Abstract
Potato late blight, caused by Phytophthora infestans, has been the most damaging disease in Turkey since 2010. In this study, 127 isolates of P. infestans were obtained from the main growing areas of Turkey between 2015 and 2017. Their phenotypic and genotypic features were revealed and presented with those of reference isolates. These isolates were categorized by their mating type, in vitro mefenoxam sensitivity, mtDNA haplotype, RG57 DNA fingerprinting patterns, simple sequence repeat (SSR) markers, and aggressiveness on a set of potato differential lines. All isolates were of the A2 mating type and mtDNA haplotype Ia, were resistant to mefenoxam, and had RG57 and SSR fingerprints similar to the 13_A2 clonal lineage reported in Europe. This is the first report of 13_A2 in Turkey. Virulence abilities against potato resistance (R) genes R1, R2, R3, R4, R6, R7, R10, and R11 were observed in most of the isolates. The mating type ratios and SSR marker analysis indicate that in Turkey, the sexual reproduction of P. infestans is limited. These results underline that the movement of asexual individuals and the generation of sub-clonal difference are the factors driving the population structure of P. infestans in Turkey.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phytophthora infestans (Mont.) de Bary, an oomycete plant pathogen, is one of the most important diseases of tomato and potato throughout the world. The pathogen has a heterothallic reproductive system and needs the interaction of two opposite mating types, designated as A1 and A2 (Judelson 1997). The late blight disease caused by the pathogen appears to be very severe especially in areas where the temperate climate prevails and there is a need for intensive use of pesticides in its control (Fry 2008). Detailed information about the population structure, diversity, and dynamics of P. infestans is important in terms of the development and updating of management strategies of the disease, as well as the breeding and improvement of resistant cultivars (Forbes 2012).
By the nineteenth century, the late blight caused the famous Great Famine in Ireland with several years of epidemics, which resulted in the deaths of more than one million people and the migration of more than one million people (Brurberg et al. 2011; Kröner et al. 2017). Since then, migration probably through exported seed tubers has enabled this pathogen to spread worldwide (Fry et al. 1993). As the latest result of these movements, in some Northern European countries in the 1980s, pathogen populations underwent quick changes. Many studies on the characterization of P. infestans have shown that new pathogen populations rapidly replace old populations of European populations, and most probably this population consists of individuals who have genetically diverse capabilities from Mexico (Fry et al. 1993). The geographical distribution of P. infestans populations in potato production areas is expressed by clonal lineages defined after an analysis of genotypic and phenotypic characterization (Cooke and Lees 2004). Virulence that originated in Europe, an effective A2 mating type, Ia mtDNA haplotype, and mefenoxam resistance with increased aggressiveness are the characteristics of clonal lineage 13_A2 (Li et al. 2012; Cooke et al. 2012; Kiiker et al. 2018), which was recorded for the first time in Germany and the Netherlands in 2004, followed by Poland in 2006, and affected 80% of the pathogen population in Great Britain in 2008 (Chmielarz et al. 2014; Rekad et al. 2017). Since its first report, this clonal lineage has maintained its presence throughout Europe, and new clonal lineages that have emerged over time have disappeared into the pathogen populations, while 13_A2 has remained. Because of its high sporulation ability and infection even at low temperatures, the 13_A2 clonal lineage replaced the populations of US-1 and other genotypes rather quickly. Indeed, such displacements have been identified in areas of potato production in China, Algeria, and Southern India (Chowdappa et al. 2015; Rekad et al. 2017; Tian et al. 2016).
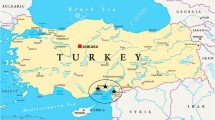
Although the cultivation area and production of potato in Turkey varies from year to year, approximately 4.5 million tons of production is provided from 140.000 ha (TUIK 2018). Plant is mainly cultivated in three regions, categorized by their weather condition and various types of agricultural practices. In central part of Turkey, plants are grown from May until October in large-acreage fields with average Turkey weather. In western Turkey, they are cultivated in intermediate sized fields and grown twice a year. In South-Eastern Turkey, they are planted in small areas in the winter. The control of late blight outbreak in Turkey has become more difficult in a manner similar to observations in Europe and the USA. In Europe and the USA, this situation is based on changes in the population structure of the pathogen (Cooke et al. 2009; Hu et al. 2012; Mariette et al. 2016). Therefore, this study was carried out in Afyon, Bolu, Bursa, Izmir, Nevşehir, Niğde, Tokat, and Trabzon provinces in order to represent the population of the pathogen in the potato production areas of the country.
Climate of major potato-growing regions in Turkey is suitable for P. infestans, and thus, pathogen constitutes a serious threat to production in the country (Gunacti et al. 2019; Tosun et al. 2007). It occurs intensely in cases where disease control measures are not taken and this limits the production of potato to a large extent. While the strategies used to control the disease were successful until 2010 and prevented the emergence of the epidemic character of the disease, after 2010, a period began in which the character of the epidemics of the pathogen was frequently seen. Today, disease control requires more intensive fungicide applications than earlier and possible failures in these applications cause yield losses up to 100%. All these issues mentioned above suggest that there is a change in the population of the pathogen after 2010 and that new clonal strains may have entered the potato production sites. As a matter of fact, the reports of the potato growers’ association indicated that the epidemics and threats caused by the disease increased gradually during this period. P. infestans isolates were collected from potato fields during 2015–2017, and their clonal lineage, mating types, race diversity, and mefenoxam susceptibility were determined in order to better understand the epidemics and their ecology caused by the individuals forming the present pathogen populations. In order to provide timely information to the growers and to create and modify the management strategies of the disease, genotypic properties were tried to be shaped by fast-performing simple sequence repeat (SSR) analyses, and these results were supported by restriction fragment length polymorphism (RFLP) with the RG-57 probe, mtDNA haplotypes, and allozyme assays. The results obtained from the study are expected to contribute to the short- and long-term disease management in Turkey.
Material and methods
Isolate collection and maintenance
Blighted material was obtained from the major producing areas represent 45% of the total cultivation area of the country during the vegetation period of 2015–2017 (Table 1; Fig. 1). The geographical location of the samples varied between growing seasons in relation to the occurrence and development of the disease. The leaves bearing a single lesion were washed with sterile distilled water and dried with the aid of filter paper. These leaves were then placed individually in Petri dishes containing 1.5% water agar for 48 h at 18 °C to stimulate sporulation. After observing sporulation with a dissecting microscope, an agar block of 3 to 4 mm2 was contacted with several sporangia, allowing their passage onto the agar block.
In order to obtain pure culture, these agar blocks were individually placed in rye A agar amended with rifampicin and ampicillin (rye A + rif + amp) and allowed to grow in the dark at 18 °C (Caten and Jinks 1968). Following incubation under a 12-h light/dark cycle for several days, mycelium was moved to fresh medium. The short-term storage of the isolates was performed on rye A agar medium at 15 °C in the dark. For long-term storage, the isolates were incubated in pea agar medium for 2 weeks at 20 °C. For each isolate, 5 discs of 10 mm diameter from these cultures were transferred to cryotubes (2.2 mL) containing 1.5 mL of 15% dimethylsulfoxide (DMSO). The cryotube vials were allowed to pre-freeze at − 80 °C for 24 h and finally transferred rapidly to − 152 °C.
Mating type tests
A 10-mm-diameter agar disc from 2 weeks of culture of the isolate to be tested was placed to one side of a pea agar–containing Petri dish, and a mycelial disc of an A1 tester (US-17 genotype, US970001) or an A2 tester (US-8 genotype, US040009) was transferred on the other side. The isolates were maintained in the dark at 18 °C for 2–3 weeks. After the two mycelial colonies contacted each other, the contact site was regularly checked for oospore formation for seven consecutive days using a microscope at × 100 magnification. If oospores were detected in a Petri dish using the A1 reference isolate, the tested isolate was classified as A2 mating type, and vice versa.
Mefenoxam sensitivity assay
Resistance to mefenoxam was tested on pea agar media amended with mefenoxam (Ridomil Gold EC, 49% active ingredient, Syngenta) at application concentrations of 5 and 100 ppm. During the experiments, isolates were maintained in the dark at 16 °C for 2 weeks. The isolates were grouped as susceptible in cases where the cultures of P. infestans on media with both 5 and 100 ppm mefenoxam were less than 40% of the diameter of the culture on the control Petri dishes without mefenoxam. The isolates were described as intermediate when the growth rate in cultures was > 40% of the control on the media with 5 ppm mefenoxam but less than 40% of the control on the media with 100 ppm mefenoxam. The isolates were defined as resistant when their cultures were bigger than 40% of the control on both types of mefenoxam Petri dishes (Bakonyi et al. 2002). P. infestans isolates used as references were as follows: resistant isolate US-8 (US040009), US-11 (US050007), and two susceptible isolates US-1 (2162 TW97-Pi-42) and US-23 (BL2009P4).
Allozyme analysis
For each isolate, allozyme genotypes were investigated in the glucose-6-phosphate isomerase (Gpi) and peptidase (Pep) loci. Mycelial mass obtained from the pea broth culture or sporangia from sporulated leaves were placed in sterile 1.5-mL microcentrifuge tubes and extracted in accordance with the procedure of Goodwin et al. (1995). Allozyme genotypes were examined by cellulose-acetate electrophoresis for Gpi and Pep loci. Reference isolates of the genotypes US-1 (Gpi 86/100, Pep 92/100) and US-8 (Gpi 100/111/122, Pep100/100) were included as controls in each acetate plate. The distance at which proteins from unknown isolates migrated was determined by comparison with reference isolates.
mtDNA haplotype
The mtDNA haplotype analyses were performed with small modifications in the method of Griffith and Shaw (1998). Two primer sets derived from the genomic DNA library of P. infestans were used: P2 (5′-TTCCCTTTGTCCTCTACCGAT-3′ and 5′-TTACGGCGGTTTAGCACATACA-3′) and P4 (5′-TGGTCATCCAGAGGTTTATGTT-3′ and 5′-CCGATACCGATACCAGCACCAA-3′). The PCR reaction of P2 was performed in a 25-μL reaction mixture containing 2.5 μL of 10× buffer, 250 μM dNTP, 1.875 μM MgCl2, 2.5 μM of each primer, 1 U Taq DNA polymerase, and 10–50 ng of template DNA. The amplification was conducted under the following cycling conditions: after 30 s of initial denaturation at 94 °C, PCR amplification was carried out for 40 cycles (94 °C for 30 s, 64 °C for 60 s, and 72 °C for 60 s and a final extension of 5 min at 72 °C). The products were digested using restriction enzyme MspI, resolved on a 1.5% agarose gel stained with ethidium bromide (1 mg mL−1), and visualized on a UV transilluminator. The reaction of P4 was performed similarly to the P2 reaction, but with 0.7 μM MgCl2. The conditions of the reactions were also similar, but the initial denaturation time was 90 s, primer annealing temperature 60 °C, and extension time 90 s. The PCR products were digested using the restriction enzyme EcoRI and visualized as described for the P2 reaction.
RFLP analysis with probe RG-57
Isolates were incubated in pea broth at 18 °C for 2 weeks. The mycelia were vacuum-filtered with Whatman No. 1 filter papers and lyophilized overnight at − 5 °C. Genomic DNA was individually isolated using Sigma’s GenElute Plant Genomic DNA Extraction Kit, and a RFLP analysis with probe RG-57 was undertaken on 127 isolates using the method specified by Goodwin et al. (1992). Transfer to a positively charged nylon membrane, hybridization with a non-radioactive RG-57 probe, and autoradiography were used in accordance with the manufacturer’s instructions (Roche Applied Science DIG High Prime Labeling and Detection Kit II). The genotype of the isolates was identified using the band patterns of the reference isolates belonging to the clonal lineages US-1, US-8, and US-18 (Forbes et al. 1998). After comparing the obtained data with the profiles of known clonal lineages, genotypic discrimination of the tested isolates was performed (Chowdappa et al. 2013; Danies et al. 2013; Runno-Paurson et al. 2009).
Multiplex microsatellite marker analysis
For the analysis of SSR marker, a total of 12 SSR primers (Pi02, Pi04, Pi4B, Pi63, Pi70, D13, G11, PinfSSR2, PinfSSR4, PinfSSR6, PinfSSR8, and PinfSSR11) were used in this study (Knapova and Gisi 2002; Lees et al. 2006; Li et al. 2010). Three separate multiplex reactions were conducted using three panels of primers. For automated fragment analysis, one primer of each locus was labeled with PET, FAM, NED, and VIC fluorescent dye according to Applied Biosystems™ (ABI). The dyes were assigned to loci in such a way that loci with the same dye had non-overlapping ranges of allele sizes. The PCR reactions were performed in a 10-μL reaction mixture consisting of 1 μL of 10× AmpliTaq (Applied Biosystems), 200 μM dNTP, 1.5 mM MgCl2, 10 pmol of each primer, and 1 U Taq DNA polymerase (Applied Biosystems). PCR amplifications were performed using a T-100 thermocycler (Bio-Rad) under the following conditions: 10 min at 95 °C; 28 cycles of 20 s at 95 °C, 25 s at 58 °C, 60 s at 72 °C; and a final extension of 20 min at 72 °C. Post-PCR processing was undertaken by dispensing 9.5 μL of size standards in high-deionized formamide mix, 0.05 μL of Genescan-500 LIZ size standard (Applied Biosystems [ABI], PN 4322682), 9.45 μL of Hi-Di formamide (ABI, PN 4311320), and 0.5 μL of each PCR product from panel 1–3 reactions into each of the 96-well ABI 3730xl plates. The PCR products were analyzed on an ABI 3730xl capillary system with POP-7 Polymer (ABI, PN 4335615). The PCR amplicons were compared with a set of size standards, and the alleles were scored accordingly (Lees et al. 2006).
Virulence assays
Virulence phenotypes of each of the 127 isolates were evaluated using a detached-leaf assay employing a combination of Black and Mastenbroek’s differential sets for S. demissum R-genes R1–R11 (R1 (Mastenbroek 43154-5), R2 (Black 1512c), R3 (Mastenbroek 4642-1), R4 (Mastenbroek 4431-5), R5 (Black 3053-18), R6 (Black XD2-21), R7 (Black 2182ef(7)), R8 (Black 2424a(5)), R9 (Black 2573), R10 (Black 3618ad(1)), and R11 (Black 5008ab(6))), and potato cv. Bintje as a susceptible control (provided by the Scottish Agricultural Science Agency) (Black et al. 1953; Malcolmson and Black 1966). The leaves were obtained by cultivating the tubers having each resistance gene of the differential set under greenhouse or controlled climate room conditions. The young leaves of which the growth process was completed were collected from the middle part of the canopy of 6–8-week-old plants and used in inoculations. Inoculations were performed in 150-mm-diameter Petri dishes containing 75 mL of water agar (1.2%). After solidification of water agar, Petri dishes were inverted to form a moisture cycle and 5 leaflets were placed (abaxial side up) on the lid of Petri dishes. Sporangial suspensions at 4.0 × 104 sporangia mL−1 concentration were prepared from 10-day-old cultures on rye B agar, and inoculation to all leaflets was performed by giving 20-μL sporangial suspension to both sides of the main vein of the leaflet. After inoculation, Petri dishes were sealed with double-layer parafilm and incubated in a 16-h-light/8-h-dark cycle at 16 °C. The virulence assays were repeated at least twice for each isolate. The effect of pathogen isolates on the potato genotypes constituting the differential set was evaluated using the following scale 7 days after inoculation: 0, no symptoms; 1, small necrotic lesion; 2, < 10% area covered; 3, 10–50% area covered; 4, 50–75% area covered; and 5, > 75% area covered. The reaction was compatible if sporulation was detected in at least six leaflets out of ten, and the cumulative score was at least 15. Compatible interactions were usually indicated by large, sporulating lesions (Runno-Paurson et al. 2009).
Data analysis
Calculation of genetic similarity of SSR patterns was performed using the statistical software R v. 3.6.1 (R Development Core Team 2019), using Jaccard’s index coefficient based on the proportion of shared alleles for all SSR primers. The cluster analysis was evaluated with unweighted pair group method algorithms (UPGMA) using R and Mega X (Kumar et al. 2018).
Results
A total of 127 P. infestans isolates from 35 locations in 17 counties in Turkey during 2015–2017 were obtained from diseased plants (Table 1). Isolates derived from the single sporangium were identified as the allozyme genotypes Gpi 100/100 and Pep 96/96, mating type A2, and mtDNA haplotype la. DNA fingerprint analyses using probe RG57 also found that all of the isolates overlapped with fingerprint characteristic of the Blue 13 clonal lineage.
SSR genotyping analysis using the set of 12 primers (Pi02, Pi4B, PiG11, Pi04, Pi63, Pi70, D13, PinfSSR11, PinfSSR2, PinfSSR4, PinfSSR6, and PinfSSR8) showed no polymorphism in 81 of the 127 isolates of P. infestans from Turkey. The notable exception to this was two uncommon genotypes that exhibited difference in one step at loci Pi02 and PiG11 when compared with the most commonly found genotypes. Minor variations at Pi02 and PiG11 loci were determined in 26 and 46 isolates, respectively, including 20.5% for Pi02 and 36.2% for PiG11 of the entire isolates tested. These 72 isolates were collected from four different provinces (Afyon, Bolu, İzmir, and Niğde). All of the remaining isolates belonged to a single multi-locus genotype. This dominant genotype was heterozygous for almost all of the analyzed loci (Pi02, 266/268/270; Pi4B, 205/213; PiG11, 154/160; Pi04, 166/170; Pi63, 273/279; D13, 134/156; PinfSSR4, 284/294; PinfSSR6, 240/244; and PinfSSR8, 260/266), except for the loci PinfSSR2 (173/173), Pi70 (192/192), and PinfSSR11 (341/341). Minor variants of this genotype were found as 268/270 at Pi02 (26 isolate) and 154/162 at PiG11 (46 isolates) (Table 2). A dendrogram based on cluster analysis of 127 isolates that have the similar simple sequence repeat genotype 13_A2 and 17 reference isolates is shown in Fig. 2. In total, 84 isolates were resistant to mefenoxam (66.1%), and 43 showed an intermediate reaction (33.9%) (Table 1).
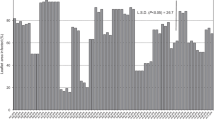
Virulence results obtained from tests showed that there were significant differences among the isolates of P. infestans in Turkey, and this collection of isolates was found to consist of 16 races. The most common race identified in the isolate population was R1.2.3.4.6.7.10.11 (54 isolates), followed by R1.2.3.4.6.7.8.10.11 (34 isolates). R1.3.4.6.7.8.10.11, R1.2.3.4.5.6.7.10.11, and R1.2.3.4.6.7.11 were represented by five isolates each. Four races were represented by a single isolate, and the remaining races by two or four isolates (Table 3). All isolates overcame resistance genes R1, R4, and R7 in the same proportion (88%). Only six isolates collected in 2015 were able to overcome resistance genes R5 and R9, and one isolate was able to overcome ten resistance genes, with the exception of R9. The number of virulence factors in each isolate was 6 to 10 in potato isolates. Of the isolates, 46 (grouped in four races) were found to have nine virulence factors. The mean numbers of virulence factors per isolate (Ci) and race (Cp) for isolates were calculated (Blandón-Díaz et al. 2012) as 8.2 and 7.8, respectively. Ci being higher than Cp in isolates indicates that complex races predominate within potato populations of P. infestans in Turkey.
Discussion
Knowledge about the degree of diversity that can be identified across a population is extremely dependent on the type of molecular marker used. Of the three types of genotypic response models used in this study, mtDNA haplotype analysis is the less descriptive for the characterization of individuals within a pathogen population; in P. infestans, it permits significant differentiation among only four genotypes. mtDNA is, however, a proper tool for determination of the different clonal lineages within the population, as clearly indicated by Ristaino et al. (2001). On the other hand, SSR and RFLP molecular techniques give a high degree of polymorphism and quickly and easily determine the frequency of genes or alleles in populations. With reducing cost of DNA sequencing and increasing availability of large sequence data sets permit the mining of this data for large numbers of SSRs. Thus, SSR genotyping has been widely used in genetic diversity analysis of P. infestans populations (Kiiker et al. 2018; Knapova and Gisi 2002; Tian et al. 2016). In this study, using as few as two SSR markers among the most polymorphic, we were able to differentiate almost all isolates. On the other hand, since all isolates from Turkey exhibit the single SSR genotype, even though we increased the number of SSR markers from 2 to 12, the SSR genotype in all 127 isolates remained the same. These data indicate that the population of P. infestans in Turkey has revealed that consists of a single clonal lineage, although RFLP analysis showed only minor variations in a few isolates.
The late blight epidemics in Turkey during 2015–2017 were important, being started by one new clonal lineage of P. infestans, 13_A2. This led to occurrence of the disease that had not been seen in the country in previous years. US-8 and US-1 genotypes detected in tomato in previous years (Tosun et al. 2007) were not found in these epidemics in Turkey. The 13_A2 lineage was first reported in Germany and the Netherlands in 2004, followed by Poland in 2006, then in Great Britain in 2008, as well as in China and was also recently reported in India, where it was most likely to have entered from Europe (Cooke et al. 2012; Dey et al. 2018; Li et al. 2013b; Rekad et al. 2017). In Southern India, the 13_A2 lineage was responsible for severe late blight epidemics in potato and tomato and displaced the previous population consisting of US-1 and other genotypes (Chowdappa et al. 2015). Due to the import of potato seeds from the Netherlands, France, Germany, and the UK, genotypic variation in P. infestans populations from Turkey was expected. In parallel with this hypothesis, it is seen that the pathogen population in Turkey A2 mating type and Ia with mtDNA haplotype consists of a single clonal lineage. The results of the study clearly revealed that the population of the pathogen consisting of a single clonal lineage was of European origin because in the analyses using the SSR marker, the allele sizes commonly obtained in Europe were determined (Cooke et al. 2012; Dey et al. 2018; Li et al. 2013a; Stroud et al. 2016).
Mefenoxam, which has the ability to cure the plant against the attacks of sensitive pathogen populations, is one of the best fungicides used successfully against P. infestans (Cohen et al. 1979). However, it is known that certain clonal lines such as US-8 and US-11 do not have sensitivity to mefenoxam. The findings of this and previous studies have shown that the reaction against mefenoxam can still be used as a reliable character in determining any clonal lineage. The use of this test combines with allozyme assays that are rapid and can be helpful in predicting mefenoxam susceptibility. Although the products containing mefenoxam have been replaced by new active substances in the last 10 years, the isolates tested in this study were found to be resistant to mefenoxam (Chmielarz et al. 2014; Cooke et al. 2011; Kiiker et al. 2018). In contrast, recent studies conducted in the USA showed that the sensitivity to mefenoxam increased in the new clonal lineages of the isolates compared with the 2000s (Danies et al. 2013). This increased sensitivity is attributed to less use of fungicide than in the past. As a result of random mutations, resistance to phenylamide group fungicides, including mefenoxam, can occur naturally among pathogen populations. However, the use of intense fungicide causes an increase in populations by creating a selection pressure in favor of these resistant individuals (Stellingwerf et al. 2018). In the period from the mid-990s until the early 2000s, the frequent and unconscious use of mefenoxam in Turkey caused the increase in the amount of resistant individuals in the total pathogen population. This has made selective pressure for the present clonal lineages of P. infestans to consist solely of resistant individuals and caused the decrease in genotypic diversity in the population as determined in previous studies (Grünwald et al. 2006). On the other hand, a high phenotypic diversity was maintained in the Turkish population of P. infestans resistant to mefenoxam.
Despite low genotypic differences, a wide difference was found in response to virulence and fungicide resistance in the pathogen population. These results are consistent with the results of a study conducted in North China. In this study, virulence showed a wide variation on the basis of isolates and some of these isolates showed virulence to all resistance (R) genes (Guo et al. 2009). However, unlike the population in China, P. infestans population in Turkey did not succeed to infect all the R genes located in potato differential sets, and racial diversity and common structure consisted of groups that generally infect 8–9 potato clones. In other parts of the world, a complex race structure and high virulence diversity have been identified (Deahl et al. 2003; Lehtinen et al. 2008; Pérez et al. 2001). Similar to other countries in the world, this complex race diversity and high virulence differences have been identified (Deahl et al. 2003; Lehtinen et al. 2008; Pérez et al. 2001). In the study conducted, all of the isolates in the culture stocks showed high virulence to potato clones bearing the resistance genes R1, R2, R4, and R7. These results are consistent with the general characteristics of the isolates in the EuroBlight database. As a matter of fact, isolates in this database are reported to show high virulence to R1, R2, R4, R7, and R11 (www.euroblight.net). A selection pressure created by the R genes found in the most commonly cultivated cultivars may have played a role in this complex racial diversity determined in P. infestans isolates. For example, the ‘Sante’ variety found among the grower preferences is known to have the resistance genes R1, R2, and R10, and these R genes may, to some extent, be responsible for the high frequency of virulence (Flier et al. 2007). The occurrence of new blight races is associated with a mutation in the non-pathogenic (Avr) gene encoding the effector protein, so that the effector can no longer be recognized by the R protein. The high virulence diversity in isolates with the same SSR multi-locus genotype can be explained by the localization of the Avr genes in a hypervariable region of the genome (Jiang et al. 2008).
In conclusion, the present P. infestans population consists of a single clonal lineage (13_A2), belonging to the A2 mating type and the Ia mtDNA haplotype. Furthermore, based on SSR profiles, this population resembles the P. infestans population of European countries that provide the seed tubers that are planted in Turkey. However, the Turkish P. infestans population exhibited a large variation in race diversity and its composition, and the individuals resistant to mefenoxam remained high in the population. This clonal lineage, as mentioned above, has been common in potato production areas of European countries since 2006. In addition, critical issues, such as pathogen survival from year to year, sources of the primary inoculum that triggered epidemic outbreaks at the beginning of the cultivation period and analysis of their competition, and the performance of the wild solanaceous species as a source of inoculum remain largely unknown and could explain the details behind the survival and persistence of this clonal lineage notwithstanding possible attacks by other genotypes.
References
Bakonyi, J., Láday, M., Dula, T., & Érsek, T. (2002). Characterisation of isolates of Phytophthora infestans from Hungary. European Journal of Plant Pathology, 108(2), 139–146.
Black, W., Mastenbroek, C., Mills, W. R., & Peterson, L. C. (1953). A proposal for an international nomenclature of races of Phytophthora infestans and of genes controlling immunity in Solanum demissum derivatives. Euphytica, 2(3), 173–179.
Blandón-Díaz, J. U., Widmark, A. K., Hannukkala, A., Andersson, B., Högberg, N., & Yuen, J. E. (2012). Phenotypic variation within a clonal lineage of Phytophthora infestans infecting both tomato and potato in Nicaragua. Phytopathology, 102(3), 323–330.
Brurberg, M. B., Elameen, A., Le, V. H., et al. (2011). Genetic analysis of Phytophthora infestans populations in the Nordic European countries reveals high genetic variability. Fungal Biology, 115(4–5), 335–342.
Caten, C. E., & Jinks, J. L. (1968). Spontaneous variability of single isolates of Phytophthora infestans. I. Cultural variation. Canadian Journal of Botany, 46(4), 329–348.
Chmielarz, M., Sobkowiak, S., Debski, K., Cooke, D. E. L., Brurberg, M. B., & Sliwka, J. (2014). Diversity of Phytophthora infestans from Poland. Plant Pathology, 63(1), 203–211.
Chowdappa, P., Kumar, N. B. J., Madhura, S., Kumar, M. S. P., Myers, K. L., Fry, W. E., Squires, J. N., & Cooke, D. E. L. (2013). Emergence of 13_A2 blue lineage of Phytophthora infestans was responsible for severe outbreaks of late blight on tomato in south-West India. Journal of Phytopathology, 161(1), 49–58.
Chowdappa, P., Kumar, N. B. J., Madhura, S., Kumar, M. S. P., Myers, K. L., Fry, W. E., & Cooke, D. E. L. (2015). Severe outbreaks of late blight on potato and tomato in South India caused by recent changes in the Phytophthora infestans population. Plant Pathology, 64(1), 191–199.
Cohen, Y., Reuveni, M., & Eyal, H. (1979). The systemic antifungal activity of Ridomil against Phytophthora infestans on tomato plants. Phytopathology, 69(6), 645–649.
Cooke, D. E. L., & Lees, A. K. (2004). Markers, old and new, for examining Phytophthora infestans diversity. Plant Pathology, 53(6), 692–704.
Cooke, L. R., Little, G., Armstrong, C., et al. (2009). Recent changes in the Phytophthora infestans population in Northern Ireland and first results from a new all-Ireland late blight project. PPO-Special Report, 13, 183–190.
Cooke, L. R., Schepers, H. T. A. M., Hermansen, A., Bain, R. A., Bradshaw, N. J., Ritchie, F., Shaw, D. S., Evenhuis, A., Kessel, G. J. T., Wander, J. G. N., Andersson, B., Hansen, J. G., Hannukkala, A., Nærstad, R., & Nielsen, B. J. (2011). Epidemiology and integrated control of potato late blight in Europe. Potato Research, 54(2), 183–222.
Cooke, D. E. L., Cano, L. M., Raffaele, S., Bain, R. A., Cooke, L. R., Etherington, G. J., Deahl, K. L., Farrer, R. A., Gilroy, E. M., Goss, E. M., Grünwald, N. J., Hein, I., MacLean, D., McNicol, J. W., Randall, E., Oliva, R. F., Pel, M. A., Shaw, D. S., Squires, J. N., Taylor, M. C., Vleeshouwers, V. G. A. A., Birch, P. R. J., Lees, A. K., & Kamoun, S. (2012). Genome analyses of an aggressive and invasive lineage of the Irish potato famine pathogen. PLoS Pathogens, 8(10), 1–14.
Danies, G., Small, I. M., Myers, K., Childers, R., & Fry, W. E. (2013). Phenotypic characterization of recent clonal lineages of Phytophthora infestans in the United States. Plant Disease, 97(7), 873–881.
Deahl, K. L., Pagani, M. C., Vilaro, F. L., Perez, F. M., Moravec, B., & Cooke, L. R. (2003). Characteristics of Phytophthora infestans isolates from Uruguay. European Journal of Plant Pathology, 109(3), 277–281.
Dey, T., Saville, A., Myers, K., Tewari, S., Cooke, D. E. L., Tripathy, S., Fry, W. E., Ristaino, J. B., & Roy, S. G. (2018). Large sub-clonal variation in Phytophthora infestans from recent severe late blight epidemics in India. Scientific Reports, 8, 4429.
Flier, W. G., Kroon, L. P. N. M., Hermansen, A., van Raaij, H. M. G., Speiser, B., Tamm, L., Fuchs, J. G., Lambion, J., Razzaghian, J., Andrivon, D., Wilcockson, S., & Leifert, C. (2007). Genetic structure and pathogenicity of populations of Phytophthora infestans from organic potato crops in France, Norway, Switzerland and the United Kingdom. Plant Pathology, 56(4), 562–572.
Forbes, G. A. (2012). Using host resistance to manage potato late blight with particular reference to developing countries. Potato Research, 55(3–4), 205–216.
Forbes, G. A., Goodwin, S. B., Drenth, A., Oyarzun, P., Ordonez, M. E., & Fry, W. E. (1998). A global marker database for Phytophthora infestans. Plant Disease, 82(7), 811–818.
Fry, W. E. (2008). Phytophthora infestans: the plant (and R gene) destroyer. Molecular Plant Pathology, 9(3), 385–402.
Fry, W. E., Goodwin, S. B., Dyer, A. T., Matuzak, J. M., Drenth, A., Tooley, P. W., Sujkowski, L. S., Koh, Y. J., Cohen, B. A., Spielman, L. J., Deahl, K. L., Inglis, D. A., & Sandlan, K. P. (1993). Historical and recent migration of Phytophthora infestans: chronology, pathways and implication. Plant Disease, 77(7), 653–661.
Goodwin, S. B., Drenth, A., & Fry, W. E. (1992). Cloning and genetic analyses of two highly polymorphic, moderately repetitive nuclear DNAs from Phytophthora infestans. Current Genetics, 22(2), 107–115.
Goodwin, S., Schneider, R., & Fry, W. E. (1995). Use of cellulose-acetate electrophoresis for rapid identification of allozyme genotypes of Phytophthora infestans. Plant Disease, 79(11), 1181–1185.
Griffith, G. W., & Shaw, D. S. (1998). Polymorphisms in Phytophthora infestans: four mitochondrial haplotypes are detected after PCR amplification of DNA from pure cultures or from host lesions. Applied and Environmental Microbiology, 64(10), 4007–4014.
Grünwald, N. J., Sturbaum, A. K., Romero Montes, G., Garray Serrano, E., Lozoya-Saldaña, H., & Fry, W. E. (2006). Selection for fungicide resistance within a growing season in field populations of Phytophthora infestans at the center of origin. Phytopathology, 96(12), 1397–1403.
Gunacti, H., Ay, T., & Can, C. (2019). Genotypic and phenotypic characterization of Phytophthora infestans populations from potato in Turkey. Phytoparasitica, 47(3), 429–439.
Guo, J., van der Lee, T., Qu, D. Y., Yao, Y. Q., Gong, X. F., Liang, D. L., Xie, K. Y., Wang, X. W., & Govers, F. (2009). Phytophthora infestans isolates from Northern China show high virulence diversity but low genotypic diversity. Plant Biology, 11(1), 57–67.
Hu, C. H., Perez, F., Donohoo, R., et al. (2012). Recent genotypes of Phytophthora infestans in eastern USA reveal clonal populations and reappearance of mefenoxam sensitivity. Plant Disease, 96(9), 1323–1330.
Jiang, R. H. Y., Tripathy, S., Govers, F., & Tyler, B. M. (2008). The RXLR effector reservoir in two Phytophthora species is dominated by a single rapidly evolving super-family with more than 700 members. Proceedings of the National Academy of Sciences of the United States of America, 105(2), 4874–4879.
Judelson, H. S. (1997). The genetics and biology of Phytophthora infestans: modern approaches to a historical challenge. Fungal Genetics and Biology, 22(2), 65–76.
Kanetis, L., Pittas, L., Tsaltas, D., & Ioannou N. (2013). Population structure of P. infestans in Cyprus and a synopsis on its Mediterranean status. EuroBlight Workshop, 12-15 May, Limassol, Cyprus.
Kiiker, R., Hansen, M., Williams, I. H., Cooke, D. E. L., & Runno-Paurson, E. (2018). Outcome of sexual reproduction in the Phytophthora infestans population in Estonian potato fields. European Journal of Plant Pathology, 152(2), 395–407.
Knapova, G., & Gisi, U. (2002). Phenotypic and genotypic structure of Phytophthora infestans populations on potato and tomato in France and Switzerland. Plant Pathology, 51(5), 641–653.
Kröner, A., Mabon, R., Corbiére, R., Montarry, J., & Andrivon, D. (2017). The coexistence of generalist and specialist clonal lineages in natural populations of the Irish Famine pathogen Phytophthora infestans explains local adaptation to potato and tomato. Molecular Ecology, 26(7), 1891–1901.
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6), 1547–1549.
Lees, A. K., Wattier, R., Shaw, D. S., Sullivan, L., Williams, N. A., & Cooke, D. E. L. (2006). Novel microsatellite markers for the analysis of Phytophthora infestans populations. Plant Pathology, 55(3), 311–319.
Lehtinen, A., Hannukkala, A., Andersson, B., Hermansen, A., Le, V. H., Naerstad, R., Brurberg, M. B., Nielsen, B. J., Hansen, J. G., & Yuen, J. (2008). Phenotypic variation in Nordic populations of Phytophthora infestans in 2003. Plant Pathology, 57(2), 227–234.
Li, Y., Govers, F., Mendes, O., Testa, A., Jacobsen, E., Huang, S. W., & van der Lee, T. A. J. (2010). A new set of highly informative SSR markers for Phytophthora infestans population analysis assembled into an efficient multiplex. Molecular Ecology Resources, 10(6), 1098–1105.
Li, Y., van der Lee, T. A., Evenhuis, A., et al. (2012). Population dynamics of Phytophthora infestans in the Netherlands reveals expansion and spread of dominant clonal lineages and virulence in sexual offspring.G3. Genes Genomes Genetics, 2(12), 1529–1540.
Li, Y., Cooke, D. E. L., Jacobsen, E., & van der Lee, T. (2013a). Efficient multiplex simple sequence repeat genotyping of the oomycete plant pathogen Phytophthora infestans. Journal of Microbiological Methods, 92(3), 316–322.
Li, Y., van der Lee, T., Zhu, J. H., Jin, G. H., Lan, C. Z., Zhu, S. X., Zhang, R. F., Liu, B. W., Zhao, Z. J., Kessel, G., Huang, S. W., & Jacobsen, E. (2013b). Population structure of Phytophthora infestans in China - geographic clusters and presence of the EU genotype Blue_13. Plant Pathology, 62(4), 932–942.
Malcolmson, J. F., & Black, W. (1966). New R genes in Solanum demissum Lindl. and their complementary races of Phytophthora infestans (Mont.) de Bary. Euphytica, 15(2), 199–203.
Mariette, N., Mabon, R., Corbiere, R., Boulard, F., Glais, I., Marquer, B., Pasco, C., Montarry, J., & Andrivon, D. (2016). Phenotypic and genotypic changes in French populations of Phytophthora infestans: are invasive clones the most aggressive? Plant Pathology, 65(4), 577–586.
Montes, M. S., Nielsen, B. J., Schmidt, S. G., Bødker, L., Kjøller, R., & Rosendahl, S. (2016). Population genetics of Phytophthora infestans in Denmark reveals dominantly clonal populations and specific alleles linked to metalaxyl-M resistance. Plant Pathology, 65(5), 744–753.
Pérez, W. G., Gamboa, J. S., Falcon, Y. V., Coca, M., Raymundo, R. M., & Nelson, R. J. (2001). Genetic structure of Peruvian populations of Phytophthora infestans. Phytopathology, 91(10), 956–965.
Rekad, F. Z., Cooke, D. E. L., Puglisi, I., Randall, E., Guenaoui, Y., Bouznad, Z., Evoli, M., Pane, A., Schena, L., Magnano di San Lio, G., & Cacciola, S. O. (2017). Characterization of Phytophthora infestans populations in northwestern Algeria during 2008-2014. Fungal Biology, 121(5), 467–477.
Ristaino, J. B., Groves, C. T., & Parra, G. R. (2001). PCR amplification of the Irish potato famine pathogen from historic specimens. Nature, 411, 695–697.
Runno-Paurson, E., Fry, W. E., Myers, K. L., Koppel, M., & Mand, M. (2009). Characterisation of Phytophthora infestans isolates collected from potato in Estonia during 2002-2003. European Journal of Plant Pathology, 124(4), 565–575.
R Development Core Team. (2019). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria.
Stellingwerf, J. S., Phelan, S., Doohan, F. M., Ortiz, V., Griffin, D., Bourke, A., Hutten, R. C. B., Cooke, D. E. L., Kildea, S., & Mullins, E. (2018). Evidence for selection pressure from resistant potato genotypes but not from fungicide application within a clonal Phytophthora infestans population. Plant Pathology, 67(7), 1528–1538.
Stroud, J. A., Shaw, D. S., Hale, M. D., & Steele, K. A. (2016). SSR assessment of Phytophthora infestans populations on tomato and potato in British gardens demonstrates high diversity but no evidence for host specialization. Plant Pathology, 65(2), 334–341.
Tian, Y. E., Yin, J. L., Sun, J. P., Ma, Y. F., Wang, Q. H., Quan, J. L., & Shan, W. X. (2016). Population genetic analysis of Phytophthora infestans in northwestern China. Plant Pathology, 65(1), 17–25.
Tosun, N., Yıldırım, A., Türküsay, H., & Tanyolaç, B. (2007). Genetic variation among Phytophthora infestans (tomato blight) isolates from western Turkey revealed by inter simple sequence repeat (ISSR) and random amplified polymorphic DNA (RAPD) markers. Pakistan Journal of Botany, 39(3), 897–902.
TUIK (2018). http://www.tuik.gov.tr. Accessed 10 Sept 2019.
Funding
This research was supported by TUBITAK (grant 116O329).
Author information
Authors and Affiliations
Contributions
MEG and WEF designed the study. MEG, NA, GÖ, KM, AÇ, WEF, UP, TY, MA, HK, DELC, and NZ carried out the experiments. MEG, WEF, NA, GÖ, and DELC contributed to the interpretation of the results. MEG took the lead in writing the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Göre, M.E., Altın, N., Yaman, T. et al. Severe outbreaks of Phytophthora infestans on potato in Turkey caused by recent changes in the pathogen population structure. Phytoparasitica 47, 693–709 (2019). https://doi.org/10.1007/s12600-019-00768-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12600-019-00768-5