochondria-associated cell death forms, namely, disulfidptosis (1771 genes), cuproptosis (14 genes), necroptosis (101 genes), autophagy (367 genes), ferroptosis (88 genes), pyroptosis (52 genes), and apoptosis (580 genes) [5, 6, 8] (Table S1).
Identification of the MCD-DEGs in LIHC
Transcriptome data of 371 LIHC patients and 276 normal samples from the TCGA and GTEx cohorts were compared using the “limma” package in the R software with adjusted p-value < 0.05 and |log2FC|> 1. We performed Cox regression analysis on the TCGA-LIHC, ICGC-LIRI, and GSE14520 cohorts to identify genes associated with prognosis, using a p-value threshold of < 0.05. The intersecting genes of the above analyses were identified as differentially expressed genes (DEGs). After intersecting the screened DEGs with genes associated with each type of mitochondrial-associated cell death, the MCD-DEGs were obtained.
Identification of the MCDS
To further identify the MCD-DEGs that are associated with the prognosis of LIHC patients, 10 machine learning algorithms were integrated, including survival support vector machine (survival-SVM), generalized boosted regression modeling (GBM), supervised principal components (SuperPC), partial least squares regression for Cox (plsRcox), CoxBoost, stepwise Cox, Ridge, the least absolute shrinkage and selection operator (LASSO), elastic network (Enet), and random survival forest (RSF) (a total of 67 combinations) [20]. To develop the consensus MCD-DEGs signature with high stability performance, the 67 machine learning algorithm combinations were fitted in the leave-one-out cross-validation (LOOCV) framework. According to Harrell’s C-index of each algorithm combination, the final suitable model was selected.
Variation analysis of the MCDS
The cBioPortal (http://www.cbioportal.org/) database [21] was used to explore the mutation landscape of MCDS in LIHC based on 348 LIHC samples (with full expression, mutation, and copy number variation data) from the TCGA database. The coexpression patterns of the MCDS in LIHC and their varied characteristics are shown as a Circos plot.
Consensus clustering and principal component analysis
Consensus clustering (CC) analysis was carried out using the R package “ConsensusClusterPlus” [22]. The algorithmic model was configured to implement agglomerative “pam” clustering, utilizing a Spearman correlation distance metric and resampling 80% of the sample population for a total of 1000 iterations. Using the R package “stats,” we conducted principal component analysis (PCA), whereby the expression data were normalized by calculating the z score, and a dimensionality reduction analysis was performed using the prcomp function to obtain an optimized low-dimensional matrix for downstream analyses.
Functional enrichment analysis
Encyclopedia of Genes and Genomes (KEGG) and Gene Oncology (GO) analyses of the DEGs were performed using the “clusterprofiler” and “ggplot2” R packages. The different HALLMARK functions between the MCDI-high and MCDI-low groups in the three cohorts were explored using the “GSVA” package of R with the “h.all. v2022.1.Hs.symbols.gm” (https://www.gsea-msigdb.org/gsea/msigdb/collections.jsp) database [23].
Construction of the nomogram model based on MCDS
MCDS and relevant clinical features were set together to construct the prognostic nomogram model to predict the overall survival (OS) of LIHC patients in three cohorts [24]. Calibration analysis, decision curve analysis (DCA), and receiver operating curve (ROC) analysis were performed using the “rms,” “survival,” and “timeROC” R packages [25, 26].
Single-cell sequencing analysis of the MCDS
Single-cell RNA-seq data from the GEO database (GSE156625) for LIHC was downloaded and analyzed using the “Seurat” package (version 4.4.1). After a series of standard quality procedures using the “NormalizeData,” “FindVariableFeatures,” “ScaleData,” “RunPCA,” “FindNeighbors,” “FindClusters,” “RunUMAP,” and “FindAllMarkers” functions, a total of 78,274 cells were included for further analysis. The “SingleR” function and the known markers from the literature were used to assign cell types [27]. The biological function of the marker genes in each cell type was identified using the “ClusterGVis” and “org.Hs.eg.db” R packages and the “enrichCluster” function [28]. Using the “inferCNV” package in R [29], epithelial cells were identified as either malignant epithelial cells or normal liver cells. The CNVs of malignant cells in the epithelial cells were calculated with fibroblasts as a reference. Pseudotime analysis was conducted using the R packages “CytoTRACE” and “Monocle2” [30, 31]. Moreover, the “AddModuleScore” function was used for the signature-specific score calculation based on the MCDS.
Tumor microenvironment analysis
The expression data of MCDS and the levels of various infiltrating immune cells in the TCGA-LIHC and ICGC-LIRI LIHC cohorts were downloaded and analyzed using the TIMER, CIBERSORT, EPIC, and MCPcounter algorithms via the “IOBR” package [32].
Immunotherapy and targeted therapy response prediction
Accumulating evidence suggests that the immune checkpoint mechanism plays a crucial role in suppressing the T-cell-mediated antitumor immune response within the tumor microenvironment [33]. To evaluate the association between the MCDS/MCDI and the level of immune checkpoint molecules in LIHC, we selected eight immune checkpoint-related transcripts, namely, CD274, CTLA4, HAVCR2, LAG3, PDCD1, PDCD1LG2, TIGIT, and SIGLEC15, and extracted their respective expression values for the corresponding cohorts. The Tumor Immune Dysfunction and Exclusion (TIDE) algorithm [34] was then used to predict the immunotherapy response between the MCDI-high and low groups, while the Genomics of Drug Sensitivity in Cancer (GDSC) database [35], accessed via https://www.cancerrxgene.org/, was utilized to predict the efficacy of sorafenib in LIHC patients with varying MCDI values.
Analysis of spatial transcriptomics
Spatial transcriptomics (ST) data pertaining to LIHC were retrieved from the Mendeley Data (https://data.mendeley.com/datasets/skrx2fz79n/1) [36] and subsequently reanalyzed utilizing the “Seurat” package (version 4.4.1). The process of quality control was meticulously executed in accordance with methodologies delineated in previously published literature [36]. Additionally, the signature score of the MCDI was incorporated into the metadata of the ST dataset employing the “AddModuleScore” function, utilizing default parameters within “Seurat.” To visualize the spatial expression patterns, the “SpatialFeaturePlot” function of the “Seurat” package (version 4.4.1) was employed. Spots exhibiting an MCDI score exceeding 0 were classified as positive.
Validation of the immunotherapy response of MCDI
Seven ST samples were included in the current study, comprised of five LIHC ICB non-responders and two responders. After calculating the MCDI score within each sample, the ratio of MCDI-positive spots could be obtained. To further validate the value of the MCDI index in predicting the response to ICB therapy, we also incorporated 298 bladder cancer patients with complete treatment response information from the IMvigor210 dataset who received anti-PD-1 treatment [37]. The MCDI was recalculated using Cox regression analysis, and the sensitivity of patients with different MCDI scores to ICB therapy was examined.
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
We used the RT-qPCR analysis to detect and compare the transcript levels of nine MCDS between normal liver samples (normal controls: NCs) and three liver cancer cell lines, including HepG2, HCCLM3, and Huh7. The total RNA was isolated with an RNA extraction kit (Fastagen), and cDNA was synthesized by using a reverse transcriptase M-MLV kit (Takara, Japan) according to the manufacturer’s instructions. RT-qPCR was performed on the ABI PRISM 7500 sequence detector (Applied Biosystems) with an RT-qPCR kit (Promega). The data were calculated by a standard curve method and normalized to the expression of the gene-encoding β-actin (human) [38].
Western blotting
The protein level of PAK1IP1 was measured by western blot analysis using previously described procedures [39]. Briefly, the samples were denatured by boiling in 1 × SDS loading buffer. Following electrophoresis on 8–12% SDS-PAGE gels, the samples were transferred onto nitrocellulose membranes. Blocking was performed using 5% non-fat dry milk in PBS with 0.1% Tween 20 for 60 min at room temperature. Subsequently, specific primary antibodies (PAK1IP1: Affinity, Df2563) were incubated with the blots overnight at 4 ℃. Washing was performed in PBS with 0.1% Tween 20, followed by incubation with HRP-conjugated secondary antibodies against rabbit. Finally, the detection was carried out using an ECL Western Blotting kit (Pierce) after washing the blots with PBS with 0.1% Tween 20. The signal intensity was determined using the Tanon 5200 s Chemiluminescent Imaging System (Tanon).
Immunohistochemistry (IHC)
To assess the prognostic significance of PAK1IP1 in LIHC patients, a 68-patient clinical cohort (cohort-1) diagnosed with LIHC was recruited, while a separate cohort (cohort-2) comprising 90 LIHC patients receiving sorafenib treatment was used to investigate the treatment effect of Sorafenib on LIHC among patients with varying levels of PAK1IP1 expression. The clinical characteristics of the enrolled LIHC patients are presented in Table S2. All participants were recruited from the Eastern Hepatobiliary Surgery Hospital (EHBH, Shanghai, China), and the ethical committee of EHBH approved the sample collection protocol.
The standard immunohistochemistry method [39] was employed to evaluate gene protein levels across the LIHC patients in both cohorts. In brief, formalin-fixed liver sections that were 4 µm thick were subjected to immunostaining for PAK1IP1. The positive area was quantified using Image-Scope software, as outlined in the manufacturer’s instructions. The primary antibodies (as used in western blotting) and the secondary antibodies (anti-rabbit) were employed according to the manufacturer’s guidelines. Hematoxylin was utilized as a counterstain for liver slides.
To minimize the bias caused by perioperative death, patients who died within 30 days were excluded from both cohorts. The optimal cutoff values of PAK1IP1 expression were determined to classify the patients into high- and low-expression groups, and subsequently, the association between PAK1IP1 expression and 5-year OS in LIHC patients was evaluated by comparing these cohorts.
Statistical analysis
Statistical analyses were performed using R software (v.4.2.1). Differences between the two groups were analyzed using either Student’s t-test (2-tailed) or Wilcoxon’s rank-sum test (Mann–Whitney U-test), while the Kruskal–Wallis test was used for comparisons among multiple groups. Survival curves were generated using Kaplan‒Meier analysis and compared using the log-rank test. Spearman’s rank test and correlation coefficient were utilized to assess and quantify the correlation between continuous variables. The threshold for statistical significance was set at p < 0.05.
Results
Identification of MCDS and model construction
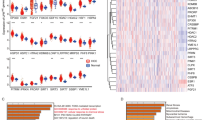
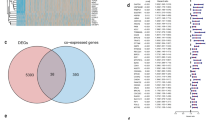
In the study, we utilized a total of 371 patients (368 with clinical information) from the TCGA database as the training cohort, while the validation cohorts comprised 243 patients from the ICGC-LIRI and 221 patients from the GSE14520. The clinical parameters of these patients from three cohorts are presented in Table S3, and the workflow of this study is shown in the Graphic abstract. To ensure the representativeness of the identified DEGs, we took the intersection of DEGs from the above three cohorts, and a total of 246 common DEGs were identified (Fig. 1A, Table S4-S7). According to the GO and KEGG enrichment analysis, the DEGs were closely associated with the response to cell cycle and DNA replication-related activity processes (Fig. 1B, Table S8). After intersecting the DEGs identified by screening of genes associated with each type of mitochondrial cell death, 22 genes related to disulfidptosis, one gene related to pyroptosis, five genes related to ferroptosis, seven genes related to apoptosis, and four genes related to autophagy were identified (Fig. 1C, Figure S1, Table S9). After removing the repeated genes, 35 MCD-related DEGs were included for further analysis (Table S9).
Identification of MCD-DEGs and the construction of consensus MCDI model. A Three LIHC cohorts were utilized to identify the DEGs, and the intersection of these genes was obtained (246). B GO and KEGG enrichment of the 246 DEGs. C The MCD gene list contains 35 genes, of which 22 genes from Disulfidptosis, one gene from Pyroptosis, five genes from Ferroptosis, seven genes from Apoptosis, and four genes from autophagy were identified. D A total of 67 kinds of prediction models via LOOCV framework and further calculated the C-index of each model in TCGA-LIHC and ICGC-LIRI datasets. E RSF algorithm presents the relative importance of the MCD-DEGs in the prognosis of LIHC. F Cross-validation of the constructed signature via LASSO regression analysis. G Univariate Cox regression analysis for the nine genes in the TCGA cohort. H The location, expression, and correlation of MCD-DEGs in the TCGA cohort
As shown in Fig. 1D, StepCox, RSF + StepCox, and LASSO + StepCox presented the highest c-index in both datasets (Table S10). To avoid the variable selection bias, we intersected the RSF and LASSO algorithms (using repeated tenfold cross-validation) to identify the most suitable MCDS signature, while the multivariate Cox regression analysis was implemented for model construction. The model exported the MCDI of each patient was calculated by adding the products of gene expression and corresponding coefficients, and the LIHC patients were divided into MCDI-high and MCDI-low subgroups based on the median value of the MCDI. According to LASSO regression analysis, nine genes were identified, including TRIM21, SPC25, CCT6A, GINS1, PAK1IP1, CBS, G6PD, SFN, and EPO (Fig. 1E, F). Based on the Cox regression analysis, a nine-gene signature was constructed (Fig. 1G). The chromosomal location and coexpression pattern of each MCD-DEG are shown in Fig. 1H. We found that almost all the MCDS were positively correlated with each other in LIHC, except for CBS (Fig. 1H). The expression levels of the MCDS were compared between LIHC samples and normal samples in the TCGA-LIHC cohort. Our results consistently showed that most MCDS (except for CBS and EPO) were overexpressed in LIHC tissues compared to normal samples (Figure S2A), and their upregulation in LIHC was significantly associated with worse clinical outcomes (Figure S2B). To further substantiate the prognostic value of the nine MCDS in the context of 3P medicine for LIHC, we initially utilized the Kaplan–Meier Plotter (https://kmplot.com/analysis/) [40] to assess the relationship between these MCDS and patient prognosis in the TCGA-LIHC cohort (Figure S3). Consistent results were observed in the ICGC-LIRI and GSE14520 validation cohorts (Figure S4). Additionally, we incorporated a newly published multi-omics cohort study of 228 HCC patients [16], analyzing the association between MCDS and clinical features of LIHC, including Barcelona Clinic Liver Cancer (BCLC) stage and recurrence-free survival (RFS) (Figure S5-7). The findings indicated that most MCDS, particularly CCT6A, G6PD, and PAK1IP1, displayed consistent behavior across all cohorts, further validating their potential as biomarkers.
In addition, we used the cBioPortal database to evaluate the alteration landscape of MCDS in LIHC. As shown in Figure S8A and B, the alteration frequency of the MCDS in LIHC is low, with PAK1IP1 possessing the highest mutation frequency (7%).
Construction of MCDI and its association with clinical characteristics of LIHC
The multivariate Cox regression analysis was performed to calculate the coefficient of each MCDS in the TCGA-LIHC cohort. The following formula was employed: MCDI = TRIM21exp × 0.15707 − − SPC25exp × 0.0087927 + CCT6Aexp × 0.061738 + GINS1exp × 0.071776 + PAK1IP1exp × 0.35471 − − CBSexp × 0.24494 + G6PDexp × 0.098825 + EPOexp × 0.15885 + SFNexp × 0.025531. A high MCDI value correlated with worse clinical outcomes, such as high tumor grade and stage and worse survival, in the three LIHC cohorts (Fig. 2A–C, G). Based on the expression data of MCDS, the TCGA-LIHC cohort was classified into two clusters (C1 and C2) according to the CC analysis (Fig. 2D). Kaplan–Meier survival analysis was utilized to explore the association between the OS time and the two clusters, and we found that patients in C1 had better survival (Fig. 2E, p < 0.001). Moreover, the alluvial diagrams showed that the majority of the C1 cluster was associated with low tumor grade, low tumor stage, and low MCDI (Fig. 2F, G).
Association between MCDI and the clinical features of LIHC. A–C Raincloud plots of the relationship between MCDI and different LIHC molecular subtypes in three cohorts. D Unsupervised clustering of MCD-DEGs in TCGA-LIHC cohort. Patients were grouped into two molecular clusters when k = 2. E Kaplan–Meier analysis of the prognosis of LIHC patients in different molecular clusters. F Alluvial diagram shows the interrelationship between MCDI groups, tumor grade, tumor stage, molecular clusters, and survival status in LIHC patients. G Heatmap of nine MCD-DEGs and clinical features
A high MCDI correlates with worse LIHC patient survival and with the enrichment of cell cycle-related pathways
Based on the calculated MCDI value, the LIHC patients in the TCGA-LIHC cohort, ICGC-LIRI cohort, and GSE14520 cohort were divided into the MCDI-high and MCDI-low groups, and a high MCDI correlated with worse clinical outcomes (Fig. 3A, B). The MCDI-high and MCDI-low groups were effectively separated in all three cohorts using PCA (Fig. 3C). In a further study, we applied GSVA to explore the differences in HALLMARK functions between the subgroups classified by the MCDI. The statistical HALLMARK pathways identified in the ICGC-LIRI cohort are shown in Fig. 3D. Notably, we found that the most common HALLMARK pathways among the three cohorts were G2M_CHECKPOINT, MITOTIC_SPINDLE, and BILE_ACID_METABOLISM (Fig. 3E). The detailed HALLMARK functions are shown in Table S911-13.
Performance of the gene signature prediction model in the internal training and external validation cohorts. A Distribution of MCDI according to the survival status and time in TCGA-LIHC, ICGC-LIRI, and GSE14520 cohorts. B Kaplan–Meier analysis of the overall survival in the MCDI-high and MCDI-low groups in the three independent cohorts. C Principal component analysis (PCA) plot based on the MCDI in the three independent cohorts. D GSEA analysis of the HALLMARK functions between two MCDI groups in the ICGC-LIRI cohort. E The interaction of enrichment HALLMARK functions among the three independent cohorts
Validation of the prognostic ability of MCDI in LIHC cohorts
We conducted univariate and multivariate Cox regression analyses to investigate whether MCDI could independently predict the prognosis of LIHC. Our results showed that MCDI was a significant risk factor in the univariate Cox regression model (HR = 2.716; 95% CI, 2.077–3.553; p < 0.001, Fig. 4A). After adjusting for potential confounding variables, the multivariate model demonstrated that MCDI was still a significant independent predictor of LIHC prognosis (HR = 2.475; 95% CI, 1.854–3.304,; p < 0.001, Fig. 4A). In a further study, we developed a nomogram model in the TCGA cohort to predict the 1-, 3-, and 5-year OS using multivariable Cox and stepwise regression analyses. The model incorporated stage, T stage, and MCDI as risk factors (Fig. 4B). Calibration curves demonstrated the excellent accuracy of this model (Fig. 4C). Furthermore, DCA revealed that the nomogram model and the MCDI outperformed the other predictors employed in this study (Fig. 4D). Based on the nomogram score, significant differences in survival were observed between the high- and low-risk groups (Fig. 4E). The prognostic function of the MCDI was also validated in the other two LIHC cohorts, and the nomogram models are displayed in Figure S9A and B. We also evaluated the area under curve (AUC) values in the three LIHC cohorts and found that the nomogram exhibited superior accuracy in predicting the OS of LIHC patients (Fig. 4D). These findings highlight the potential clinical utility of the nomogram model and MCDI in identifying LIHC patients at high risk of mortality, facilitating personalized treatment strategies, and improving patient outcomes. Furthermore, we compared the performance of the constructed nomogram index and the MCDI against a trivial model using only age, sex, tumor stage, and tumor grade as predictors. The DCA curves over 1, 3, and 5 years demonstrate that our model consistently performs better than these individual predictors in all LIHC cohorts (Figure S10A-C).
Construction and evaluation of the prognostic nomogram model. A Univariate and multivariate Cox regression analysis for the MCDI and relevant clinical features in the TCGA-LIHC cohort. B A prognostic nomogram was developed to predict the survival of LIHC patients. C Calibration plots showing the probability of 1-, 3-, and 5-year overall survival in the TCGA-LIHC cohort. D Decision curve analysis (DCA) of a nomogram predicting the 3-year survival in TCGA-LIHC cohort. E Kaplan–Meier analyses for the two LIHC groups based on the nomogram score. F Time-dependent receiver operator characteristic (ROC) analysis of nomogram in TCGA-LIHC, ICGC-LIRI, and GSE14520 cohorts
Single-cell analysis suggested MCDS score correlates with the development of LIHC
To further investigate the expression and distribution of MCDS at a single-cell resolution, we analyzed scRNA-seq data for LIHC (GSE156625). After applying several standard quality control procedures to the data, a total of 78,234 cells were included in our analysis (Figure S11A-E). The cells were classified into 28 clusters and annotated into eight cell types (Fig. 5A). The marker genes of each cell type are shown in Fig. 5B, C, and the top five biological processes enriched for each cell type’s marker genes are also presented. We noticed that MCDS were enriched in epithelial, fibroblast, and NK cells (Figure S11F and G). We used inferCNV, an algorithm that detects cells with significant copy number variations by sorting expressed genes based on their chromosomal locations and applying a sliding window of 100 genes within each chromosome to the relative expression values, to identify malignant epithelial cells (Fig. 5D). Accordingly, the epithelial cells were separated into normal epithelial cells (cluster 8 and 21) and malignant cells (cluster 7: low malignancy; cluster 15, 20, 22, and 23: middle malignancy; cluster 14, 16, and 24: high malignancy) (Fig. 5E). Subsequently, we predicted the differentiation states of epithelial cells using CytoTRACE and Monocle 2 pseudotime trajectory analyses, and we observed a clear lineage trajectory stemming from normal cells to malignant cells and from low/middle malignant cells to high malignant cells (Fig. 5F, G). Based on the “AddModuleScore” function, the MCDS score in each cell was calculated, and the expression levels of the MCDS and the MCDS score in various cell types are shown in Figure S4F and G. Of notice, we observed a significant correlation between MCDS score and the malignant degree of tumor cells (Fig. 5H).
Exploration of MCDI with scRNA. A Louvain clustering of 78,234 cells identifies 28 mega-clusters in the liver. B The heatmap of the average expression of top ten DEGs among eight cell types of liver tumor cells. The top five biological processes enriched for each cell type’s marker genes were presented. C Expression of genes correlated with stromal, epithelial, and immune cell types. D The hierarchical heatmap showing large-scale CNVs in liver cancer epithelial cells. E UMAP (Uniform Manifold Approximation and Projection) plot showing the three sub-clusters of malignant cells. F Predicted ordering by CytoTRACE, which orders the epithelial cells based on their developmental potential from normal hepatocyte to malignant cells. Boxplots show the median and interquartile range and whiskers extend from min to max. G Trajectory analysis by Monocle 2 combining three malignant sub-clusters. H Comparison of the MCD-DEGs score between the sub-clusters with low/middle and high degrees of malignancy
MCDI can predict immunotherapy response in LIHC
The association between the levels of MCDS/MCDI and the enrichment scores of various immune cells were measured using various algorithms. Interestingly, we found that most MCDS expression as well as MCDI was positively associated with the infiltration of B cells, CD4+ T cells, CD8+ T cells, neutrophil, macrophage, and myeloid dendritic cells in the TCGA-LIHC cohort (Fig. 6A, Table S14). The interaction between MCDS and the immune cell infiltration in LIHC was facilitated in the ICGC-LIRI cohort (Table S15). As shown in Fig. 6B, we noticed that the levels of most MCDS and MCDI were positively associated with the expression levels of immune checkpoint molecules both in TCGA-LIHC and ICGC-LIRI cohorts. Moreover, a high MCDI was also associated with a high TIDE score both in all three LIHC cohorts, suggesting that patients with a high MCDI may derive less therapeutic benefit from immunotherapy than those with a low MCDI (Fig. 6C).
Association of the MCD-DEGs and the immune features of LIHC A Heatmap presents the association between the MCD-DEGs/MCDI and the infiltration of various immune cells. B Association between the levels of MCD-DEGs/MCDI and the expression of immune checkpoint monocles. The differences of the seven immune checkpoint monocles expression in MCDI-high and MCDI-low groups. C Comparison of the TIDE score between MCDI-high and low groups in TCGA-LIHC, ICGC-LIRI, and GSE14520 cohorts, respectively
To validate the utility of MCDI in predicting the response of LIHC patients to ICB therapy, we conducted ST analysis on tumor sections from seven LIHC patients undergoing anti-PD-1 treatment, categorized into non-responders (n = 5) and responders (n = 2), as depicted in Fig. 7A. Notably, we observed that the counts of Unique Molecular Identifiers (UMIs) in ICB non-responders (#P11_T and #P3_T) were higher than those in responders (#P7_T and #P9_T), suggesting a more complex TME in the non-responders (Fig. 7B). Further, considering that each spot contained multiple cells, we proposed a signature-based strategy to evaluate the enrichment degree of different cell types in each spot (Fig. 7C) [41]. It can be seen that the malignant cells exhibit a high degree of patient heterogeneity, while stromal and immune cells demonstrate co-localization (Fig. 7C). After calculating the MCDI score in each ST sample, we found both the MCDI score and the MCDI-positive cell ratio in ICB non-responders were higher than that in ICB responders (Fig. 7D–H, Figure S12).
Validation of MCDI in predicting the ICB treatment response. A, B UMAP (Uniform Manifold Approximation and Projection) plots showing spots from all sections, color-coded according to their sample source, with annotations for the number of expressed unique molecular identifiers (nUMI) and genes (nGene), respectively. C A UMAP plot illustrating cell type enrichment scores across all sections, where each dot represents a spot from ST sections, and the intensity of the color denotes the enrichment score and spot density of the corresponding samples. D Dot plots highlighting the average expression levels of MCD-DEGs and the MCDI score across each ST sample. E–H Left: spatial distribution of cell types for each patient across different sections. Middle: dot plots displaying the average expression of the top 3 differentially expressed markers in specified cell types. Right: the distribution and quantification of MCDI score within each sample. In the dot plots, the size of each dot indicates the proportion of cells expressing the marker in each cell group, while the color signifies the intensity of expression
To further elucidate the predictive value of the MCDI in determining patient responses to ICB treatment, we incorporated an additional bladder cancer cohort (the IMvigor210 cohort). As illustrated in Fig. 7I, patients exhibiting higher MCDI scores were associated with reduced OS and a diminished response to immunotherapy.
In vitro experiments validation of the clinical significance and targeted therapy response of MCDS in LIHC
We utilized RT-qPCR analysis to validate the differential mRNA expression of multiple MCDS between normal liver cells and liver cancer cell lines. The PCR primer sequences for each MCDS are shown in Table S16. As depicted in Fig. 8A to F, the mRNA expression levels of CCT6A, TRIM21, SPC25, GINS1, G6PD, and PAK1IP1 were found to be significantly higher in liver cancer cells than in normal liver cells. Conversely, the expression levels of EPO and CBS were decreased in liver cancer cells, which was consistent with the bioinformatic analysis results (Fig. 8G and H). However, it is noteworthy that the expression level of SFN was found to be decreased in liver cancer cells, which was not concordant with the bioinformatic findings (Fig. 8I).
In vitro experiments and in-house clinical cohorts’ validation of the significance of MCD-DEGs in LIHC. A–I RT-qPCR analysis of MCD-DEGs mRNA expression in normal liver cells (NC), HepG2, HCCLM3, and Huh7 cells. Data in (A–I) are presented as mean ± SD (n = 3 per group) and p-values by two-tailed unpaired Student’s t-test are indicated. J Association between MCD-DEGs/MCDI and the sorafenib IC50 values in GDSC database. K Western blot analysis of PAK1IP1 protein expression in NC, HepG2, HCCLM3, and Huh7 cells. L Kaplan–Meier curves of 5-year Overall Survival (OS) of LIHC between PAK1IP1-high and PAK1IP1-low groups from clinical cohort 1. M Kaplan–Meier curves of 5-year OS of LIHC between PAK1IP1-high and PAK1IP1-low groups from the sorafenib cohort (clinical cohort 2)
Given that sorafenib is commonly prescribed as an initial treatment for advanced-stage LIHC patients, we leveraged the publicly accessible GDSC database to investigate the correlation between sorafenib IC50 (Sorafenib_30 in GDSC version 1 and Sorafenib_1085 in GDSC version 2) and MCDS expression in individuals with LIHC. Strikingly, the levels of seven of the MCDS and the MCDI were negatively associated with the sorafenib IC50 in LIHC, implying that patients with a high MCDI might respond favorably to sorafenib treatment (Fig. 8J).
Since PAK1IP1 presents the highest correlation with sorafenib IC50 level in LIHC, we thus measured the protein levels of PAK1IP1 in three different liver cancer cell lines and in our EHBH cohorts and assessed its potential utility in stratifying individuals who may respond to sorafenib treatment (Fig. 8K). Our results, as illustrated in Fig. 8L and M, demonstrated that higher expression of PAK1IP1 was associated with poorer survival outcomes in cohort 1, while in the sorafenib-treated cohort (cohort 2), elevated expression levels of PAK1IP1 was associated with improved survival outcomes.
Discussion
The novelty of this study within the context of 3P medicine
To our knowledge, this is the first study that comprehensively explored seven diverse patterns of mitochondrial-associated PCD, identified nine MCD-DEGs in LIHC, and constructed a MCDI signature in LIHC in the framework of the 3P medicine approach. By integrating predictive analytics, preventive strategies, and personalized approaches, we have validated the MCDI signature’s performance in two external cohorts (ICGC-LIRI and GSE14520), underscoring the predictive power of our model. This aligns with the preventive aspect of 3P medicine by potentially identifying early biomarkers that could aid in early detection and prevention of disease progression. Moreover, the clinical applicability of our nomogram model, which combines the MCDI with relevant clinical parameters, exemplifies the personalized approach of 3P medicine, offering a tailored prognostic assessment and therapeutic guidance for LIHC patients. Importantly, our findings demonstrated a significant correlation between the MCDI and the tumor microenvironment as well as drug sensitivity in LIHC. These results emphasize the potential value of the MCDI in stratifying patients for better personalized therapy.
The prognostic value of MCDS in LIHC
PCD involves a series of biological or pathological processes that participate in normal physiological processes and disease development by regulating tissue homeostasis, stress response, and cell proliferation, which is significantly associated with immune responses against many human cancers [7, 11, 42]. Previous studies have reported the role of multiple PCD patterns in several human cancers [42,43,44]; however, the genetic landscape, the multimolecular mechanism, and the immune function of MCD-related genes in LIHC remain incompletely understood. Herein, we constructed a MCDI signature based on nine MCDS (TRIM21, SPC25, CCT6A, GINS1, PAK1IP1, CBS, G6PD, SFN, and EPO) to predict the prognosis and personalized therapy response of individual LIHC patients, exemplifying the predictive capabilities of 3P medicine. This model not only enhances our understanding of the genetic landscape of MCD-related genes in LIHC but also provides a foundation for the development of targeted preventive and therapeutic strategies.
Several molecules, including TRIM21, SPC25, GINS1, CBS, G6PD, EPO, and SFN, have been extensively studied in various cancers; nonetheless, how these molecules play their roles and the related regulatory mechanisms in LIHC remains incompletely known. To verify our bioinformatic findings, RT-qPCR was applied to compare the expression of MCDS between normal liver cells and three liver cancer cell lines. Consistently, the results were in line with expectations except for SFN at the mRNA level, which needs further verification at the protein level. SPC25, a component of the NDC80 complex, has been reported to be associated with a poor prognosis in LIHC [45]. GINS1, a protein complex involved in DNA replication, plays a role in cancer cell growth, migration, invasion, and chemoresistance in several cancers, such as glioma, lung, and colon cancers [46, 47]. CBS is an enzyme involved in methionine metabolism, the downregulation of which was associated with poor survival and high tumor-infiltrating Tregs in LIHC [48]. G6PD belongs to a pivotal enzyme in glucose metabolism, playing a crucial role in cancer cell reprogramming and malignant transformation [49]. EPO, an apoptosis-related gene, has been studied in chemotherapy-induced anemia in cancer patients and related to adverse outcomes [50]. The SFN protein is widely involved in various cellular biological processes, including cell proliferation, apoptosis, signal transduction, and cell cycle regulation. It has been reported that the upregulation of SFN associates with the progression and poor prognosis of LIHC [51], while the other report probes that SFN could regulate the expression of CDK1 and CCNB1 to inhibit the growth of LIHC cells [52]. PAK1IP1 is one of the adaptor proteins of PAK1 which is closely involved in cancer cell survival, proliferation, migration, and invasion of LIHC [53], while the role of PAK1IP1 in the carcinogenesis and prognosis of LIHC remains unclear and requires further study.
CCT6A is a component of the chaperonin complex involved in protein folding and quality control, which has been implicated in various cellular processes, including cytoskeleton organization, cell signaling, and gene expression. TRIM21 has been extensively studied concerning immune response, intracellular signaling, and protein ubiquitination, and its overexpression was found to be a prognostic risk factor [54]. Although it has been reported that higher CCT6A and TRIM21 expression contribute to LIHC cell proliferation and a poorer prognosis [55], the underlying mechanisms remain unclear. Besides, the value of CCT6A and TRIM21 in stratifying LIHC patients to improve personalized therapy has not been evaluated. In the current study, we found that most MCDS were overexpressed in LIHC patients and that such upregulation was associated with worse survival. Based on the formula determined via multivariate Cox regression analysis, the MCDI for each LIHC patient was calculated, and the MCDI-based nomogram presented with great clinical performance, suggesting that the MCDI could be a potential prognostic biomarker in LIHC patients.
The clinical translational significance of MCDS in LIHC within the 3P medicine framework
An increasing number of studies have suggested that tumor cells may influence T-cell infiltration into the TME and the function of immune regulatory cells through their interactions with the TME, which is relevant to cancer immunotherapy [2, 56]. The significant correlations between MCDS/MCDI levels and immune cell infiltration into the TME highlight the complex interplay between tumor genetics and the immune microenvironment. Our study revealed significant positive correlations between MCDS/MCDI levels and the presence of infiltrating immune cells, underscoring a profound genetic-immunological interaction. Notably, patients with elevated MCDI levels demonstrated increased expression of immune checkpoint molecules, suggesting a heightened potential for immune evasion or immunosuppression. In line with this, the TIDE score, indicative of potential resistance to immunotherapy, was also elevated in patients with high MCDI levels. These observations were consistently validated by results from both the LIHC spatial transcriptomics cohort and the IMvigor210 bladder cancer cohort. These findings are particularly relevant to the preventive and personalized arms of 3P medicine, as they suggest potential avenues for modulating immune responses in LIHC. As the first-line treatment of advanced LIHC, the response to sorafenib could primarily affect the clinical outcome of LIHC patients [3]. Herein, we found that the levels of the MCDS and MCDI were negatively associated with the sorafenib IC50 in the TCGA-LIHC cohort, suggesting that MCDI-high patients might benefit from sorafenib treatment compared with MCDI-low patients. Given the observed significant correlation between PAK1IP1 levels and the sorafenib IC50 in LIHC, the clinical significance of PAK1IP1 in LIHC was further verified in our enrolled clinical cohorts, suggesting its great potential in predicting the prognosis and sorafenib sensitivity in LIHC. This aligns with the preventive and personalized goals of 3P medicine, where the identification of biomarkers and the stratification of patient populations can lead to more targeted and effective therapeutic strategies. However, further studies are warranted to explore the underlying molecular mechanisms through which PAK1IP1 contributes to the prognosis of LIHC.
Limitations
Several limitations of the study need to be recognized despite the great potential value of our model in the three LIHC cohorts. Firstly, the retrospective nature of our study may introduce inherent biases, which could affect the generalizability of our findings. Secondly, the utility of our model in clinical decision-making has yet to be validated in phase 3 randomized controlled trials, a necessary step to confirm its efficacy and reliability in clinical settings. Furthermore, we acknowledge the need for further validation of our model in larger, multicentric clinical cohorts. This should include comparisons of different cross-validation methods such as LOOCV, holdout, tenfold, and stratified cross-validation, as well as different versions of the C-index to thoroughly assess the impact on model prediction capabilities. Additionally, it is imperative to conduct further research and seek validation through well-designed prospective clinical trials and sophisticated bioinformatics analyses to accurately determine the model’s predictive efficacy, particularly with regard to its ability to forecast differences in progression-free survival in the context of immunotherapy treatment. Moreover, our validation of the differential expression of the MCDS was limited to in vitro experiments and the protein level analysis of PAK1IP1. This approach does not fully explore the prognostic or predictive value of the identified genes, and future studies should extend these findings through more comprehensive molecular and cellular experiments. Such studies could include knockdown, overexpression, and rescue experiments aligned with sorafenib response assessments in both cell lines and animal models, as well as potential mechanistic studies identifying relevant upstream and downstream molecules.
Conclusion
In alignment with the predictive aspect of 3P medicine, this study leverages multi-omics to develop a novel predictive model, the MCDI, which is based on mitochondria-associated cell death genes. By focusing on the prognostic potential of these genes and their response to immunotherapy and targeted therapy, this research aims to provide a more precise and individualized prognosis and treatment response prediction for LIHC patients. The identification of several biomarkers, particularly PAK1IP1, may serve as a cornerstone for clinical utility in predicting prognosis and guiding personalized treatment decision-making, thereby fulfilling the preventive and personalized goals of 3P medicine.
PPPM innovation highlights
Working hypothesis in the framework of PPPM
Predictive, preventive, and personalized medicine (PPPM) represents an advanced healthcare concept, focusing on predicting patient prognosis and guiding therapeutic strategies based on novel technologies, including public databases and multi-omics [57,58,59]. This approach facilitates the development of precise preventive interventions and the provision of customized healthcare solutions [60]. To achieve its goals, PPPM relies on identifying a range of molecular biomarkers through extensive multi-omics analyses. These biomarkers are crucial for early detection of individuals requiring intervention, predicting patient survival, and determining the effectiveness of targeted therapies and immunotherapies. This knowledge allows for the personalization of treatment approaches. Utilizing these biomarkers, PPPM aims to customize therapeutic interventions based on each individual’s unique molecular profile, optimizing clinical results, minimizing side effects, and ushering in a new era of precision in patient care [57,58,59,60]. Moreover, bioinformatics plays a pivotal role in the discovery of these disease markers and in transitioning from conventional medical practices to a more PPPM-oriented model. In this context, we postulate that the mitochondrial cell death (MCD) is intricately linked to the progression and prognosis of LIHC. Its pronounced association with immune responses and sensitivity to targeted therapy underscores its potential in guiding personalized treatment strategies. Through a synthesis of multi-omics data and advanced machine learning techniques, our research endeavors to develop a MCDI, which aims to refine the accuracy of prognosis estimations, enable the implementation of targeted preventive measures, and foster the development of personalized therapeutic strategies within the 3P medicine framework.
Expert recommendations and innovation towards 3P medicine
LIHC ranks as the third leading cause of cancer-related deaths worldwide, with both its incidence and mortality rates continuing to rise [1]. The evolution from conventional medical practices towards the 3P medicine framework is required. The advent and integration of multi-omics technologies have catalyzed a transformative approach to biomarker discovery across an array of biological dimensions, including bulk, single-cell, and spatial transcriptomics, as well as DNA, proteins, and metabolites. This technological revolution holds the promise of enhancing the ability in predicting and preventing oncological diseases through the identification of novel diagnostic and prognostic biomarkers.
In the context of LIHC, the application of multi-omics methodologies under the framework of 3P medicine offers a comprehensive exploration of the disease’s molecular architecture [61]. By integrating multi-omics profiles, clinicians can obtain critical information for treatment selection, which can facilitate the development of targeted interventions and personalized therapy strategies.
Predictive medical approach
In this study, we pioneer the exploration of the potential application of MCD in LIHC. To this end, we have developed an innovative approach that integrates multi-omics data with a large-scale machine learning framework. Subsequently, we identified a MCDI characterized by its high predictive accuracy and significant potential for clinical translation. By leveraging the MCDI, it becomes possible to predict the survival outcomes of LIHC patients across both training and validation cohorts, thus establishing a foundation for the use of MCDI in forecasting adverse clinical outcomes. We also introduce an MCDI-integrated nomogram as a practical tool for quantifying the expected survival rates of patients in clinical settings. Based on our findings, we suggest emphasizing the potential value of MCD in predicting the prognosis and treatment responses of LIHC patients, with the aim of improving clinical outcomes and optimizing the cost-effectiveness of treatments.
Targeted prevention
Despite significant advancements in the fields of cancer diagnostics and therapeutics, the majority of patients with LIHC are diagnosed at advanced stages of the disease. This circumstance effectively precludes the option for curative interventions, culminating in suboptimal clinical outcomes. Recognizing individuals at risk of developing LIHC and implementing early interventions is crucial for alleviating the burden of LIHC. Adopting advanced preventive measures for individuals at high risk represents a more cost-effective approach. Our study leverages novel machine learning techniques to stratify LIHC patients based on their risk levels. This enables clinicians to devise personalized intervention strategies for each risk category, including personalized screening schedules and detailed health guidance recommendations, to facilitate early detection and prevent disease progression. The biomarker signature developed in this study serves as a simple, economical, and broadly feasible tool for assessing the individual risk of patients, thereby preventing the onset and progression of LIHC.
Personalized medicine
LIHC manifests marked heterogeneity and harbors a complex TME, characterized by distinct molecular features, diverse natural progression, and varied biological behaviors [2, 4, 36, 56]. This complexity poses significant challenges for clinicians in the management of cancer patients. Notably, while conventional histopathological evaluations and comparable therapeutic approaches may lead to remission in certain patients, others may suffer premature mortality. Therefore, it is quite necessary to identify specific patient groups that are superior to targeted therapy or immunotherapy. Our research emphasizes the crucial importance of multi-omics approaches in advancing personalized medicine for LIHC under the 3P medicine framework, aiming to improve patient outcomes and minimize unnecessary treatment expenditures. Through the integration of genomics, single-cell, spatial, and bulk transcriptomics data of LIHC, we discovered that patients in the high-risk category exhibited an increased potential for immune evasion or immunosuppression and showed resistance to anti-PD-1 treatment. However, they might benefit from sorafenib treatment. Given the variability in treatment responses among patients with different molecular backgrounds, these multi-omics strategies offer a molecular basis for the tailored selection of treatment. Leveraging the MCDI, clinicians can stratify LIHC patients into two subgroups with distinct molecular features, thereby providing essential information for personalized treatment decisions and advancing the practice of 3P medicine in LIHC management.
How does the presented innovation go beyond the state of the art contributing to the paradigm shift from reactive medicine to PPPM?
The presented innovation significantly advances the field of LIHC treatment by introducing a novel signature, namely MCDI, which is a multi-omic, machine learning-driven tool for prognosis prediction and therapy response assessment. This work goes beyond the state of the art by actively contributing to the paradigm shift from reactive medicine to predictive, preventive, and personalized medicine in several key ways:
-
(1)
Predictive aspect: The MCDI model leverages a comprehensive multi-omic approach, integrating data from bulk, single-cell, and spatial transcriptomics, to predict patient outcomes. By identifying critical genes associated with MCD and constructing a prognostic index, this model provides a predictive tool that can forecast survival outcomes and therapy response in LIHC patients with high accuracy. This predictive capability is a significant step towards the PPPM approach, as it allows for early intervention and tailored treatment strategies based on individual patient profiles.
-
(2)
Preventive aspect: The study’s identification of MCD-related genes that are associated with LIHC progression offers potential targets for early detection and intervention. By understanding the molecular mechanisms that drive cancer growth and metastasis, this research can inform the development of preventive strategies that may reduce the incidence of LIHC or delay its progression. The MCDI could be used to stratify patients into risk categories, enabling personalized screening and preventive measures for those at the highest risk.
-
(3)
Personalized medicine: The integration of the MCDI with clinical parameters in a nomogram model exemplifies the personalization aspect of PPPM. By providing a quantitative basis for predicting individual patient responses to personalized therapies, including sorafenib and ICB treatment, the MCDI facilitates the selection of the most effective treatment for each patient. This personalized approach can facilitate clinicians for decisions making and alleviate the burden of LIHC.
-
(4)
Integration of advanced technologies: The use of machine learning algorithms to analyze and interpret complex multi-omic data represents a cutting-edge approach that is becoming increasingly important in biomarker discovery and drug development. This integration of advanced technologies into the PPPM framework is a significant innovation that enhances the precision and reliability of prognostic models.
In summary, our research demonstrates the potential of PPPM strategies in the field of LIHC by showcasing the MCDI as a transformative tool that aligns with the PPPM philosophy. This data-driven, individualized approach to LIHC management provides valuable insights into the progress in prediction, prevention, and personalized medicine. The MCDI has the potential on guiding personalized therapy strategies selection, enhance treatment efficacy, and ultimately contribute to better clinical outcomes in LIHC patients.
Data availability
All the data used in this study were collected in this article and supplemental materials.
Availability of data and materials
TCGA datasets enrolled in this study are openly available in the National Cancer Institute GDC Data Portal (https://portal.gdc.cancer.gov/). TCGA data are displayed under the Project IDs “TCGA-LIHC.” GEO datasets are publicly available in the National Center for Biotechnology Information Portal (https://www.ncbi.nlm.nih.gov/geo/), including GSE14520 and GSE156625. ICGC-LIRI dataset is openly available in ICGC (https://dcc.icgc.org/releases/current/Projects). The spatial transcriptomics of LIHC cohorts can be obtained from Mendeley Data (skrx2fz79n). IMvigor210 bladder cancer cohort can be accessed from http://research-pub.gene.com/IMvigor210CoreBiologies.
Code availability
Analyses were conducted using R (version 4.2.1) and GraphPad (GraphPad Prism 8.0). The codes used to support the findings of this study is available from the corresponding author on reasonable request.
Abbreviations
- AUC:
-
Area under the curve
- BCLC:
-
Barcelona Clinic Liver Cancer
- CC:
-
Consensus clustering
- CI:
-
Confidence interval
- CNV:
-
Copy number variation
- DCA:
-
Decision curve analysis
- EPO:
-
Erythropoietin
- G6PD:
-
Glucose-6-phosphate dehydrogenase
- GDSC:
-
Genomics of Drug Sensitivity in Cancer
- GEO:
-
Gene Expression Omnibus
- GINS1:
-
GINS complex subunit 1 (Sld5 homolog)
- GO:
-
Gene ontology
- GSVA:
-
Gene set variation analysis
- HR:
-
Hazard ratio
- IHC:
-
Immunohistochemistry
- ICGC:
-
International Cancer Genome Consortium
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LASSO:
-
Least absolute shrinkage and selection operator
- LOOCV:
-
Leave-one-out cross-validation
- LIHC:
-
Liver hepatocellular carcinoma
- MCD:
-
Mitochondria-associated cell death
- MCD-DEGs:
-
Mitochondria-associated cell death differentially expressed genes
- MCDI:
-
Mitochondrial cell death index
- MCDS:
-
Mitochondria-associated cell death signature genes
- NC:
-
Normal controls
- OS:
-
Overall survival
- PAK1IP1:
-
P21-activated kinase 1 interacting protein 1
- PCA:
-
Principal component analysis
- PPPM:
-
Predictive, preventive, and personalized medicine
- RFS:
-
Recurrence-free survival
- RT-qPCR:
-
Reverse transcription-quantitative polymerase chain reaction
- ROC:
-
Receiver operating characteristic
- RSF:
-
Random survival forest
- TIDE:
-
Tumor Immune Dysfunction and Exclusion
- TIMER:
-
Tumor Immune Estimation Resource
- UMAP:
-
Uniform Manifold Approximation and Projection
- SD:
-
Standard deviation
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. https://doi.org/10.3322/caac.21660.
Wang Z, Wang Y, Gao P, Ding J. Immune checkpoint inhibitor resistance in hepatocellular carcinoma. Cancer Lett. 2023;555:216038.
Reiss KA, Yu S, Mamtani R, Mehta R, D’Addeo K, Wileyto EP, Taddei TH, Kaplan DE. Starting dose of sorafenib for the treatment of hepatocellular carcinoma: a retrospective, multi-institutional study. J Clin Oncol. 2017;35(31):3575–81. https://doi.org/10.1200/JCO.2017.73.8245.
Hu D, Wang Y, Shen X, Mao T, Liang X, Wang T, Shen W, Zhuang Y, Ding J. Genetic landscape and clinical significance of cuproptosis-related genes in liver hepatocellular carcinoma. Genes Dis. 2024;11.
Liu X, Nie L, Zhang Y, Yan Y, Wang C, Colic M, Olszewski K, Horbath A, Chen X, Lei G, et al. Actin cytoskeleton vulnerability to disulfide stress mediates disulfidptosis. Nat Cell Biol. 2023;25(3):404–14. https://doi.org/10.1038/s41556-023-01091-2.
Tsvetkov P, Coy S, Petrova B, Dreishpoon M, Verma A, Abdusamad M, Rossen J, Joesch-Cohen L, Humeidi R, Spangler RD, et al. Copper induces cell death by targeting lipoylated tca cycle proteins. Science. 2022;375(6586):1254–61. https://doi.org/10.1126/science.abf0529.
Bock FJ, Tait S. Mitochondria as multifaceted regulators of cell death. Nat Rev Mol Cell Biol. 2020;21(2):85–100. https://doi.org/10.1038/s41580-019-0173-8.
Tang D, Kang R, Berghe TV, Vandenabeele P, Kroemer G. The molecular machinery of regulated cell death. Cell Res. 2019;29(5):347–64. https://doi.org/10.1038/s41422-019-0164-5.
Martinou JC, Youle RJ. Mitochondria in apoptosis: bcl-2 family members and mitochondrial dynamics. Dev Cell. 2011;21(1):92–101. https://doi.org/10.1016/j.devcel.2011.06.017.
Tuzlak S, Kaufmann T, Villunger A. Interrogating the relevance of mitochondrial apoptosis for vertebrate development and postnatal tissue homeostasis. Genes Dev. 2016;30(19):2133–51. https://doi.org/10.1101/gad.289298.116.
Gao W, Wang X, Zhou Y, Wang X, Yu Y. Autophagy, ferroptosis, pyroptosis, and necroptosis in tumor immunotherapy. Signal Transduct Target Ther. 2022;7(1):196. https://doi.org/10.1038/s41392-022-01046-3.
Napoletano F, Baron O, Vandenabeele P, Mollereau B, Fanto M. Intersections between regulated cell death and autophagy. Trends Cell Biol. 2019;29(4):323–38. https://doi.org/10.1016/j.tcb.2018.12.007.
Russell RC, Guan KL. The multifaceted role of autophagy in cancer. EMBO J. 2022;41(13):e110031. https://doi.org/10.15252/embj.2021110031.
Stockwell BR, Friedmann AJ, Bayir H, Bush AI, Conrad M, Dixon SJ, Fulda S, Gascon S, Hatzios SK, Kagan VE, et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell. 2017;171(2):273–85. https://doi.org/10.1016/j.cell.2017.09.021.
Sun X, Ou Z, Chen R, Niu X, Chen D, Kang R, Tang D. Activation of the p62-keap1-nrf2 pathway protects against ferroptosis in hepatocellular carcinoma cells. Hepatology. 2016;63(1):173–84. https://doi.org/10.1002/hep.28251.
Chen L, Zhang C, Xue R, Liu M, Bai J, Bao J, Wang Y, Jiang N, Li Z, Wang W, et al. Deep whole-genome analysis of 494 hepatocellular carcinomas. Nature. 2024;627(8004):586–93. https://doi.org/10.1038/s41586-024-07054-3.
Zhang J, Bajari R, Andric D, Gerthoffert F, Lepsa A, Nahal-Bose H, Stein LD, Ferretti V. The international cancer genome consortium data portal. Nat Biotechnol. 2019;37(4):367–9. https://doi.org/10.1038/s41587-019-0055-9.
Roessler S, Jia HL, Budhu A, Forgues M, Ye QH, Lee JS, Thorgeirsson SS, Sun Z, Tang ZY, Qin LX, et al. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 2010;70(24):10202–12. https://doi.org/10.1158/0008-5472.CAN-10-2607.
Liu J, Sun G, Pan S, Qin M, Ouyang R, Li Z, Huang J. The cancer genome atlas (tcga) based m(6)a methylation-related genes predict prognosis in hepatocellular carcinoma. Bioengineered. 2020;11(1):759–68. https://doi.org/10.1080/21655979.2020.1787764.
Liu Z, Liu L, Weng S, Guo C, Dang Q, Xu H, Wang L, Lu T, Zhang Y, Sun Z, et al. Machine learning-based integration develops an immune-derived lncrna signature for improving outcomes in colorectal cancer. Nat Commun. 2022;13(1):816. https://doi.org/10.1038/s41467-022-28421-6.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cbioportal. Sci Signal. 2013;6(269):l1. https://doi.org/10.1126/scisignal.2004088.
Wilkerson MD, Hayes DN. Consensusclusterplus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26(12):1572–3. https://doi.org/10.1093/bioinformatics/btq170.
Hanzelmann S, Castelo R, Guinney J. Gsva: gene set variation analysis for microarray and rna-seq data. BMC Bioinform. 2013;14(7):1–5. https://doi.org/10.1186/1471-2105-14-7.
Balachandran VP, Gonen M, Smith JJ, Dematteo RP. Nomograms in oncology: more than meets the eye. Lancet Oncol. 2015;16(4):e173–80. https://doi.org/10.1016/S1470-2045(14)71116-7.
Blanche P, Dartigues JF, Jacqmin-Gadda H. Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks. Stat Med. 2013;32(30):5381–97. https://doi.org/10.1002/sim.5958.
Alba AC, Agoritsas T, Walsh M, Hanna S, Iorio A, Devereaux PJ, Mcginn T, Guyatt G. Discrimination and calibration of clinical prediction models: users’’ guides to the medical literature. JAMA. 2017;318(14):1377–84. https://doi.org/10.1001/jama.2017.12126.
Sharma A, Seow J, Dutertre CA, Pai R, Bleriot C, Mishra A, Wong R, Singh G, Sudhagar S, Khalilnezhad S, et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell. 2020;183(2):377–94. https://doi.org/10.1016/j.cell.2020.08.040.
Zhang J. Clustergvis: one-step to cluster and visualize gene expression matrix. 2022
Patel AP, Tirosh I, Trombetta JJ, Shalek AK, Gillespie SM, Wakimoto H, Cahill DP, Nahed BV, Curry WT, Martuza RL, et al. Single-cell rna-seq highlights intratumoral heterogeneity in primary glioblastoma. Science. 2014;344(6190):1396–401. https://doi.org/10.1126/science.1254257.
Trapnell C, Cacchiarelli D, Grimsby J, Pokharel P, Li S, Morse M, Lennon NJ, Livak KJ, Mikkelsen TS, Rinn JL. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol. 2014;32(4):381–6. https://doi.org/10.1038/nbt.2859.
Gulati GS, Sikandar SS, Wesche DJ, Manjunath A, Bharadwaj A, Berger MJ, Ilagan F, Kuo AH, Hsieh RW, Cai S, et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science. 2020;367(6476):405–11. https://doi.org/10.1126/science.aax0249.
Zeng D, Ye Z, Shen R, Yu G, Wu J, Xiong Y, Zhou R, Qiu W, Huang N, Sun L, et al. Iobr: multi-omics immuno-oncology biological research to decode tumor microenvironment and signatures. Front Immunol. 2021;12:687975. https://doi.org/10.3389/fimmu.2021.687975.
Topalian SL, Drake CG, Pardoll DM. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell. 2015;27(4):450–61. https://doi.org/10.1016/j.ccell.2015.03.001.
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, Li Z, Traugh N, Bu X, Li B, et al. Signatures of t cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–8. https://doi.org/10.1038/s41591-018-0136-1.
Yang W, Soares J, Greninger P, Edelman EJ, Lightfoot H, Forbes S, Bindal N, Beare D, Smith JA, Thompson IR, et al. Genomics of Drug Sensitivity in Cancer (gdsc): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013;41:D955-61. https://doi.org/10.1093/nar/gks1111.
Liu Y, Xun Z, Ma K, Liang S, Li X, Zhou S, Sun L, Liu Y, Du Y, Guo X, et al. Identification of a tumour immune barrier in the hcc microenvironment that determines the efficacy of immunotherapy. J Hepatol. 2023;78(4):770–82. https://doi.org/10.1016/j.jhep.2023.01.011.
Mariathasan S, Turley SJ, Nickles D, Castiglioni A, Yuen K, Wang Y, Kadel EI, Koeppen H, Astarita JL, Cubas R, et al. Tgfbeta attenuates tumour response to pd-l1 blockade by contributing to exclusion of t cells. Nature. 2018;554(7693):544–8. https://doi.org/10.1038/nature25501.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative pcr and the 2(-delta delta c(t)) method. Methods. 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.1262.
Xiang DM, Sun W, Ning BF, Zhou TF, Li XF, Zhong W, Cheng Z, Xia MY, Wang X, Deng X, et al. The hlf/il-6/stat3 feedforward circuit drives hepatic stellate cell activation to promote liver fibrosis. Gut. 2018;67(9):1704–15. https://doi.org/10.1136/gutjnl-2016-313392.
Menyhart O, Nagy A, Gyorffy B. Determining consistent prognostic biomarkers of overall survival and vascular invasion in hepatocellular carcinoma. R Soc Open Sci. 2018;5(12):181006. https://doi.org/10.1098/rsos.181006.
Wu R, Guo W, Qiu X, Wang S, Sui C, Lian Q, Wu J, Shan Y, Yang Z, Yang S, et al. Comprehensive analysis of spatial architecture in primary liver cancer. Sci Adv. 2021;7(51):eabg3750. https://doi.org/10.1126/sciadv.abg3750.
Hu D, Zhang T, Yan Z, Wang L, Wang Y, Meng N, Tu B, Teng Y, Li Z, Lou X, et al. Multimolecular characteristics of cell-death related hub genes in human cancers: a comprehensive pan-cancer analysis. Cell Cycle. 2022;21(22):2444–54. https://doi.org/10.1080/15384101.2022.2101337.
Zou Y, Xie J, Zheng S, Liu W, Tang Y, Tian W, Deng X, Wu L, Zhang Y, Wong CW, et al. Leveraging diverse cell-death patterns to predict the prognosis and drug sensitivity of triple-negative breast cancer patients after surgery. Int J Surg. 2022;107:106936. https://doi.org/10.1016/j.ijsu.2022.106936.
Pan H, Pan J, Li P, Gao J. Characterization of panoptosis patterns predicts survival and immunotherapy response in gastric cancer. Clin Immunol. 2022;238:109019. https://doi.org/10.1016/j.clim.2022.109019.
Yang J, Huang Y, Song M, Pan Q, Zhao J, He J, Ouyang D, Yang C, Han Y, Tang Y, et al. Spc25 promotes proliferation and stemness of hepatocellular carcinoma cells via the dna-pk/akt/notch1 signaling pathway. Int J Biol Sci. 2022;18(14):5241–59. https://doi.org/10.7150/ijbs.71694.
Yang H, Liu X, Zhu X, Zhang M, Wang Y, Ma M, Lv K. Gins1 promotes the proliferation and migration of glioma cells through usp15-mediated deubiquitination of top2a. iScience. 2022;25(9):104952. https://doi.org/10.1016/j.isci.2022.104952.
Nagahama Y, Ueno M, Miyamoto S, Morii E, Minami T, Mochizuki N, Saya H, Takakura N. Psf1, a dna replication factor expressed widely in stem and progenitor cells, drives tumorigenic and metastatic properties. Cancer Res. 2010;70(3):1215–24. https://doi.org/10.1158/0008-5472.CAN-09-3662.
Zhou YF, Song SS, Tian MX, Tang Z, Wang H, Fang Y, Qu WF, Jiang XF, Tao CY, Huang R, et al. Cystathionine beta-synthase mediated prrx2/il-6/stat3 inactivation suppresses tregs infiltration and induces apoptosis to inhibit hcc carcinogenesis. J Immunother Cancer. 2021;9(8).https://doi.org/10.1136/jitc-2021-003031.
Yang HC, Wu YH, Yen WC, Liu HY, Hwang TL, Stern A, Chiu DT. The redox role of g6pd in cell growth, cell death, and cancer. Cells. 2019;8(9):1055. https://doi.org/10.3390/cells8091055.
Arcasoy MO. Erythropoiesis-stimulating agent use in cancer: preclinical and clinical perspectives. Clin Cancer Res. 2008;14(15):4685–90. https://doi.org/10.1158/1078-0432.CCR-08-0264.
Yang L, Zhang Z, Sun Y, Pang S, Yao Q, Lin P, Cheng J, Li J, Ding G, Hui L, et al. Integrative analysis reveals novel driver genes and molecular subclasses of hepatocellular carcinoma. Aging (Albany NY). 2020;12(23):23849–71. https://doi.org/10.18632/aging.104047.
Hui Y, Zeng H, Feng Y, Qin W, Chen P, Huang L, Zhong W, Lin L, Lv H, Qin X. Regulatory role of sfn gene in hepatocellular carcinoma and its mechanism. Biotechnol Bioprocess Eng. 2021;26(3):375–83. https://doi.org/10.1007/s12257-020-0292-2.
Wong LL, Lam IP, Wong TY, Lai WL, Liu HF, Yeung LL, Ching YP. Ipa-3 inhibits the growth of liver cancer cells by suppressing pak1 and nf-kappab activation. PLoS ONE. 2013;8(7):e68843. https://doi.org/10.1371/journal.pone.0068843.
Pan JA, Sun Y, Jiang YP, Bott AJ, Jaber N, Dou Z, Yang B, Chen JS, Catanzaro JM, Du C, et al. Trim21 ubiquitylates sqstm1/p62 and suppresses protein sequestration to regulate redox homeostasis. Mol Cell. 2016;61(5):720–33. https://doi.org/10.1016/j.molcel.2016.02.007.
Ying Z, Tian H, Li Y, Lian R, Li W, Wu S, Zhang HZ, Wu J, Liu L, Song J, et al. Cct6a suppresses smad2 and promotes prometastatic tgf-beta signaling. J Clin Invest. 2017;127(5):1725–40. https://doi.org/10.1172/JCI90439.
Pinter M, Scheiner B, Peck-Radosavljevic M. Immunotherapy for advanced hepatocellular carcinoma: a focus on special subgroups. Gut. 2021;70(1):204–14. https://doi.org/10.1136/gutjnl-2020-321702.
Liu J, Shi Y, Zhang Y. Multi-omics identification of an immunogenic cell death-related signature for clear cell renal cell carcinoma in the context of 3p medicine and based on a 101-combination machine learning computational framework. EPMA J. 2023;14(2):275–305. https://doi.org/10.1007/s13167-023-00327-3.
Ge Z, Feng P, Zhang Z, Liang Z, Chen R, Li J. Identification of novel serum protein biomarkers in the context of 3p medicine for intravenous leiomyomatosis: a data-independent acquisition mass spectrometry-based proteomics study. EPMA J. 2023;14(4):613–29. https://doi.org/10.1007/s13167-023-00338-0.
Kurysheva NI, Rodionova OY, Pomerantsev AL, Sharova GA, Golubnitschaja O. Machine learning-couched treatment algorithms tailored to individualized profile of patients with primary anterior chamber angle closure predisposed to the glaucomatous optic neuropathy. EPMA J. 2023;14(3):527–38. https://doi.org/10.1007/s13167-023-00337-1.
Lu M, Zhan H, Liu B, Li D, Li W, Chen X, Zhou X. N6-methyladenosine-related non-coding rnas are potential prognostic and immunotherapeutic responsiveness biomarkers for bladder cancer. EPMA J. 2021;12(4):589–604. https://doi.org/10.1007/s13167-021-00259-w.
Goldstein E, Yeghiazaryan K, Ahmad A, Giordano FA, Frohlich H, Golubnitschaja O. Optimal multiparametric set-up modelled for best survival outcomes in palliative treatment of liver malignancies: unsupervised machine learning and 3 pm recommendations. EPMA J. 2020;11(3):505–15. https://doi.org/10.1007/s13167-020-00221-2.
Funding
This work was supported by a grant from the National Natural Science Foundation of China (82002588, 32300731), the Natural Science Foundation project of Shanghai (23ZR1477200), the State Key Laboratory of the Cancer Biology Project (CBSKL2022ZDKF05) and the Shanghai Key Laboratory of Cell Engineering (14DZ2272300).
Author information
Authors and Affiliations
Contributions
All authors searched the literature, designed the study, interpreted the findings, and revised the manuscript. DTH, XS, and PG carried out data management and statistical analysis and drafted the manuscript. DTH, XS, PG, TTM, YC, WFS, YGZ, and JD helped with cohort identification and data management. YGZ and JD contributed to the critical revision of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All participants were enrolled from the Eastern Hepatobiliary Surgery Hospital (EHBH), and the sample collection procedure was approved by the ethics committee of EHBH.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Dingtao Hu, Xu Shen, and Peng Gao share co-first authorship.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, D., Shen, X., Gao, P. et al. Multi-omic profiling reveals potential biomarkers of hepatocellular carcinoma prognosis and therapy response among mitochondria-associated cell death genes in the context of 3P medicine. EPMA Journal 15, 321–343 (2024). https://doi.org/10.1007/s13167-024-00362-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13167-024-00362-8