Abstract
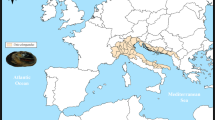
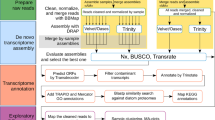
The genetic basis for bivalves’ adaptation and evolution is not well understood. Even few studies have focused on the mechanism of molluscan molecular evolution between the coastal intertidal zone and deep-sea environment. In our studies, we first conducted the transcritpome assembly of Modiolus modiolus mussels living in coastal intertidal zones. Also, we conducted transcriptome comparison analyses between M. modiolus and Bathymodiolus platifrons living in hydrothermal vents and cold methane/sulfide-hydrocarbon seeps. De novo assemblies of the clean reads yielded a total of 182 476 and 156 261 transcripts with N50 values of 1 769 and 1 545 in M. modiolus and B. platifrons. A total of 27 868 and 23 588 unigenes were identified, which also displayed the similar GO representation patterns. Among the 10 245 pairs of putative orthologs, we identified 26 protein-coding genes under strong positive selection (Ka/Ks>1) and 12 genes showing moderate positive selection (0.5<Ka/Ks<1). Most of those genes are predicted to be involved in stress resistance. Overall, our study first provides the transcriptomic database for M. modiolus. Transcriptome comparison illustrates the genome evolution between M. modiolus and B. platifrons, and provides an important foundation for future studies on these two species.
Article PDF
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
References
Barry J P, Buck K R, Kochevar R K, et al. 2002. Methane–based symbiosis in a mussel, Bathymodiolus platifrons, from cold seeps in Sagami Bay, Japan. Invertebrate Biology, 121(1): 47–54
Bettencourt R, Roch P, Stefanni S, et al. 2007. Deep sea immunity: unveiling immune constituents from the hydrothermal vent mussel Bathymodiolus azoricus. Marine Environmental Research, 64(2): 108–127
Canesi L, Gallo G, Gavioli M, et al. 2002. Bacteria–hemocyte interactions and phagocytosis in marine bivalves. Microscopy Research and Technique, 57(6): 469–476
Dame R F. 2011. Ecology of Marine Bivalves: An Ecosystem Approach. 2nd ed. Boca Raton: CRC Press
Dinesen G E, Morton B. 2014. Review of the functional morphology, biology and perturbation impacts on the boreal, habitat–forming horse mussel Modiolus modiolus (Bivalvia: Mytilidae: Modiolinae). Marine Biology Research, 10(9): 845–870
Duperron S, Guezi H, Gaudron S M, et al. 2011. Relative abundances of methane–and sulphur–oxidising symbionts in the gills of a cold seep mussel and link to their potential energy sources. Geobiology, 9(6): 481–91
Egas C, Pinheiro M, Gomes P, et al. 2012. The Transcriptome of Bathymodiolus azoricus gill reveals expression of genes from endosymbionts and free–living deep–sea bacteria. Marine Drugs, 10(8): 1765–1783
Fujiwara Y, Takai K, Uematsu K, et al. 2000. Phylogenetic characterization of endosymbionts in three hydrothermal vent mussels: influence on host distributions. Marine Ecology Progress Series, 208: 147–155
Grabherr M G, Haas B J, Yassour M, et al. 2011. Full–length transcriptome assembly from RNA–Seq data without a reference genome. Nature Biotechnology, 29(7): 644–652
Jones W J, Won Y J, Maas P A Y, et al. 2006. Evolution of habitat use by deep–sea mussels. Marine Biology, 148(4): 841–851
Kanehisa M, Araki M, Goto S, et al. 2008. KEGG for linking genomes to life and the environment. Nucleic Acids Research, 36: D480–D484
Kavembe G D, Franchini P, Irisarri I, et al. 2015. Genomics of adaptation to multiple concurrent stresses: insights from comparative transcriptomics of a cichlid fish from one of earth’s most extreme environments, the Hypersaline Soda Lake Magadi in Kenya, East Africa. Journal of Molecular Evolution, 81(3–4): 90–109
Li Bo, Dewey C N. 2011. RSEM: accurate transcript quantification from RNA–Seq data with or without a reference genome. BMC Bioinformatics, 12: 323
Li Li, Stoeckert C J Jr, Roos D S. 2003. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Research, 13(19): 2178–2189
Li Qi, Zhao Xuelin, Kong Lingfeng, et al. 2013. Transcriptomic response to stress in marine bivalves. ISJ–Invertebrate Survival Journal, 10(1): 84–93
Mao Xizeng, Cai Tao, Olyarchuk J G, et al. 2005. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics, 21(19): 3787–3793
Nakamura–Kusakabe I, Nagasaki T, Kinjo A, et al. 2016. Effect of sulfide, osmotic, and thermal stresses on taurine transporter mRNA levels in the gills of the hydrothermal vent–specific mussel Bathymodiolus septemdierum. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 191: 74–79
Riesgo A, Andrade S C S, Sharma P P, et al. 2012. Comparative description of ten transcriptomes of newly sequenced invertebrates and efficiency estimation of genomic sampling in nonmodel taxa. Frontiers in Zoology, 9: 33
Saavedra C, Bachère E. 2006. Bivalve genomics. Aquaculture, 256(1–4): 1–14
Schuster S C. 2008. Next–generation sequencing transforms today’s biology. Nature Methods, 5(1): 16–18
Sun Jin, Zhang Yu, Xu Ting, et al. 2017. Adaptation to deep–sea chemosynthetic environments as revealed by mussel genomes. Nature Ecology & Evolution, 1(5): 0121
Toubiana M, Gerdol M, Rosani U, et al. 2013. Toll–like receptors and MyD88 adaptors in Mytilus: complete cds and gene expression levels. Developmental & Comparative Immunology, 40(2): 158–166
Vitti J J, Grossman S R, Sabeti P C. 2013. Detecting natural selection in genomic Data. Annual Review of Genetics, 47(1): 97–120
Wang Zhong, Gerstein M, Snyder M. 2009. RNA–Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics, 10(1): 57–63
Wang Shan, Hou Rui, Bao Zhenmin, et al. 2013. Transcriptome sequencing of Zhikong scallop (Chlamys farreri) and comparative transcriptomic analysis with Yesso scallop (Patinopecten yessoensis). PLoS One, 8(5): e63927
Wang Hailiang, Sun Li. 2017. Comparative metagenomics reveals insights into the deep–sea adaptation mechanism of the microorganisms in Iheya hydrothermal fields. World Journal of Microbiology Biotechnology, 33: 86
Wang Shi, Zhang Jinbo, Jiao Wenqian, et al. 2017. Scallop genome provides insights into evolution of bilaterian karyotype and development. Nature Ecology & Evolution, 1(5): e0120
Wong Y H, Sun Jin, He Lisheng, et al. 2015. High–throughput transcriptome sequencing of the cold seep mussel Bathymodiolus platifrons. Scientific Reports, 5: 16597
Yang Ziheng. 2007. PAML 4: phylogenetic analysis by maximum likelihood. Molecular Biology and Evolution, 24(8): 1586–1591
Young M D, Wakefield M J, Smyth G K, et al. 2010. Gene ontology analysis for RNA–seq: accounting for selection bias. Genome Biology, 11(2): R14
Zhang Guofang, Fang Xiaodong, Guo Ximing, et al. 2012. The oyster genome reveals stress adaptation and complexity of shell formation. Nature, 490(7418): 49–54
Zhao Xuelin, Yu Hong, Kong Lingfeng, et al. 2014. Comparative transcriptome analysis of two oysters, Crassostrea gigas and Crassostrea hongkongensis provides insights into adaptation to hypo–osmotic conditions. PLoS One, 9(11): e111915
Zheng Ping, Wang Minxiao, Li Chaolun, et al. 2017. Insights into deep–sea adaptations and host–symbiont interactions: a comparative transcriptome study on Bathymodiolus mussels and their coastal relatives. Molecular Ecology, 26(19): 5133–5148
Acknowledgements
The authors thank the scientists and crew of the R/V Kexue for their assistance in specimen collecting.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Foundation item: The Strategic Priority Research Program of the Chinese Academy of Sciences under contract No. XDB06010101; the Technological Innovation Project financially supported by Qingdao National Laboratory for Marine Science and Technology under contract No. 2015ASKJ02-03; Shandong Provincial Natural Science Foundation, China under contract No. ZR2016DQ13; the Earmarked Fund for Modern Agro-industry Technology Research System under contract No. CARS-48; the Taishan Scholars Climbing Program of Shandong; the project funded by China Postdoctoral Science Foundation.
Rights and permissions
About this article
Cite this article
Meng, J., Yang, M., Xu, F. et al. Transcriptome assembly of Modiolus modiolus and comparative analysis with Bathymodiolus platifrons. Acta Oceanol. Sin. 37, 38–45 (2018). https://doi.org/10.1007/s13131-018-1232-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13131-018-1232-2