Abstract
Orthoptera have some of the largest genomes of all insects. At the same time, the architecture of their genomes remains poorly understood. Comparative cytological data across a wide range of taxa, even for basic parameters such as chromosome number, may provide important insights into the evolution of these genomes and help answer the question of why some species attained such large genome sizes. We collected and compiled more than 1,000 records of chromosome numbers of 339 genera (13.8% of 2,452 known genera) and 769 species (6.2% of 12,250 known species) of Caelifera, the suborder of Orthoptera that includes those taxa with short antennae. Within the family Acrididae, most of the records come from the subfamilies Oedipodinae (N = 325), Melanoplinae (N = 192) and Gomphocerinae (N = 254). Out of the 621 investigated species of Acrididae, 459 (73.9%) shared a chromosome number of 2n♂ = 23. Chromosome numbers of 2n♂ = 17 (12.2%) and 2n♂ = 21 (9.9%) were less common. The remaining 4.0% of species exhibited different chromosome numbers between 2n♂ = 8 (6 + XY) and 2n♂ = 27. Plotted on a phylogenetic tree, our results confirm that chromosome numbers, especially in the largest grasshopper family Acrididae, are highly conserved with a basic count of 2n♂ = 23 (22 + X0), sometimes reduced to, e.g., 2n♂ = 17 (16 + X0) in some genera of the slant-faced grasshopper subfamily Gomphocerinae. Species with divergent chromosome numbers occur in many of the groups we studied, but are not a systematic trait and have evolved multiple times independently. Our study supports the view that chromosome numbers are much more stable across the investigated Caelifera compared to Ensifera, the second suborder of Orthoptera that includes the long antennae bush crickets and crickets. Our results significantly extend our knowledge on the diversity of this character in Caelifera.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prior to the genomic era, cytogenetic studies provided the foundation for our understanding of animal genome organization (Bugrov, 1988, 1996; Bugrov & Vysotskaya, 1981; Confalonieri et al., 1998; Gokhman & Kuznetsova, 2006; King, 1995; Kirkpatrick, 2010; Vandergast et al., 2017). While genetic and genomic sequencing have become far more popular fields of research, cytogenetic studies still provide important information about the genomic organization of a species and clues to the evolution of whole groups of taxa (White, 1973). They have been used to address a variety of systematic, evolutionary and phylogenetic questions in plants and animals and have helped to improve our understanding of speciation (Charlesworth, 2004; Charlesworth & Charlesworth, 2005; Grzywacz et al., 2019; Navarro & Barton, 2003). Comparative cytogenetics implements relatively simple studies of chromosome numbers and morphology, but it may also include more complex analyses of various banding patterns or highly specified gene probes with fluorescent staining (White & Solt, 1978; Zhong et al., 1996; Gokhman & Kuznetsova, 2006; Bishop, 2010). While these complex methods allow fine-scale analyses on the level of populations, comparative studies of chromosome numbers may give us insight into the higher levels of evolutionary processes.
Grasshoppers of the family Acrididae (Orthoptera: Caelifera) have been the target of intense cytogenetic studies (Cigliano et al., 2021). This group has been suggested to be relatively uniform in their chromosome number, with some exceptions (Hewitt, 1979; John & Hewitt, 1966). While a diploid chromosome number of 2n♂ = 23 (22 + X0) is considered the basic plan for Acrididae (Hewitt, 1979), different kinds of rearrangements, especially Robertsonian fusions, led to a reduction in chromosome number in some groups of Caelifera (e.g., many Eurasian Gomphocerinae have 2n♂ = 17 (16 + X0) chromosomes). McClung (1917) considered this variation in the number of chromosomes to be a matter of rearrangements of chromatin rather than a result of the loss or gain of individual chromosomes. Besides this, some variation in the sex determining system has led to variation in chromosome number. In general, loss of the Y chromosome led to the highly conserved sex chromosome pattern of X0♂/XX♀ found in most species. Due to several chromosome rearrangements (autosomes and sex chromosomes), some species evolved several alternative sex determining systems, e.g., neo-XY♂/XX♀ or even neo-X1X2Y♂/neo-X1X1X2X2♀ or X1X20♂/X1X1X2X2♀ (Palacios-Gimenez et al., 2013, 2018; Castillo et al., 2010; Hewitt, 1979; White, 1973) leading to some variation in chromosome number and providing a possible basis for reproductive isolation in some species groups.
Despite some exceptions, in comparison with its sister group, Ensifera (katydids, crickets and allies), the variation in chromosome number is relatively lower in Caelifera. Also, in general, the chromosome number appears to be lower in Caelifera compared to most Ensifera, as Warchałowska-Śliwa (1998) reported a basic number of 2n♂ = 31 (30 + XO) chromosomes in males of most of the investigated subfamilies of the family Tettigoniidae. Interestingly, genome sizes are, regardless of the chromosome number, much smaller in Ensifera compared to Caelifera, which may suggest some duplication events at the advent of the diversification of Caelifera (Husemann et al., 2021; Mao et al., 2020). A recent meta-analysis of Polyneoptera showed that many interacting factors underlie chromosome variation (Sylvester et al., 2020). Warchałowska-Śliwa (1998) summarized the cytogenetic information of about 400 species of Tettigoniidae with the aim of tracing the evolution of chromosome number in that ensiferan family. Such a systematic review and analysis of chromosome number and evolution is lacking for the diverse caeliferan family Acrididae. Hence, with the aim of closing this gap, we provide new karyotype data of 36 species (and additional estimates of 8 previously investigated species) of Acrididae and assembled a dataset of 1,284 records of chromosome numbers for Caelifera representing 339 genera (13.8% of 2,452 known genera) and 769 species (6.2% of 12,250 known species), including 1,108 records of Acrididae. We provide an overview of the variability of karyotypes for several subfamilies of Acrididae and map chromosome numbers on the most recent phylogeny of Caelifera (Song et al., 2018) in order to get an insight into the evolution of chromosome number in this diverse group of grasshoppers.
Materials and methods
Material examined
We collected male grasshoppers belonging to 16 species of Oedipodinae by sweep net sampling on field trips between 2014 and 2016 (SI 1).Voucher specimens were deposited at the entomological collection of the Zoological Museum Hamburg, Germany (ZMH). David B. Weissman and David Lightfoot have collected and analyzed western US grasshoppers over the years and we included 28 unreported results herein (DBW, unpubl. Data).
Cytogenetic analyses
We dissected and fixed testes of the collected specimens in the field in a solution consisting of three parts of ethanol–acetic acid (3:1, v/v). Specimens were fixed in 99.9% ethanol after dissection. The samples were subsequently stored in a freezer at -20 °C until further processing. NU conducted chromosome analysis: Testes were stained with an alcohol–carmine solution for several hours, before being transferred to glass slides for squash preparation, chromosome counts and microscopic imaging (see Lightfoot et al., 2011; Ueshima & Rentz, 1979). DBW material was analyzed as in Rentz & Weissman (1984).
Chromosome mapping and ancestral state reconstruction
We added our newly generated data to a large dataset based on previously published data: We screened the literature and additionally included all unique records from two online databases: www.bchrom.csic.es and www.coleoguy.github.io/karyotypes. In total, we gathered 1,284 records of chromosome numbers of Caelifera, including the 1,108 records of Acrididae used in our analyses. Throughout the manuscript, we only show the male chromosome numbers if the normal X0 sex determining system is realized. In cases of deviating sex chromosome configurations, these are noted.
We visualized the distribution of male chromosome numbers of Acrididae as a histogram in R using the packages ggplot2 (Wickham, 2016), ggpubr (Kassambara, 2020), scales (Wickham & Seidel, 2020) and cowplot (Wilke, 2019). All subfamilies were colored to display differences in numbers between taxa.
We mapped male chromosome numbers on the most recent phylogeny of the group (Song et al., 2018) using the R packages BiocManager (Morgan, 2019), phytools (Revell, 2012), vctrs (Wickham et al., 2020), ggplot2 (Wickham, 2016), ggtree (Yu et al., 2018, 2017), gtable (Wickham & Pedersen, 2019), grid (R Core Team, 2020), ggstance (Henry et al., 2020) and tidyverse (Wickham et al., 2019). A parsimony-based ancestral state reconstruction of chromosome number was done in Mesquite v. 3.51 (The Mesquite Project Team., 2019) based on a character matrix approach. We chose this reconstruction method due to missing data of several groups within the tree and several chromosome number configurations occurring only single times throughout the evaluated taxa. Parsimony approaches for ancestral state reconstruction are known to be as accurate in state reconstruction of deep and shallow nodes as likelihood approaches (Holland et al., 2020). We performed all R analyses in R version 3.6.3 (R Core Team, 2020).
Results
Cytogenetic analyses
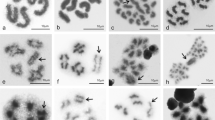
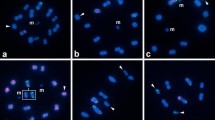
All species newly analyzed here had a karyotype of 2n♂ = 23 chromosomes with no variation or heteromorphism in any of the specimens studied (Fig. 1, SI 1) with the exception of Teicophrys californiae Descamps, 1977, which had 2n♂ = 17. All chromosomes were acrocentric or telocentric.
Histogram displaying A: the distribution of chromosome numbers across Caelifera and B: chromosome numbers across the different subfamilies of Acrididae. Chromosome numbers are shown as relative frequencies in percent; here, just subfamilies with more than ten records are shown as separated units. Subfamilies with lower sample sizes are aggregated in the unit other (Calliptaminae, Coptacrinae, Euryphyminae, Hemiacridinae, Marelliinae, Pauliniinae, Pezotettiginae, Proctolabinae, Rhytidochrotinae, Spathosterninae and Tropidopolinae)
Review of chromosome numbers in Caelifera
We assembled records of chromosome numbers for 1,284 records of Caelifera, including 1,108 records for Acrididae. The data include 769 species in 339 genera of Caelifera (SI 1). We found multiple records for many species, some of which documented variation in the chromosome count of some species. Specifically, we found documented differences for Miramella alpina (Kollar, 1833), Bucephalacris bohlsii (Giglio-Tos, 1898), Circotettix coconino Rehn, 1921, C. crotalum Rehn, 1921, C. undulatus thalassinus Saussure, 1884, Trimerotropis cyaneipennis Bruner, 1889, T. gracilis gracilis (Thomas, 1872), T. ochraceipennis (Blanchard, 1851), T. sparsa (Thomas, 1875) and Podisma pedestris (Linnaeus, 1758) with 2n♂ = 21 and 2n♂ = 23 individuals reported; Scyllinula humilis (Blanchard, 1851), Dichroplus maculipennis (Blanchard, 1851) and Leiotettix sanguineus Bruner, 1906 with 2n♂ = 22 (20 + XY) and 2n♂ = 23; Orphulella punctata (De Geer, 1773) and Chortoicetes terminifera (Walker, 1870) with 2n♂ = 17 and 2n♂ = 23; Leiotettix politus Rehn, 1913 with 2n♂ = 13 and 2n♂ = 14 (12 + XY); Dichroplus pratensis Bruner, 1900 with 2n♂ = 18 (16 + XY) and 2n♂ = 22 (20 + XY); Oedipoda schochii Brunner von Wattenwyl, 1884 2n♂ = 23 and 2n♂ = 25; Gomphocerus sibiricus (Linnaeus, 1767) 2n♂ = 17 and 2n♂ = 19; Dichroplus vittatus Bruner, 1900 2n♂ = 18 (16 + XY) and 2n♂ = 20 (18 + XY); and Dichroplus fuscus (Thunberg, 1815) 2n♂ = 19 and 2n♂ = 23. Such discrepancies within a species should be reinvestigated as these differences, if reconfirmed, potentially represent cryptic species situations.
For trait mapping and ancestral state reconstruction, we also included some more general records at the family or subfamily level with missing species identification based on the data from White (1973). Within the family Acrididae, most of the records come from the subfamilies Oedipodinae (N = 325), Melanoplinae (N = 192) and Gomphocerinae (N = 254). Out of the 621 investigated species of Acrididae, 459 (73.9%) shared a chromosome number of 2n♂ = 23. Chromosome numbers of 2n♂ = 17 (12.2%) and 2n♂ = 21 (9.9%) were less common. The remaining 4.0% of species exhibited different chromosome numbers between 2n♂ = 8 (6 + XY) and 2n♂ = 27. A chromosome number of 2n♂ = 17 was found mostly in Gomphocerinae (Stenobothrini, Gomphocerini and European Chrysochraontini), while 2n♂ = 21 was found mostly in Melanoplinae and some Oedipodinae (Trimerotropis Stål, 1873 and Circotettix Scudder, 1876). The lowest number of chromosomes of all Caelifera studied so far was found in Dichroplus silveiraguidoi Liebermann, 1956 with 2n♂ = 8 (6 + XY) (Mesa et al., 1982). A high number with 2n♂ = 25 was found in Oedipoda schochii Brunner von Wattenwyl, 1884 (Türkoglu & Koca, 2002) and Conometopus sulcaticollis (Blanchard, 1851) (Mesa et al., 1982). The highest number with 2n♂ = 27 was found in Dichroplus intermedius Ronderos, 1976 (Türkoglu & Koca, 2002). Caelifera show more deviation from the typical 2n♂ = 23 than Acrididae alone. The number of 2n♂ = 19 is common within Pamphagidae (e.g., Melanotmethis Uvarov, 1943, Pezotmethis Uvarov, 1943, Strumiger Zubovski, 1896) and several Eumastacidae genera (e.g., Phytomastax Bey-Bienko, 1949, Gomphomastax Brunner von Wattenwyl, 1898, Clinomastax Bey-Bienko, 1949) (e.g., Bugrov, 1986, 1988, 1996; Bugrov et al., 1991; Vysotskaya, 1983; White, 1968). Interestingly the Tetriginae genera are known to be more variable in their chromosome number configuration. Here, several species of the genus Tetrix have a common chromosome number of 2n♂ = 13 (Bugrov, 1996).
Mapping and ancestral state reconstruction
We mapped the chromosome number on the phylogeny provided by Song et al. (2018) (Fig. 2) and performed ancestral state reconstruction. In their phylogeny, Song et al. (2018) showed that Acrididae roughly form four monophyletic groups (Clades A to D). Our analysis shows that three of these groups (A to C) show different degrees of polymorphism in chromosome number, whereas the fourth group (Clade D) appears monomorphic with a consistent chromosome number of 2n♂ = 23. However, we were not able to obtain chromosome numbers for all taxa included in the phylogeny.
Male chromosome numbers mapped on the phylogeny of Acrididae constructed by Song et al. (2018). Mapping and ancestral state reconstruction of chromosome number with Mesquite
Clade A comprises Marelliinae, Pauliniinae, Leptysminae, Ommatolampidinae (polyphyletic) and Rhytidochrotinae. Leptysminae most commonly showed the typical 2n♂ = 23, but chromosome numbers also comprised 2n♂ = 19 in the genus Tetrataenia Stål, 1873, 2n♂ = 13 or 21 in Leptysma Stål, 1873 and 2n♂ = 21 in Stenopola Stål, 1873. In Ommatolampidinae, chromosome numbers varied among the tribes Abracrini (2n♂ = 19: Jodacris Giglio-Tos, 1897, Sitalces Stål, 1878; 2n♂ = 21: Eujivarus Bruner, 1911, Abracris Walker, 1870, Omalotettix Bruner, 1906), Pycnosarcini (2n♂ = 17: Pycnosarcus Bolívar, 1906, Lagidacris Amédégnato & Descamps, 1979) and Clematodini (2n♂ = 21: Bucephalacris Giglio-Tos, 1894).
Clade B comprises mostly paraphyletic subfamilies: Hemiacridinae, Tropidopolinae, Oedipodinae, Coptacrinae (monophyletic), Gomphocerinae and Acridinae. Chromosome number deviation from 2n♂ = 23 was only recorded for Oedipodinae (Trimerotropis, Circotettix: 2n♂ = 21; Oedipoda: 2n♂ = 25; Machaerocera Saussure, 1859 2n♂ = 16 (14 + XY); Chortoicetes Brunner von Wattenwyl, 1893: 2n♂ = 17) and Gomphocerinae (Gomphocerus Thunberg, 1815, Neopodismopsis Bey-Bienko, 1932 syn. Chloealtis Harris, 1841: 2n♂ = 19 Euchorthippus Tarbinsky, 1926, Euthystira Fieber, 1852, Eclipophleps Tarbinsky, 1927, Chorthippus Fieber, 1852, Gomphocerus, Mongolotettix Rehn, 1928, Myrmeleotettix Bolívar, 1914, Omocestus Bolívar, 1878, Chloealtis Harris, 1841, Podismopsis Zubovski, 1900, Stenobothrus Fischer, 1853: 2n♂ = 17; Mermiria Stål, 1873, Scyllinula Carbonell, 1995 2n♂ = 22 (20 + XY)).
Clade C represents the most variable group within the dataset. The group contains several genera of the paraphyletic subfamilies Hemiacridinae, Oxyinae, Copiocerinae and the monophyletic Melanoplinae, Proctolabinae and Spathosterninae. Chromosome number varied between 2n♂ = 17 and 2n♂ = 25 in the Melanoplinae genus Hesperotettix Scudder, 1876 and between 2n♂ = 8 (6 + XY) and 2n♂ = 23 within the genus Dichroplus Stål, 1873. Further variation within the subfamily was recorded for several genera of Dichroplini (2n♂ = 8 (6 + XY), 2n♂ = 13 – 16 (14 + XY), 2n♂ = 18 (16 + XY) – 23, 2n♂ = 22 (20 + XY), 2n♂ = 27) and Podismini (2n♂ = 21 – 23, 2n♂ = 25). Except for the genera Anapodisma and Coscineuta Stål, 1873 (2n♂ = 21) and some genera of the Copiocerinae (Aleuas Stål, 1878: 2n♂ = 20 (18 + XY), 2n♂ = 22 (20 + XY), 2n♂ = 19; Bucephalacris: 2n♂ = 21; Zygoclistron Rehn, 1905: 2n♂ = 20 (18 + XY), all remaining subfamilies showed the common chromosome number of 2n♂ = 23.
Ancestral state reconstruction (Fig. 2) suggests an ancestral chromosome number of 2n♂ = 23 for Acrididae. Changes in chromosome number across the phylogeny in most cases represent single species.
Discussion
Based on our dataset, we confirm a high stability of chromosome number in Acrididae with almost three quarters (73.9%) of all records reporting a number of 2n♂ = 23. This is in line with previous findings of White (1973), who suggested that two-thirds of all species have this karyotype, and findings of Aswathanarayana & Ashwath (2006) who even suggested that 90% of Acrididae share this configuration. Our study therefore confirmed the traditional view of Acrididae as a prime example of karyotypic conservatism (White, 1973), but provides a more comprehensive analysis.
Due to the rather monomorphic chromosome number configuration, many studies investigate additional chromosomal characteristics like the number of chromosome arms (e.g., Vysotskaya, 1993; Bugrov & Vysotskaya, 1981), C-banding patterns (e.g., Souza & Melo, 2007; Bugrov et al., 1991; Vysotskaya & Bugrov, 1987) or even chiasmata frequency (e.g., Gusachenko et al., 1992; Cano et al., 1986; Riva et al., 1984). However, the additional characteristics of the chromosomes were out of the scope of this study and we focused on the numbers alone.
Nevertheless, despite high degree of conservation of the chromosome number configuration of 2n♂ = 23, some groups exhibited deviations from this typical number: Several tribes of Gomphocerinae (i.e., Stenobothrini, Gomphocerini and Chrysochraontini) share a number of 2n♂ = 17, while several Tetriginae species show a configuration of 2n♂ = 13 (Bugrov, 1996). Many Pamphagidae genera show a general chromosome number of 2n♂ = 19 (e.g., Bugrov, 1986, 1996). Coleman (1948) suggested that this reduction in the chromosome number was the result of centric fusions, also known as Robertsonian translocations (Cabrero & Camacho, 1986). It is difficult to assess whether the event of chromosome number reduction occurred a single time or gradually in multiple events because the currently available phylogenetic data include only few taxa with this reduced chromosome number. As no intermediate forms have been found in any closely related groups, it may be possible that the reduction occurred in a single step as, for example, also suggested in Oxyopidae spiders (Stávale et al., 2011). However, intermediates may also be of meiotic disadvantage potentially explaining their absence.
A reduction to 2n♂ = 21 is fairly widespread in some genera of Melanoplinae, e.g., Hesperotettix and Dichroplus, and the Oedipodinae Trimerotropis and Circotettix. The North American tribe Trimerotropini was the subject of intense cytogenetic studies and showed variation in chromosome number between 2n♂ = 21 and 2n♂ = 23. Within this tribe, species of Trimerotropis and Circotettix show geographic variation in chromosome number, and several evolutionary scenarios have been developed, potentially explaining these differing chromosome numbers (Confalonieri et al., 1998; Confalonieri & Bidau, 1986; Evans, 1954; White, 1949; Coleman, 1948). White (1949) suggested that the ancestral state is the typical 2n♂ = 23 and proposed that the fusion of two acrocentric chromosomes to a metacentric chromosome has produced the decreased karyotype of 2n♂ = 21 in species of Circotettix and Trimerotropis. The metacentric chromosomes in other Trimerotropini genera originated probably by pericentric inversions (Evans, 1954), rather than translocations as suggested for the Gomphocerinae (Coleman, 1948). The effect of this chromosomal polymorphism for reproductive isolation remains debated: natural hybrids with 2n♂ = 22 have been observed in several crosses; yet, sperm quality strongly suffered in many cases suggesting some degree of hybrid sterility (Shaw et al., 1998; John et al., 1983; John & Weissman, 1977; Evans, 1954).
While deviations in chromosome numbers across most of the investigated taxa seem rather species-specific and hence have little systematic value, this appears different in some European Gomphocerinae, which share in general the reduced number of 2n♂ = 17 (except for e.g., Eremippus mistshenkoi Stebaev, 1965 2n♂ = 19 (Bugrov et al., 1993); Chorthippus hammarstroemi (Miram, 1907) 2n♂ = 21 (Kiknadze & Vyotskaya, 1970) or Stenobothrus eurasius 2n♂ = 16 (XY) (Bugrov et al., 1991)), and in the American Trimerotropini. In the latter, species have even been divided into three cytogenetically distinct groups (Sections A to C; Weissman & Rentz, 1980; White, 1949, 1951) differing in their chromosome number and morphology. It has been suggested that these differences may contribute to reproductive isolation and, therefore, speciation (e.g., Shaw et al., 1998). The two main chromosomal Sections A and B were also recovered in phylogenetic reconstructions using mitochondrial and nuclear genes (Husemann et al., 2012) and hence represent a useful systematic character. This still has to be evaluated in the Gomphocerinae.
In turn, some genera are particularly diverse in their chromosome constitutions, foremost the Melanoplinae genera Dichroplus and Hesperotettix. Chromosome numbers vary between 2n♂ = 18 (16 + XY) and 26 (24 + XY) within the genus Hesperotettix (McClung, 1917) and between 2n♂ = 8 (6 + XY) and 2n♂ = 27 in Dichroplus (Castillo et al., 2017; Mesa et al., 1982). Interestingly, there have been several studies performed on the chromosome number variation of the species Podisma sapporensis and Podisma pedestris in hybridization zones (e.g., Warchałowska-Śliwa et al., 2008; Bella et al., 1991). These studies show that reproductive isolation systems exist in hybrids, but the variation is most likely based on Robertsonian translocations between a sex chromosome and an autosome, and several chromosome rearrangements. Further, they show a clear differentiation into X0 and neo-XY chromosome races and complex chromosomal polymorphism in contact zones, which could permit the differentiation of several chromosomal races (Warchałowska-Śliwa et al., 2008).
Conclusion
Overall, a basic chromosome number of 2n♂ = 23 was observed across the whole Acrididae phylogeny and hence in all four clades described by Song et al. (2018). No subfamily with a number consistently diverging from the standard 2n♂ = 23 was recovered in the tree; but, some taxon-specific chromosome number variation appears to be present in Gomphocerinae and Trimerotropini. We conclude that the chromosome number in Caelifera, and specifically in Acrididae, is rather constant and phylogenetically less informative compared to several groups of Ensifera, which show more variation (e.g., Eneopterinae with range from 2n♂ = 9 (6 + XXY) (Palacios-Gimenez et al., 2017) up to 2n♂ = 57 in Rhaphidophoridae (Vandergast et al., 2017 and references therein)). The reasons for this need to be further explored in the future.
Availability of data and material
The authors declare the availability of data in the supplementary section of the manuscript. Further material is available in the Zoological Museum Hamburg (ZMH).
Code availability
Not applicable.
References
Aswathanarayana, N. V., & Ashwath. (2006). Structural polymorphism and C-banding pattern in a few Acridid grasshoppers. Cytologia, 71(3), 223–228.
Bella, J. L., Westerman, M., Lopez-Fernandez, C., De la Torre, J., Rubio, J. M., & Gosalvez, J. (1991). Sex chromosome and autosome divergence in Podisma (Orthoptera) in western Europe. Genetics Selection Evolution, 23(1), 1–9.
Bishop, R. (2010). Applications of fluorescence in situ hybridization (FISH) in detecting genetic aberrations of medical significance. Bioscience Horizons, 3(1), 85–95.
Bugrov, A. G. (1986). Neo-XY sex chromosome determination in grasshoppers Asiomethis heptapotamicus heptapotamicus (Zub.) and Atrichomethis semenovi (Zub.) (Orthoptera: Pamphagidae) (in Russian). Tsitologia, 117–119.
Bugrov, A. G. (1988). Karyotypes and the phylogeny of the grasshoppers of Asian region of the USSR (in Russian). Ph.D. Thesis, Biological Institute, Siberian Branch of the USSR Acadamy of Sciences. Novosibirsk.
Bugrov, A. G. (1996). Karyotypes of the short-horned Orthopteran insects (Orthoptera, Caelifera) from Russia, Kazakhstan, Central Asia, and the Caucasus. Folia Biologica (Kraków), 44, 15–26.
Bugrov, A. G., Gusachenko, A. M., & Vysotskaya, L. V. (1991). Karyotypes and C-heterochromatin regions of grasshoppers of the tribe Gomphocerini (Orthoptera: Acrididae: Gomphocerinae) in the USSR fauna (in Russian). Zoological Journal, 55–63.
Bugrov, A. G., Sergeev, M. G., & Vyotskaya, L. V. (1993). The phylogenetic status of the grasshopper genus Eremippus Uv. (Orthoptera: Acrididae). Cytogenetic analysis (in Russian). Karyosystematics of the Invertebrate Animals, Zoological Institute Press. St. Petersburg, 2, 18–21.
Bugrov, A. G., & Vysotskaya, L. V. (1981). Karyotypical properties of some grasshoppers (Orthoptera: Acridoidea) from Siberia, Middle Asia and Far East (in Russian). Ecology Novosibirsk University Press, 3–12.
Cabrero, J., & Camacho, J. P. M. (1986). Cytogenetic studies in Gomphocerine grasshoppers. I. Comparative analysis of chromosome C-banding pattern. Heredity, 56(3), 365–372.
Cano, M. I., Henriques-Gil, N., Arana, P., & Santos, J. L. (1986). The relationship between chiasma frequency and bivalent length: Effects of genotype and supernumerary chromosomes. Heredity, 56(3), 305–310.
Castillo, E. R., Marti, D. A., & Bidau, C. J. (2010). Sex and neo-sex chromosomes in Orthoptera: a review. Journal of Orthoptera Research, 213–231.
Castillo, E. R., Taffarel, A., Maronna, M. M., Cigliano, M. M., Palacios-Gimenez, O. M., Cabral-de-Mello, D. C., & Martí, D. A. (2017). Phylogeny and chromosomal diversification in the Dichroplus elongatus species group (Orthoptera, Melanoplinae). Plos One, 12(2), e0172352.
Charlesworth, D. (2004). Plant evolution: Modern sex chromosomes. Current Biology, 14(7), R271–R273.
Charlesworth, D., & Charlesworth, B. (2005). Sex chromosomes: Evolution of the weird and wonderful. Current Biology, 15(4), R129–R131.
Cigliano, M. M., Braun, H., Eades, D. C., & Otte, D. (2021). Homepage: Orthoptera Species File. http://orthoptera.speciesfile.org/HomePage/Orthoptera/HomePage.aspx
Coleman, L. C. (1948). The cytology of some western species of Trimerotropis (Acrididae). Genetics, 33(6), 519.
Confalonieri, V. A., & Bidau, C. J. (1986). The B-chromosomes of two species of Cylindrotettix (Leptysminae, Acrididae). Genetica, 68(2), 87–95.
Confalonieri, V. A., Sequeira, A. S., Todaro, L., & Vilardi, J. C. (1998). Mitochondrial DNA and phylogeography of the grasshopper Trimerotropis pallidipennis in relation to clinal distribution of chromosome polymorphisms. Heredity, 81(4), 444–452.
De Souza, M. J., & De Melo, N. F. (2007). Chromosome study in Schistocerca (Orthoptera-Acrididae-Cyrtacanthacridinae): Karyotypes and distribution patterns of constitutive heterochromatin and nucleolus organizer regions (NORs). Genetics and Molecular Biology, 30(1), 54–59.
Evans, W. L. (1954). Cytology of the grasshopper genus Circotettix. The American Naturalist, 88(838), 21–32.
Gokhman, V. E., & Kuznetsova, V. G. (2006). Comparative insect karyology: Current state and applications. Entomological Review, 86(3), 352–368.
Grzywacz, B., Tasuta, H., Bugrov, A.G. Warchałowska-Śliwa, E. (2019). Genetic markers reveal a reinforcement of variation in the tension zone between chromosome races in the brachypterous grasshopper Podisma sapporensis Shir. on Hokkaido Island. Scientific Reports, 9, 16860.
Gusachenko, A. M., Warchałowska-Śliwa, E., Maryańska-Nadachowska, A., Bugrov, A. G., & Vystotskaja, L. V. (1992). Cytogenetic analysis of populations of Chorthippus albomarginatus. DE GEER/Acrididae: Orthoptera. Folia Biologica, 1(40), 27–31.
Henry, L., Wickham, H., & Chang, W. (2020). ggstance: Horizontal “ggplot2” Components. https://CRAN.R-project.org/package=ggstance
Hewitt, G. M. (1979). Animal Cytogenetics. Volume 3. Insecta 1: Orthoptera, Grasshoppers and cCrickets. Animal Cytogenetics. Volume 3. Insecta 1: Orthoptera, Grasshoppers and Crickets.
Husemann, M., Namkung, S., Habel, J. C., Danley, P. D., & Hochkirch, A. (2012). Phylogenetic analyses of band-winged grasshoppers (Orthoptera, Acrididae, Oedipodinae) reveal convergence of wing morphology. Zoologica Scripta, 41(5), 515–526. https://doi.org/10.1111/j.1463-6409.2012.00548.x
Husemann, M., Sadílek, D., Dey, L.-S., Hawlitschek, O., & Seidel, M. (2021). New genome size estimates for band-winged and slant-faced grasshoppers (Orthoptera: Acrididae: Oedipodinae, Gomphocerinae) reveal the so far largest measured insect genome. Caryologia, 73, 111–120.
John, B., & Hewitt, G. M. (1966). Karyotype stability and DNA variability in the Acrididae. Chromosoma, 20(2), 155–172.
John, B., Lightfoot, D. C., & Weissman, D. B. (1983). The meiotic behaviour of natural F1 hybrids between Trimerotropis suffusa Scudder and T. cyaneipennis Bruner (Orthoptera: Oedipodinae). Canadian Journal of Genetics and Cytology, 25(5), 467–477.
John, B., & Weissman, D. B. (1977). Cytogenetic components of reproductive isolation in Trimerotropis thalassica and T. occidentalis. Chromosoma, 60(2), 187–203.
Kassambara, A. (2020). ggpubr: “ggplot2” Based Publication Ready Plots. https://rpkgs.datanovia.com/ggpubr/
Kiknadze, I. I., & Vyotskaya, L. V. (1970). Measurements of DNA mass per nucleus in the grasshopper species with different numbers of chromosomes (in Russian). Tsitologia, 12, 1100–1108.
King, M. (1995). Species evolution: The role of chromosome change. Cambridge University Press.
Kirkpatrick, M. (2010). How and why chromosome inversions evolve. PLoS Biology, 8(9), e1000501.
Lightfoot, D. C., Weissman, D. B., & Ueshima, N. (2011). Phymonotus jacintotopos: A new genus and species of shield-backed katydid (Orthoptera: Tettigoniidae: Tettigoniinae: Nedubini) from the San Jacinto Mountains of California, USA. Zootaxa, 2937(1), 49–65.
Mao, Y., Zhang, N., Nie, Y., Zhang, X., Li, X., & Huang, Y. (2020). Genome size of 17 species from Caelifera (Orthoptera) and determination of internal standards with very large genome Size in insecta. Frontiers in Physiology, 11, 1321.
McClung, C. E. (1917). The multiple chromosomes of Hesperotettix and Mermiria (Orthoptera). Journal of Morphology, 29(2), 519–605.
Mesa, A., Ferreira, A., & Carbonell, C. S. (1982). Cariologia De Los Acridoideos Neotropicales: Estado Actual De Su Conocimiento y Nuevas Contribuciones, 18, 507–526.
Morgan, M. (2019). BiocManager: Access the bioconductor project package repository. R Package Version, 1(30), 10.
Navarro, A., & Barton, N. H. (2003). Chromosomal speciation and molecular divergence–accelerated evolution in rearranged chromosomes. Science, 300(5617), 321–324.
Palacios-Gimenez, O. M., Castillo, E. R., Martí, D. A., & Cabral-de-Mello, D. C. (2013). Tracking the evolution of sex chromosome systems in Melanoplinae grasshoppers through chromosomal mapping of repetitive DNA sequences. BMC Evolutionary Biology, 13(1), 1–12.
Palacios-Gimenez, O. M., Dias, G. B., De Lima, L. G., Ramos, É., Martins, C., & Cabral-de-Mello, D. C. (2017). High-throughput analysis of the satellitome revealed enormous diversity of satellite DNAs in the neo-Y chromosome of the cricket Eneoptera surinamensis. Scientific Reports, 7(1), 1–11.
Palacios-Gimenez, O. M., Milani, D., Lemos, B., Castillo, E. R., Martí, D. A., Ramos, E., et al. (2018). Uncovering the evolutionary history of neo-XY sex chromosomes in the grasshopper Ronderosia bergii (Orthoptera, Melanoplinae) through satellite DNA analysis. BMC Evolutionary Biology, 18(1), 1–10.
R Core Team. (2020). R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. https://www.R-project.org/
Rehn, J. A. (1921). Descriptions of new and critical notes upon previously known forms of North American Oedipodinae (Orthoptera; Acrididae): Second paper. Transactions of the American Entomological Society (1890-), 47(3), 171–197.
Rentz, D. F., & Weissman, D. B. (1984). Five new species of the bandwinged grasshopper genus Trimerotropis Stål (Orthoptera: Oedipodinae). Pan-Pacific Entomologist, 60(3), 227–237.
Revell, L. J. (2012). phytools: An R package for phylogenetic comparative biology (and other things). Methods in Ecology and Evolution, 3(2), 217–223.
Riva, E., Fox, D. P., Giraldez, R., & Santos, J. L. (1984). Chiasma frequency and distribution in the presence and absence of supernumerary chromosome segments in the grasshopper, Euchorthippus pulvinatus gallicus. Heredity, 53(1), 101–106.
Shaw, D. D., Lightfoot, D. C., & Weissman, D. B. (1998). Chromosomes as isolating mechanisms in Trimerotropine grasshoppers. Journal of Orthoptera Research, 157–163.
Song, H., Mariño-Pérez, R., Woller, D. A., & Cigliano, M. M. (2018). Evolution, diversification, and biogeography of grasshoppers (Orthoptera: Acrididae). Insect Systematics and Diversity, 2(4), 3.
Stávale, L. M., Schneider, M. C., Brescovit, A. D., & Cella, D. M. (2011). Chromosomal characteristics and karyotype evolution of Oxyopidae spiders (Araneae, Entelegynae). Genetics and Molecular Research, 752–763.
Sylvester, T., Hjelmen, C. E., Hanrahan, S. J., Lenhart, P. A., Johnston, J. S., & Blackmon, H. (2020). Lineage-specific patterns of chromosome evolution are the rule not the exception in Polyneoptera insects. Proceedings of the Royal Society B, 287(1935), 20201388.
The Mesquite Project Team. (2019). Documentation for Mesquite: a modular system for evolutionary analysis. http://www.mesquiteproject.org
Türkoglu, Ş., & Koca, S. (2002). Chromosomes of Oedipoda schochi schochi and Acrotylus insbricus (Orthoptera, Acrididae, Oedipodinae). Karyotypes and C-and G-band patterns. Turkish Journal of Zoology, 26(3), 327–332.
Ueshima, N., & Rentz, D. F. (1979). Chromosome systems in the North American Decticinae with reference to Robertsonian changes (Orthoptera: Tettigoniidae). Cytologia, 44(3), 693–714.
Vandergast, A. G., Weissman, D. B., Wood, D. A., Rentz, D. C., Bazelet, C. S., & Ueshima, N. (2017). Tackling an intractable problem: Can greater taxon sampling help resolve relationships within the Stenopelmatoidea (Orthoptera: Ensifera)? Zootaxa, 4291(1), 1–33.
Vysotskaya, L. V. (1993). Cytological study of grasshoppers belonging to the family Acrididae (Orthoptera) (in Russian). Ph.D. Thesis, Institute Cytology and Genetics, Siberian Branch of the USSR Academy of Sciences. Novosibirsk.
Vysotskaya, L. V., & Bugrov, A. G. (1987). Comparative karyological analysis of the tribe Bryodemini (Orthoptera: Acrididae: Oedipodinae) in the fauna USSR (in Russian). Zoological Journal, 66, 1189–1195.
Warchałowska-Śliwa, E. (1998). Karyotype characteristics of katydid orthopterans (Ensifera, Tettigoniidae) and remarks on their evolution at different taxonomic levels. Folia Biologica (Kraków), 46, 143–176.
Warchałowska-Śliwa, E., Tatsuta, H., Akimoto, S.-I., Maryańska-Nadachowska, A., Kowalczyk, M., & Bugrov, A. (2008). Geographical patterns of chromosomal differentiation in the brachypterous grasshopper Podisma sapporensis (Orthoptera: Acrididae). European Journal of Entomology, 105(2), 185–196.
Weissman, D. B., & Rentz, D. C. F. (1980). Cytological, morphological, and crepitational characteristics of the trimerotropine (Aerochoreutes, Circotettix, and Trimerotropis) grasshoppers (Orthoptera; Oedipodinae). Transactions of the American Entomological Society, 253–272.
White, G. A., & Solt, M. (1978). Chromosome numbers in Crambe, Crambella, and Hemicrambe 1. Crop Science, 18(1), 160–161.
White, M. J. D. (1949). A cytological survey of wild populations of Trimerotropis and Circotettix (Orthoptera, Acrididae). I. The chromosomes of twelve species. Genetics, 34(5), 537–563.
White, M. J. D. (1951). Cytogenetics of orthopteroid insects. Advances in Genetics, 4, 267–330.
White, M. J. D. (1968). Karyotypes and nuclear size in the spermatogenesis of grasshoppers belonging to the subfamilies Gomphomastacinae, Chininae and Biroellinae (Orthoptera, Eumastacidae). Caryologia, 21(2), 167–179.
White, M. J. D. (1973). Animal Cytology and Evolution (3rd ed.). Cambridge University Press.
Wickham, H. (2016). ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York. https://ggplot2.tidyverse.org
Wickham, H., Averick, M., Bryan, J., Chang, W., McGowan, L. D., François, R., et al. (2019). Welcome to the Tidyverse. Journal of Open Source Software, 4(43), 1686.
Wickham, H., Henry, L., & Vaughan, D. (2020). vctrs: Vector Helpers. https://CRAN.R-project.org/package=vctrs
Wickham, H., & Pedersen, T. L. (2019). gtable: Arrange “Grobs” in Tables. https://CRAN.R-project.org/package=gtable
Wickham, H., & Seidel, D. (2020). Scales: Scale functions for visualization. R Package Version, 1(1), 1.
Wilke, C. (2019). cowplot: Streamlined Plot Theme and Plot Annotations for “ggplot2.” https://CRAN.R-project.org/package=cowplot
Yu, G., Lam, T. T. Y., Zhu, H., & Guan, Y. (2018). Two methods for mapping and visualizing associated data on phylogeny using ggtree. Molecular Biology and Evolution, 35(12), 3041–3043.
Yu, G., Smith, D. K., Zhu, H., Guan, Y., & Lam, T. T. Y. (2017). ggtree: An R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods in Ecology and Evolution, 8(1), 28–36.
Zhong, X. B., de Jong, J. H., & Zabel, P. (1996). Preparation of tomato meiotic pachytene and mitotic metaphase chromosomes suitable for fluorescence in situ hybridization (FISH). Chromosome Research, 4(1), 24–28.
Acknowledgements
We thank Dr. Axel Hochkirch (University of Trier) for providing samples from the Canary Islands. Many thanks to Michael G. Sergeev who provided us with the literature. Thanks to two anonymous reviewers for helpful and insightful comments on a previous version of the manuscript. We further thank the Orthopterists’ Society for grants to MH and LSD, personal funding from Heinrich-Böll-Stiftung to LSD and Dr. Elisabeth Hintelmann for further funding. This work benefited from expertise sharing and discussions within the DFG priority program SPP 1991.
Funding
Open Access funding enabled and organized by Projekt DEAL. Orthopterists’ Society grant to MH and LSD; Hintelmann prize to MH; personal funding of Heinrich Böll Stiftung to LSD.
Author information
Authors and Affiliations
Contributions
Martin Husemann took part in conceptualization and supervision; Martin Husemann, Lara-Sophie Dey, David Sadílek, Norihiro Ueshima and David Weissman were involved in methodology; Martin Husemann, Lara-Sophie Dey and Norihiro Ueshima carried out formal analysis and investigation, and acquired the funding and resources; Martin Husemann and Lara-Sophie Dey wrote and prepared the original draft; Martin Husemann, Lara-Sophie Dey, David Sadílek, Norihiro Ueshima, Oliver Hawlitschek, Hojun Song and David Weissman performed writing, review and editing; and Lara-Sophie Dey designed the layout.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors declare their consent to publication of the scientific article.
Conflicts of interest
The authors declare no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Husemann, M., Dey, LS., Sadílek, D. et al. Evolution of chromosome number in grasshoppers (Orthoptera: Caelifera: Acrididae). Org Divers Evol 22, 649–657 (2022). https://doi.org/10.1007/s13127-022-00543-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-022-00543-1