Abstract
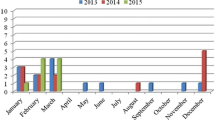
In this study, the prevalence of various enteric viruses in Mytilus galloprovincialis (Mediterranean mussel) belonging to class A and class B mollusc-harvesting areas in the Campania region in southern Italy was evaluated. One hundred and eight mussels were analysed using real-time reverse transcription PCR during a 2-year collection period (2014–2015) to detect the following viruses: human norovirus (genogroups I and II), rotavirus, astrovirus, sapovirus, aichivirus, hepatitis A virus and hepatitis E virus. Overall, 50.93% of mussels were contaminated by at least one of the tested viruses. Of these virus-positive mussels, 63.63% were contaminated by two or more viruses. In 2014, only three of the eight investigated viruses were detected: astrovirus, sapovirus and aichivirus, whereas in 2015, seven of the eight viruses were detected (only hepatitis E virus was not identified). Astrovirus was the most frequently detected virus in both sampling periods. In 2014, sapovirus was detected at the same frequency as astrovirus (16.00%), followed by aichivirus (8%). In 2015, astrovirus (32.53%) was most frequently detected, followed by norovirus GII (26.50%), sapovirus (18.07%), hepatitis A virus (16.87%), rotavirus (16.87%), aichivirus (13.25%) and norovirus GI (12.05%).This study describes, for the first time, the presence of aichivirus and sapovirus in mussels in Italy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sanitary controls of bivalve molluscs are based, in compliance with the European Union (EU) regulations (Reg. EC 854/2004, Reg. EC 2073/2005), on the detection of bacterial indicators such as Escherichia coli and do not require detection of any virus, neglecting the well-known role of molluscs as a vehicle for transmission of enteric viruses (Varela et al. 2016; Polo et al. 2010). Several studies showed that bacteria are not representative of the level of viral contamination, because viruses have a higher environmental resistance and are more resistant to depuration (Fusco et al. 2013; Polo et al. 2014b; Varela et al. 2016). For this reason, mussels cannot be guaranteed to be virus free even when EU standards on microbiological quality are met. Indeed, bivalves, because of their filter-feeding nature, retain and concentrate viruses present in their growing areas polluted by faecal contamination, thus becoming a risky food especially when eaten raw or undercooked (Rizzo et al. 2007; Alfano-Sobsey et al. 2012). It is widely reported in the literature that bivalves, such as oysters and mussels, are often contaminated by enteric viruses (Henigman et al. 2015; Mesquita et al. 2016; Varela et al. 2016), and their consumption is frequently involved in the occurrence of viral outbreaks in human (Rizzo et al. 2007; Le Guyader et al. 2008; Alfano-Sobsey et al. 2012; Rajko-Nenow et al. 2014; Boxman et al. 2016). Therefore, the scope of this study was to assess the presence of enteric viruses in Mytilus galloprovincialis (Mediterranean mussel) sampled from various class A and class B production areas in the Campania region in southern Italy. The following viruses were tested: astrovirus (AsV), aichivirus (AiV), norovirus genogroups I and II (NoVGI, NoVGII), rotavirus (RV), sapovirus (SaV), hepatitis A virus (HAV) and hepatitis E virus (HEV). Among the investigated viruses, NoV is the most frequently associated with foodborne gastroenteritis (Le Guyader et al. 2006, 2008) and is often linked to the consumption of contaminated shellfish. HAV, although much less frequently detected, causes a more serious pathology, which can also be fatal (Polo et al. 2014a, 2015). AsV, RV and SaV are considered important causes of gastroenteritis in children younger than 5 years of age (Rovida et al. 2013), although an increasing number of cases involving adults have been described in the last several years (Le Guyader et al. 2008; Rovida et al. 2013; Arena et al. 2014). These viruses have been detected in the faeces of symptomatic/asymptomatic individuals, as well as in environmental samples (e.g., surface water, untreated and treated wastewaters, river water, sewage and bivalve molluscs), suggesting that they are widespread in the environment (Le Guyader et al. 2008; van Maarseveen et al. 2010; Rovida et al. 2013; Iritani et al. 2014; Varela et al. 2016; Ruggeri et al. 2015). Little information is available on the epidemiology of AiV, several studies reported the detection of AiV in environmental samples such as sewage and surface water (Di Martino et al. 2013; Lodder et al. 2013) as well as in faecal specimens from patients affected by gastroenteritis (Ambert-Balay et al. 2008). HEV can be shed in animal and human faeces and has been detected in sewage samples (La Rosa et al. 2010) and less frequently in mussels (Mesquita et al. 2016; Diez-Valcarce et al. 2012; Donia et al. 2012). The present study showed a wide occurrence of enteric viruses in the analysed samples, confirming that the use of E. coli alone as an indicator of faecal contamination is insufficient to guarantee the microbiological safety of mussels. These findings may be helpful to evaluate the use of the tested viruses as potential indicators of faecal contamination of mussels.
Materials and Methods
Sampling
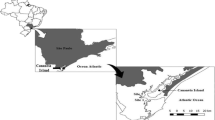
A total of 108 mussels (Mytilus galloprovincialis) were collected from the Campania region in southern Italy from local veterinary services over a 24-month period (January 2014–December 2015). Twenty-five batches of mussels were collected in 2014 and 83 in 2015. Sampling was carried out in both class A (values of E. coli ≤230 Most Probable Number, (MPN)/100 g of flesh and intravalvular liquid, Reg. EC 854/2004) and class B (values of E. coli ≤4600 MPN/100 g) areas, as classified by the EU regulation.
Viral Recovery and RNA Extraction
Viral DNA and RNA extraction was carried out according to the ISO/TS 15216-1:2013 using mengovirus clone (vMC0) as a nucleic acid extraction efficiency control (Costafreda et al. 2006). Two grams of hepatopancreas (approximately 10 mussels) was recovered by dissection from pooled mussels. Each pooled hepatopancreas sample was spiked with 10 µl of vMC0 stock (1.6 × 103 tissue culture infectious dose (TCID50)/ml), then 2 ml of proteinase K solution (0.1 mg/ml) (Qiagen) was added and the sample was shaken (320 rpm for 1 h/37 °C) and incubated (15 min/60 °C). Samples were subsequently centrifuged (3000×g/5 min), and the supernatant immediately used for nucleic acid extraction. Nucleic acids were extracted using the MagMAX Express automatic system (Applied Biosystems) with the viral RNA/DNA Kit according to the manufacturers’ instructions. Nucleic acids were eluted in 90 µl of elution buffer containing 40 unit/µl RNase inhibitor (Promega) and immediately analysed by real-time reverse transcription PCR (RT-qPCR) or stored at −80 °C until use.
Real-Time RT-qPCR for Virus Detection
Real-time reverse transcription quantitative polymerase chain reaction (RT-qPCR) was performed on a 7500 Fast Real-Time PCR system thermocycler (Applied Biosystems). The RNA UltraSense reaction kit (Thermo Fisher Scientific) was used in reactions of 25 µl final volume, with 5 µl of RNA extract as the target. The extraction efficiency (R) for each sample was determined as previously reported (Polo et al. 2015) by comparing the threshold cycle value of vMC0 control with the threshold cycle value of vMC0 in the sample. Samples with R > 5% were considered correctly extracted. Each viral target was analysed in triplicate with primers (Tema Ricerca) and probe (Thermo Fisher Scientific) specific for the virus tested. HAV, NoV genogroup I (GI) and NoV genogroup II (GII) were detected using the protocol specified in the ISO/TS 15216-1:2013. The other viral targets were analysed as previously described: HEV (Jothikumar et al. 2006), AiV (Kitajima et al. 2013) and RV (Zeng et al. 2008). Detection of AsV and SaV was performed using commercial kits, Astrovirus RT-PCR kit and Sapovirus RT-PCR kit (ceeramTOOLS, bioMérieux).
Results were analysed as follows: when at least one replicate (out of three performed) showed a C T value ≤40, the sample was considered positive. Samples negative or with C T values >40 were retested tenfold diluted to evaluate possible presence of inhibitor. Quantification was carried out using standard curves constructed by amplifying serial dilutions of the quantified standard positive control (standard plasmid DNA from 1–1 × 106 copies/reaction). Log genome copies were plotted against CT number and results were expressed as the number of genome copies per gram of digestive tissue (copies/g DT). Amplification efficiency (E) for each standard curve was calculated as previously described (Amoroso et al. 2011). Among the viral targets tested, only AiV was not quantified since only one field positive stool sample was available. Quantitative analysis was carried out separately on positive samples.
HEV-positive control was a kind gift of the Federal Research Institute for Animal Health “FLI” (Germany). HAV, NoVGI, NoVGII and RV plasmids were kindly provided by the Italian National Reference Laboratory for monitoring of viral contamination of bivalve molluscs (Istituto Superiore di Sanità, Rome, Italy). Astrovirus and Sapovirus Q standards were purchased from ceeramTOOLS (bioMérieux). An AiV positive control (a human stool positive sample) was kindly provided by the Centre National de Référence des Virus Entériques (Dijon Cedex, France).
Sequencing and phylogenetic analysis
All positive samples exhibiting a threshold cycle ≤38 underwent sequence analysis. Samples with 38 < CT ≤ 40 were considered positive, but not sufficiently concentrated to allow for sequencing. Positive samples were amplified by conventional RT-PCR using the OneStep RT-PCR kit (Qiagen), with different primer mixtures for each virus, according to previously described protocols (Gouvea et al. 1990; Vinje and Koopmans 1996; Le Guyader et al. 1996; Noel et al. 1995; Varela et al. 2016). RT-PCR amplicons were purified using Exonuclease I and shrimp alkaline phosphatase (ExoSAP-IT, Affymetrix, USB). Nucleotide sequencing of amplified genes was performed as previously described (Amoroso et al. 2013). Sequences were analysed and corrected with Chromas Pro2.23 software (Technelysium, Queensland, Australia). Phylogenetic analysis was performed using MEGA6 software (www.megasoftwares.com) (Tamura et al. 2013). Phylogenetic trees were built using the maximum likelihood and neighbour joining (for NoV) methods using the Tamura-Nei substitution model, as suggested by Mega 6 Model Test. Sequences of the viral strains identified in this study were deposited in the GenBank database as follows: RVA: KX027300–KX027306; AsV: KX027307 and KX027308; SaV: KX467788. NoV sequences were shorter than 200 bp and could not be submitted to the GenBank database.
Statistical Analysis
All statistical analyses were carried out using IBM® SPSS® Statistics 21 software (IBM Corp., Armonk, NY, USA). Pearson’s Chi-squared test, with one degree of freedom, was performed to evaluate correlations among viruses. P values <0.05 were considered statistically significant.
Results
Virus Detection by Real-Time RT-qPCR
All RNA extractions conducted in this study achieved acceptable recovery rates (>5%). In 2014, the viruses identified in the 25 samples collected were AsV, SaV and AiV (Table 1), while in the 83 samples collected in 2015, all tested viruses were detected in at least one sample excluding HEV, with AsV as the most frequently detected virus (Table 1). Overall, our results showed that 50.92% of mussels (55/108) were contaminated by at least one of the viruses tested. Among the 55 samples, 20 were contaminated with one virus (18.51% of the total), while 35 samples were contaminated with at least two viruses; of the latter, 16 samples (14.81% of the total) were contaminated with 3 or more viruses. In the 2-year study period, the most frequently detected virus was AsV (28.70% of positive samples), followed by NoVGII (20.37% of positive samples). The least frequently detected virus was NoVGI (9.26%). NoV (GI and/or GII) was detected in 25 out of 108 samples, and HAV was detected in 14 samples. With respect to class production areas, a 42.10% positivity was observed in samples belonging to class A (8 positive samples out of 19 analysed), a 52.81% positivity (47/89 samples) in samples collected in class B. The difference in prevalence of virus positivity observed between mussels harvested in class A and class B, does not correspond to differences in quantitative amount of virus detected.
Standard Curves and Virus Quantification
For each virus, a standard curve was constructed based on the average cycle threshold values of three replicates against the amount of plasmid/reaction. The efficiencies of amplification (E = 10−1/S–1, where S is the slope of the linear regression curve) for each standard curve were as follows: EHAV = 1.18 (S = −2.96, r 2 = 0.99); E NoVGI = 1.11 (S = −3.07, r 2 = 0.99); E NoVGII = 1.04 (S = −3.23, r 2 = 0.99); EAsV = 0.95 (S = −3.43, r 2 = 0.99); E SaV = 0.96 (S = −3.43, r 2 = 0.99); and E RV = 1.13 (S = −3.04, r 2 = 0.99). The estimated limit of detection for each target virus was 5 copies/reaction.
Quantitative Real-time RT-qPCR was used to estimate the genome copies of positive samples. The mean quantities for most of the viruses ranged between 102 and 103 genome copies/g DT; only NoVGII and RV were found at higher levels, reaching 107 genome copies/g DT (3 samples, threshold cycles: 17.71, 18.09 and 19.13) and 105 genome copies/g DT (10 samples), respectively. NoVGII yielded the highest average level (1.77 × 107 genome copies/g DT) with quantities ranging from 1.18 × 102 genome copies/g DT to 5.82 × 107 genome copies/g DT. RV showed an average quantity of 1.34 × 105 genome copies/g DT, followed by AsV (5.58 × 102 genome copies/g DT), SaV (4.04 × 102 genome copies/g DT), HAV (2.31 × 102 genome copies/g DT) and NoVGI (1.02 × 102 genome copies/g DT). Notably, the average value for NoVGII was much lower (0.57 × 102) when excluding the three highest positive samples from calculations.
Phylogenetic Analysis of RV, AsV, NoV and SaV
Seven RV strains identified by real-time RT-PCR were further characterised by sequence and phylogenetic analyses. These seven strains showed approximately 97.5–99.5% nucleotide identity among one another. Six of them were identified as rotavirus A (RVA) belonging to genotype G1, lineage I, and clustered in two different branches. One strain was identified as belonging to genotype G3. The strains demonstrated the highest nucleotide identities with RVA strains previously characterised in humans in Southeast Asia (India, Indonesia, Japan, Thailand and Bangladesh). These strains were less related to G1 RVA strains previously detected in the same area of Italy from both faecal and environmental samples. Only 3 out of 22 NoVGII-positive samples yielded good end-point PCR results and were therefore additionally investigated by sequence analysis. The NoVGII strains belonged to genotypes GII.4 and GII.Pe. Phylogenetic analysis showed a strict correlation (97.4%) between the strains detected in mussels and NoV strains detected in human samples previously reported in Italy and Europe. Two AsV-positive samples were sequenced and compared with other sequences available in the GenBank database. These two strains were classified as human AsV genotype 1 and genotype 2, respectively, and shared 98.5% nucleotide identity with human AsV strains. One SaV-positive sample by real-time RT-PCR was also found to be SaV-positive by nested PCR. This strain was classified as human SaV genotype GI.1 and shared 99% nucleotide identity with other human SaV strains.
Correlation Among Viruses
Using Pearson’s Chi-squared test, a possible correlation among the tested viruses was evaluated (Table 2). The statistical analyses revealed the highest correlations (>0.30) among presence of hepatitis A and rotavirus or astrovirus (040 and 0.31, respectively); and among NoVGI and NoVGII, RV and AsV (0.52, 0.43 and 0.36, respectively). Similar correlation (<0.30) was observed between all the other viruses. Furthermore, the simultaneous occurrence of other viruses in HAV-positive samples was evaluated. Since none of the samples collected in 2014 were HAV-positive, only samples harvested in 2015 were included in the analyses. Interestingly, all 14 HAV-positive samples (named HAV.P) were positive for at least one other virus (Table 3). Specifically, five HAV.P samples were positive for one other virus (AsV and RV); four HAV.P samples were positive for two other viruses (NoVGII/SaV; NoVGII/RV; and NoVGI/AsV); and five HAV.P samples were positive for three or more viruses.
Discussion
In the present study, the occurrence of enteric viruses in 108 mussels, collected in the gulf of Naples, was analysed. As reported in previous studies conducted in Europe (La Rosa et al. 2012; Fusco et al. 2013; Suffredini et al. 2014), this study confirmed that NoVs are more frequently widespread in mussels compared with HAV. NoV detected shared the highest degree of nucleotide identity with genotypes GII.4 and GII.Pe. These genotypes, prevalent worldwide, are frequently associated with the occurrence of human outbreaks and have been less frequently detected in mussels, where detection of genogroup I is more common (Kittigul et al. 2016).
In agreement with previous studies (Gabrieli et al. 2007; Le Guyader et al. 2008), several mussel samples were contaminated by AsV. Two AsV-positive samples, were classified by sequencing as human AsV (HAsV) genotypes 1 and 2. HAsV-1 is predominant in Italy, while the presence of HAsV-2 is rare and only recently detected in Italy (De Grazia et al. 2011).
NoV and RV were not detected in 2014. This is an unexpected result since both viruses are frequently detected in mussels, especially during the winter season. There is no explanation for this result, since the same samples were positive for SaV, HAsV and AiV confirming the occurrence of human faecal contamination.
A possible correlation among the viruses detected was also evaluated, since presence of more than one virus in the same samples was frequent. Correlations between the presence of HAV and the presence of RV, AsV, NoVGI and NoVGII were observed. The occurrence of NoVGI and NoVGII was strictly correlated with the presence of AsV, RV and SaV. Further studies are needed to confirm the simultaneous occurrence of these viruses in mussels, to evaluate whether a quantitative correlation also exists.
Interestingly, the study highlighted the presence of two emerging viruses in mussels associated with human gastroenteritis: SaV and AiV and described the first identification of SaV in mussels harvested in Italy. Identification of SaV in shellfish remains rare, although in the last years the number of human cases has increased. SaV has been previously reported in Moroccan shellfish (Benabbes et al. 2013), individual clams in Japan (Iizuka et al. 2013) and Japanese oysters meeting the legal requirements for raw consumption (Ueki et al. 2010). Recently, SaV was identified for the first time also in shellfish in Europe, in particular mussels, clams and cockles harvested in Spain (Varela et al. 2016).
AiV has been detected in oysters associated with gastroenteritis outbreaks in Japan (Iritani et al. 2014), in shellfish in Tunisia (Sdiri-Loulizi et al. 2010) and in oysters associated with the occurrence of a gastroenteritis outbreak in France (Le Guyader et al. 2008). In the present study, presence of AiV in mussels was reported (Mytilus galloprovincialis), and, to our knowledge, this finding represents the first identification of AiV in mussels worldwide. Both AiV and SaV were detected in mussels at a moderate frequency (12.04 and 17.59%, respectively), suggesting the circulation of these viruses in the environment and their possible active role in human gastroenteritis.
Mussels contaminated by viruses were harvested in both the production class areas. Even though the number of samples investigated in the two classes is different (19 and 89, respectively), it is a major concern that the different positivity observed (42.10% class A and 52.81% class B) is not so marked as expected, showing a warring level of contamination also in samples harvested in class A area. Overall data raise questions about the safety of mussels analysed, since those belonging to class A are considered ready for consumption and those from class B require a process of depuration, which is known to be ineffective against viruses (Da Silva et al. 2007; Maalouf et al. 2010; Polo et al. 2014b).
These findings confirm that use the presence of E. coli to evaluate faecal pollution in mussels is not sufficient to guarantee the absence of viral contamination. Similarly, other studies reported the occurrence of NoV- and HAV-positive molluscs harvested from areas considered to be in compliance with the legal requirements (Baker et al. 2011; Guillois-Becel et al. 2009; Le Guyader et al. 2010). To guarantee the safety of consumable shellfish, the current legislation on the use of bacterial indicators alone should be revised, as previously suggested (Baker et al. 2011; Le Guyader et al. 2010; Varela et al. 2016). In the interim, an effective strategy for the protection of consumer’s health could involve surveillance and increased research of enteric viruses in mussels.
References
Alfano-Sobsey, E., Sweat, D., Hall, A., Breedlove, F., Rodriguez, R., Greene, S., et al. (2012). Norovirus outbreak associated with undercooked oysters and secondary household transmission. Epidemiology and Infection, 140(2), 276–282.
Ambert-Balay, K., Lorrot, M., Bon, F., Giraudon, H., Kaplon, J., Wolfer, M., et al. (2008). Prevalence and genetic diversity of Aichi virus strains in stool samples from community and hospitalized patients. Journal of Clinical Microbiology, 46(4), 1252–1258.
Amoroso, M. G., Corrado, F., De Carlo, E., Lucibelli, M. G., Martucciello, A., Guarino, A., et al. (2013). Bubaline herpesvirus 1 associated with abortion in a mediterranean water buffalo. Research in Veterinary Science, 94(3), 813–816.
Amoroso, M. G., Salzano, C., Cioffi, B., Napoletano, M., Garofalo, F., Guarino, A., et al. (2011). Validation of a Real–time PCR assay for fast and sensitive quantification of Brucella spp. in water buffalo milk. Food Control, 22, 1466–1470.
Arena, C., Amoros, J. P., Vaillant, V., Ambert-Balay, K., Chikhi-Brachet, R., Jourdan-Da Silva, N., et al. (2014). Acute diarrhea in adults consulting a general practitioner in France during winter: Incidence, clinical characteristics, management and risk factors. BMC Infectious Diseases, 14, 574-014–574-044.
Baker, K., Morris, J., McCarthy, N., Saldana, L., Lowther, J., Collinson, A., et al. (2011). An outbreak of norovirus infection linked to oyster consumption at a UK restaurant. Journal of Public Health (Oxford, England), 33(2), 205–211.
Benabbes, L., Ollivier, J., Schaeffer, J., Parnaudeau, S., Rhaissi, H., Nourlil, J., et al. (2013). Norovirus and other human enteric viruses in Moroccan shellfish. Food and Environmental Virology, 5(1), 35–40.
Boxman, I. L., Verhoef, L., Vennema, H., Ngui, S. L., Friesema, I. H., Whiteside, C., et al. (2016). International linkage of two food-borne hepatitis A clusters through traceback of mussels, the Netherlands, 2012. Euro Surveillance: Bulletin European Sur Les Maladies Transmissibles = European Communicable Disease Bulletin, 21(3), 30113-7917.ES.2016.21.3.30113.
Costafreda, M. I., Bosch, A., & Pinto, R. M. (2006). Development, evaluation, and standardization of a real-time TaqMan reverse transcription-PCR assay for quantification of hepatitis A virus in clinical and shellfish samples. Applied and Environmental Microbiology, 72(6), 3846–3855.
Da Silva, A. K., Le Saux, J. C., Parnaudeau, S., Pommepuy, M., Elimelech, M., & Le Guyader, F. S. (2007). Evaluation of removal of noroviruses during wastewater treatment, using real-time reverse transcription-PCR: Different behaviors of genogroups I and II. Applied and Environmental Microbiology, 73(24), 7891–7897.
De Grazia, S., Platia, M. A., Rotolo, V., Colomba, C., Martella, V., & Giammanco, G. M. (2011). Surveillance of human astrovirus circulation in Italy 2002–2005: Emergence of lineage 2c strains. Clinical Microbiology & Infection, 17(1), 97–101.
Di Martino, B., Di Profio, F., Ceci, C., Di Felice, E., & Marsilio, F. (2013). Molecular detection of aichi virus in raw sewage in Italy. Archives of Virology, 158(9), 2001–2005.
Diez-Valcarce, M., Kokkinos, P., Soderberg, K., Bouwknegt, M., Willems, K., de Roda-Husman, A. M., et al. (2012). Occurrence of human enteric viruses in commercial mussels at retail level in three European countries. Food and Environmental Virology, 4(2), 73–80.
Donia, D., Dell’Amico, M. C., Petrinca, A. R., Martinucci, I., Mazzei, M., Tolari, F., et al. (2012). Presence of hepatitis E RNA in mussels used as bio-monitors of viral marine pollution. Journal of Virological Methods, 186(1–2), 198–202.
Fusco, G., Aprea, G., Galiero, G., Guarino, A., & Viscardi, M. (2013). Escherichia coli, Salmonella spp., hepatitis A virus and norovirus in bivalve molluscs in southern italy. Veterinaria Italiana, 49(1), 55–58.
Gabrieli, R., Macaluso, A., Lanni, L., Saccares, S., Di Giamberardino, F., Cencioni, B., et al. (2007). Enteric viruses in molluscan shellfish. The New Microbiologica, 30(4), 471–475.
Gouvea, V., Glass, R. I., Woods, P., Taniguchi, K., Clark, H. F., Forrester, B., et al. (1990). Polymerase chain reaction amplification and typing of rotavirus nucleic acid from stool specimens. Journal of Clinical Microbiology, 28(2), 276–282.
Guillois-Becel, Y., Couturier, E., Le Saux, J. C., Roque-Afonso, A. M., Le Guyader, F. S., Le Goas, A., et al. (2009). An oyster-associated hepatitis A outbreak in France in 2007. Euro Surveillance: Bulletin Europeen Sur Les Maladies Transmissibles = European Communicable Disease Bulletin, 14(10), 19144.
Henigman, U., Biasizzo, M., Vadnjal, S., Toplak, I., Gombac, M., Steyer, A., et al. (2015). Molecular characterisation of noroviruses detected in mussels (Mytilus galloprovincialis) from harvesting areas in slovenia. The New Microbiologica, 38(2), 225–233.
Iizuka, S., Takai-Todaka, R., Ohshiro, H., Kitajima, M., Wang, Q., Saif, L. J., et al. (2013). Detection of multiple human sapoviruses from imported frozen individual clams. Food and Environmental Virology, 5(2), 119–125.
Iritani, N., Kaida, A., Abe, N., Kubo, H., Sekiguchi, J., Yamamoto, S. P., et al. (2014). Detection and genetic characterization of human enteric viruses in oyster-associated gastroenteritis outbreaks between 2001 and 2012 in osaka city, japan. Journal of Medical Virology, 86(12), 2019–2025.
Jothikumar, N., Cromeans, T. L., Robertson, B. H., Meng, X. J., & Hill, V. R. (2006). A broadly reactive one-step real-time RT-PCR assay for rapid and sensitive detection of hepatitis E virus. Journal of Virological Methods, 131(1), 65–71.
Kitajima, M., Hata, A., Yamashita, T., Haramoto, E., Minagawa, H., & Katayama, H. (2013). Development of a reverse transcription-quantitative PCR system for detection and genotyping of aichi viruses in clinical and environmental samples. Applied and Environmental Microbiology, 79(13), 3952–3958.
Kittigul, L., Thamjaroen, A., Chiawchan, S., Chavalitshewinkoon-Petmitr, P., Pombubpa, K., & Diraphat, P. (2016). Prevalence and molecular genotyping of noroviruses in market oysters, mussels, and cockles in bangkok, Thailand. Food and Environmental Virology, 8(2), 133–140.
La Rosa, G., Fratini, M., Spuri Vennarucci, V., Guercio, A., Purpari, G., & Muscillo, M. (2012). GIV noroviruses and other enteric viruses in bivalves: A preliminary study. The New Microbiologica, 35(1), 27–34.
La Rosa, G., Pourshaban, M., Iaconelli, M., Vennarucci, V. S., & Muscillo, M. (2010). Molecular detection of hepatitis E virus in sewage samples. Applied and Environmental Microbiology, 76(17), 5870–5873.
Le Guyader, F. S., Bon, F., DeMedici, D., Parnaudeau, S., Bertone, A., Crudeli, S., et al. (2006). Detection of multiple noroviruses associated with an international gastroenteritis outbreak linked to oyster consumption. Journal of Clinical Microbiology, 44(11), 3878–3882.
Le Guyader, F. S., Krol, J., Ambert-Balay, K., Ruvoen-Clouet, N., Desaubliaux, B., Parnaudeau, S., et al. (2010). Comprehensive analysis of a norovirus-associated gastroenteritis outbreak, from the environment to the consumer. Journal of Clinical Microbiology, 48(3), 915–920.
Le Guyader, F. S., Le Saux, J. C., Ambert-Balay, K., Krol, J., Serais, O., Parnaudeau, S., et al. (2008). Aichi virus, norovirus, astrovirus, enterovirus, and rotavirus involved in clinical cases from a french oyster-related gastroenteritis outbreak. Journal of Clinical Microbiology, 46(12), 4011–4017.
Le Guyader, F., Neill, F. H., Estes, M. K., Monroe, S. S., Ando, T., & Atmar, R. L. (1996). Detection and analysis of a small round-structured virus strain in oysters implicated in an outbreak of acute gastroenteritis. Applied and Environmental Microbiology, 62(11), 4268–4272.
Lodder, W. J., Rutjes, S. A., Takumi, K., & de Roda Husman, A. M. (2013). Aichi virus in sewage and surface water, the Netherlands. Emerging Infectious Diseases, 19(8), 1222–1230.
Maalouf, H., Zakhour, M., Le Pendu, J., Le Saux, J. C., Atmar, R. L., & Le Guyader, F. S. (2010). Distribution in tissue and seasonal variation of norovirus genogroup I and II ligands in oysters. Applied and Environmental Microbiology, 76(16), 5621–5630.
Mesquita, J. R., Oliveira, D., Rivadulla, E., Abreu-Silva, J., Varela, M. F., Romalde, J. L., et al. (2016). Hepatitis E virus genotype 3 in mussels (Mytilus galloprovinciallis), Spain. Food Microbiology, 58, 13–15.
Noel, J. S., Lee, T. W., Kurtz, J. B., Glass, R. I., & Monroe, S. S. (1995). Typing of human astroviruses from clinical isolates by enzyme immunoassay and nucleotide sequencing. Journal of Clinical Microbiology, 33(4), 797–801.
Polo, D., Álvarez, C., Díez, J., Darriba, S., Longa, Á., & Romalde, J. L. (2014a). Viral elimination during commercial depuration of shellfish. Food Control, 43, 206–212.
Polo, D., Avarez, C., Vilarino, M. L., Longa, A., & Romalde, J. L. (2014b). Depuration kinetics of hepatitis A virus in clams. Food Microbiology, 39, 103–107.
Polo, D., Varela, M. F., & Romalde, J. L. (2015). Detection and quantification of hepatitis A virus and norovirus in Spanish authorized shellfish harvesting areas. International Journal of Food Microbiology, 193, 43–50.
Polo, D., Vilarino, M. L., Manso, C. F., & Romalde, J. L. (2010). Imported mollusks and dissemination of human enteric viruses. Emerging Infectious Diseases, 16(6), 1036–1038.
Rajko-Nenow, P., Keaveney, S., Flannery, J., McIntyre, A., & Dore, W. (2014). Norovirus genotypes implicated in two oyster-related illness outbreaks in ireland. Epidemiology and Infection, 142(10), 2096–2104.
Rizzo, C., Di Bartolo, I., Santantonio, M., Coscia, M. F., Monno, R., De Vito, D., et al. (2007). Epidemiological and virological investigation of a norovirus outbreak in a resort in Puglia, Italy. BMC Infectious Diseases, 7, 135.
Rovida, F., Campanini, G., Piralla, A., Adzasehoun, K. M., Sarasini, A., & Baldanti, F. (2013). Molecular detection of gastrointestinal viral infections in hospitalized patients. Diagnostic Microbiology and Infectious Disease, 77(3), 231–235.
Ruggeri, F. M., Bonomo, P., Ianiro, G., Battistone, A., Delogu, R., Germinario, C., et al. (2015). Rotavirus genotypes in sewage treatment plants and in children hospitalized with acute diarrhea in Italy in 2010 and 2011. Applied and Environmental Microbiology, 81(1), 241–249.
Sdiri-Loulizi, K., Hassine, M., Aouni, Z., Gharbi-Khelifi, H., Sakly, N., Chouchane, S., et al. (2010). First molecular detection of aichi virus in sewage and shellfish samples in the Monastir region of Tunisia. Archives of Virology, 155(9), 1509–1513.
Suffredini, E., Lanni, L., Arcangeli, G., Pepe, T., Mazzette, R., Ciccaglioni, G., et al. (2014). Qualitative and quantitative assessment of viral contamination in Bivalve molluscs harvested in Italy. International Journal of Food Microbiology, 184, 21–26.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., & Kumar, S. (2013). MEGA6: Molecular evolutionary genetics analysis version 6.0. Molecular Biology Evolution, 30(12), 2725–2729.
Ueki, Y., Shoji, M., Okimura, Y., Miyota, Y., Masago, Y., Oka, T., et al. (2010). Detection of sapovirus in oysters. Microbiology and Immunology, 54(8), 483–486.
van Maarseveen, N. M., Wessels, E., de Brouwer, C. S., Vossen, A. C., & Claas, E. C. (2010). Diagnosis of viral gastroenteritis by simultaneous detection of adenovirus group F, astrovirus, rotavirus group A, norovirus genogroups I and II, and sapovirus in two internally controlled multiplex real-time PCR assays. Journal of Clinical Virology, 49(3), 205–210.
Varela, M. F., Polo, D., & Romalde, J. L. (2016). Prevalence and genetic diversity of human sapoviruses in shellfish from commercial production areas in Galicia, Spain. Applied and Environmental Microbiology, 82(4), 1167–1172.
Vinje, J., & Koopmans, M. P. (1996). Molecular detection and epidemiology of small round-structured viruses in outbreaks of gastroenteritis in the Netherlands. The Journal of Infectious Diseases, 174(3), 610–615.
Zeng, S. Q., Halkosalo, A., Salminen, M., Szakal, E. D., Puustinen, L., & Vesikari, T. (2008). One-step quantitative RT-PCR for the detection of rotavirus in acute gastroenteritis. Journal of Virological Methods, 153(2), 238–240.
Funding
This work was supported by Grant RF-2011-02350023 from Italian Ministry of Health.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Fusco, G., Di Bartolo, I., Cioffi, B. et al. Prevalence of Foodborne Viruses in Mussels in Southern Italy. Food Environ Virol 9, 187–194 (2017). https://doi.org/10.1007/s12560-016-9277-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12560-016-9277-x