Abstract
Cellular DNA is constantly exposed to endogenous or exogenous factors that can induce lesions. Several types of lesions have been described that can result from UV/ionizing irradiations, oxidative stress, or free radicals, among others. In order to overcome the deleterious effects of such damages, i.e., mutagenicity or cytotoxicity, cells possess a highly complex DNA repair machinery, involving repair enzymes targeting specific types of lesions through dedicated cellular pathways. In addition, DNA is highly compacted in the nucleus, the first level of compaction consisting of ~ 147 DNA base pairs wrapped around a core of histones, the so-called nucleosome core particle. In this complex environment, the DNA structure is highly constrained, and fine-tuned mechanisms involving remodeling processes are required to expose the DNA to repair enzymes and to facilitate the damage removal. However, these nucleosome-specific mechanisms remain poorly understood, and computational methods emerged only recently as powerful tools to investigate DNA damages in such complex systems as the nucleosome. In this mini-review, we summarize the latest advances brought out by computational approaches in the field, opening new exciting perspectives for the study of DNA damage and repair in the nucleosome context.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA is continuously altered by different factors, from both endogenous and exogenous sources, that can be harmful to genome integrity. The cell’s defense relies on sophisticated repair pathways that involve recognition, scission, and ligation enzymes for bringing the DNA structure back to its canonical state. Depending on the damage to be repaired, different pathways can be activated that constitute the DNA damage response (DDR). Indeed, many different types of lesions have been described in vivo, and recognition enzymes are capable of distinguishing them, specifically activating the nucleobase excision repair (NER) pathway for bulky lesions (e.g., for UV-induced pyrimidine dimers), the base excision repair (BER) for more discrete nucleobase modifications (e.g., for oxidation, abasic sites, single-strand breaks), the mismatch repair (MMR) for base pair mismatches, the ribonucleotide excision repair (RER) for misincorporated ribonucleotides, and the non-homologous end-joining (NHEJ) or the homologous recombination (HR) for double-strand breaks, among others.
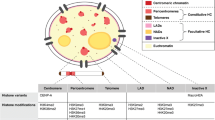
The mechanisms underlying the different repair pathways are now broadly understood, and both experimental and computational methods shed light on how the repair enzymes can detect the structural signature of the different lesions among the billions of canonical base pairs. Sensor proteins scan the genome, localize specific lesions, and trigger specific signaling pathways involving the activation of downstream kinases and effector proteins to halt the cell cycle and activate DNA repair. Noteworthy, dysregulation or mutations in DNA repair proteins can contribute to the development of diseases such as cancer and serious genetic diseases (e.g., Cockayne syndrome). While the molecular mechanisms driving lesion formation and repair have been well described in B-DNA, in the nucleus, the DNA undergoes a drastic compaction and is subjected to strong structural constraints which in turn modulates damage induction and repair (see Fig. 1). Indeed, the first level of DNA compaction involves its wrapping around an octamer of histone proteins, resulting in the so-called nucleosome core particle that comprises ~ 147 base pairs (McGinty and Tan 2015). Noteworthy, positional demarcations of the DNA sequence around the nucleosome structure, named superhelical locations (SHL), are conventionally used and refer to the coiling of the DNA strand around the histone octamer core with respect to the dyad position (SHL 0).
a DNA compaction levels in the nucleus. b Structure of a nucleosome, adapted from Smirnova et al. (2024). Superhelical locations (SHL) are marked in magenta. c Equilibrium between open and closed states of the chromatin. d Examples of DNA lesions (DPC, DNA–protein crosslink)
In this context, DDR is much more complex as the nucleosome has to undergo disassembly to expose DNA to repair proteins. As a matter of fact, little is known about the nucleosome-specific DDR mechanisms, which appear to involve epigenetic processes through the modulation of histone post-translational modifications (PTM), histone variant exchange, and the action of chromatin remodelers, as reviewed elsewhere (Polo and Almouzni 2015; Hauer and Gasser 2017; Smerdon et al. 2023). Many pending questions remain to be investigated to build up a better understanding of each sequential step of the damage recognition and repair, as well as the specificities of lesion formation in the context of the nucleosome which remain poorly defined.
In the last decades, computational studies of the nucleosome dynamics and its interactions with partner proteins or other nucleosomes multiplied thanks to outstanding advances in computational resources. Showcasing state-of-the-art techniques, these studies provided breakthrough insights into the nucleosome architecture and epigenetic processes. They allowed to capture motions taking place on microsecond to millisecond timescales such as nucleosomal DNA dynamics, inter-nucleosome interactions, ionic strength, and sequence effects. They also highlighted the importance of inter-NCP linker length and environment and the role of histone tails and their epigenetic marks (Collepardo-Guevara et al. 2015; Bowerman and Wereszczynski 2016; Lequieu et al. 2016; Armeev et al. 2019, 2021; Kono and Ishida 2020; Alvarado et al. 2021; Peng et al. 2021). Over the past 5 years, simulations of DNA damages within the nucleosome have multiplied, providing important first insights at the atomic scale improving our understanding of DDR in this context.
In this review, we put the emphasis on the insights from computational studies to unravel the complex mechanisms driving DNA damage formation and repair in the nucleosome. We describe how computational approaches have contributed to the field with respect to the damage type modeled and discuss the broad perspectives for future computer-based investigations. The increasing amounts of such investigations hold the promise of outstanding advances in the field, shedding light on the fine-tuned molecular mechanisms associated with DNA damage recognition and processing within the nucleosome.
Double-strand breaks
DNA double-strand breaks (DSB) are highly lethal damages consisting in the double cleavage of the DNA backbone on the two facing strands within a range of 10 base pairs, which can result from exposure to irradiation, chemical agents, or reactive oxygen species (ROS). Interestingly, it has been shown that the nucleosome compaction acts as a protective shield against DSB formation, especially around the dyad axis where the DNA-histone interactions are the strongest (Brambilla et al. 2020). Likewise, investigations of clustered oxidative lesions repair revealed that the nucleosome embedding hampers the endogenous formation of DSB during base excision repair (Cannan et al. 2014). However, DNA–protein crosslinks induced by histone tail reaction at clustered abasic sites favor the formation of DSB at specific superhelical location (SHL) on the nucleosome core particle (Banerjee et al. 2017). These results highlight the complexity DSB formation mechanisms within the nucleosome structure, most certainly driven by specific mechanical constraints.
The influence of DNA compaction is also observed in DSB repair (Seeber et al. 2013). Indeed, PTM (adenosine diphosphate (ADP)-ribosylation, acetylation, ubiquitination, phosphorylation, etc.), chromatin remodelers (poly(ADP-ribose) polymerase (PARP), inositol requiring 80 (INO80) complexes, etc.), and histone variants exchange (H2A.Z, H2A.X, etc.) must act in precisely orchestrated processes to ensure the nucleosome unwrapping for further exposure of the damage site to downstream repair enzymes (Gursoy-Yuzugullu et al. 2016; Karch et al. 2017; Sharma et al. 2019).
In this context, the dynamical release of DSB DNA ends from the histone core has been revealed by Cleri et al. (2018) relying on microsecond-range classical molecular dynamic (MD) simulations and umbrella sampling-based free energy calculations. The authors described how DSB cannot spontaneously fully detach from the histone core because of small free energy barriers of few kcal/mol to sequentially break the DNA–protein interactions around the lesion site. These calculations suggest a ~ 5-nm limitation of the distance that the DNA ends can easily reach with respect to the histone core center, imposed by the constraining action of the histone tails that prevents further opening. They revealed that the lesion mechanical signature is highly influenced by its interaction with the close-by histone tails at different locations: with inward/outward orientations at SHL − 2.5/ − 3, at the dyad (SHL 0), and at the DNA entry point (SHL − 7). Overall, they described the DSB as rather strongly attached to the histone core, with small fluctuations presuming the possible local detachment of short DNA sections on > 100 µs timescales that are nonetheless extremely constrained by interactions with the histone tails. This is in line with the necessity of remodelers and/or histone tails PTM to achieve any larger-scale release of the damaged DNA for exposure to downstream repair enzymes. Noteworthy, they performed molecular stress calculations showing that the circular bend of the nucleosomal DNA constitutes an internal stress that might be the driving force for spontaneous local opening, which might also be crucial for nucleosome remodeling processes.
These results constitute the first insights into the atomic-scale structural details of a DSB within the nucleosome environment, which provides important data towards the better understanding of these deleterious lesions recognition and processing. This is unfortunately the only computational study we could find for DSB, and such approaches could help gathering extensive atomic-scale data to further delineate the mechanisms behind the interplay between DSB repair and epigenetic marks, the DSB recognition by sensor proteins, or even their structural signature in higher-scale architectures.
Oxidative lesions
DNA oxidative lesions comprise a wide range of chemical species, due to the direct oxidation of DNA or the reactions with oxidative molecules such as ROS (Cadet et al. 2017). Due to its relatively low ionization potential, guanine is the most sensitive nucleobase towards oxidation. The radical cation guanine can evolve towards different products (see Fig. 2), the most frequent one being the 8-oxoguanine, whose redox potential is even lower. 8-Oxoguanine thus can be reoxidized to bulkier lesions such as the spiroiminodihydantoin and the guanidinohydantoin products. Oxidized nucleobases and their products can also react with the biological environment to form intrastrand or interstrand crosslinks with nucleobases, protein residues, or molecules.
The study of nucleosomal oxidation must face the issue of the competition between the radical reactivity of the oxidized DNA and the hole transfer along the strand in a very complex system. The DNA double helix is a semiconductor polymer whose electronic properties depend on its sequence, but also on its surrounding electrostatic field (Arnold et al. 2016; Peluso et al. 2019). The histone core and the flexible histone tails, which are rich in positively charged residues that can interact with the nucleobases, impact the redox properties of the nucleosomal DNA (Davis et al. 2012). In addition, the tyrosine residues can quench the charge transfer by attracting the radical hole. Experimentally, this leads to difficulties to predict and control the location of the damages and suppose the use of dedicated sequences with guanines tracts embedded in dA-dT-rich sequences (Sun et al. 2019; Wen et al. 2023) and specific cofactors (Núñez et al. 1999).
Computationally, it means that the highly complex dynamical behavior of the nucleosome must be taken into account to correctly describe the combinatorial effects (sequence, mechanical constraints, electrostatic environment) while the redox and charge transfers properties must be calculated at the quantum level, which dramatically increases the computational cost. Voityuk and Davis (2007) determined the redox properties of nucleosomal guanines by Neglect of Diatomic Differential Overlap with Gaussian-type orbital (NDDO-G) calculations to decipher the impact of the mechanical distortion due to the super helicoidal wrapping and of the proximal water or protein environment. They found that the inter-base pair parameters with the 3′-neighbor and their fluctuations have the biggest impact on the redox properties of the guanines, even more than the proximity of the guanidinium group of an arginine. Despite overlooking of the histone tail dynamic impact on the DNA, this study highlights the major role played by the local chromatin structure and its modulation in the formation of genomic oxidative damages.
Beyond the first oxidation of the guanine, the characteristics of 8-oxoguanine can be also explored by means of computational methods, as already described in B-DNA (Lee et al. 2012; Dršata et al. 2013; La Rosa and Zacharias 2016). Kılıç et al. (2023) have recently published a work in which they describe the impact of the nucleosome core on the 8-oxoguanine redox properties. In addition to microsecond timescale classical simulations, they used hybrid quantum mechanics/molecular mechanics (QM/MM) approaches to determine the oxidation free energy of the 8-oxoguanine site in the framework of the Marcus theory. They observed few structural changes in the nucleobase or its close neighboring before or after oxidation. They suggest that the detection of oxidative damages in compacted chromatin is driven by a DNA-mediated hole transfer from BER proteins, in agreement with some experimental data reviewed in Arnold et al. (2016). Nevertheless, recent studies focusing on the 8-oxoguanine DNA glycosylase 1 (OGG1) protein describe a detection mechanism based on conformational recognition and further flipping of the damages (Lukina et al. 2017; Shigdel et al. 2020; Jiang et al. 2021; Cintori et al. 2023), which can be extended to nucleosome thanks to local unwrapping and interaction with histone tails (Bilotti et al. 2017; Ren et al. 2021).
Overall, the DNA packing in chromatin can affect DNA redox properties and its propensity to undergo various oxidative lesions which can further evolve in bulkier, more mutagenic damages. The role of histone tails in both oxidation, damage formation, and repair has been observed experimentally but remains sparsely explored computationally as it requires the quantum description of the hole, which can become dramatically expensive in such a complex system. Fast QM/MM approaches for charge transfer calculation (Kubař and Elstner 2013; Wolter et al. 2017) and the development of dedicated machine learning algorithms (Krämer et al. 2020; Bag et al. 2020; Lee et al. 2022) open the door for the computational mapping of the DNA oxidation hotspots in the nucleosome.
DNA–protein crosslinks
DNA–protein crosslinks (DPC) are a diverse class of lesions which can be induced by a variety of processes (ionizing irradiation, ROS, cellular metabolism, etc.). They result from the covalent linkage between the DNA and a protein and constitute deleterious, bulky damages. Several repair pathways can remove them that mostly involve the processing of the three distinct DPC components: the covalent linkage hydrolysis, the excision of the damaged nucleotide, and the proteolytic repair (Zhang et al. 2020; Weickert and Stingele 2022). Interestingly, recent studies suggested a possible beneficial role of DPC in genome protection mechanisms, highlighting the high complexity of this lesion type (Kühbacher and Duxin 2020).
In the nucleosome, DPC can arise from the linkage between histone proteins and DNA and exhibit contrasted roles in both the initiation or the prevention of DNA repair depending on the circumstances (Zhou et al. 2012; Weng and Greenberg 2015; Ren et al. 2021). The Greenberg’s group has released a very large body of studies shedding light on histone-DNA crosslinking resulting from histone linkage with a variety of lesions, all elegantly summarized in a recent review (Ren et al. 2022). Their work delineated the basis of DNA-histone adduct formation and their possible implication in DNA damage and repair.
In recent computational studies, we described the dynamical behavior of single and clustered abasic sites in the nucleosome and mapped their interactions with proximal histone tails. Based on classical MD simulations starting from an X-ray structure featuring symmetric tetrahydrofuran at SHL 4.5 and − 4.5 (Osakabe et al. 2017), we described the dynamical signatures of canonical abasic sites at these positions (Bignon et al. 2020). We revealed the monotonous behavior of these isolated lesions in the constrained DNA helix, which differs from B-DNA and from control simulations with their tetrahydrofuran counterpart. Importantly, we were able to capture the key interactions between the histone H2A N-terminal tail and the lesions sites, pinpointing H2A K13 as a crucial residue. Such interaction might be important for the formation of DPC, as H2A is responsible for ~ 77% of these at SHL 4.7 (Sczepanski et al. 2013).
We later investigated the same features in the case of clustered abasic sites at the SHL 1.5 mutational hotspot (Bignon et al. 2021). Owing to MD simulations of the nucleosome bearing 2-base pair-distant abasic sites at this location, we highlighted how their cumulative effect resulted in a much stronger distortion of the DNA helix than what was observed for isolated lesions. We also described fast, stable interactions with histones H3 and H4 tails depending on the lesion position, again mapping the key amino acids that could be involved in DPC formation, which was in excellent agreement with the experimental observations by Greenberg et al. (see Fig. 3).
a Schematic representation of an abasic site (AP) isolated (left) or involved in a DPC (right). b Mapping of the interactions between histone H4 basic residues and clustered abasic sites (positions 89 and 207) in the nucleosome from MD simulations. Adapted from Bignon et al. (2021)
The combination of experimental and theoretical approaches is of utmost importance to scrutinize the nucleosome reactivity and map the interactions preceding DPC formation, as shown by the recent work by some of us with the Greenberg’s group (Wen et al. 2023). In this study, a GGGGGCGCGGGGG sequence was inserted at different positions in the nucleosome and oxidized, leading to water trapping and DPC formation with histone tails. While electrophoresis and mass spectrometry experiments were used for DPC detection and quantification, our MD simulations allowed us to describe the contact panorama between the different tails and the DNA. The very good agreement between computational and experimental data unveiled the role of histone tails lysines in the formation of DPC with oxidized DNA, and the parameters that favor the occurrence of these damages: accessibility of the guanine, proximity of H2B, H4, and H3 histone tails, and no quenching by tyrosine residues. We have also demonstrated that a well-located GGG tract is sufficient to form a DPC with a histone tail.
These results constitute first insights into the structural and chemical mechanisms preceding DPC formation in the nucleosome. They set the grounds for a large-scale mapping of tail interactions with DPC-inducing DNA lesions, and further simulations of formed DPC in the nucleosome will provide important data to capture their structural signature and unravel their repair mechanisms.
Photolesions
DNA photolesions mostly occur upon UV irradiation through thymines. Their sequence-dependent formation and repair has been recently assessed in 4096 oligonucleotide sequences (Kufner et al. 2023). Both cyclobutane pyrimidine dimers (CPD) and pyrimidine 6–4 pyrimidone (6-4PP) are associated with distortions of the B-DNA helix, which can be large and very dynamic (see Fig. 4). It is now possible to map, at unprecedented resolution (single-nucleotide for the yeast genome using the CPD-seq method), their distribution and subsequent repair in the nucleosome (Mao et al. 2017). The levels of CPD along a nucleosome obey a hallmark ~ 10.3 base pair periodicity (Stark et al. 2022), a specific “photo-footprint” where the CPD formation is maximal as the phosphate backbone lies far from the histone surface (“outward” rotational setting). This periodicity was retrieved by MD simulations that allowed to quantify conformational coupling for all thymine pairs possibly prone to dimerization (Nayis et al. 2023). Zacharias and coworkers also relied on time-dependent density functional theory (TDDFT) calculations to assess HOMO → LUMO transitions that prefigure the dimerization, thus refining the Kohler’s criterion for CPD formation in B-DNA situating pre-reactive conformation with C5-C6 distance below 4.17 Å and a torsion angle as low as 36°. Beyond the nucleosome, new insights into chromatin influence on CPD formation across the genome could call for coupling all-atom molecular dynamic simulations with coarse-grained approaches.
a Thymine positions over an α-satellite nucleosome (front and top views). Formation yields obey a 10.3 base pair periodicity, with more propensity when they face outwards (outer) the histone core than inwards (in). b Structure of the two main DNA photolesions: cyclobutane pyrimidine dimers (CPD) and pyrimidine 6–4 pyrimidone (6-4PP)
The second most frequent DNA photolesion, 6-4PP, follows a very contrasted pattern and forms more randomly around the nucleosome, with a predominance for linker DNA. It surmised that an increased thymine–thymine flexibility is needed to form the 6-4PP defect, which can be assessed by all-atom MD simulations. In turn, similar simulations with ad hoc parameters and a coarse-grained post-processing situated the role of the nucleosome architecture in tuning the conformational landscape of lesions positioned at SHL3.5 and 4.0, also highlighting the influence of the histones H2A and H2B tails in tuning conformational states of the lesion (Matoušková et al. 2020). Such approaches offer the possibility, not exploited yet, to further decompose the effect of the histone tails whose N-terminal residues were found to play a non-negligible role.
Beyond the formation of CPD and 6-4PP, the interaction with photosensitizers or the formation of photochemical cross-links within the histones (Sperling and Sperling 1978) could be investigated by QM/MM-MD protocols used successfully for B-DNA (Dumont et al. 2015; Bignon et al. 2015, 2017). The recent publication of the ultraviolet-damaged DNA binding (UV-DDB) repair complex bound to the nucleosome with 6-4PP also offers interesting opportunities for computational investigations of photolesion repair mechanisms (Matsumoto et al. 2019).
DNA adducts
DNA adducts result from the covalent binding of endogenous or exogenous chemicals to the nucleobases, which can lead to mutations and carcinogenesis. Depending on the type of the adduct, it can hinder DNA replication and cell division, provoke the incorporation of base mismatches during replication, or result in the formation of abasic sites and the subsequent strand breaks (Marnett and Burcham 1993). Because of their cytotoxic potential, DNA adducts have gathered the interest of scientists for the design of anti-cancer therapeutics such as platinum drugs (Makovec 2019). The latest developments of experimental approaches such as mass spectrometry-based adductomics allow to scrutinize the exposome and to map DNA adducts across the genome (Boysen and Nookaew 2022; Cui and Wang 2022).
Formation and repair of DNA adducts have been investigated in the nucleosome context revealing specificities linked to the DNA mechanical constraints. For instance, the NER pathway can repair platinum − DNA adducts in the nucleosome in a process modulated by histone PTM, yet with only 10–30% efficiency compared to B-DNA (Smerdon et al. 2023). The specificities of the nucleosome environment appear to be crucial for the design of novel anticancer drugs, allowing to develop molecules with stereochemistry-based site selectivity (Chua et al. 2015). Indeed, the nucleosome architecture offers DNA locations with specific, distinct structural behaviors that drive the selective formation of DNA adducts. It has been shown that covalent adducts by benzo[a]pyrene diol epoxide (BPDE, see Fig. 5) are favored on nucleosome locations where the minor groove width fluctuations are larger (i.e., far from the dyad) (Thrall et al. 1994). Besides, the effect of different DNA adducts can dramatically differ from one another: nucleosome disassembly induced by ethidium bromide (McMurray and Van Holde 1986), DNA rotation upon echinomycin or distamycin binding (Low et al. 1986), and even nucleosome-favorable DNA structural deformations induced by BPDE adducts (minor groove width widening, bending) (Mann et al. 1997).
a 3D structure of the benzo[a]pyrene diol epoxide (BPDE) compound. b Crystal structure of a BPDE-dA adduct in B-DNA, with a facing dG mismatch (Schwartz et al. 1997). c Structures of a S 5′-8-cyclo-2′-dG (top) and ( +)-cis-anti-B[a]P-dG (bottom) adducts in the nucleosome from MD simulations. Adapted with permission from Cai et al. (2015). Copyright 2015 American Chemical Society
In the past few years, Broyde, Geacintov, and collaborators have published an extensive series of classical MD-based investigations reporting the conformational behavior of benzo[a]pyrene (B[a]P)-derived adducts within the nucleosome, highlighting the role of histone tails in the DDR activation. They first reported the conformational behavior of different lesions (R and S 5′-8-cyclo-2′-dG and the ( +)-cis-anti-B[a]P-dG) located near the nucleosome dyad, underlining the more pronounced structural destabilization induced by the bulkier B[a]P-dG lesion compared to the 5′-8 cyclopurine damages, also characterized by a weaker histone-DNA interaction energy estimated by molecular mechanics/Poisson-Boltzmann surface area (MMPBSA) calculations (Cai et al. 2015) (see Fig. 5).
Following up, they compared the NER processing rates of BPDE-dG adducts (cis and trans-N2-guanine) and 5′-8 cyclopurine lesions in a joint experimental-theoretical study. They shed light on the importance of the lesion-induced DNA structural perturbation which also depends on the lesion rotational positioning (Shafirovich et al. 2019). Importantly, they showed that while the BPDE-dG adducts are quite efficiently processed by NER in both positions, the 5′-8 cyclopurine structural signature is much less pronounced in the nucleosome than in B-DNA, which results in a decreased repair efficiency. They also confirmed this trends for the 10R and 10S ( +)-cis-anti-B[a]P-N2-dG adducts and the 5′R-8-cyclo-2′-deoxyguanosine lesion, suggesting that the intrinsic structural signature of the lesions are re-defined in the nucleosomal embedding, driving their recognition and processing by repair enzymes (Cai et al. 2020).
They also mapped the interactions of B[a]P-dG derivatives with the histone, first revealing that these lesions can induce the entrapment of a nearby histone tail (H2B tail for an adduct at SHL3.0) that could hamper their correct epigenetic marking, most probably impacting their cellular functions (Fu et al. 2016, 2017). Later on, they showed how histone H3 and H4 acetylation could modulate the exposure of B[a]P-dG adducts for facilitating NER initiation (Cai et al. 2018; Fu et al. 2021). For adducts positioned at SHL − 1, interactions with the H4 tail appeared to be modified upon poly-acetylation of H4 K5, K8, K12, and K16, which induce a flipping of the facing dC nucleobase, further captured by the H3 tail. The authors suggested that the ability of the H3 tail to concomitantly bind the NER-triggering protein XPC and capture the dC partner of the adduct could play a key role in triggering the DDR. For adducts at SHL 6.25, they further revealed that the H3K56 acetylation could amplify the structural distortions at the B[a]P-dG site, which might be an important step for facilitating the access of repair enzymes to the damage.
Overall, these extensive computational studies provide outstanding insights into adduct formation, structural signature, positional effect, and interplay with histone tails. They demonstrate how powerful computational methods can be to delineate the atomistic details of DNA damage formation and repair in the nucleosome. Such investigations are of utmost importance to push the field further on. They provide solid grounds to foster more studies about other endogenous or exogenous DNA adduct formation and repair mechanisms, and for the development of next-generation anticancer drugs taking into account the nucleosome environment.
Ribonucleotide misincorporation
During DNA synthesis by DNA polymerases, faulty incorporation of ribonucleotides can occur and provoke replicative stress and genome instability if left unrepaired (McElhinny et al. 2010). This very common type of replication error is efficiently recognized and cleaved on its 5′ end by the ribonuclease (Rnase) HII, triggering downstream DDR events (Sparks et al. 2012). At first, such misincorporation was thought to essentially lead to deleterious consequences, but new evidences shed light onto its functional role and utility in specific situations linked to DNA damage repair (Dalgaard 2012; Potenski and Klein 2014).
The structural impact of ribonucleotides in hundreds of base pair long-free DNA has been studied by atomic force microscopy, describing a shortening of the molecule induced by the ribose-induced backbone compaction and a decrease of its persistence length (Meroni et al. 2017). While even a small percentage of ribonucleotide in the DNA helix could reduce the formation of nucleosomes (Hovatter and Martinson 1987), a 552-fold decrease of repair rates by the Rnase HII in the nucleosome compared to B-DNA has been highlighted in vitro (Ren et al. 2019), underlining the deleterious effect of such lesion.
Computational investigations brought out new insights into the structural impact of a single ribonucleotide incorporation in the nucleosome (Fu et al. 2019). Positioning the ribonucleotide at different positions (near SHL 0, SHL 2.5, and SHL 5.5), with different rotational orientations (inwards or outwards), they described the distortions induced by the damage on the DNA structure and on DNA-histone interactions. They revealed that the inward position of the damage exhibits a C3′-endo ribose puckering that is stabilized by the interactions with the histone core and propagates to the adjacent deoxyribonucleotides, hence inducing an important structural perturbation in a 2-base pair radius, with a maximum effect at SHL 2.5.
The molecular mechanisms of single ribonucleotide repair in the nucleosome remains to be unraveled, but these computational investigations constitute the first insights into their structural signature at different SHL and orientations. Importantly, they suggested a strong role of histone-DNA contacts in the amplification of the ribonucleotide-induced structural distortion, which could be of utmost importance for triggering the DDR. Future combined experimental and theoretical studies will be crucial to shed light on the ribonucleotide misincorporation repair in the nucleosome context, which most probably involves PTM and remodeling events.
Conclusions and outlook
In the last decade, computational approaches have emerged as an important way to accumulate new knowledge about DNA damage and repair in the nucleosome context, at atomic or even electronic scales. The particular DNA constraints imposed by the nucleosome structure imply different formation/repair mechanisms and site specificities than what is observed in B-DNA. Indeed, the formation of lesions will be favored at specific SHL and rotational orientations, while their repair can necessitate DNA unwrapping by chromatin remodelers, most probably involving specific signaling by histone PTM.
Classical MD simulations of nucleosome core particles bearing DNA damages have multiplied over the past 5 years, highlighting a real effort from the community to improve our knowledge of DNA damage and repair in this complex architecture. Such investigations shed light on the specific structural signature of a variety of DNA lesions (DNA adducts, photolesions, DPC, DSB, abasic sites, ribonucleotides) in the nucleosome. These tools allow the mapping, at the atomic scale, of canonical and PTM-bearing histone tail interactions with the lesion sites, which provide invaluable information about how they influence the lesion-induced local DNA distortion and about which residues could be involved in the formation of DPC. The increasing computational power also allowed the very recent release of costly QM/MM studies, providing unprecedented evidences of the intrinsic reactivity of DNA nucleobases (guanines) and histone residues in the nucleosome context.
Overall, the recent accumulation of computational efforts to model the dynamics and reactivity of damaged nucleosomes portends future cutting-edge work. The combination of experimental and theoretical approaches provides complementary data to describe these complex processes. Importantly, the growing amounts of experimental structures featuring the binding mode of repair enzymes and DDR-related chromatin remodelers or histone PTM writer/erasers to the nucleosome provide important starting points for further modeling investigations (Eustermann et al. 2018; Matsumoto et al. 2019; Weaver et al. 2022; Zheng et al. 2023; Smirnova et al. 2024). Besides, the latest developments towards affordable classical MD simulations of multi-million atom systems (Antolínez et al. 2024) and machine learning–based approaches (Jackson et al. 2023) constitute interesting methodological perspectives for future investigations. The use of coarse-graining approaches, which have shown their worth for the study of undamaged multi-nucleosome systems, could also bring out new perspectives on DNA damage signature and structural effect on higher-scale architectures. We believe that the computational efforts described in this review highlight the start of a growing interest for atomic- and electronic-scale modeling of DNA damages in the nucleosome, which forecasts a new exciting era for uncovering precise mechanisms modulating their formation, structural signature, and repair.
Data availability
No original data was produced for this review article. All data presented in the figures are available from corresponding references.
Code availability
Not applicable.
References
Alvarado W, Moller J, Ferguson AL, de Pablo JJ (2021) Tetranucleosome interactions drive chromatin folding. ACS Cent Sci 7:1019–1027. https://doi.org/10.1021/acscentsci.1c00085
Antolínez S, Jones PE, Phillips JC, Hadden-Perilla JA (2024) AMBERff at scale: multimillion-atom simulations with AMBER force fields in NAMD. Biophys J 123:422a. https://doi.org/10.1016/j.bpj.2023.11.2563
Armeev GA, Gribkova AK, Pospelova I et al (2019) Linking chromatin composition and structural dynamics at the nucleosome level. Curr Opin Struct Biol 56:46–55. https://doi.org/10.1016/j.sbi.2018.11.006
Armeev GA, Kniazeva AS, Komarova GA et al (2021) Histone dynamics mediate DNA unwrapping and sliding in nucleosomes. Nat Commun 12:2387. https://doi.org/10.1038/s41467-021-22636-9
Arnold AR, Grodick MA, Barton JK (2016) DNA charge transport: from chemical principles to the cell. Cell Chem Biol 23:183–197. https://doi.org/10.1016/j.chembiol.2015.11.010
Bag S, Aggarwal A, Maiti PK (2020) Machine learning prediction of electronic coupling between the guanine bases of DNA. J Phys Chem A 124:7658–7664. https://doi.org/10.1021/acs.jpca.0c04368
Banerjee S, Chakraborty S, Jacinto MP et al (2017) Probing enhanced double-strand break formation at abasic sites within clustered lesions in nucleosome core particles. Biochemistry 56:14–21. https://doi.org/10.1021/acs.biochem.6b01144
Bignon E, Gattuso H, Morell C et al (2015) DNA photosensitization by an “insider”: photophysics and triplet energy transfer of 5-methyl-2-pyrimidone deoxyribonucleoside. Chem A Eur J 21:11509–11516. https://doi.org/10.1002/chem.201501212
Bignon E, Marazzi M, Besancenot V et al (2017) Ibuprofen and ketoprofen potentiate UVA-induced cell death by a photosensitization process. Sci Rep 7:8885. https://doi.org/10.1038/s41598-017-09406-8
Bignon E, Claerbout VEP, Jiang T et al (2020) Nucleosomal embedding reshapes the dynamics of abasic sites. Sci Rep 10:17314. https://doi.org/10.1038/s41598-020-73997-y
Bignon E, Gillet N, Jiang T et al (2021) A dynamic view of the interaction of histone tails with clustered abasic sites in a nucleosome core particle. J Phys Chem Lett 12:6014–6019. https://doi.org/10.1021/acs.jpclett.1c01058
Bilotti K, Kennedy EE, Li C, Delaney S (2017) Human OGG1 activity in nucleosomes is facilitated by transient unwrapping of DNA and is influenced by the local histone environment. DNA Repair 59:1–8. https://doi.org/10.1016/j.dnarep.2017.08.010
Bowerman S, Wereszczynski J (2016) Effects of MacroH2A and H2A.Z on nucleosome dynamics as elucidated by molecular dynamics simulations. Biophys J 110:327–337. https://doi.org/10.1016/j.bpj.2015.12.015
Boysen G, Nookaew I (2022) Current and future methodology for quantitation and site-specific mapping the location of DNA adducts. Toxics 10:45. https://doi.org/10.3390/toxics10020045
Brambilla F, Garcia-Manteiga JM, Monteleone E et al (2020) Nucleosomes effectively shield DNA from radiation damage in living cells. Nucleic Acids Res 48:8993–9006. https://doi.org/10.1093/nar/gkaa613
Cadet J, Davies KJA, Medeiros MH et al (2017) Formation and repair of oxidatively generated damage in cellular DNA. Free Radical Biol Med 107:13–34. https://doi.org/10.1016/j.freeradbiomed.2016.12.049
Cai Y, Kropachev K, Terzidis MA et al (2015) Differences in the access of lesions to the nucleotide excision repair machinery in nucleosomes. Biochemistry 54:4181–4185. https://doi.org/10.1021/acs.biochem.5b00564
Cai Y, Fu I, Geacintov NE et al (2018) Synergistic effects of H3 and H4 nucleosome tails on structure and dynamics of a lesion-containing DNA: binding of a displaced lesion partner base to the H3 tail for GG-NER recognition. DNA Repair (amst) 65:73–78. https://doi.org/10.1016/j.dnarep.2018.02.009
Cai Y, Geacintov NE, Broyde S (2020) Variable impact of conformationally distinct DNA lesions on nucleosome structure and dynamics: implications for nucleotide excision repair. DNA Repair 87:102768. https://doi.org/10.1016/j.dnarep.2019.102768
Cannan WJ, Tsang BP, Wallace SS, Pederson DS (2014) Nucleosomes suppress the formation of double-strand DNA breaks during attempted base excision repair of clustered oxidative damages. J Biol Chem 289:19881–19893. https://doi.org/10.1074/jbc.M114.571588
Chua EYD, Davey GE, Chin CF et al (2015) Stereochemical control of nucleosome targeting by platinum-intercalator antitumor agents. Nucleic Acids Res 43:5284–5296. https://doi.org/10.1093/nar/gkv356
Cintori L, Di Guilmi A-M, Canitrot Y et al (2023) Spatio-temporal dynamics of the DNA glycosylase OGG1 in finding and processing 8-oxoguanine. DNA Repair 129:103550. https://doi.org/10.1016/j.dnarep.2023.103550
Cleri F, Landuzzi F, Blossey R (2018) Mechanical evolution of DNA double-strand breaks in the nucleosome. PLoS Comput Biol 14:e1006224. https://doi.org/10.1371/journal.pcbi.1006224
Collepardo-Guevara R, Portella G, Vendruscolo M et al (2015) Chromatin unfolding by epigenetic modifications explained by dramatic impairment of internucleosome interactions: a multiscale computational study. J Am Chem Soc 137:10205–10215. https://doi.org/10.1021/jacs.5b04086
Cui Y, Wang Y (2022) Mass spectrometry-based DNA adductomics. TrAC Trends Anal Chem 157:116773. https://doi.org/10.1016/j.trac.2022.116773
Dalgaard JZ (2012) Causes and consequences of ribonucleotide incorporation into nuclear DNA. Trends Genet 28:592–597. https://doi.org/10.1016/j.tig.2012.07.008
Davis WB, Bjorklund CC, Deline M (2012) Probing the effects of DNA–protein interactions on DNA hole transport: the N-terminal histone tails modulate the distribution of oxidative damage and chemical lesions in the nucleosome core particle. Biochemistry 51:3129–3142. https://doi.org/10.1021/bi201734c
Dršata T, Kara M, Zacharias M, Lankaš F (2013) Effect of 8-oxoguanine on DNA structure and deformability. J Phys Chem B 117:11617–11622. https://doi.org/10.1021/jp407562t
Dumont E, Wibowo M, Roca-Sanjuán D et al (2015) Resolving the benzophenone DNA-photosensitization mechanism at QM/MM level. J Phys Chem Lett 6:576–580. https://doi.org/10.1021/jz502562d
Eustermann S, Schall K, Kostrewa D et al (2018) Structural basis for ATP-dependent chromatin remodelling by the INO80 complex. Nature 556:386–390. https://doi.org/10.1038/s41586-018-0029-y
Fu I, Cai Y, Zhang Y et al (2016) Entrapment of a histone tail by a DNA lesion in a nucleosome suggests the lesion impacts epigenetic marking: a molecular dynamics study. Biochemistry 55:239–242. https://doi.org/10.1021/acs.biochem.5b01166
Fu I, Cai Y, Geacintov NE et al (2017) Nucleosome histone tail conformation and dynamics: impacts of lysine acetylation and a nearby minor groove benzo[a]pyrene-derived lesion. Biochemistry 56:1963–1973. https://doi.org/10.1021/acs.biochem.6b01208
Fu I, Smith DJ, Broyde S (2019) Rotational and translational positions determine the structural and dynamic impact of a single ribonucleotide incorporated in the nucleosome. DNA Repair 73:155–163. https://doi.org/10.1016/j.dnarep.2018.11.012
Fu I, Geacintov NE, Broyde S (2021) Molecular dynamics simulations reveal how H3K56 acetylation impacts nucleosome structure to promote DNA exposure for lesion sensing. DNA Repair (amst) 107:103201. https://doi.org/10.1016/j.dnarep.2021.103201
Gursoy-Yuzugullu O, House N, Price BD (2016) Patching broken DNA: nucleosome dynamics and the repair of DNA breaks. J Mol Biol 428:1846–1860. https://doi.org/10.1016/j.jmb.2015.11.021
Hauer MH, Gasser SM (2017) Chromatin and nucleosome dynamics in DNA damage and repair. Genes Dev 31:2204–2221. https://doi.org/10.1101/gad.307702.117
Hovatter KR, Martinson HG (1987) Ribonucleotide-induced helical alteration in DNA prevents nucleosome formation. Proc Natl Acad Sci USA 84:1162–1166. https://doi.org/10.1073/pnas.84.5.1162
Jackson NE, Savoie BM, Statt A, Webb MA (2023) Introduction to machine learning for molecular simulation. J Chem Theory Comput 19:4335–4337. https://doi.org/10.1021/acs.jctc.3c00735
Jiang T, Monari A, Dumont E, Bignon E (2021) Molecular Mechanisms associated with clustered lesion-induced impairment of 8-oxoG recognition by the human glycosylase OGG1. Molecules 26:6465. https://doi.org/10.3390/molecules26216465
Karch KR, Langelier M-F, Pascal JM, Garcia BA (2017) The nucleosomal surface is the main target of histone ADP-ribosylation in response to DNA damage. Mol BioSyst 13:2660–2671. https://doi.org/10.1039/C7MB00498B
Kılıç M, Diamantis P, Johnson SK et al (2023) Redox-based defect detection in packed DNA: insights from hybrid quantum mechanical/molecular mechanics molecular dynamics simulations. J Chem Theory Comput 19:8434–8445. https://doi.org/10.1021/acs.jctc.3c01013
Kono H, Ishida H (2020) Nucleosome unwrapping and unstacking. Curr Opin Struct Biol 64:119–125. https://doi.org/10.1016/j.sbi.2020.06.020
Krämer M, Dohmen PM, Xie W et al (2020) Charge and exciton transfer simulations using machine-learned Hamiltonians. J Chem Theory Comput 16:4061–4070. https://doi.org/10.1021/acs.jctc.0c00246
Kubař T, Elstner M (2013) A hybrid approach to simulation of electron transfer in complex molecular systems. J R Soc Interface 10:20130415. https://doi.org/10.1098/rsif.2013.0415
Kufner CL, Krebs S, Fischaleck M et al (2023) Sequence dependent UV damage of complete pools of oligonucleotides. Sci Rep 13:2638. https://doi.org/10.1038/s41598-023-29833-0
Kühbacher U, Duxin JP (2020) How to fix DNA-protein crosslinks. DNA Repair 94:102924. https://doi.org/10.1016/j.dnarep.2020.102924
La Rosa G, Zacharias M (2016) Global deformation facilitates flipping of damaged 8-oxo-guanine and guanine in DNA. Nucl Acids Res 44:9591–9599. https://doi.org/10.1093/nar/gkw827
Lee MH, Brancolini G, Gutiérrez R et al (2012) Probing charge transport in oxidatively damaged DNA sequences under the influence of structural fluctuations. J Phys Chem B 116:10977–10985. https://doi.org/10.1021/jp2091544
Lee AJ, Rackers JA, Bricker WP (2022) Predicting accurate ab initio DNA electron densities with equivariant neural networks. Biophys J 121:3883–3895. https://doi.org/10.1016/j.bpj.2022.08.045
Lequieu J, Córdoba A, Schwartz DC, de Pablo JJ (2016) Tension-dependent free energies of nucleosome unwrapping. ACS Cent Sci 2:660–666. https://doi.org/10.1021/acscentsci.6b00201
Low CML, Drew HR, Waring MJ (1986) Echinomycin and distamycin induce rotation of nucleosome core DNA. Nucl Acids Res 14:6785–6801. https://doi.org/10.1093/nar/14.17.6785
Lukina MV, Koval VV, Lomzov AA et al (2017) Global DNA dynamics of 8-oxoguanine repair by human OGG1 revealed by stopped-flow kinetics and molecular dynamics simulation. Mol BioSyst 13:1954–1966. https://doi.org/10.1039/C7MB00343A
Makovec T (2019) Cisplatin and beyond: molecular mechanisms of action and drug resistance development in cancer chemotherapy. Radiol Oncol 53:148–158. https://doi.org/10.2478/raon-2019-0018
Mann DB, Springer DL, Smerdon MJ (1997) DNA damage can alter the stability of nucleosomes: effects are dependent on damage type. Proc Natl Acad Sci USA 94:2215–2220. https://doi.org/10.1073/pnas.94.6.2215
Mao P, Wyrick JJ, Roberts SA, Smerdon MJ (2017) UV-induced DNA damage and mutagenesis in chromatin. Photochem Photobiol 93:216–228. https://doi.org/10.1111/php.12646
Marnett LJ, Burcham PC (1993) Endogenous DNA adducts: potential and paradox. Chem Res Toxicol 6:771–785. https://doi.org/10.1021/tx00036a005
Matoušková E, Bignon E, Claerbout VEP et al (2020) Impact of the Nucleosome histone core on the structure and dynamics of DNA-containing pyrimidine–pyrimidone (6–4) photoproduct. J Chem Theory Comput 16:5972–5981. https://doi.org/10.1021/acs.jctc.0c00593
Matsumoto S, Cavadini S, Bunker RD et al (2019) DNA damage detection in nucleosomes involves DNA register shifting. Nature 571:79–84. https://doi.org/10.1038/s41586-019-1259-3
McElhinny SAN, Kumar D, Clark AB et al (2010) Genome instability due to ribonucleotide incorporation into DNA. Nat Chem Biol 6:774–781. https://doi.org/10.1038/nchembio.424
McGinty RK, Tan S (2015) Nucleosome structure and function. Chem Rev 115:2255–2273. https://doi.org/10.1021/cr500373h
McMurray CT, Van Holde KE (1986) Binding of ethidium bromide causes dissociation of the nucleosome core particle. Proc Natl Acad Sci USA 83:8472–8476. https://doi.org/10.1073/pnas.83.22.8472
Meroni A, Mentegari E, Crespan E et al (2017) The incorporation of ribonucleotides induces structural and conformational changes in DNA. Biophys J 113:1373–1382. https://doi.org/10.1016/j.bpj.2017.07.013
Nayis A, Liebl K, Zacharias M (2023) Coupling of conformation and CPD damage in nucleosomal DNA. Biophys Chem 300:107050. https://doi.org/10.1016/j.bpc.2023.107050
Núñez ME, Hall DB, Barton JK (1999) Long-range oxidative damage to DNA: effects of distance and sequence. Chem Biol 6:85–97. https://doi.org/10.1016/S1074-5521(99)80005-2
Osakabe A, Arimura Y, Matsumoto S et al (2017) Polymorphism of apyrimidinic DNA structures in the nucleosome. Sci Rep 7:41783. https://doi.org/10.1038/srep41783
Peluso A, Caruso T, Landi A, Capobianco A (2019) The dynamics of hole transfer in DNA. Molecules 24:4044. https://doi.org/10.3390/molecules24224044
Peng Y, Li S, Onufriev A et al (2021) Binding of regulatory proteins to nucleosomes is modulated by dynamic histone tails. Nat Commun 12:5280. https://doi.org/10.1038/s41467-021-25568-6
Polo SE, Almouzni G (2015) Chromatin dynamics after DNA damage: the legacy of the access–repair–restore model. DNA Repair 36:114–121. https://doi.org/10.1016/j.dnarep.2015.09.014
Potenski CJ, Klein HL (2014) How the misincorporation of ribonucleotides into genomic DNA can be both harmful and helpful to cells. Nucleic Acids Res 42:10226–10234. https://doi.org/10.1093/nar/gku773
Ren M, Cheng Y, Duan Q, Zhou C (2019) Transesterification reaction and the repair of embedded ribonucleotides in DNA are suppressed upon the assembly of DNA into nucleosome core particles. Chem Res Toxicol 32:926–934. https://doi.org/10.1021/acs.chemrestox.9b00059
Ren M, Shang M, Wang H et al (2021) Histones participate in base excision repair of 8-oxodGuo by transiently cross-linking with active repair intermediates in nucleosome core particles. Nucleic Acids Res 49:257–268. https://doi.org/10.1093/nar/gkaa1153
Ren M, Greenberg MM, Zhou C (2022) Participation of histones in DNA damage and repair within nucleosome core particles: mechanism and applications. Acc Chem Res 55:1059–1073. https://doi.org/10.1021/acs.accounts.2c00041
Schwartz JL, Rice JS, Luxon BA et al (1997) Solution structure of the minor conformer of a DNA duplex containing a dG mismatch opposite a benzo[a]pyrene diol epoxide/dA adduct: glycosidic rotation from syn to anti at the modified deoxyadenosine. Biochemistry 36:11069–11076. https://doi.org/10.1021/bi971306u
Sczepanski JT, Zhou C, Greenberg MM (2013) Nucleosome core particle-catalyzed strand scission at abasic sites. Biochemistry 52:2157–2164. https://doi.org/10.1021/bi3010076
Seeber A, Hauer M, Gasser SM (2013) Nucleosome remodelers in double-strand break repair. Curr Opin Genet Dev 23:174–184. https://doi.org/10.1016/j.gde.2012.12.008
Shafirovich V, Kolbanovskiy M, Kropachev K et al (2019) Nucleotide excision repair and impact of site-specific 5’,8-cyclopurine and bulky DNA lesions on the physical properties of nucleosomes. Biochemistry 58:561–574. https://doi.org/10.1021/acs.biochem.8b01066
Sharma D, De Falco L, Padavattan S et al (2019) PARP1 exhibits enhanced association and catalytic efficiency with γH2A.X-nucleosome. Nat Commun 10:5751. https://doi.org/10.1038/s41467-019-13641-0
Shigdel UK, Ovchinnikov V, Lee S-J et al (2020) The trajectory of intrahelical lesion recognition and extrusion by the human 8-oxoguanine DNA glycosylase. Nat Commun 11:4437. https://doi.org/10.1038/s41467-020-18290-2
Smerdon MJ, Wyrick JJ, Delaney S (2023) A half century of exploring DNA excision repair in chromatin. J Biol Chem 299:105118. https://doi.org/10.1016/j.jbc.2023.105118
Smirnova E, Bignon E, Schultz P et al (2024) Binding to nucleosome poises human SIRT6 for histone H3 deacetylation. eLife 12:RP87989. https://doi.org/10.7554/eLife.87989
Sparks JL, Chon H, Cerritelli SM et al (2012) RNase H2-initiated ribonucleotide excision repair. Mol Cell 47:980–986. https://doi.org/10.1016/j.molcel.2012.06.035
Sperling J, Sperling R (1978) Photochemical cross-linking of histones to DNA in nucleosomes. Nucl Acids Res 5:2755–2774. https://doi.org/10.1093/nar/5.8.2755
Stark B, Poon GMK, Wyrick JJ (2022) Molecular mechanism of UV damage modulation in nucleosomes. Comput Struct Biotechnol J 20:5393–5400. https://doi.org/10.1016/j.csbj.2022.08.071
Sun H, Zheng L, Yang K, Greenberg MM (2019) Positional dependence of DNA hole transfer efficiency in nucleosome core particles. J Am Chem Soc 141:10154–10158. https://doi.org/10.1021/jacs.9b03686
Thrall BD, Mann DB, Smerdon MJ, Springer DL (1994) Nucleosome structure modulates benzo[a]pyrenediol epoxide adduct formation. Biochemistry 33:2210–2216. https://doi.org/10.1021/bi00174a030
Voityuk AA, Davis WB (2007) Hole transfer energetics in structurally distorted DNA: the nucleosome core particle. J Phys Chem B 111:2976–2985. https://doi.org/10.1021/jp066470i
Weaver TM, Hoitsma NM, Spencer JJ et al (2022) Structural basis for APE1 processing DNA damage in the nucleosome. Nat Commun 13:5390. https://doi.org/10.1038/s41467-022-33057-7
Weickert P, Stingele J (2022) DNA–protein crosslinks and their resolution. Annu Rev Biochem 91:157–181. https://doi.org/10.1146/annurev-biochem-032620-105820
Wen T, Kermarrec M, Dumont E, et al (2023) DNA–histone cross-link formation via hole trapping in nucleosome core particles. J Am Chem Soc jacs.3c08135. https://doi.org/10.1021/jacs.3c08135
Weng L, Greenberg MM (2015) Rapid histone-catalyzed DNA lesion excision and accompanying protein modification in nucleosomes and nucleosome core particles. J Am Chem Soc 137:11022–11031. https://doi.org/10.1021/jacs.5b05478
Wolter M, Elstner M, Kleinekathöfer U, Kubař T (2017) Microsecond Simulation of electron transfer in DNA: bottom-up parametrization of an efficient electron transfer model based on atomistic details. J Phys Chem B 121:529–549. https://doi.org/10.1021/acs.jpcb.6b11384
Zhang H, Xiong Y, Chen J (2020) DNA–protein cross-link repair: what do we know now? Cell Biosci 10:3. https://doi.org/10.1186/s13578-019-0366-z
Zheng L, Tsai B, Gao N (2023) Structural and mechanistic insights into the DNA glycosylase AAG-mediated base excision in nucleosome. Cell Discov 9:62. https://doi.org/10.1038/s41421-023-00560-0
Zhou C, Sczepanski JT, Greenberg MM (2012) Mechanistic studies on histone catalyzed cleavage of apyrimidinic/apurinic sites in nucleosome core particles. J Am Chem Soc 134:16734–16741. https://doi.org/10.1021/ja306858m
Funding
NG has received support from the Agence Nationale de la Recherche (grant number ANR-20-CE29-0002–01).
Author information
Authors and Affiliations
Contributions
EB conceptualized the work. NG, ED, and EB performed the literature search and wrote the review.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Materials availability
No new materials were presented in this review.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gillet, N., Dumont, E. & Bignon, E. DNA damage and repair in the nucleosome: insights from computational methods. Biophys Rev 16, 345–356 (2024). https://doi.org/10.1007/s12551-024-01183-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-024-01183-9