Abstract
X-ray crystallography visualizes the world at the atomic level. It has been used as the most powerful technique for observing the three-dimensional structures of biological macromolecules and has pioneered structural biology. To determine a crystal structure with high resolution, it was traditionally required to prepare large crystals (> 200 μm). Later, synchrotron radiation facilities, such as SPring-8, that produce powerful X-rays were built. They enabled users to obtain good quality X-ray diffraction images even with smaller crystals (ca. 200–50 μm). In recent years, one of the most important technological innovations in structural biology has been the development of X-ray free electron lasers (XFELs). The SPring-8 Angstrom Compact free electron LAser (SACLA) in Japan generates the XFEL beam by accelerating electrons to relativistic speeds and directing them through in-vacuum, short-period undulators. Since user operation started in 2012, we have been involved in the development of serial femtosecond crystallography (SFX) measurement systems using XFEL at the SACLA. The SACLA generates X-rays a billion times brighter than SPring-8. The extremely bright XFEL pulses enable data collection with microcrystals (ca. 50–1 μm). Although many molecular analysis techniques exist, SFX is the only technique that can visualize radiation-damage-free structures of biological macromolecules at room temperature in atomic resolution and fast time resolution. Here, we review the achievements of the SACLA-SFX Project in the past 5 years. In particular, we focus on: (1) the measurement system for SFX; (2) experimental phasing by SFX; (3) enzyme chemistry based on damage-free room-temperature structures; and (4) molecular movie taken by time-resolved SFX.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Measurement system for SFX
X-ray free electron lasers (XFELs) possess the unique features of extreme peak brightness, ultrashort duration, and high spatial coherence of X-ray pulses. Following the construction of the world’s first XFEL facility, the Linac Coherent Light Source (LCLS) in the USA in 2009, the SPring-8 Angstrom Compact free electron LAser (SACLA; Ishikawa et al. 2012) was built in Harima Science Garden City, Hyogo, Japan in 2011 (Fig. 1a). In the SACLA, the pulse duration is in the range of 2–10 fs and the pulse repetition rate is 30 Hz or 60 Hz, depending on the operation mode. A novel crystallographic technique, serial femtosecond crystallography (SFX), utilizing the unique features of XFELs, has opened a new area of structural biology (Chapman et al. 2011). SFX enables data collection from many randomly oriented microcrystals streamed with an injector device across the XFEL beam, which allows single-pulse diffraction from a microcrystal within femtosecond exposure time at ambient temperature (Fig. 1b). Thereby, SFX allows a “diffraction-before-destruction” approach, by circumventing radiation damage (including photoreduction) of the sample, because the diffraction process is terminated on a time-scale shorter than that of the damage process (Barty et al. 2012; Neutze et al. 2000). In addition, time-resolved SFX, which is an applied technique of SFX, is an innovative technology that enables us to observe how proteins move while functioning (Tenboer et al. 2014).
The X-ray free electron laser (XFEL), a new technological innovation in structural biology. a The XFEL facility, the SPring-8 Angstrom Compact free electron LAser (SACLA), is built next to SPring-8 in Harima Science Garden City, Hyogo, Japan. b The serial femtosecond crystallography (SFX) measurement scheme at the SACLA. A pump laser can be used for data collection in time-resolved SFX
SFX overcomes displacement and distortion of protein structure caused by crystal freezing and radiation damage during data collection, which are problems in conventional X-ray crystallography with in-house diffractometers, as well as synchrotron facilities. Using microcrystals for SFX has further advantages. Compared with large crystals, crystal disorder induced by osmotic shock during soaking manipulation is small in microcrystals. A higher surface-to-volume ratio of the crystal allows rapid diffusion of compounds. Thus, microcrystals are more suitable for ligand or heavy atom derivatization by soaking than large crystals. In addition, large crystals may grow into polycrystals and it takes time and effort to adjust the crystal orientation at the time of data collection in conventional crystallography, but microcrystals easily form monocrystals, and data can be conveniently collected in SFX without worrying about the orientation.
In the SACLA, the measurement system for SFX consists of components including the detector, sample chamber, injector, and data acquisition system; this system can be routinely applied to many kinds of experiments for different users (Mafuné et al. 2016; Nakane et al. 2016b; Tono et al. 2015). In the SFX technique, protein structures are determined by merging the X-ray diffraction images from thousands (Sugahara et al. 2017) to a half million (Batyuk et al. 2016) microcrystals measured in an hour to a day. It is noteworthy that the SACLA-SFX Project has significantly contributed to developments of methods for reducing the consumption of soluble protein samples. A liquid-jet injector we initially used at the SACLA needed hundreds of milligrams of sample to solve one protein structure (Tono et al. 2015). Since not every protein can be crystallized in large quantities, it was, therefore, a problem that the injector with large sample consumption could not be applied to every sample in general purpose. At that time, by a group working at the LCLS, a lipidic cubic phase (LCP) injector was invented for membrane proteins crystallized in LCP (Weierstall et al. 2014). We extended the scope of the LCP injector to samples of soluble proteins. When the viscosity of the sample is increased by adding a highly viscous medium, the flow rate of the stream extruded from the injector to the XFEL becomes extremely slow. Thus, the sample consumption required for structure determination can be considerably reduced. After screening dozens of media, we established the microcrystal-carrier matrix method and reduced sample consumption to as little as 1 mg (Sugahara et al. 2015). Intriguingly, the method can be applied not only to soluble proteins but also to membrane proteins crystallized in bicelles (Nakane et al. 2016a) and detergent micelles (Suga et al. 2017). Moreover, the method was effective for stabilization of the sample stream and accurate measurement during data collection for time-resolved SFX (Nango et al. 2016).
Experimental phasing by SFX
In X-ray crystallography, structure factors are Fourier transformed to yield real-space electron density maps. Here, both the amplitude and the phase of structure factors are necessary. However, one cannot directly measure the phase of diffracted waves, only the amplitude. As reviewed by Scapin (2013), most structures are, nowadays, solved by the molecular replacement method, which estimates phases from structures of homologous proteins. The determination of novel structures requires experimental (de novo) phasing, where phases are calculated from isomorphous differences between native and derivative crystals and/or anomalous differences between Bijvoet pairs. Readers are referred to Taylor (2010) for a general review of these methods. Such differences are typically very small, less than a few percent for anomalous differences in most cases. Thus, great care must be taken in both instrumentation and data processing to accurately capture them (Liu and Hendrickson 2015). Experimental phasing is more challenging with SFX. In contrast to conventional crystallography, where one or a few crystals are used to complete a dataset, data from tens of thousands of crystals are merged in SFX. Variations in crystals lead to lower precision; they include inhomogeneity of structures, a variety of heavy atom occupancies, and differences in crystal size and morphology. Fluctuations of XFEL pulse intensities and wavelength spectra are also sources of errors. Moreover, all measurements using XFELs are partial reflections because crystals hardly rotate during an exposure. Thus, they represent only varying fractions (called the partiality) of full reflections. In Monte Carlo integration, these random variables are integrated out by merging a huge number of observations (Kirian et al. 2011).
In 2014, gadolinium single-wavelength anomalous diffraction (SAD) phasing of lysozyme at the LCLS (Barends et al. 2014) was reported as the first successful experimental phasing with SFX. This system exhibited a large anomalous difference between Friedel pairs, or a Bijvoet ratio, of more than 10%. Thus, it was still unknown if one could phase more realistic targets with smaller anomalous difference signals. In 2015, we reported a native sulfur/chloride SAD phasing of lysozyme at the SACLA (Nakane et al. 2015). The native SAD phasing uses anomalous difference signals from sulfur atoms in cysteine and methionine residues. It, therefore, does not require heavy atom derivatization. However, anomalous differences from sulfur are much smaller than those from heavier atoms. Our system contained ten sulfur atoms and one chloride anion from the solvent. The calculated Bijvoet ratio was 1.6% at 7.0 keV (1.771 Å). Note that the precision of Monte Carlo integration improves in proportion to the square root of the multiplicity (MacKay 1998). That is, the detection of 1% differences requires 100 times more data than the detection of 10% differences. Moreover, success is not guaranteed in the presence of systematic biases. Our protocol for phasing uses CrystFEL (White et al. 2012) for integration and merging and a popular phasing suite, SHELXD for substructure determination and SHELXE for phase calculation and density modification (Sheldrick 2010). We test extensive combinations of SHELX parameters, i.e., high resolution cutoff, solvent content, and the number of sites for calculation. We found that iterations of density modification and automatic chain tracing in SHELXE can improve very noisy initial maps hardly interpretable by human eyes to high-quality maps that can be completed by Buccaneer (Cowtan 2006). Eventually, we succeeded in phasing using 130,000 indexed images.
This protocol was also successfully applied to the phasing of copper-containing nitrite reductase crystals (Fukuda et al. 2016a). They contained six copper atoms in the asymmetric unit and the calculated Bijvoet ratio was 1.7% at 10.8 keV (1.418 Å). Similar strategies were also adopted by others (Batyuk et al. 2016). In 2016, two more sulfur SAD phasings were reported from the LCLS, using 125,000 indexed images for thaumatin (Nass et al. 2016) and 500,000 indexed images for a membrane protein, A2a receptor (Batyuk et al. 2016). The calculated Bijvoet ratios were 2.1% and 1.9%, respectively. These studies jointly established experimental phasing with SFX, even from weak anomalous signals. Nonetheless, the required numbers of indexed images were so high that a huge amount of purified proteins (e.g., ~ 5 mg for sulfur SAD of lysozyme by Nakane et al. 2015) and long beam time were necessary. To enable phasing from fewer images, two approaches are possible: one is to increase the signal and the other is to improve the data processing algorithms.
To increase the signal, we developed a new derivatization method for membrane proteins (Nakane et al. 2016a) based on the observation that detergent molecules and lipids are often found on the surface of membrane proteins. We synthesized an iodine-labeled detergent named HAD13a by an amido-coupling of caprylic acid and the so-called magic triangle, or I3C, which contains three iodine atoms bound to a benzene ring. We soaked this compound into bacteriorhodopsin (bR) crystals grown in bicelles, collected diffraction images at 7.0 keV, and succeeded in SAD phasing from 23,000 indexed images. Two molecules of HAD13a were found in the asymmetric unit. They resulted in a large Bijvoet ratio of 10.5%. If we were to apply sulfur SAD phasing instead of iodine SAD phasing without HAD13a, the Bijvoet ratio would be only 0.9%. By combining a native dataset without HAD13a, we could perform a single isomorphous replacement with anomalous scattering (SIRAS) phasing from 3000 native and 4000 derivative indexed images. This reduction in the number of required images from 23,000 to 7000 in total was probably caused by isomorphous difference signals and anomalous difference signals providing complementary phase information. Thus, it is recommended that one collects a native dataset as well as derivative dataset(s) for efficient use of precious beamtime. We also confirmed that HAD13a can bind to crystals grown in the LCP and provides anomalous difference signals.
Researchers at the SACLA have also succeeded in SIRAS phasing using mercury (Yamashita et al. 2015) and SAD phasing using mercury or selenomethionine (Yamashita et al. 2017). Here, it was also demonstrated that indexing ambiguity in SFX does not prevent experimental phasing if treated properly. Although derivatization is necessary and sometimes non-trivial, heavier atoms provide stronger anomalous signals than sulfur atoms. In the most favorable case, one could phase from only 3000 indexed images by praseodymium SAD phasing (Sugahara et al. 2017). Selenium SAD phasing was also successful at the LCLS (Hunter et al. 2016).
Improvements in SFX data processing algorithms are underway. Indeed, the gadolinium SAD phasing of lysozyme initially required 60,000 indexed images (Barends et al. 2014) but only 7000 indexed images with newer programs and optimized processing protocols (Nass et al. 2016). Refinement of detector geometry and crystal orientations is now available in most programs (Ginn et al. 2015b; Kabsch 2014; White et al. 2016). Scaling by a multiplicative scale factor and a B factor can compensate for variations in crystals and XFEL pulses to some extent. It is sometimes useful to apply per-image resolution cutoff. Programs such as PRIME (Uervirojnangkoorn et al. 2015), cppxfel (Ginn et al. 2015a), and cxi.merge (Sauter 2015) can apply post-refinement to improve diffraction parameters and correct for the partiality. PRIME was used in the multiple isomorphous replacement with anomalous scattering (MIRAS) phasing of BinAB (Colletier et al. 2016). Efforts to improve data processing programs benefit not only experimental phasing but also other applications, especially time-resolved studies, where small differences between excited and ground state structures are analyzed. To support these efforts, we have uploaded raw diffraction images to the CXIDB (Maia 2012), so that others can reprocess our data for research and educational purposes. Since datasets differ in hit-finding thresholds, signal-to-noise levels, space group symmetry, asymmetric unit contents, and heterogeneities, comparison of the number of necessary images among different datasets is not very instructive. Instead, it is hoped that new algorithms will be tested on common, public datasets.
Enzyme chemistry based on damage-free room-temperature structures
As a model protein to study damage-free structures of metalloenzymes, we used copper-containing nitrite reductase (CuNiR), which is one of the key enzymes in microorganism denitrification and catalyzes one electron reduction of nitrite to nitric oxide (NO2 − + 2H+ + e− → NO + H2O). Typical CuNiRs are homotrimeric proteins containing two distinct Cu sites in each monomer. Type 1 Cu (T1Cu) with a Cys-Met-His2 ligand set is an electron acceptor, whereas type 2 Cu (T2Cu) with a His3 ligand set is a catalytic center located between two monomers. The two Cu sites are linked by an electron pathway (Cys-His bridge) and a sensor loop, which is thought to control electron distribution between T1Cu and T2Cu. Two conserved residues, Asp98 and His255 (Alcaligenes faecalis numbering), are located above the T2Cu site and bridged by one water molecule designated as bridging water (Fig. 2a). They are essential for the activity of CuNiRs because they assist proton transfer (PT) to NO2 − (Boulanger et al. 2000; Boulanger and Murphy 2001; Kataoka et al. 2000). Although electron transfer (ET) from T1Cu to T2Cu can occur in the resting state (RS) (Wijma et al. 2006), it is dramatically accelerated in the presence of NO2 − (Kobayashi et al. 1999; Leferink et al. 2011). One explanation for this gating-like phenomenon is that the binding of NO2 − raises the redox potential of T2Cu; however, the pH dependence of intramolecular ET in the presence of NO2 − cannot be explained by such a mechanism (Kobayashi et al. 1999). Instead, it was proposed by Kobayashi et al. that Cu-reduction-induced structural change of His255 is responsible for the gating-like mechanism, though no structural evidence had been obtained. Because it has been proven that intramolecular ET in CuNiR is accompanied by PT (Brenner et al. 2009; Leferink et al. 2011), one can speculate that intramolecular ET contributes to PT to NO2 − and that the structural change of His255 is involved in the proton-coupled ET (PCET) reaction.
SFX and synchrotron radiation crystallography (SRX) structures of Alcaligenes faecalis copper-containing nitrite reductase (CuNiR) (AfNiR). a The T2Cu site of the SFX resting state (RS) structure (PDB ID: 4YSC). b Structural difference of His255 between the SRX RS structure (yellow, PDB ID: 4YSE) and the SFX RS structure (green, PDB ID: 4YSC). Water molecules and Cu atoms are shown as spheres. The dashed yellow lines show hydrogen bonds. c Superposition of the SFX nitrite complex (NC) structure (red, PDB ID: 5D4I) on the SRX NC structure (blue, PDB ID: 5D4H)
Although crystal structures of CuNiRs have been determined by synchrotron radiation crystallography (SRX) for more than 20 years, SRX data might be affected by some problems related to the high radiation dose delivered to the crystals: (i) synchrotron X-rays cause photoreduction of metalloproteins as well as radiation damage to amino acid residues (Berglund et al. 2002; Schlichting et al. 2000) and the Cu centers in CuNiR are rapidly reduced by synchrotron X-rays (Antonyuk and Hough 2011; Hough et al. 2008); (ii) after the photoreduction of T2Cu, NO2 − is then reduced to produce NO and water, which complicate the interpretation of the electron density; and (iii) cryogenic manipulations for reducing radiation damages in SRX can change the structural population of amino acid residues (Fraser et al. 2009, 2011) and enzyme–substrate complexes (Fukuda and Inoue 2015; Keedy et al. 2014). Therefore, we applied SFX with XFELs, which enables damage-free structural determination of metalloproteins (Johansson et al. 2013; Kern et al. 2012) and evaluation of the native conformational population at room temperature (Liu et al. 2013).
We determined the SFX and SRX structures of Alcaligenes faecalis CuNiR (AfNiR) in the RS and the nitrite complex (NC) form and analyzed them (Fukuda et al. 2016a). Compared with the imidazole ring of enzymatically important His255 in the SFX RS structure, that in the SRX RS structure rotated about 20° and, hence, the H-bond partners of His255 were switched from the carbonyl O atom of Glu279 to the hydroxyl O atom of Thr280 (Fig. 2b). Interestingly, this pair of Glu (or Gln) and Thr (or Ser) is conserved in CuNiRs. Combining SFX and SRX, we also showed that the imidazole ring of His244 in Geobacillus thermodenitrificans CuNiR, which corresponds to His255 in AfNiR, slightly rotated as a result of photoreduction (Fukuda et al. 2016b). Therefore, the structural change of this enzymatically important His is thought to be caused by the reduction of Cu, as was predicted previously (Kobayashi et al. 1999). Activity assay with a T280V mutant of AfNiR further supported that the rotated state of His255 is a transient conformation important for the CuNiR activity because the activity of the T280V mutant was significantly lower than that of the wild-type enzyme. Since Val280 in the T280V mutant lacks the hydroxyl O atom that can form an H-bond with His255, the rotation of His255 is inhibited in this mutant. On the basis of these findings, we proposed that the switching of the H-bond partners of His255 may facilitate the PT reaction from His255 to bridging water. Because the hydroxyl O atom of Thr280 is less negatively charged than the carbonyl O atom of Gln279, the Nδ1 atom of His255 makes a longer and weaker H-bond with Thr280. As a result, the H atom is more attracted to the Nδ1 atom of His255 and a proton on the Nε2 atom is transferred to bridging water. It is noteworthy that recent computational chemistry supports indirect PT from His255 to NO2 − via bridging water, which is coupled with T2Cu reduction (Li et al. 2015). While T2Cu in the SRX NC structure was coordinated by NO2 − that showed the near face-on mode as was observed in known CuNiR structures (Antonyuk et al. 2005; Tocheva et al. 2004), NO2 − featuring a more vertical binding mode was observed at T2Cu in the SFX NC structure (Fig. 2c). A recent “multiple structures serially from one crystal” experiment also revealed that the low-dose SRX NC structure of Achromobacter cycloclastes CuNiR showed a conformation of NO2 − similar to our vertical binding mode (Horrell et al. 2016). Therefore, the near face-on mode which has been observed in reported SRX structures of CuNiRs is thought to correspond to the binding mode when T2Cu is photoreduced. The structural change of NO2 − upon Cu reduction may partly explain why PCET in CuNiR is affected by the presence of NO2 −. Because the N atom of NO2 − becomes closer to His255 through its conformational change from vertical to near face-on, the rotated conformation of the imidazole ring of His255 might be preferred due to steric hindrance and, hence, hinder reverse PT from bridging water to His255. To summarize, comparison between SFX and SRX structures could provide a more detailed description of the reaction mechanism in CuNiR; therefore, we think that this approach can also be applied to other metalloenzymes.
Molecular movie taken by time-resolved SFX
A protein’s structure is closely related to its function. X-ray crystallography has contributed to ternary structure determination for precise understanding of how proteins work. However, most protein structures obtained by conventional crystallography are in a static state before or after the molecule functions. While time-resolved crystallographic methods, including Laue diffraction and trapping of reaction intermediates (Moffat 2001), have been used to observe the conformational changes in proteins that are at work, there are several difficulties, such as radiation damage from synchrotron sources, the time resolution, and other technical issues. SFX, using intense, femtosecond X-ray pulses from XFELs, allows the determination of protein structures at room temperature using diffraction patterns from microcrystals before the onset of radiation damage. The technique is of great utility for time-resolved experiments in measuring ultrafast reactions in proteins when combined with the pump-probe approach, in which a brief laser pulse initiates a light-dependent reaction in the molecule. Light-triggered reaction in microcrystals has the advantages of synchronization of the reaction and efficient optical excitation.
As the first results using pump-probe time-resolved SFX, two studies of photosystem II (Kern et al. 2014; Kupitz et al. 2014) were reported, but their interpretation remains provisional, in part because of the low resolution (4.5 to 5 Å). Tenboer et al. (2014) showed structural changes of photoactive yellow protein (PYP) at delay times of 10 ns and 1 μs with high resolution (1.6 Å). Pande et al. (2016) revealed a snapshot series of PYP intermediates over a time range from 100 fs to 3 ps. Barends et al. (2015) reported ultrafast structural changes in a myoglobin–CO complex upon photolysis in a time range from ~ 100 fs to 150 ps. In these experiments, ultrafast reactions were observed using a liquid-jet injector. Since microcrystals travel very fast in the jet (typical flow speed ca. 10 m/s), the range of available delay times is currently limited. On ultrafast time scales, protein motions are local and often involve only chromophore or amino acid side chains. In contrast, motions on slower time scales (nanoseconds to milliseconds) might involve larger conformational changes in proteins at work.
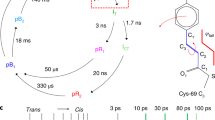
Recently, a pump-probe time-resolved SFX setup using a nanosecond laser has been developed at the SACLA to observe such protein movements on a slower time scale (Kubo et al. 2017). Pump conditions, including wavelength and focal size, are readily adjustable using an optical-fiber-based setup, so users can collect as much data as possible in their limited XFEL beam time. In the setup, an excitation (pump) laser illuminates microcrystals from two directions to excite molecules in crystal lattices with high efficiency. Using the setup, as the first result, we successfully observed structural changes in bR, a light-driven proton pump derived from Halobacterium salinarum (Nango et al. 2016). The protein consists of seven transmembrane helices and contains a retinal chromophore that is covalently bound to Lys216. The all-trans retinal undergoes isomerization to the 13-cis configuration on absorption of light, causing structural change in the protein via several intermediates. The structural change results in PT from the cytoplasmic side to the extracellular side, and the protein recovers the initial state, completing the photocycle. Considerable effort has been made to understand how structural changes in bR transport a proton uphill against a transmembrane potential (Haupts et al. 1999; Neutze et al. 2002; Wickstrand et al. 2015). Cryotrapping studies of bR from many laboratories provided structural information about structural changes during the photocycle. Despite the success, these results have been criticized because of radiation damage (Borshchevskiy et al. 2011, 2014; Matsui et al. 2002) and disagreement among the structures from various groups (Wickstrand et al. 2015). We circumvented these concerns by recording a three-dimensional movie of structural changes in bR at room temperature at 2.1 Å resolution using time-resolved SFX at the SACLA. We observed conformational changes in bR at 13 time points from nanoseconds to milliseconds following photoactivation (Fig. 3). Our data revealed that an initially twisted retinal displaces Trp182 and Leu93 towards the cytoplasm and allows a water molecule (W452) to order between Leu93, Thr89, and the Schiff base (SB) on the retinal in the L-state. Hydrogen-bonding interactions from the protonated SB, which is a proton donor to Wat452 or Thr89, create a pathway for PT to a proton acceptor, Asp85. This observation explains how the SB makes contact with Asp85 despite being turned towards the cytoplasmic side by photoisomerization. We also found that, once a proton is transferred, the hydrogen-bonding interaction between Asp85 and Thr89 is lost, which breaks the connectivity to the extracellular side of the protein. Our results clarified how structural changes in bR achieve unidirectional proton transport.
Four snapshots from a molecular movie of bacteriorhodopsin (bR) taken by time-resolved SFX. Structures of bR at four selected delay time points—16 ns, 760 ns, 36.2 μs, and 1725 μs—are drawn as ribbon models. Blue and yellow meshes represent positive and negative |F obs|light–|F obs|dark difference Fourier electron density maps, respectively
New insight into water oxidation in photosystem II has been proposed recently based on the results of time-resolved SFX at the SACLA (Suga et al. 2017). Now, many users perform time-resolved SFX experiments using the setup at the SACLA and capture the movement of various reacting proteins. In the LCLS, a different type of reaction trigger, mixing with a ligand or a substrate, has been used for visualizing structural changes in riboswitch RNA (Stagno et al. 2017) and in β-lactamase (Kupitz et al. 2017). Time-resolved SFX techniques will open a new era of study of protein structural dynamics and provide unprecedented data for understanding protein functions.
Outlook for dynamic structural biology
This review highlights some recent achievements of the SACLA-SFX Project. The SFX measurement methods developed in the project will be applicable to a wide range of target biological macromolecules in structural biology using XFEL facilities around the world. Following the LCLS and the SACLA, newly constructed XFEL facilities—European XFEL in Germany, Pohang Accelerator Laboratory (PAL)-XFEL in South Korea, and SwissFEL in Switzerland—start user operations in 2017 or 2018. Data processing programs for SFX are making remarkable progress each year. Development of more sophisticated programs will accelerate the structure determination and increase the resolution of determined structures. Especially, molecular movies of many kinds of enzymes will be taken by time-resolved SFX by visualizing whole pictures of three-dimensional structural change that occur from moment to moment during catalytic reactions, to elucidate the dynamic structure–function relationships of enzymes.
Together with the rise of SFX, the evolution of cryo-electron microscopy (Cryo-EM) technology in recent years has revolutionized structural biology. In the past, the resolution of single-particle analysis was about 10 Å, but today, it has reached 1.8 Å with the advent of direct electron detectors and advanced software (Merk et al. 2016). It is becoming possible to observe unstable protein complexes and study the conformational landscape of proteins (Dashti et al. 2014, 2017). XFEL and Cryo-EM techniques can provide complementary information on protein structures and dynamics. The breakthroughs in XFEL crystallography and Cryo-EM will bring about dramatic progress in our understanding of life phenomena at the molecular level, and lead to innovation in medicine, for example, by aiding drug design that targets dynamic protein–protein interactions.
References
Antonyuk SV, Hough MA (2011) Monitoring and validating active site redox states in protein crystals. Biochim Biophys Acta 1814(6):778–784. https://doi.org/10.1016/j.bbapap.2010.12.017
Antonyuk SV, Strange RW, Sawers G, Eady RR, Hasnain SS (2005) Atomic resolution structures of resting-state, substrate- and product-complexed Cu-nitrite reductase provide insight into catalytic mechanism. Proc Natl Acad Sci U S A 102(34):12041–12046. https://doi.org/10.1073/pnas.0504207102
Barends TR, Foucar L, Botha S, Doak RB, Shoeman RL, Nass K et al (2014) De novo protein crystal structure determination from X-ray free-electron laser data. Nature 505(7482):244–247. https://doi.org/10.1038/nature12773
Barends TRM, Foucar L, Ardevol A, Nass K, Aquila A, Botha S et al (2015) Direct observation of ultrafast collective motions in CO myoglobin upon ligand dissociation. Science 350(6259):445–450. https://doi.org/10.1126/science.aac5492
Barty A, Caleman C, Aquila A, Timneanu N, Lomb L, White TA et al (2012) Self-terminating diffraction gates femtosecond X-ray nanocrystallography measurements. Nat Photonics 6:35–40. https://doi.org/10.1038/nphoton.2011.297
Batyuk A, Galli L, Ishchenko A, Han GW, Gati C, Popov PA et al (2016) Native phasing of x-ray free-electron laser data for a G protein-coupled receptor. Sci Adv 2(9):e1600292. https://doi.org/10.1126/sciadv.1600292
Berglund GI, Carlsson GH, Smith AT, Szöke H, Henriksen A, Hajdu J (2002) The catalytic pathway of horseradish peroxidase at high resolution. Nature 417(6887):463–468. https://doi.org/10.1038/417463a
Borshchevskiy VI, Round ES, Popov AN, Büldt G, Gordeliy VI (2011) X-ray-radiation-induced changes in bacteriorhodopsin structure. J Mol Biol 409(5):813–825. https://doi.org/10.1016/j.jmb.2011.04.038
Borshchevskiy V, Round E, Erofeev I, Weik M, Ishchenko A, Gushchin I et al (2014) Low-dose X-ray radiation induces structural alterations in proteins. Acta Crystallogr D Biol Crystallogr 70:2675–2685. https://doi.org/10.1107/S1399004714017295
Boulanger MJ, Murphy MEP (2001) Alternate substrate binding modes to two mutant (D98N and H255N) forms of nitrite reductase from Alcaligenes faecalis S-6: structural model of a transient catalytic intermediate. Biochemistry 40(31):9132–9141. https://doi.org/10.1021/bi0107400
Boulanger MJ, Kukimoto M, Nishiyama M, Horinouchi S, Murphy MEP (2000) Catalytic roles for two water bridged residues (Asp-98 and His-255) in the active site of copper-containing nitrite reductase. J Biol Chem 275(31):23957–23964. https://doi.org/10.1074/jbc.M001859200
Brenner S, Heyes DJ, Hay S, Hough MA, Eady RR, Hasnain SS, Scrutton NS (2009) Demonstration of proton-coupled electron transfer in the copper-containing nitrite reductases. J Biol Chem 284(38):25973–25983. https://doi.org/10.1074/jbc.M109.012245
Chapman HN, Fromme P, Barty A, White TA, Kirian RA, Aquila A et al (2011) Femtosecond X-ray protein nanocrystallography. Nature 470(7332):73–77. https://doi.org/10.1038/nature09750
Colletier JP, Sawaya MR, Gingery M, Rodriguez JA, Cascio D, Brewster AS et al (2016) De novo phasing with X-ray laser reveals mosquito larvicide BinAB structure. Nature 539(7627):43–47. https://doi.org/10.1038/nature19825
Cowtan K (2006) The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr D Biol Crystallogr 62(Pt 9):1002–1011. https://doi.org/10.1107/S0907444906022116
Dashti A, Schwander P, Langlois R, Fung R, Li W, Hosseinizadeh A et al (2014) Trajectories of the ribosome as a Brownian nanomachine. Proc Natl Acad Sci U S A 111(49):17492–17497. https://doi.org/10.1073/pnas.1419276111
Dashti A, Hail DB, Mashayekhi G, Schwander P, des Georges A, Frank J et al (2017) Conformational dynamics and energy landscapes of ligand binding in RyR1. bioRxiv 167080. https://doi.org/10.1101/167080
Fraser JS, Clarkson MW, Degnan SC, Erion R, Kern D, Alber T (2009) Hidden alternative structures of proline isomerase essential for catalysis. Nature 462(7273):669–673. https://doi.org/10.1038/nature08615
Fraser JS, van den Bedem H, Samelson AJ, Lang PT, Holton JM, Echols N et al (2011) Accessing protein conformational ensembles using room-temperature X-ray crystallography. Proc Natl Acad Sci U S A 108(39):16247–16252. https://doi.org/10.1073/pnas.1111325108
Fukuda Y, Inoue T (2015) High-temperature and high-resolution crystallography of thermostable copper nitrite reductase. Chem Commun (Camb) 51(30):6532–6535. https://doi.org/10.1039/c4cc09553g
Fukuda Y, Tse KM, Nakane T, Nakatsu T, Suzuki M, Sugahara M et al (2016a) Redox-coupled proton transfer mechanism in nitrite reductase revealed by femtosecond crystallography. Proc Natl Acad Sci U S A 113(11):2928–2933. https://doi.org/10.1073/pnas.1517770113
Fukuda Y, Tse KM, Suzuki M, Diederichs K, Hirata K, Nakane T et al (2016b) Redox-coupled structural changes in nitrite reductase revealed by serial femtosecond and microfocus crystallography. J Biochem 159(5):527–538. https://doi.org/10.1093/jb/mvv133
Ginn HM, Brewster AS, Hattne J, Evans G, Wagner A, Grimes JM et al (2015a) A revised partiality model and post-refinement algorithm for X-ray free-electron laser data. Acta Crystallogr D Biol Crystallogr 71(Pt 6):1400–1410. https://doi.org/10.1107/S1399004715006902
Ginn HM, Messerschmidt M, Ji X, Zhang H, Axford D, Gildea RJ et al (2015b) Structure of CPV17 polyhedrin determined by the improved analysis of serial femtosecond crystallographic data. Nat Commun 6:6435. https://doi.org/10.1038/ncomms7435
Haupts U, Tittor J, Oesterhelt D (1999) Closing in on bacteriorhodopsin: progress in understanding the molecule. Annu Rev Biophys Biomol Struct 28:367–399. https://doi.org/10.1146/annurev.biophys.28.1.367
Horrell S, Antonyuk SV, Eady RR, Hasnain SS, Hough MA, Strange RW (2016) Serial crystallography captures enzyme catalysis in copper nitrite reductase at atomic resolution from one crystal. IUCrJ 3(Pt 4):271–281. https://doi.org/10.1107/S205225251600823X
Hough MA, Antonyuk SV, Strange RW, Eady RR, Hasnain SS (2008) Crystallography with online optical and X-ray absorption spectroscopies demonstrates an ordered mechanism in copper nitrite reductase. J Mol Biol 378(2):353–361. https://doi.org/10.1016/j.jmb.2008.01.097
Hunter MS, Yoon CH, DeMirci H, Sierra RG, Dao EH, Ahmadi R et al (2016) Selenium single-wavelength anomalous diffraction de novo phasing using an X-ray-free electron laser. Nat Commun 7:13388. https://doi.org/10.1038/ncomms13388
Ishikawa T, Aoyagi H, Asaka T, Asano Y, Azumi N, Bizen T et al (2012) A compact X-ray free-electron laser emitting in the sub-angstrom region. Nat Photonics 6(8):540–544. https://doi.org/10.1038/nphoton.2012.141
Johansson LC, Arnlund D, Katona G, White TA, Barty A, DePonte DP et al (2013) Structure of a photosynthetic reaction centre determined by serial femtosecond crystallography. Nat Commun 4:2911. https://doi.org/10.1038/ncomms3911
Kabsch W (2014) Processing of X-ray snapshots from crystals in random orientations. Acta Crystallogr D Biol Crystallogr 70(Pt 8):2204–2216
Kataoka K, Furusawa H, Takagi K, Yamaguchi K, Suzuki S (2000) Functional analysis of conserved aspartate and histidine residues located around the type 2 copper site of copper-containing nitrite reductase. J Biochem 127(2):345–350
Keedy DA, van den Bedem H, Sivak DA, Petsko GA, Ringe D, Wilson MA et al (2014) Crystal cryocooling distorts conformational heterogeneity in a model Michaelis complex of DHFR. Structure 22(6):899–910. https://doi.org/10.1016/j.str.2014.04.016
Kern J, Alonso-Mori R, Hellmich J, Tran R, Hattne J, Laksmono H et al (2012) Room temperature femtosecond X-ray diffraction of photosystem II microcrystals. Proc Natl Acad Sci U S A 109(25):9721–9726. https://doi.org/10.1073/pnas.1204598109
Kern J, Tran R, Alonso-Mori R, Koroidov S, Echols N, Hattne J et al (2014) Taking snapshots of photosynthetic water oxidation using femtosecond X-ray diffraction and spectroscopy. Nat Commun 5:4371. https://doi.org/10.1038/ncomms5371
Kirian RA, White TA, Holton JM, Chapman HN, Fromme P, Barty A et al (2011) Structure–factor analysis of femtosecond microdiffraction patterns from protein nanocrystals. Acta Crystallogr A 67(Pt 2):131–140. https://doi.org/10.1107/S0108767310050981
Kobayashi K, Tagawa S, Deligeer, Suzuki S (1999) The pH-dependent changes of intramolecular electron transfer on copper-containing nitrite reductase. J Biochem 126(2):408–412
Kubo M, Nango E, Tono K, Kimura T, Owada S, Song CY et al (2017) Nanosecond pump–probe device for time-resolved serial femtosecond crystallography developed at SACLA. J Synchrotron Radiat 24:1086–1091. https://doi.org/10.1107/S160057751701030x
Kupitz C, Basu S, Grotjohann I, Fromme R, Zatsepin NA, Rendek KN et al (2014) Serial time-resolved crystallography of photosystem II using a femtosecond X-ray laser. Nature 513(7517):261–265. https://doi.org/10.1038/nature13453
Kupitz C, Olmos JL Jr, Holl M, Tremblay L, Pande K, Pandey S et al (2017) Structural enzymology using X-ray free electron lasers. Struct Dyn 4(4):044003. https://doi.org/10.1063/1.4972069
Leferink NGH, Han C, Antonyuk SV, Heyes DJ, Rigby SEJ, Hough MA et al (2011) Proton-coupled electron transfer in the catalytic cycle of Alcaligenes xylosoxidans copper-dependent nitrite Reductase. Biochemistry 50(19):4121–4131. https://doi.org/10.1021/bi200246f
Li Y, Hodak M, Bernholc J (2015) Enzymatic mechanism of copper-containing nitrite reductase. Biochemistry 54(5):1233–1242. https://doi.org/10.1021/bi5007767
Liu Q, Hendrickson WA (2015) Crystallographic phasing from weak anomalous signals. Curr Opin Struct Biol 34:99–107. https://doi.org/10.1016/j.sbi.2015.08.003
Liu W, Wacker D, Gati C, Han GW, James D, Wang D et al (2013) Serial femtosecond crystallography of G protein-coupled receptors. Science 342(6165):1521–1524. https://doi.org/10.1126/science.1244142
MacKay DJC (1998) Introduction to monte carlo methods. NATO ASI Ser D Behav Soc Sci 89:175–204
Mafuné F, Miyajima K, Tono K, Takeda Y, Kohno JY, Miyauchi N et al (2016) Microcrystal delivery by pulsed liquid droplet for serial femtosecond crystallography. Acta Crystallogr D Struct Biol 72:520–523. https://doi.org/10.1107/S2059798316001480
Maia FRNC (2012) The coherent X-ray imaging data bank. Nat Methods 9(9):854–855. https://doi.org/10.1038/nmeth.2110
Matsui Y, Sakai K, Murakami M, Shiro Y, Adachi S, Okumura H et al (2002) Specific damage induced by X-ray radiation and structural changes in the primary photoreaction of bacteriorhodopsin. J Mol Biol 324(3):469–481. https://doi.org/10.1016/S0022-2836(02)01110-5
Merk A, Bartesaghi A, Banerjee S, Falconieri V, Rao P, Davis MI et al (2016) Breaking Cryo-EM resolution barriers to facilitate drug discovery. Cell 165(7):1698–1707. https://doi.org/10.1016/j.cell.2016.05.040
Moffat K (2001) Time-resolved biochemical crystallography: a mechanistic perspective. Chem Rev 101(6):1569–1581. https://doi.org/10.1021/cr990039q
Nakane T, Song C, Suzuki M, Nango E, Kobayashi J, Masuda T et al (2015) Native sulfur/chlorine SAD phasing for serial femtosecond crystallography. Acta Crystallogr D Biol Crystallogr 71(Pt 12):2519–2525. https://doi.org/10.1107/S139900471501857X
Nakane T, Hanashima S, Suzuki M, Saiki H, Hayashi T, Kakinouchi K et al (2016a) Membrane protein structure determination by SAD, SIR, or SIRAS phasing in serial femtosecond crystallography using an iododetergent. Proc Natl Acad Sci U S A 113(46):13039–13044. https://doi.org/10.1073/pnas.1602531113
Nakane T, Joti Y, Tono K, Yabashi M, Nango E, Iwata S et al (2016b) Data processing pipeline for serial femtosecond crystallography at SACLA. J Appl Crystallogr 49:1035–1041. https://doi.org/10.1107/S1600576716005720
Nango E, Royant A, Kubo M, Nakane T, Wickstrand C, Kimura T et al (2016) A three-dimensional movie of structural changes in bacteriorhodopsin. Science 354(6319):1552–1557. https://doi.org/10.1126/science.aah3497
Nass K, Meinhart A, Barends TR, Foucar L, Gorel A, Aquila A et al (2016) Protein structure determination by single-wavelength anomalous diffraction phasing of X-ray free-electron laser data. IUCrJ 3(Pt 3):180–191. https://doi.org/10.1107/S2052252516002980
Neutze R, Wouts R, van der Spoel D, Weckert E, Hajdu J (2000) Potential for biomolecular imaging with femtosecond X-ray pulses. Nature 406(6797):752–757. https://doi.org/10.1038/35021099
Neutze R, Pebay-Peyroula E, Edman K, Royant A, Navarro J, Landau EM (2002) Bacteriorhodopsin: a high-resolution structural view of vectorial proton transport. Biochim Biophys Acta Biomembr 1565(2):144–167. https://doi.org/10.1016/S0005-2736(02)00566-7
Pande K, Hutchison CDM, Groenhof G, Aquila A, Robinson JS, Tenboer J et al (2016) Femtosecond structural dynamics drives the trans/cis isomerization in photoactive yellow protein. Science 352(6286):725–729. https://doi.org/10.1126/science.aad5081
Sauter NK (2015) XFEL diffraction: developing processing methods to optimize data quality. J Synchrotron Radiat 22(Pt 2):239–248. https://doi.org/10.1107/S1600577514028203
Scapin G (2013) Molecular replacement then and now. Acta Crystallogr D Biol Crystallogr 69(Pt 11):2266–2275
Schlichting I, Berendzen J, Chu K, Stock AM, Maves SA, Benson DE et al (2000) The catalytic pathway of cytochrome p450cam at atomic resolution. Science 287(5458):1615–1622
Sheldrick GM (2010) Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr D Biol Crystallogr 66(Pt 4):479–485. https://doi.org/10.1107/S0907444909038360
Stagno JR, Liu Y, Bhandari YR, Conrad CE, Panja S, Swain M et al (2017) Structures of riboswitch RNA reaction states by mix-and-inject XFEL serial crystallography. Nature 541(7636):242–246. https://doi.org/10.1038/nature20599
Suga M, Akita F, Sugahara M, Kubo M, Nakajima Y, Nakane T et al (2017) Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL. Nature 543(7643):131–135. https://doi.org/10.1038/nature21400
Sugahara M, Mizohata E, Nango E, Suzuki M, Tanaka T, Masuda T et al (2015) Grease matrix as a versatile carrier of proteins for serial crystallography. Nat Methods 12(1):61–63. https://doi.org/10.1038/nmeth.3172
Sugahara M, Nakane T, Masuda T, Suzuki M, Inoue S, Song CY et al (2017) Hydroxyethyl cellulose matrix applied to serial crystallography. Sci Rep 7:703. https://doi.org/10.1038/s41598-017-00761-0
Taylor GL (2010) Introduction to phasing. Acta Crystallogr D Biol Crystallogr 66(Pt 4):325–338
Tenboer J, Basu S, Zatsepin N, Pande K, Milathianaki D, Frank M et al (2014) Time-resolved serial crystallography captures high-resolution intermediates of photoactive yellow protein. Science 346(6214):1242–1246. https://doi.org/10.1126/science.1259357
Tocheva EI, Rosell FI, Mauk AG, Murphy ME (2004) Side-on copper-nitrosyl coordination by nitrite reductase. Science 304(5672):867–870. https://doi.org/10.1126/science.1095109
Tono K, Nango E, Sugahara M, Song C, Park J, Tanaka T et al (2015) Diverse application platform for hard X-ray diffraction in SACLA (DAPHNIS): application to serial protein crystallography using an X-ray free-electron laser. J Synchrotron Radiat 22(Pt 3):532–537. https://doi.org/10.1107/S1600577515004464
Uervirojnangkoorn M, Zeldin OB, Lyubimov AY, Hattne J, Brewster AS, Sauter NK et al (2015) Enabling X-ray free electron laser crystallography for challenging biological systems from a limited number of crystals. Elife 4:e05421. https://doi.org/10.7554/eLife.05421
Weierstall U, James D, Wang C, White TA, Wang D, Liu W et al (2014) Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography. Nat Commun 5:3309. https://doi.org/10.1038/ncomms4309
White TA, Kirian RA, Martin AV, Aquila A, Nass K, Barty A et al (2012) CrystFEL: a software suite for snapshot serial crystallography. J Appl Crystallogr 45:335–341. https://doi.org/10.1107/S0021889812002312
White TA, Mariani V, Brehm W, Yefanov O, Barty A, Beyerlein KR et al (2016) Recent developments in CrystFEL. J Appl Crystallogr 49(Pt 2):680–689. https://doi.org/10.1107/S1600576716004751
Wickstrand C, Dods R, Royant A, Neutze R (2015) Bacteriorhodopsin: would the real structural intermediates please stand up? Biochim Biophys Acta Gen Subj 1850(3):536–553. https://doi.org/10.1016/j.bbagen.2014.05.021
Wijma HJ, Jeuken LJC, Verbeet MP, Armstrong FA, Canters GW (2006) A random-sequential mechanism for nitrite binding and active site reduction in copper-containing nitrite reductase. J Biol Chem 281(24):16340–16346. https://doi.org/10.1074/jbc.M601610200
Yamashita K, Pan D, Okuda T, Sugahara M, Kodan A, Yamaguchi T et al (2015) An isomorphous replacement method for efficient de novo phasing for serial femtosecond crystallography. Sci Rep 5:14017. https://doi.org/10.1038/srep14017
Yamashita K, Kuwabara N, Nakane T, Murai T, Mizohata E, Sugahara M et al (2017) Experimental phase determination with selenomethionine or mercury-derivatization in serial femtosecond crystallography. IUCrJ 4:639–647. https://doi.org/10.1107/S2052252517008557
Acknowledgements
We appreciate all the members of the SACLA-SFX Project, especially Dr. Keitaro Yamashita for the valuable comments on the manuscript. This work was supported by the X-ray Free Electron Laser Priority Strategy Program of the Ministry of Education, Culture, Sports, Science and Technology in Japan and partially by the Strategic Basic Research Programs of the Japan Science and Technology Agency.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Eiichi Mizohata declares that he has no conflict of interest. Takanori Nakane declares that he has no conflict of interest. Yohta Fukuda declares that he has no conflict of interest. Eriko Nango declares that she has no conflict of interest. So Iwata declares that he has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
This article is part of a Special Issue on ‘Biomolecules to Bio-nanomachines - Fumio Arisaka 70th Birthday’ edited by Damien Hall, Junichi Takagi and Haruki Nakamura.
Rights and permissions
About this article
Cite this article
Mizohata, E., Nakane, T., Fukuda, Y. et al. Serial femtosecond crystallography at the SACLA: breakthrough to dynamic structural biology. Biophys Rev 10, 209–218 (2018). https://doi.org/10.1007/s12551-017-0344-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-017-0344-9