Abstract
Chemokine-like receptors (CMKLRs) are multi-functional receptors with roles in regulating leukocyte and inflammation. Currently, two members of the CMKLRs, CMKLR1 and CMKLR2, have been found in salmonids, indicating that the CMKLRs had expanded from an ancestor gene in teleost fish. In the present study, the third member of the CMKLRs, defined as CMKLR3, was identified and cloned in rainbow trout. The trout CMKLR3 possessed conserved features of the CMKLR family including seven transmembrane regions, a dynein regulatory complex (DRC) motif, and two cysteine residues, but shared low sequence identities with fish CMKLR1 and CMKLR2 (24‒38 %), which was confirmed by phylogenetic tree analysis. Trout CMKLR3 was highly expressed in body kidney, head kidney, and IgM+ B cells, indicating its functional role in regulating leukocytes. We were able to modulate the expression of trout CMKLR3 in vivo by bacterial and parasitic infections but it remained apathetic to virus infection, and it was also successfully modulated in vitro by peptidoglycan and cytokines (IFN-γ and IL-6). Our results suggest that trout CMKLR3 is regulated in a complex way and has an important regulatory role in inflammatory responses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chemokine receptors belong to the family of G-protein-coupled seven-transmembrane receptors, which predominantly express on the surface of leukocytes and play crucial regulatory roles in the movement of leucocytes by binding with their specific ligands (chemokines). According to the chemokine classes that they bind, the various chemokine receptors have been named as follows: CCR, which binds to CC-chemokines, CXCR, which binds to CXC chemokines, XCR, which binds to XC chemokines, and CX2CR, which binds to CX3C chemokines; in all cases, X is any amino acid and C is cysteine [1]. After binding with their ligands, the chemokine/receptor system may trigger multiple intracellular signaling events involving in gene transcription, cytoskeleton rearrangement, and chemotaxis [2, 3].
Except for the classical chemokine receptors, there are some atypical chemokine receptors (ACKRs) that also are seven-transmembrane molecules and can bind to chemokines but cannot induce the classical chemokine receptor signaling pathway [4–6]. Human ACKRs include at least five members such as ACKR1 (Duffy antigen receptor for chemokines, DARC), ACKR2 (D6 and CC chemokine-binding protein 2 [CCBP2]), ACKR3 (CXC-chemokine receptor 7 [CXCR7], and RDC1), ACKR4 (CC-chemokine receptor-like 1 [CCRL1], CCXCKR and CCR11), and CCRL2 (chemokine receptor on activated macrophages [CRAM]) [7]. ACKRs play important roles in the regulation of classical chemokine/receptor responses. ACKR1 can bind a large number of CXC and CC chemokines, and functions as a chemokine sink [7, 8]. ACKR2 can bind at least 12 inflammatory CC chemokines and plays a role in the regulation of inflammatory responses [5, 6]. CCRL2 can bind chemokine CCL5 and CCL19 to recycle and reduce local concentration of these two chemokines, and functions in the regulation of dendritic cell (DC) trafficking and other immune responses [9–11]. CCRL2 can also bind chemoattractant chemerin and functions in the regulation of inflammatory responses [12, 13].
Competitive binding to the ligand chemerin with CCRL2 brings into focus another seven-transmembrane molecular, chemokine-like receptor (CMKLR1), also termed as ChemR23 [14]. CCRL2 binds chemerin and increases the local chemerin concentration, and then presents it to CMKLR1 on adjacent cells, resulting in development of IgE-dependent, mast cell-dependent cutaneous anaphylaxis [13]. In addition to cooperation with CCRL2, CMKLR1 is also involved in regulating leukocytes and inflammatory processes. Several leukocytes including immature DCs, macrophages, monocytes, and CD4 + T lymphocytes can express it [14]. CMKLR1 expressed on macrophages and DCs can bind the active chemerin and induce cell migration [15, 16]. In-vivo studies using CMKLR-deficient mice showed that the CXCL1, interleukin (IL)-6, tumor necrosis factor (TNF), and IL-1β were reduced even under the lipopolysaccharide (LPS) induction [17]. Mammalian CMKLR1 can also serve as a co-receptor for some simian immunodeficiency virus (SIV) clones and human immunodeficiency virus (HIV)-1 strain [18]. These all indicated that CMKLR1 is a multi-functional receptor [19].
In fish, a number of chemokine receptors had been identified and some of them are fish-specific (e.g., CXCR3b, CCR12) [20, 21]. Current studies mainly focus on the classic chemokine receptors of fish but a few have looked at the atypical chemokine receptors. CMKLRs in teleost fish expanded from a common ancestor, as consequently two members (CMKLR1 and CMKLR2) were found in salmonids and three members (CMKLR1-3) in Northern pike Esox lucius, which may be the result of genome duplication [22]. To the best of our knowledge, no reports focus on the function of CMKLRs of fish species. Thus, in this study, the third member of the CMKLR family, CMKLR3, is identified for the first time from rainbow trout Oncorhynchus mykiss, and its expressions in tissues or cells under diseases or stimulation are characterized to broaden our knowledge on CMKLRs in fish, and to provide the basis for clearly understanding their functions.
Materials and methods
Fish
Rainbow trout (average of 100 g) were obtained from the Mill of Elrich Trout Fishery (Aberdeenshire, UK). Fish were maintained in 1-m-diameter aerated fiberglass tanks with a re-circulating water system at 14 ± 1 °C and fed twice daily with standard commercial pellets (EWOS). Prior to the experiments, fish were acclimated for at least 2 weeks.
Total RNA extraction and cDNA synthesis
Total RNA was independently extracted from 17 selected tissues including caudal fin, adipose fin, thymus, gills, brain, scales, skin, muscle, liver, spleen, head kidney, body kidney, intestine, heart, blood, and adipose tissue using Trizol Reagent (Invitrogen, CA) following the manufacturer’s instruction. The cDNA was synthesized from total RNA using the BioScript™ Reverse Transcriptase kit (Bioline, UK).
Cloning of trout CMKLR3 and sequence analysis
The trout CMKLR1 protein sequence was used as a bait to search the trout expressed sequence tag (EST) database by tBLASTn software, and an EST (704 bp) that contained a 5′-untranslated region (5′-UTR) and an incomplete open reading frame (ORF) were identified. This EST was further used as query to search the trout genome database (http://www.genoscope.cns.fr/trout) using BLASTn software. A scaffold (MMSRT116B_scaff_1281_1) of 4980 base pairs (bp) in length was obtained. This genome sequence was analyzed using GenScan to get the potential 5′-UTR, 3′-UTR, and ORF. Primers were designed within the predicted 5′-UTR and 3′-UTR, and polymerase chain reaction (PCR) amplification was done using head kidney cDNA samples as a template. A single band was obtained and the PCR product was sequenced by Eurofins Biotec. The primers used for gene clone are listed in Table 1.
The predicted amino acid sequences were deduced from the nucleotide sequence using Translate at the ExPASy Molecular Biology Server (http://www.expasy.org/tools). Multiple sequences alignment was conducted using the ClustalW program [23]. Transmembrane domain was predicted by the SMART7 program [24]. A phylogenetic tree was constructed by the neighbor-joining methods using MEGA 6 software and set with a bootstrap of 10,000 times [25].
Tissue distribution of trout CMKLR3 transcripts
Seventeen tissues as mentioned above were collected from six healthy fish, and RNA preparation, cDNA synthesis, and real-time PCR analysis were performed as described previously [26, 27]. The expression levels of trout CMKLR3 were normalized to that of EF-1α and expressed as arbitrary units. Primers used for real-time PCR are listed in Table 1.
Gene expression of trout CMKLR3 in IgM+ cells isolated from rainbow trout tissues
In trout, two different subpopulations of B cells have been identified, IgM+/IgD+/IgT− (IgM+ cells) and IgM−/IgD−/IgT+ (IgT+ cells). Since the expression of many chemokine receptors had been found in IgM+ B cells [28], we chose the IgM + cells isolated from the blood, spleen, head kidney, gills, and intestine of rainbow trout to examine the expression pattern of CMKLR3 in this study. The IgM+ cells from different tissues were isolated as described previously [28]. The expression of CMKLR3 in the cells was measured using the same method for those in tissues.
Gene expression of trout CMKLR3 after bacterial, parasitic, and viral infections
Bacterial infection: Forty-eight fish (average of 100 g) were randomly divided into two groups, a challenge group and a control group. The fish in the challenge group were injected intraperitoneally (i.p) with a 0.5-ml suspension (1 × 106 cfu in PBS) of Yersinia ruckeri (strain MT3072), the causative agent of red mouth disease [29], and fish in the control group were injected i.p. with the same amount of PBS followed by rearing at 15 ± 1 °C with a re-circulating water system. Head kidney tissues of six fish from each group were collected at 6, 24, 48, and 72 h post-injection, respectively. Real-time PCR quantification of the expression of CMKLR3 was done as described previously [26] and expressed as fold change relative to the time-matched controls.
Parasitic infection: The proliferative kidney disease (PKD) caused by the myxozoan parasite, Tetracapsuloides bryosalmonae, is a disease with major economic impact on salmonid aquaculture [30]. In the present study, the caudal kidney tissues were sampled in late July at a water temperature of 15-16 °C from fish (average of 100 g) provided by a commercial trout farm in Southern England. The different clinical stages of the fish were determined using the kidney swelling index from Clifton-Hadley et al. [31]. Expression of trout CMKLR3 in the caudal kidney of the fish in different stages of PKD was studied by real-time PCR quantification.
Virus infection: Fish weighing 14.7 g were bath challenged with the viral hemorrhagic septicemia virus (VHSV) at 14 °C in an aquarium facility [32]. The infection of VHSV was confirmed by real-time PCR using primers of the VHSV N gene as described previously [33]. The kidney was sampled from six of the challenged fish or from phosphate buffered saline (PBS) mock-challenged fish at 1 and 3 days post-challenge. Expression of trout CMKLR3 was analyzed by real-time PCR and expressed in terms of fold change relative to the time-matched controls.
Gene expression of trout CMKLR3 in head kidney primary macrophages
Since CMKLR1 could be expressed and regulated by several cytokines in human macrophages, the primary trout head kidney macrophages were isolated to study the expression modulation of trout CMKLR3. The primary head kidney macrophages were prepared as in a previous study [34], and stimulated with peptidoglycan (PGN, 5 μg/ml, Invivogen) and recombinant cytokines including rIFN-γ (20 ng/ml) [26], rIL-6 (100 ng/ml) [34], and rTNF-α (10 ng/ml) [35] for 2, 4, 8, and 24 h. RNA extraction and real-time PCR were performed according to the method described above.
Statistical analysis
All data were expressed as mean ± SEM. Statistical analysis was analyzed using SPSS Statistics package 19.0 (SPSS Inc., Chicago, Illinois). The data from the infection studies were analyzed using one-way analysis of variance (ANOVA) and the least significant difference (LSD) post-hoc test, and the data from in-vitro experiments were analyzed by a paired-sample t test. Differences with p < 0.05 were considered statistically significant.
Results
Cloning and sequence analysis of trout CMKLR3
The amplified trout CMKLR3 cDNA (GenBank accession No. KM516349) was 1381 bp in length, which contained a 5′-UTR of 150 bp, an ORF of 1146 bp encoding for 381 amino acids, and a 3′-UTR of 85 bp. The trout CMKLR3 gene was perfectly mapped to the genomic scaffold 1281, which contained a single intron of 1221 bp in the 5′-UTR. There was a stop codon (TGA) in the 5′-UTR upstream of the start codon (ATG) (Fig. S1), indicating that the complete ORF of this gene had been obtained. In addition, the CMKLR3 of Atlantic salmon was obtained by using trout CMKLR3 as a query to search the salmon genome database (http://salmondb.cmm.uchile.cl), whose ORF was 1098 bp in length encoding for 365 amino acids and located on genomic contig AGKD00000000.4 (Fig. S2).
Trout CMKLR3 shared respectively 93.5 and 75.8 % amino acid sequence identities with Atlantic salmon and Northern pike CMKLR3, which were higher than those with fish CMKLR1 and CMKLR2 (24–38 %), and mammalian CMKLR1 (40.5–41.8 %) (Table 2). Multiple sequence alignment revealed that trout CMKLR3 contained several conserved CMKLRs family features. Firstly, trout CMKLR3 possessed the G-coupled protein seven-transmembrane regions (TM), which divided the molecular into an extracellular amino-terminal, three extracellular regions (ECL), three intracellular regions (ICL), and a cytoplasmic carboxyl region (Figs. 1, 2). Secondly, unlike the classical chemokine receptors, which contain four cysteine residues (one in the N-terminal and three in the ECLs) [1, 4, 6], trout CMKLR3 only possessed two cysteine residues in the ECL1 and ECL2 region. The same case was also found in other CMKLRs family members. Thirdly, all CMKLRs shared a dynein regulatory complex (DRC) motif at the beginning of ICL2 instead of DRY motif at the same position in the classical chemokine receptors. Lastly, the N-terminal or C-terminal of CMKLRs contained several residues known to be important for activation or function of chemokine receptors, e.g., N-glycosylation sites and tyrosine O-sulfation sites at the N-terminal (Fig. 1).
Multiple alignment of CMKLRs. The multiple alignment of CMKLRs was produced using ClustalW and the conserved amino acids were shaded using BOXSHADE (version 3.21). The N-terminus, seven-transmembrane domains (TM1-7), three extracellular loops (ECL1-3), three intracellular loops (ICL1-3) and C-terminus are marked above the alignment. The two conserved cysteine residues in ECL1 and ECL2 are indicated by a white triangle below the alignment. The potential N-liked glycosylation site at the N-terminal and conserved tyrosine residues are marked in the black box and the dotted box, respectively. The DRC motif in the ICL2 region is marked by a black dot under the sequences. The accession numbers for sequences used in this alignment are given in Fig. 3
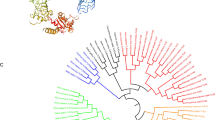
To confirm the sequence identities, a phylogenetic tree was constructed based on a multiple alignment of vertebrates’ CMKLRs (Fig. 3). Clearly, the tree was divided into four main clades. The CMKLR1 of mammals, birds, amphibians, and reptiles formed one clade, while fish CMKLR1 and CMKLR2 formed respective separate clades. Trout CMKLR3 was well clustered with the clade of fish CMKLR3 with a convincing bootstrap value (100 %).
An unrooted phylogenetic tree of CMKLRs. The tree was constructed using the neighbor-joining method by MEGA 6 software. The evolutionary distances were computed using the JTT matrix-based method and pairwise deletion option. The GenBank accession number of each sequence was given after the species name and molecular type
Tissue distribution of trout CMKLR3 expression
CMKLR3 expressions in 17 tissues of healthy trout were examined using real-time PCR. Results showed that trout CMKLR3 was ubiquitous, with the highest expression in caudal kidney and head kidney (p < 0.05) and the lowest in the liver and tail fins (p < 0.05). A high expression level of trout CMKLR3 was also found in spleen, blood, muscle, gills, intestines, gonad, and scales (Fig. 4).
Gene expression of trout CMKLR3 in IgM+ cells isolated from rainbow trout tissues
Since CMKLRs are involved in cell migration, the expression of CMKLR3 in cells will characterize the cell migration pattern. Thus, we analyzed the transcription of trout CMKLR3 in sorted IgM + cells by real-time PCR. Results showed that trout CMKLR3 was expressed in IgM+ cells derived from kidney, blood, spleen, gills, and intestine: it was lowest in blood IgM+ cells, moderate in head kidney and gills IgM+ cells, and highest in intestine IgM+ cells (Fig. 5).
Gene expression of trout CMKLR3 after bacterial, parasitic, and viral infections
The expression of trout CMKLR3 after infection with bacteria (Y. ruckeri), a parasite (T. bryosalmonae), and a virus (VHSV) were also investigated by real-time PCR. After infection of trout with Y. ruckeri, the expression of trout CMKLR3 in the head kidney of challenge group fish was significantly increased at 24 and 48 h post-infection when compared with that in the control group (p < 0.05) (Fig. 6a). In fish with clinical PKD, CMKLR3 expression was significantly decreased in the kidney of the fish with a PKD swelling grade of 1-2, but not changed in the fish with a swelling grade of 1, 2, and 3 (Fig. 6b), compared with those in the uninfected fish (the swelling grade 0). Following VHSV bath exposure, trout CMKLR3 remained apathetic at 1 and 3 days post-infection (Fig. 6c).
Modulation of rainbow trout O. mykiss CMKLR3 expression by bacterial (a), parasitic (b), and VHSV (c) infection. a Rainbow trout were i.p. injected with Y. ruckeri or PBS. The gene expression in the head kidney of the challenge group fish is expressed as a fold change relative to those of the control fish (means + SEM of five fish). b Kidneys from rainbow trout infected with Tetracapsuloides bryosalmonae (kidney swelling index 1, 1–2, 2, and 3) or from uninfected (index 0) fish were collected during a spontaneous infection. Results are presented as a fold change relative to the control fish (means + SEM). The numbers of fish analyzed were 11, 5, 9, 10, and 9, representing the index 0 (uninfected control), 1, 1–2, 2, and 3, respectively. c The kidney was sampled from six VHSV-challenged fish or PBS mock-challenged fish at 1 and 3 days post-challenge. Expression of trout CMKLR3 is expressed as a fold change relative to the time-matched controls (means + SEM). The significance of LSD post-hoc tests after one-way ANOVA between infected and control fish is shown above the bars: * p < 0.05 and **p < 0.01
Gene expression of trout CMKLR3 in head kidney primary macrophage
The expression of trout CMKLR3 in the head kidney primary macrophages stimulated with several known macrophage stimulants including PGN and recombinant cytokines (rIL-6, rIFN-γ, and rTNF-α) were analyzed by real-time PCR (Fig. 7). No changes were observed for trout CMKLR3 after rTNF-α stimulation. PGN upregulated CMKLR3 expression at 4 h, and the level remained high at 8 h and 24 h post-stimulation. Both rIL-6 and rIFN-γ downregulated CMKLR3 expression from 2 to 8 h post-stimulation, but no effect on the expression was observed at 24 h post-stimulation.
Modulation of rainbow trout O. mykiss CMKLR3 expression in primary head kidney macrophage. Four-day-old primary head kidney macrophages were stimulated with PGN or recombinant trout cytokines (rIL-6, rTNF-α and rIFN-γ) for 2, 4, 8, and 24 h. The gene expression of CMKLR3 is expressed as a fold change calculated as the mean expression levels in stimulated cells normalized to that of time-matched controls. The means + SEM of the cells isolated from four fish are shown. The paired sample t-tests between stimulated and time-matched control samples were analyzed with SPSS package 19.0: * p < 0.05 and **p < 0.01
Discussion
Two members of the CMKLR family had been characterized in salmonids in a previous study [22]. In the present study, the third member of the CMKLRs, which was termed CMKLR3, was identified and functionally analyzed in rainbow trout. The newly identified CMKLR3 possessed conserved structure features of the CMKLRs family that were revealed by sequence alignment analysis, including seven-transmembrane structure, DRC motif, and two cystine residues in the ECL1 and ECL2 regions (Figs. 1, 2). Meanwhile, trout CMKLR3 shared more than 75.8 % sequence identities with reported fish CMKLR3, which were higher than that with CMKLR1 (40.5–41.8 %) and CMKLR2 (24–38 %) (Table 2). In addition, phylogenetic tree analysis revealed that trout CMKLR3 clustered together with fish CMKLR3 into a clade with a convincing bootstrap value (100 %), which was clearly separated from the clade of fish CMKLR1 and CMKLR2 (Fig. 3). We also identified CMKLR3 in Atlantic salmon, which was also well clustered into the CMKLR3 clade (Fig. S2). Thus, it was reasonable to name this newly identified gene as CMKLR3. The finding of CMKLR3 in fish revealed that CMKLRs had been greatly expanded in teleost fish.
Mammalian CMKLR1 was an orphan receptor involved in signaling pathway regulation, cell migration, and inflammation [19]. The information on fish CMKLRs was scarce. In Atlantic salmon, CMKLR1, CMKLR2a, and ΨCMKLR2b had been identified, which might be the result of genome duplication [22]. Atlantic salmon CMKLR1 and CMKLR2a were highly expressed in head kidney, kidney, and spleen [22]. In the present study, we found that trout CMKLR3 was ubiquitously expressed in 17 tested tissues (Fig. 4), indicating that this receptor might play role(s) in a broad range of tissues. The high expression of trout CMKLR3 in the head kidney and spleen suggested its functional role(s) in regulating leukocytes [14]. Further expression analysis of trout CMKLR3 in IgM + B cells isolated from the intestine, gills, head kidney, and spleen revealed that it might play roles in the regulation of B lymphocytes.
We further examined whether trout CMKLR3 was modulated in bacterial, parasite, and viral infection to confirm its role in inflammatory responses. Y. ruckeri is the causative agent of enteric redmouth disease in rainbow trout and other fish species [29]. It has been found that several pro-inflammatory cytokines, e.g., IL-1β, IL-6, IL-8, TNF-α, and IFN-γ [36, 37], and some chemokine receptors, e.g., CXCR2-3 [27] and CCBP2 [38], were upregulated in immune organs of trout after Y. ruckeri infection. In the present study, we found that trout CMKLR3 was upregulated at 4, 8, and 24 h post-infection (Fig. 6a), suggesting its role in the regulation of the inflammatory response during bacterial infection. PKD of salmonid fish is caused by myxozoan Tetracapsuloides bryosalmonae, which targets the kidney of infected fish where it causes a chronic lymphoid immunopathology and induces an anti-inflammatory phenotype [30]. The downregulation of trout CMKLR3 at early stages of PKD disease (grade 1–2) (Fig. 6b) indicated its negative role in regulating anti-inflammatory responses in the kidney. Interestingly, although it is known that a number of chemokine receptors, e.g., CCR7, CCR9, CXCR3B and CXCR4, are upregulated during VHSV infection [39], trout CMKLR3 was refractory to VHSV infection in this study (Fig. 6c). Mammalian CMKLR1 seemed restrictedly to serve as a co-receptor for some SIV clones and HIV-1 strain [18, 22]. Whether the unchanged CMKLR3 expression in trout is due to the strain of the virus or without function in response to virus infection need further investigation.
Similar to mammalian CMKLRs, trout CMKLR3 was also expressed in macrophages and could be regulated by several cytokines and pathogen-associated molecular patterns (PAMPs) (Fig. 7). In primary head kidney macrophages, trout CMKLR3 was upregulated by PGN stimulation, confirming its regulating role in a host inflammatory response to bacterial infection. Furthermore, trout CMKLR3 was downregulated by IFN-γ stimulation. Similar results were also observed in mammalian CMKLR1. IFN-γ is produced by activated T cells, NK cells, and NKT cells, which is the hallmark of Th1 responses and can regulate both innate and cell-mediated immune responses [40]. The downregulation of CMKLR3 by IFN-γ stimulation demonstrated that CMKLR3 might be involved in the regulation of T cell development and activation in fish. In fish, IL-6 had been proved to be an anti-inflammatory cytokine that can modulate a number of cytokines and chemokine receptors [41]. The downregulation of trout CMKLR3 following IL-6 stimulation suggested the negative role of CMKLR3 in regulating anti-inflammatory response. In addition, trout CMKLR3 remained unchanged following TNF-α stimulation. TNF-α had a different role in the regulation of CMKLRs in different cell types. Mammalian CMKLR1 was upregulated by TNF-α in macrophages but was downregulated in monocytes [42]. The unchanged status of CMKLR3 after TNF-α revealed that fish CMKLRs might have a fish-specific regulated pattern under TNF-α stimulation. The different effects of cytokines on the expression of CMKLR3 also suggest that CMKLR3 can be regulated in a complex way. Further studies on the expression of CMKLRs in other cell types are needed to elucidate the regulation of the CMKLR network.
References
Allen SJ, Crown SE, Handel TM (2007) Chemokine: receptor structure, interactions, and antagonism. Annu Rev Immunol 25:787–820
Bennett LD, Fox JM, Signoret N (2011) Mechanisms regulating chemokine receptor activity. Immunology 134:246–256
Zweemer AJ, Toraskar J, Heitman LH, Ijzerman AP (2014) Bias in chemokine receptor signaling. Trends Immunol 35:243–252
Cancellieri C, Caronni N, Vacchini A, Savino B, Borroni EM, Locati M, Bonecchi R (2013) Review: structure-function and biological properties of the atypical chemokine receptor D6. Mol Immunol 55:87–93
Graham GJ, Locati M (2013) Regulation of the immune and inflammaroty by the ‘atypical’ chemokine receptor D6. J Pathol 229:168–175
Nibbs RJ, Graham GJ (2013) Immune regulation by atypical chemokine receptors. Nat Rev Immunol 13:815–829
Bachelerie F, Ben-Baruch A, Burkhardt AM, Combadiere C, Farber JM, Graham GJ, Horuk R, Sparre-Ulrich AH, Locati M, Luster AD, Mantovani A, Matsushima K, Murphy PM, Nibbs R, Nomiyama H, Power CA, Proudfoot AE, Rosenkilde MM, Rot A, Sozzani S, Thelen M, Yoshie O, Zlotnik A (2014) International Union of Pharmacology. LXXXIX. Update on the extended family of chemokine receptors and introducing a new nomenclature for atypical chemokine receptors. Pharmacol Rev 66:1–79
Zlotnik A, Yoshie O (2012) The chemokine superfamily revisited. Immunity 36:705–716
Fan P, Kyaw H, Su K, Zeng Z, Augustus M, Carter KC, Li Y (1998) Cloning and characterization of a novel human chemokine receptor. Biochem Biophys Res Commun 243:264–268
Hartmann TN, Leick M, Ewers S, Diefenbacher A, Schraufstatter I, Honczarenko M, Burger M (2008) Human B cells express the orhpan chemokine receptor CRAM-A/B in a maturation-stage-dependent and CCL5-modulated manner. Immunology 125:252–262
Leick M, Catusse J, Follo M, Nibbs RJ, Hartmann TN, Veelken H, Burger M (2009) CCL9 is a specific ligand of the constitutively recycling atyphical human chemokine receptor CRAM-B. Immunology 129:536–546
Yoshimura T, Oppenheim JJ (2008) Chemerin reveals its chimeric nature. J Exp Med 205:2187–2190
Zabel BA, Nakae S, Zuniga L, Kim JY, Ohyama T, Alt C, Pan J, Suto H, Soler D, Allen SJ, Handel TM, Galli SJ, Butcher EC (2008) Mast cell-expressed orphan receptor CCRL2 binds chemerin and is required for optimal induction of IgE-mediated passive cutaneous anaphylaxis. J Exp Med 205:2207–2220
Gantz I, Konda Y, Yang YK, Miller DE, Dierick HA, Yamada T (2006) Molecular cloning of a novel receptor (CMKLR1) with homology to the chemotactic factor receptors. Cytogenet Cell Genet 74:286–290
Wittamer V, Franssen JD, Vulcano M, Mirjolet JF, Poul EL, Migeotte I, Brézillon S, Tyldesley R, Blanpain C, Detheux M, Mantovani A, Sozzani S, Vassart G, Parmentier M, Communi D (2003) Specific recruitment of antigen-presenting cells by chemerin, a novel processed ligand from human inflammatory fluids. J Exp Med 198:977–985
Zabel BA, Silverio AM, Butcher EC (2005) Chemokine-like receptor 1 expression and chemerin-directed chemotaxis distinguish plasmacytoid from myeloid dendritic cells in human blood. J Immunol 174:244–251
Luangsay S, Wittamer V, Bondue B, Henau OD, Rouger L, Brait M, Franssen JD, Nadai PD, Huaux F, Parmentier M (2009) Mouse ChemR23 is expressed in dendritic cell subsets and macrophages, and mediates an anti-inflammatory activity of chemerin in a lung disease model. J Immunol 183:6489–6499
Samson M, Ediinger AL, Stordeur P, Rucker J, Verhasselt V, Sharron M, Govaerts C, Mollereau C, Vassart G, Doms RW, Parmentier M (1998) ChemR23, a putative chemoattractant receptor, is expressed in monocyte-derived dendritic cells and macrophages and is a coreceptor for SIV and some primary HIV-1 strains. Eur J Immunol 28:1689–1700
Yoshimura T, Oppenheim JJ (2011) Chemokine-like receptor 1 (CMKLR1) and chemokine (C-C motif) receptor-like 2 (CCRL2); Two multifunctional receptor with unusual properties. Exp Cell Res 317:674–684
Nomiyama H, Osada N, Yoshie O (2011) A family tree of vertebrate chemokine receptors for a unified nomenclature. Dev Comp Immunol 35:705–715
Zou J, Redmond AK, Qi ZT, Dooley H, Secombes CJ (2015) The CXC chemokine receptors of fish: insights into CXCR evolution in the vertebrates. Gen Comp Endocrinol 215:117–131
Grimholt U, Hauge H, Hauge AG, Leong J, Koop BF (2015) Chemokine receptors in Atlantic salmon. Dev Comp Immunol 49:79–80
Thompson J, Higgins D, Gibson T (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Letunic I, Doerks T, Bork P (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res 40:D302–D305
Kumar S, Tamura K, Nei M (1994) Mega-molecular evolutionary genetics analysis software for microcomputers. Comput Appl Biosci 10:189–191
Wang TH, Diaz-Rosales P, Costa MM, Compbell S, Snow M, Collet B, Martin SA, Secombes CJ (2011) Functional characterization of a nonmammalian IL-21: rainbow trout Oncorhynchus mykiss IL-21 upregulates the expression of the Th cell signature cytokines IFN-gamma, IL-10 and IL-22. J Immunol 186:708–821
Xu QQ, Li RG, Monte MM, Jiang YS, Nie P, Holland JW, Secombes CJ, Wang TH (2014) Sequence and expression analysis of rainbow trout CXCR2, CXCR3a and CXCR3b aids interpretation of lineage-specific conversion, loss and expansion of these receptors during vertebrate evolution. Dev Comp Immunol 45:201–213
Abós B, Castro R, Pignatelli J, Luque A, González L, Tafalla C (2013) Transcriptional heterogeneity of IgM + cells in rainbow trout (Oncorhynchus mykiss) tissues. PLoS ONE 8:e82737
Kumar G, Menanteau-Ledouble S, Saleh M, EI-Matbouli M (2015) Yesinia ruckeri, the causative agent of enteric redmouth disease in fish. Vet Res 46:103
Gorgoglione B, Wang TH, Secombes CJ, Holland JW (2013) Immune gene expression profiling of proliferative kidney disease in rainbow trout Oncorhynchus mykiss reveals a dominance of anti-inflammatory, antibody and T helper cell-like activities. Vet Res 44:55
Clifton-Hadley RS, Bucker D, Richards RH (1987) A study of the sequential clinical and pathological changes during proliferative kidney disease in rainbow trout, Salmo gairdneri Richardson. J Fish Dis 10:335–352
Montero J, Garcia J, Ordas MC, Casanova I, Gonzalez A, Villena A, Coll J, Tafalla C (2011) Specific regulation of the chemokine response to viral hemorrhagic septicemia virus at the entry site. J Virol 85:4046–4056
Cuesta A, Tafalla C (2009) Transcription of immune genes upon challenge with viral hemorrhagic septicemia virus (VHSV) in DNA vaccinated rainbow trout (Oncorhynchus mykiss). Vaccine 27:280–289
Costa MM, Maehr T, Diaz-Rosales P, Secombes CJ, Wang TH (2011) Bioactivity studies of rainbow trout (Oncorhynchus mykiss) interleukin-6: effects on macrophage growth and antimicrobial peptide gene expression. Mol Immunol 48:1903–1916
Hong S, Li RG, Secombes CJ, Wang TH (2013) Two types of TNF-α exist in teleost fish: phylogeny, expression, and bioactivity analysis of type-II TNF-α3 in rainbwo trout Oncorhynchus mykiss. J Immunol 191:5959–5972
Harun NO, Wang TH, Secombes CJ (2011) Gene expression profiling in naïve and vaccinated rainbow trout after Yersinia ruckeri infection: insights into the mechanisms of protection seen in vaccinated fish. Vaccine 29:4388–4399
Raida MK, Holten-Andersen L, Buchmann K (2011) Association between Yersinia ruckeri infection, cytokine expression and survival in rainbow trout (Oncorhynchus mykiss). Fish Shellfish Immunol 30:1257–1264
Qi ZT, Jiang YS, Holland JW, Nie P, Secombes CJ, Wang TH (2015) Identification and expression analysis of an atypical chemokine receptor-2 (ACKR2)/CC chemokine binding protein-2 (CCBP2) in rainbow trout (Oncorhynchus mykiss). Fish Shellfish Immunol 44:389–398
Aquilino C, Castro R, Fisher U, Tafalla C (2014) Transcriptomic response in rainbow trout gills upon infection with viral hemorrhagic septicemia virus (VHSV). Dev Comp Immunol 44:12–20
Savan R, Ravichandran S, Collins JR, Sakai M, Young HA (2009) Structural conservation of interferon γ among vertebrates. Cytokine Growth Factor Rev 20:115–124
Costa MM, Maehr T, Diaz-Rosales P, Secombes CJ, Wang T (2011) Bioactivity studies of rainbow trout (Oncorhynchus mykiss) interleukin-6: effects on macrophage growth and antimicrobial peptide gene expression. Mol Immunol 48:1903–1916
Arita M, Bianchini F, Aliberti J, Sher A, Chiang N, Hong S, Zhang R, Petasis NA, Serhan CN (2005) Stereochemical assignment, antiinflammatory properties, and receptors for the omega-3 lipid mediator resolvin E1. J Exp Med 201:713–722
Acknowledgments
This work was supported by the Scottish Funding Council for the MASTS (the Marine Alliance for Science and Technology for Scotland) (grant reference HR09011) and partially by grants from the National Natural Science Foundation of China (31302221, 31272666, and 31470130). ZQ was financially supported by the “Qinglan” project of Jiangsu Province and the overseas training plan for young and middle-aged teachers and principals of colleges and universities in Jiangsu Province, China. We thank Professor Mingxian Chang, State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, the Chinese Academy of Sciences, for her kind comments on the phylogenetic tree analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qi, Z., Zhang, Q., Holland, J.W. et al. Characterization and expression analysis of chemokine-like receptor 3 gene in rainbow trout Oncorhynchus mykiss . Fish Sci 82, 613–622 (2016). https://doi.org/10.1007/s12562-016-0997-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12562-016-0997-5