Abstract
The ability to form biofilms in natural isolate Bacillus subtilis 168 and mutants with deleted genes of regulatory proteins AbrB, DegU, CcpA, and SpoOA, constructed on its basis, was investigated to elucidate the pathways regulating biofilm formation in B. subtilis. The B. subtilis 168 wild-type forms a biofilms in the liquid medium with maximum at 48th hour of culture growth. pH optimum for the biofilm formation in the wild-type strain is in the range of 7.4–8.0. Temperature optimum was in the range of 22 to 45 °C. The level of biofilm formation for all regulatory mutants was lower than that in the wild-type for 40–50 %. Temperature and pH optima for the mutant strains are the same as for the wild-type strain—7.4–8 pH and temperature of 22–45 °C.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Biofilms are communities of surface-associated microorganisms encased in a self-produced extracellular matrix. Biofilm formation is a nearly universal bacterial trait, and biofilms are found on almost all natural and artificial surfaces [1]. Biofilm-associated infections are generally hard to treat because of the ability of biofilm-encased bacteria to resist a wide variety of external insults, including antibiotic treatments [2, 3]. Biofilms are associated with various human diseases, such as endocarditis valve, cystic fibrosis, periodontitis, otitis media, biliary tract infection, etc., and colonize medical devices and implants, causing recurrent infections [4]. Exopolysaccharide biofilm structure is common to all species of bacteria capable of their formation [5]. This suggests a common genetic mechanisms regulating the formation of biofilms. In this connection, of particular interest is the identification of global regulatory systems involved in the process of creating a biofilm structure. Bacillus subtilis is a motile, Gram-positive bacterium widely used in studies of biofilm formation as a model system [6–8]. To elucidate the pathways regulating biofilm formation of B. subtilis, we investigated the natural isolate of B. subtilis 168, and constructed on the basis of its mutant strains with knockout genes of regulatory proteins: AbrB (global transcriptional regulator), DegU (two-component response regulator of signal transduction system DegS-DegU), CcpA (regulator of carbon catabolism), and SpoOA (protein regulator of sporulation). SpoOA is a central transcriptional regulator that controls the expression of over 100 genes, including those necessary for biofilm matrix gene expression and sporulation [9]. Intermediate levels of phosphorylated form SpoOA ∼ P result in matrix gene expression, and higher levels induce sporulation genes. In this way, when SpoOA is initially phosphorylated, biofilm formation is induced as a result of matrix gene expression. As the biofilm matures, SpoOA ∼ P accumulates in certain cells and activates sporulation [8, 10]. Phosphorylated form of SpoOA also represses a biofilm matrix gene repressor—AbrB [11]. AbrB directly binds to DNA to repress transcription from promoters involved in plethora of cellular processes including those needed for biofilm formation [12, 13]. Their combined presence within the cell provides fine tune to the regulation of biofilm formation and to ensure the coordinated expression of all of the matrix genes [14]. DegU is a global regulator in B. subtilis that is involved in the control of a variety of cellular processes such as competence, motility, and secretion of degradative enzymes [15]. There are evidences that the degU mutant is defective in submerged biofilm formation [16], and colony biofilm formation is also defective in a DegU mutant due to the loss of the surface hydrophobicity protein [12, 17]. CcpA was a regulatory protein of catabolic control [18]. It is involved in regulating the expression of many genes and operons, including some transport genes, and catabolic and anabolic genes of carbon, nitrogen, and phosphate metabolism [19].
2 Materials and Methods
The strains used in the study are shown in the table.
Strains | Mutation description | Source |
|---|---|---|
Bacillus subtilis 168 | Natural isolate (wild-type) | Professor J. Stuelke, University of Göttingen, Germany |
Bacillus subtilis 168 abrB | abrB gene knockout (global transcriptional regulator) | Bacillus Genetic Stock Center (BGSC) Professor D. Zeigler, Ohio University, USA |
Bacillus subtilis 168 degU | degU gene knockout (two-component response regulator of signal transduction system DegU-DegU) | Dr. Jan Maarten van Dijl, University of Groningen, The Netherlands |
Bacillus subtilis 168 ccpA | ccpA gene knockout (global regulator of carbohydrate metabolism) | Bacillus Genetic Stock Center (BGSC) Professor D. Zeigler, Ohio University, USA |
Bacillus subtilis 168 spoOA | spoOA gene knockout (global regulator of sporulation) | Bacillus Genetic Stock Center (BGSC) Professor D. Zeigler, Ohio University, USA |
Cultivation of the strains was performed on a synthetic-E medium, whose composition is described in [20]. Seed served 16-h inoculum (1 % v/v). Bacterial growth was monitored by the change in the optical density of the culture at 600 nm. Biomass was expressed in absorbance units. Spore formation was determined by counting cells and spores by Peshkov microscopy method mode microscope Carl Zeiss Jena (Germany) in 1600 times magnification in four visual fields. The amount of free spores was expressed as a percentage of the total number of vegetative and sporulating cells.
Biofilm formation defined by the method set incubation with crystal violet (CV) [21] with modification [22]. For the analysis of experimental data, Microsoft Excel program was used. Using data of the four independent experiments, we described and compared attributes. The results were considered statistically significant at the standard deviation of σ ≤ 10 %.
3 Results and Discussion
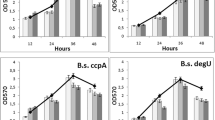
B. subtilis 168 is a genetically unmodified natural isolate. We investigated the dynamics of its growth and sporulation (Fig. 1). In liquid medium (37 °C, pH 7.4), culture enters the stationary phase of growth for 42–44 h. By this time, the amount of free spores in the culture is about 10–15 %.
Also, we investigated the dynamics of the biofilm formation of B. subtilis 168 on a liquid medium (37 °C, pH 7.4) (Fig. 2). Maximum formation of biofilm structures was observed at the 48-h culture growth.
To explore ways of regulating biofilm formation of B. subtilis 168, we used mutant strains with knockout genes of regulatory proteins AbrB, DegU, CcpA, and SpoOA.
We determined the difference in the levels of wild and mutant strains in biofilm formation in the spectrum of different pH (Fig. 3) and temperature (Fig. 4).
The medium pH ranging 5 to 9.5 for optimal biofilm formation for all strains ranged from 7.4 to 8.0. Under these pH levels, biofilm wild strain was by an average of 50 % higher than the level of each of the mutants (Fig. 3).
In the temperature ranging from 4 to 50 °C for optimal biofilm formation for all strains ranged from 22 to 45 °C. Its biofilm layer within a wild strain of an average of 40–50 % was higher than the level of each of the mutants (Fig. 4)
In the absence of each of the test levels of regulatory proteins, biofilm formation decreased by almost half with respect to a wild strain with a full set of genes. The optimal pH and temperature ranges remain unchanged. This means that the global regulation systems play a role in the formation of biofilms. However, none of the systems is not critical for the formation of biofilms.
4 Conclusions
Natural isolate of B. subtilis 168 in the liquid medium forms a biofilm with a maximum at 48 h of culture growth. Optimum pH for the formation of biofilms wild strain is in the range of 7.4–8.0. Temperature optimum is in the range of 22 to 45 °C. This corresponds to the natural conditions of the habitat of B. subtilis in the rhizosphere. The level of biofilm formation regulatory mutants in genes abrB, degU, ccpA, and spoOA was on average of 40–50 % lower than the wild-type level; thus, the global regulatory system controls the formation of biofilms, while none of the systems does not affect the process critically. Temperature and pH optima for the mutant strains are the same as for the wild-type strain—7.4–8 pH and temperature of 22–45 °C.
References
Stewart, P. S., & Franklin, M. J. (2008). Physiological heterogeneity in biofilms. Nature Reviews Microbiology, 6, 199–210.
Anderson, G. G., & O’Toole, G. A. (2008). Innate and induced resistance mechanisms of bacterial biofilms. Current Topics in Microbiology and Immunology, 322, 85–105.
Bryers, J. D. (2008). Medical biofilms. Biotechnology and Bioengineering, 100, 1–18.
Hancock, V., & Klemm, P. (2007). Global gene expression profiling of asymptomatic bacteriuria Escherichia coli during biofilm growth in human urine. Infection and Immunity, 75(2), 966–976.
Donlan, R. M. (2002). Biofilms: microbial life on surfaces. Emerging Infectious Diseases, 8, 881–890.
Branda, S. S., Gonzalez-Pastor, J. E., Ben-Yehuda, S., Losick, R., Kolter, R. (2001). Fruiting body formation by Bacillus subtilis. Proceedings of the National Academy of Sciences of the United States of America, 98, 11621–11626.
Hamon, M. A., & Lazazzera, B. A. (2001). The sporulation transcription factor Spo0A is required for biofilm development in Bacillus subtilis. Molecular Microbiology, 42, 1199–1209.
Vlamakis, H., Chai, Y., Beauregard, P., Losick, R., Kolter, R. (2013). Sticking together: building a biofilm the Bacillus subtilis way. Nature Reviews Microbiology, 11(3), 157–168.
Fujita, M., Gonzalez-Pastor, J. E., Losick, R. (2005). High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis. Journal of Bacteriology, 187, 1357–1368.
Chastanet, A., Vitkup, D., Yuan, G.-C., Norman, T. M., Liu, J. S., Losick, R. M. (2010). Broadly heterogeneous activation of the master regulator for sporulation in Bacillus subtilis. PNAS, 107(18), 8486–8491.
Strauch, M., Webb, V., Spiegelman, G., Hoch, J. A. (1990). The SpoOA protein of Bacillus subtilis is a repressor of the abrB gene. Proceedings of the National Academy of Sciences of the United States of America, 87, 1801–1805.
Verhamme, D. T., Murray, E. J., Stanley-Wall, N. R. (2009). DegU and Spo0A jointly control transcription of two loci required for complex colony development by Bacillus subtilis. Journal of Bacteriology, 191, 100–108.
Chumsakul, O., Takahashi, H., Oshima, T., Hishimoto, T., Kanaya, S., Ogasawara, N., et al. (2011). Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation. Nucleic Acids Research, 39, 414–428.
Chu, F., Kearns, D. B., McLoon, A., Chai, Y., Kolter, R., Losick, R. (2008). A novel regulatory protein governing biofilm formation in Bacillus subtilis. Molecular Microbiology, 68, 1117–1127.
Murray, E. J., Kiley, T. B., Stanley-Wall, N. R. (2009). A pivotal role for the response regulator DegU in controlling multicellular behaviour. Microbiology, 155, 1–8.
Stanley, N. R., & Lazazzera, B. A. (2005). Defining the genetic differences between wild and domestic strains of Bacillus subtilis that affect poly-gamma-dl-glutamic acid production and biofilm formation. Molecular Microbiology, 57, 1143–1158.
Kobayashi, K., & Iwano, M. (2012). BslA(YuaB) forms a hydrophobic layer on the surface of Bacillus subtilis biofilms. Molecular Microbiology, 85, 51–66.
Miwa, Y., Nakata, A., Ogiwara, A., Yamamoto, M., Fujita, Y. (2000). Evaluation and characterization of catabolite-responsive elements (cre) of Bacillus subtilis. Nucleic Acids Research, 28, 1206–1210.
Fujita, Y. (2009). Carbon catabolite control of the metabolic network in Bacillus subtilis Biosci. Biotechnology and biochemistry, 74(2), 245–259.
Morikawa, M., Kagihiro, S., Haruki, M., Takano, K., Branda, S., Kolter, R., et al. (2006). Biofilm formation by a Bacillus subtilis strain that produces γ-polyglutamate. Microbiology, 152, 2801–2807.
O’Toole, G. A., Pratt, L. A., Watnick, P. I., Newman, D. K., Weaver, V. B., Kolter, R. (1999). Genetic approaches to study of biofilms. Methods in Enzymology, 310, 91–109.
Merritt, J.H., Kadouri, D.E., O’Toole, G.A. (2005) Growing and analyzing static biofilms. Curr Protoc Microbiol. Chap.1:Unit 1B.1. doi:10.1002/9780471729259.mc01b01s00.
Acknowledgments
This work was supported by the subsidy allocated to the Kazan Federal University for the state assignment in the sphere of scientific activities. This work was performed in accordance with the Russian Government Program of Competitive Growth of the Kazan Federal University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Din, L., Rudakova, N. & Sharipova, M. Factors Influencing the Formation of Biofilms on Bacilli Model Systems. BioNanoSci. 6, 571–574 (2016). https://doi.org/10.1007/s12668-016-0271-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12668-016-0271-4