Abstract
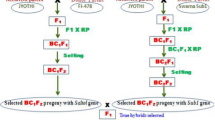
Salt stress causes considerable damage to rice with a consequent reduction in grain yield, however, conventional breeding for this stress is time-consuming and costly. Recently, marker-assisted breeding has shown enormous potential to accelerate breeding of stress tolerant varieties because of its precision, time saving, and cost effectiveness. The present study was carried out to transfer Saltol, a major QTL on chromosome 1 associated with salinity tolerance, from FL478, a tolerant genotype, into IR64, a popular lowland variety through marker-assisted backcrossing (MABC). This technique considerably enhanced the recovery rate of the recurrent parent genome within three backcross generations, which could have saved several backcrosses compared with conventional schemes to achieve the same results. By using this technique, up to 99.7% of the recurrent parent genome was recovered at BC3F2 generation, saving at least three backcrosses compared with conventional breeding schemes. Salinity tolerance of IR64-Saltol lines was evaluated using saline culture solution adjusted to electrical conductivity of 12 dS m-1 using NaCl. Based on selected physiological and growth parameters, the new Saltol introgression lines showed a significantly higher tolerance of salinity than their recurrent parent IR64. The results of this study confirm the benefits of using molecular markers in plant breeding to enhance tolerance of abiotic stresses.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Babu RJ, Weir BS, Zeng ZB. 2004. Intergrating marker-assisted selection in crop breeding-prospects and challenges. Curr. Sci. 87(5): 607–619
Bingpong IK, Manneh B, Sock M, Diaw F, Amoah NKA, Ismail AM, Gregorio G, Singh RK, Wopereis M. 2016. Improving salt tolerance of lowland rice cultivar “Rassi” through maker-added backcross breeding in West Africa. Plant Sci. 242: 288–299
Bo S, Zhang K, Wei-Min D, Ye L, Li-Quing F, Jia-Ming D, Kang-Le Z. 2007. QTL mapping of Chlorophyll contents in rice. Agr. Sci. China 6(1): 17–24
Bohra JS, Dorffling K. 1993. Potassium nutrition of rice (Oryza sativa L.) varieties under NaCl salinity. Plant Soil 152: 299–303
Castillo EG, Tuong TP, Ismail AM, Inubushi K. 2007. Response to salinity in rice: comparative effects of osmotic and ionic tresses. Plant Prod. Sci. 10(2): 159–170
Chen S, Xu CG, Lin XH, Zhang Q. 2001. Improving bacterial blight resistance of ‘6078’, an elite restorer line of hybrid rice, by molecular marker-assisted selection. Plant Breed. 120: 133–137
Collard BCY, Jahufer MZZ, Brouwer JB, Pang ECK. 2005. An introduction to markers, quantitative trail loci (QTL) mapping and marker-assisted selection for crop improvement: the basis concept. Euphytica 142: 169–196
Collard BCY, Mackill DJ. 2008. Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Phil. Trans. R. Soc. B 363: 557–572
Elahi CM, Serai F, Rasul NM, Das KC, Biswas K, Salam MA, Gomosta AR, Tumingbang E, Adorada D, Gregorio G, et al. 2004. Breeding rice for salinity tolerance using the Pokkali allele: finding a linked DNA marker. In: AS Islam AS, ed., In vitro Culture, Transformation and Molecular markers for Crop improvement. Science Publisher. Inc. USA, 157–170
Eshed Y, Zamir D. 1995. An introgression line population of Lycopersicon pennellii in the cultivated tomato enables the identification and fine mapping of yield-associated QTL. Genetics 141: 1147–1162
Flowers TJ, Yeo AR. 1981. Variability in the resistance of sodium chloride salinity within rice (Oryza sativa L.) varieties. New Phytol. 81: 363–373
Frisch M, Bohn M, Melchinger AE. 1999. Minimum sample size and optimal positioning of flanking markers in marker-assisted backcrossing for transfer of a target gene. Crop Sci. 39: 967–975
Gregorio GB. 1997. Tagging salinity tolerance genes in rice using amplified fragment length polymorphism (AFLP). Ph.D. thesis, University of the Philippines, Los Baños. pp 118
Gregorio GB, Senadhira D, Mendoza RD, Manigbas RD, Rozas NL, Guerta CQ. 2002. Progress in breeding for salinity and associated abiotic stress in rice. Field Crops Res. 76: 91–101
Haq TU, Akhtar J, Gorham J, Steele KA, Khalid M. 2008. Genetic mapping of QTLs, controlling shoot fresh and dry weight under salt stress in rice (Oryza sativa L.) cross between Co39 x Moroberekan. Pakistan. J. Bot. 40(6): 2369–2381
Hospital F, Chacosset A. 1997. Marker-assisted introgression of quantitative trait loci. Genetics 147: 1469–1485
IRRI. 1996. Standard Evaluation System Manual. International Rice Research Institute, Los Banos, Philippines
Ismail AM, Heuer S, Thomson MJ, Wissuwa M. 2007. Genetic and genomic approaches to develop rice germplasm for problem soils. Plant Mol. Biol. 65: 547–570
Koyama ML, Levesley A, Koebner RMD, Flowers TJ, Yeo AR. 2001. Quantitative trait loci for component physiological traits determining salt tolerance in rice. Plant Physiol. 125: 406–422
Le HL, Ta HL, Tran DX, Le HH, Ismail AM, Tran DK. 2012. Molecular breeding to improve salt tolerance of rice (Oryza sativa L.) in the Red River Delta of Vietnam. Int. J. Plant Genet. doi:10.1155/2012/949038
Lee KS, Choi WY, Ko JC, Kim TS, Gregorio GB. 2003. Salinity tolerance of japonica and indica rice (Oryza sativa L.) at the seedling stage. Planta 216: 1043–1046
Liu J, Liu D, Tao W, Li W, Wang S, Chen P, Cheng S, GAO D. 2000. Molecular marker-facilitated pyramiding of different genes for powdery mildew resistance in wheat. Plant Breed. 119: 21–24
Luu T.N.H, Luu MC, Ismail AM, Le HH. 2012. Introgression the salinity tolerance QTLs Saltol into AS996, the Elite rice variety of Vietnam. Am. J. Plant Sci. 3: 981–987
Masood MS, Seiji Y, Shinwari ZK, Anwar R. 2004. Mapping quantitative trait loci (QTLs) for salt tolerance in rice (Oryza sativa) using RFLPs. Parkistan. J. Bot. 36(4): 825–834
McCouch SR, Teylelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L. 2002. Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res. 9(6): 199–207
Monna L, Lin HX, Kojima S, Sasaki T. 2002. Genetic dissection of a genomic region for a quantitative trait locus, Hd3, into two loci, Hd3a and Hd3b, controlling heading date in rice. Theor. Appl. Genet. 104: 772–778
Moradi F, Ismail AM. 2007. Responses of photosynthesis, chlorophyll fluorescence and ROS scavenging system to salt stress during seedling and reproductive stages in rice. Ann. Bot. 99: 1161–1173
Moradi F, Ismail AM, Gregorio G, Egdane J. 2003. Salinity tolerance of rice during reproductive development and association with tolerance at seedling stage. Indian J. Plant Physiol. 8:105–116
Neeraja CN, Raghirang-Rodriguez R, Pamplona A, Heuer S, Collard BC Y, Septiningsih EM, Vergara G, Sanchez D, Xu K, Ismail AM, Mackill DJ. 2007. A marker-assisted backcross approach for developing submergence-tolerant rice cultivars. Theor. Appli. Genet. 115: 767–776
Platten JD, Egdane J, Ismail AM. 2013. Salinity tolerance, Na+ exclusion and allele mining of HKT1;5 in Oryza sativa and O. glaberrima: many sources, many genes, one mechanism? BMC Plant Biol. 13: 32. http://www.biomedcentral.com/1471-2229/13/32
Rahman MA, Thomson MJ, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail AM. 2016. Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization. Ann. Bot. 117:1083–1097
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX. 2005. A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat. Genet. 37: 1141–1146
Semagn K, Bjornstad A, Ndjiondjop MN. 2006. Progress and prospects of marker assisted backcrossing as a tool in crop breeding programs. African J. Biotech. 5(25): 2588–2603
Siangliw JL, Jongdee B, Pantuwan G, Toojinda T. 2007. Developing KDML105 backcross introgression lines using marker-assisted selection for QTLs associated with drought tolerant in rice. Sci. Asia 33: 207–214
Singh RK, Gregorio GB, Jain RK. 2007. QTL mapping for salinity tolerance in rice. Physiol. Mol. Biol. Plants. 13(2): 87–99
Thomson MJ, de Ocampo M, Egdane J, Katimbang M, Rahman MA, Singh RK, Gregorio GB, Ismail AM. 2007. QTL Mapping and Marker-assisted Backcrossing for Improved Salinity Tolerance in Rice. BioAsia (www.bioasia-2007.com), Supplement Papers: 6–12
Thomson MJ, de Ocampo M, Egdane J, Rahman MA, Sajie AG, Adorada DL, Tuminbang-Raiz E, Blumwald Eduardo, Seraj ZI, Singh RK, Gregorio GB, Ismail AM. 2010b. Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 3: 148. doi:10.1007/s12284-010-9053-8
Thomson MJ, Edwards JD, Septiningsih EM, Harringon SE, McCouch SR. 2006. Substitution mapping of dth 1.1, a flowering-time quantitative trait locus (QTL) associated with transgressive variation in rice, reveals multiple sub-QTL. Genetics 172: 2501–2514
Thomson MJ, Ismail AM, McCouch SR, Mackill DJ. 2010a. Marker-Assisted Breeding. In A Pareek, SK Sopory, HJ Bohnert, Govindjee, eds., Abiotic stress Adaptation in Plants: Physiological, Molecular and Genomic Foundation, Springer Science, 451–469
Thomson MJ, Tai TH, McClung AM, Lai XH, Hinga M, Lobos KB, Xu Y, Martinez CP, McCouch SR. 2003. Mapping quantitative trait loci for yield, yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor. Appl. Genet. 107: 479–493
Vanaja T, Neema VP, Mammootty KP, Balakrishnan PC, Jayaprakash NB. 2015. The first high yielding saline tolerant rice variety suited to the Kaipad tidal farming ecosystem of Kerala, India, and suited for flood prone and water-scarce environments: “Ezhome-1”. J. Organics 2: 21–32
Xu K and Mackill DJ. 1996. A major locus for submergence tolerance mapped on rice chromosome 9. Mol. Breed. 2: 219–224
Yeo AR, Yeo ME, Flowers SA, Flowers TJ. 1990. Screening of rice (Oryza sativa L.) genotypes for physiological characters contributing to salinity resistance, and their relationship to overall performance. Theor. Appl. Genet. 79: 377–384
Yoshida S, Forno D, Cock J, Gomez K. 1976. Laboratory manual for Physiological studies of Rice. International Rice Research Institute, Manila, Philippines
Zhou PH, Tan YF, He YQ, Xu CG, Zhang Q. 2003. Simultaneous improvement for four quality traits of Zhensha 97, an elite parent of hybrid rice, by molecular marker-assisted selection. Theor. Appl. Genet. 106: 326–331
Zhu JK. 2007. Plant salt stress. Encyclopedia of Life Science, John Wiley & Son
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ho, V.T., Thomson, M.J. & Ismail, A.M. Development of salt tolerant IR64 near isogenic lines through marker-assisted breeding. J. Crop Sci. Biotechnol. 19, 373–381 (2016). https://doi.org/10.1007/s12892-016-0049-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12892-016-0049-9