Abstract
Eukaryotic cell surfaces are decorated with a complex array of glycoconjugates that are usually capped with sialic acids, a large family of over 50 structurally distinct nine-carbon amino sugars, the most common member of which is N-acetylneuraminic acid. Once made available through the action of neuraminidases, bacterial pathogens and commensals utilise host-derived sialic acid by degrading it for energy or repurposing the sialic acid onto their own cell surface to camouflage the bacterium from the immune system. A functional sialic acid transporter has been shown to be essential for the uptake of sialic acid in a range of human bacterial pathogens and important for host colonisation and persistence. Here, we review the state-of-play in the field with respect to the molecular mechanisms by which these bio-nanomachines transport sialic acids across bacterial cell membranes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eukaryotic cell surfaces are decorated with complex glycoconjugates, such as glycoproteins, glycolipids and lipopolysaccharides. Negatively charged sialic acids are often found as the terminal sugar of these cell surface glycoconjugates, where they mediate a diverse array of cellular interactions, recognition and adhesion processes. Sialic acids comprise a large family of over 50 structurally distinct nine-carbon amino sugars (Fig. 1), the most common member of which is known as N-acetylneuraminic acid. Derivatives of this molecule carry various substituents at the amino or hydroxyl groups across the molecule (see the review by Vimr et al. 2004).
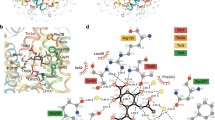
Structure of sialic acids and overview of bacterial sialic acid utilisation. a The most common sialic acid, N-acetylneuraminic acid. An asterisk represents positions that are modified to give variation. These include acetyl groups or the less common lactyl, phosphate, succinyl and methyl groups. b Lactone derivative, the third highest detected sialyl metabolite in the human gut. c 9-O-Acetylated derivative, the second highest detected sialyl metabolite in the human gut. d N-Glycolylneuraminic acid. Differing at the C-5 acetamido group, this sialyl metabolite is common in most mammals, but is not synthesised by humans
Human mucus-rich environments, such as the respiratory or gastrointestinal tracts, contain sialic acid-coated glycoconjugates and bacteria that colonise these environments have evolved mechanisms to use host-derived sialic acid for a competitive advantage (see the review by Lewis and Lewis 2012). Almagro-Moreno and Boyd (2009) found that sialic acid catabolism is largely confined to commensal or pathogenic bacteria and that most of these species colonise sialic acid-rich areas, such as the respiratory or gastrointestinal tracts, suggesting a link between sialic acid uptake/utilisation and survival in vivo. Under normal physiological conditions, sialic acid is covalently bound to glycoconjugates, and, by definition, is unavailable for utilisation by bacteria. The concentration of free sialic acid in the human respiratory and gastrointestinal tract is not well characterised, although in human serum, for example, the concentration of sialic acid is 1.6–2.2 mM, while the free concentration is ~ 0.5–3 μM, representing a relative free abundance of less than 0.2% (Sillanaukee et al. 1999).
To make sialic acid available, bacterial pathogens release it from the glycoconjugate using a neuraminidase enzyme, which hydrolyses the linkage between the sialic acid molecule and the subterminal sugar of the glycoconjugate (reviewed by Lewis and Lewis 2012). Neuraminidases are produced either endogenously by the host in response to inflammation or exogenously by neuraminidase expressing bacteria present in the sialylated niche (see the review by Vimr et al. 2004 and references therein).
Once available, bacterial pathogens utilise host-derived sialic acids in one of two ways. Firstly, using a strategy known as molecular mimicry, pathogens can incorporate sialic acid into their cell surface macromolecules, to essentially circumvent the host’s innate immune response (Bouchet et al. 2003; Vimr et al. 2004; Severi et al. 2007). At least four methods of cell surface sialylation have evolved among bacterial pathogens, including precursor scavenging, de novo biosynthesis, donor scavenging and trans-neuraminidase activity (reviewed by Vimr et al. 2004). Secondly, sialic acid can be metabolised as a source of carbon, nitrogen and energy (Vimr and Troy 1985; Olson et al. 2013). The utilisation of host-derived sialic acid provides an alternative nutrient source for bacterial pathogens and a possible selective advantage in the human host. The enzymology and structural biology of sialic acid catabolism is well established (for example, see North et al. 2013, 2014a, b, 2016; Caing-Carlsson et al. 2017), although some mechanistic details are yet to be elucidated.
A functional sialic acid transporter is essential for the uptake of sialic acid in a range of human bacterial pathogens, including Escherichia coli (Vimr and Troy 1985), Haemophilus influenzae (Severi et al. 2005) and Salmonella enterica (Severi et al. 2010). This suggests that these organisms have a dedicated transporter that is the sole route for sialic acid uptake. Moreover, disruption of the genes encoding sialic acid transporters impairs outgrowth of Salmonella enterica serovar Typhimurium and Clostridium difficile during post-antibiotic expansion (Ng et al. 2013), and of E. coli during intestinal inflammation (Huang et al. 2015). In vivo mouse models demonstrate that sialic acid import and utilisation is critical for pathogen colonisation and persistence of E. coli (Chang et al. 2004), Vibrio cholerae (Almagro-Moreno and Boyd 2009), V. vulnificus (Jeong et al. 2009) and Streptococcus agalactiae (Pezzicoli et al. 2012). Humans synthesise N-acetyl neuraminic acid and have dedicated membrane transporters to deploy it onto their surface, although these share little homology to the bacterial transporters. Thus, developing novel inhibitors that target these transporters is a plausible strategy for blocking bacterial pathogens. Here, we review the literature regarding bacterial sialic acid import. To date, there are no structural data to explain how these transporters mediate the transport of specific sialic acids across the cytoplasmic membrane, a key limitation in the field. Understanding this will be critical for the design of future antibiotics against these targets.
Import of sialic acid across the outer membrane in Gram-negative bacteria
For Gram-negative bacteria, sialic acid must first cross the outer cell membrane before it can be imported into the cell (Fig. 2). This can occur by simple diffusion through non-specific porins, such as the outer membrane proteins OmpF and OmpC via facilitated diffusion through specific channels, such as the N-acetylneuraminic acid-inducible outer membrane channel NanC, or using active transport mediated by TonB-dependent outer membrane protein receptors.
Sialic acid transporter types. Within Gram-negative bacteria, sialic acid must first cross the outer cell membrane before it can be imported into the cell. This can occur by simple diffusion through the non-specific porins, OmpF and OmpC (grouped together). Specific channels, such as NanC, allow for the facilitated diffusion of sialic acid, whereas the NanOU system allows for the active transport of sialic acid. This is mediated by an extracellular uptake protein NanU, which binds sialic acid, the β-barrel NanO and the TonB complex, which energises the transport. Once in the periplasmic space, the processing enzymes NanM (yellow square) and NanS (yellow triangle) are known to mutarotate between the anomeric states of sialic acid and deacetylate the 9 position of 9-O-acetylated derivative, respectively. Some transporter systems require a periplasmic or cell surface-associated substrate binding protein (green circles) which interacts with the membrane component of the respective transporter. Finally, to traverse the inner membrane, bacterial pathogens have evolved multiple mechanisms of sialic acid transport which vary between species. ABC transporters are primary transporters that couple solute translocation with the hydrolysis of ATP by the ATPase domains. TRAP, NanT and SiaT transporters are secondary transporters that couple solute translocation with an electrochemical gradient. It is unclear whether TRAP sialic acid transporters are proton- or sodium ion-dependent, but NanT sialic acid transporters have been shown to be proton-dependent, while SiaT sialic acid transporters have been shown to be sodium ion-dependent. The stoichiometry of ions required for transport is not well understood. YjhB is a putative permease, which is similar in topology to NanT, although it lacks the central hydrophilic domain. While the other transporters are believed to transport N-acetylneuraminic acid, YjhB is suggested to transport less common derivatives. For simplicity, sialic acids are depicted as orange circles, while the membranes are in grey
Porins form water-filled open cavities and allow molecules to traverse the outer membrane largely via passive diffusion (Galdiero et al. 2012). OmpF and OmpC, commonly referred to as general porins, facilitate non-specific diffusion of a range of small solutes (less than 600 Da) across the outer membrane, driven largely by concentration gradients between the extracellular environment and periplasmic space. However, for molecules that are present at low extracellular concentrations, such as sialic acids, substrate-specific channels or active transporters are required for efficient uptake.
The best studied sialic acid porin is NanC from E. coli. The crystal structure reveals two parallel sequences of basic amino acids that line the pore and are likely to be important for the translocation of negatively charged sialic acid(s) (Condemine et al. 2005; Wirth et al. 2009). NanC is suggested to be specific for sialic acid, although it is not known whether different types of sialic acid, other acidic sugars or even sialic acid oligomers can diffuse through the pore. The exact role of the two electropositive tracks in translocation is unclear.
The recently characterised NanOU system from Bacteroidetes species is the first example of a more complex system of active transport across the outer membrane. This system is comprised of a β-barrel protein predicted to be of the TonB-dependent receptor family, the outer membrane permease NanO and an extracellular neuraminate uptake protein, NanU (Fig. 2). The crystal structure of NanU from B. fragilis has been determined, and functional studies demonstrate that it binds sialic acid with high affinity (K D ~ 0.4 μM) (Phansopa et al. 2014). Bacteroides fragilis NanU associates with the outer membrane and enhances transport through NanO at low N-acetylneuraminic acid concentrations. Efficient transport requires a functional TonB–ExbB–ExbD complex, which couples proton-motive force with transport to drive import.
Once sialic acids traverse the outer cell membrane, they arrive at the periplasmic space. Here, two enzymes are known to carry out the further processing of sialic acid for its subsequent transport across the cytoplasmic membrane (Fig. 2).
The E. coli nanCMS operon encodes the specific channel NanC, a sialate mutarotase (NanM) and a sialate O-acetyl esterase (NanS). NanM is responsible for accelerating the conversion of the α-sialic acid anomer to the more thermodynamically stable β-sialic acid anomer (Severi et al. 2008), which is presumably the anomer that is transported by dedicated cytoplasmic membrane transporters. NanS is essential for the growth of E. coli on 9-O-acetylated sialic acids, a derivative present in human mucin oligosaccharides (Robbe-Masselot et al. 2009; Steenbergen et al. 2009).
Prior to import across the cytoplasmic membrane, periplasmic binding proteins (also referred to as extracytoplasmic solute receptors) bind sialic acid within the periplasm and deliver it to the cytoplasmic TRAP and ABC membrane transporters, conferring specificity and maintaining high affinity within the system. Structures of periplasmic binding proteins from the TRAP transport systems reveal significant conformational changes upon sialic acid binding (Johnston et al. 2008), which may be controlled by a proton-linked pathway, leading to the suggestion that protons may also be involved in sialic acid transport by the TRAP transporters (Gangi Setty et al. 2014).
Import of sialic acid across the cytoplasmic membrane
Bacterial pathogens have evolved multiple mechanisms of sialic acid transport across the cytoplasmic membrane, which vary between species (Almagro-Moreno and Boyd 2009). To date, four unique transporter families have been recognised, including those from the ATP binding cassette (ABC), tripartite ATP-independent periplasmic (TRAP), major facilitator superfamily (MFS) and sodium solute symporter (SSS) transporter families (Fig. 2).
ATP binding cassette (ABC) transporters for sialic acid transport
The ABC superfamily of transporters is one of the largest families of transporters, with representatives in all phyla (Jones and George 2004). Containing uptake and efflux systems, this superfamily is made up of numerous families and subfamilies that vary in their substrate specificity (Saurin et al. 1999). They transport a variety of substrates across both extracellular and intracellular membranes, including ions, sugars, amino acids, peptides, polysaccharides and even proteins. ABC transporters are known as primary transporters because they utilise the energy generated from ATP hydrolysis to transport solutes across a membrane (Higgins 1992).
ABC importers and exporters are present in prokaryotes. ABC importers are classified into three groups, including canonical ABC importers of type I and type II classes, and the more recently classified type III class of ABC importers, known as the “energy-coupling factor” transporters (Eitinger et al. 2011). Except for type III ABC importers, all ABC transporters are composed of four domains: two transmembrane domains that are similar in structure, forming the translocation pathway, and two cytoplasmic nucleotide binding domains that hydrolyse ATP (Higgins 1992). These four domains can exist as pairwise but individual subunits, or as fused transmembrane domains and/or nucleotide binding domains. For ABC exporters, the transmembrane domains and NBD domains are fused, while for importers, the core transmembrane domains and nucleotide binding domains are generally separate chains. The type I and type II ABC transporters contain an additional periplasmic or cell surface-associated binding protein that bind substrates with high affinity and deliver them to the transmembrane domains. Since the first high-resolution structure of the E. coli vitamin B12 ABC transporter (Locher et al. 2002), a handful of other structures have been solved, some of which are in complex with their substrate binding proteins (for example, Oldham et al. 2008; Hohl et al. 2012; Woo et al. 2012; Shintre et al. 2013).
The first sialic acid ABC transport system was identified and characterised in Haemophilus ducreyi (Post et al. 2005). The components that make up the H. ducreyi ABC transporter include a periplasmic substrate binding protein (SatA), an integral membrane permease domain (SatB) and an ATPase domain (SatD). A SatC subunit contains an integral membrane permease domain fused with an ATPase domain, as in many bacterial ABC exporters. This architecture is unique and the structure of SatBCD will further contribute to our understanding of this diverse membrane transporter superfamily. Recently, a similar SatABCD system has been identified in Corynebacterium glutamicum, a soil bacterium, as well as C. diphtheriae, C. ulcerans and C. pseudotuberculosis (Holder et al. 2011; Gruteser et al. 2012). An ABC transporter system for sialic acid has also been identified in S. pneumoniae, with a more typical structural organisation (Marion et al. 2011a, b). Here, the transporter has two transmembrane permeases (SatBC), and the two ATPase domains are a shared ATPase (msmK) that is responsible for energising multiple carbohydrate transporters (Marion et al. 2011a). Although the import of sialic acid by ABC transporters is not well characterised in other organisms, sequence alignment predicts that S. agalactiae, S. gordonii, S. pyogenes and S. sanguinis also utilise an ABC-type transport system for sialic acid (Almagro-Moreno and Boyd 2009).
Tripartite ATP-independent periplasmic (TRAP) transporters for sialic acid transport
TRAP transporters constitute a large family of specific solute transporters, all of which transport organic acids. Surprisingly, they were only identified in the late 1990s (Forward et al. 1997), long after the discovery of other bacterial transporters. For drug discovery, TRAP transporters are of particular interest because they are widespread across bacteria and archaea, but are not found in eukaryotes (Kelly and Thomas 2001).
Solutes transported by the TRAP family include a range of C4-dicarboxylates, α-keto acids, aromatic substrates and amino acids (Mulligan et al. 2011). TRAP transporters are secondary transporters that use an electrochemical gradient to facilitate solute transport (Kelly and Thomas 2001). Secondary transporters couple the movement of an ion down an electrochemical gradient with the movement of another ion or molecule against a concentration and/or electrochemical gradient. Unlike primary transporters, there is no direct coupling of ATP to allow movement of an ion or molecule across the membrane. By definition, secondary transporters operate in both directions, depending on the direction and magnitude of the concentration and/or electrochemical gradient (Poolman and Konings 1993; Severi et al. 2010).
TRAP transporters comprise three subunits: the substrate binding protein, a large membrane spanning subunit and a small membrane spanning subunit.
Much of the work to date has focused on the soluble substrate binding proteins, many of which have been crystallised and their affinity to ligands well characterised (Johnston et al. 2007; Gangi Setty et al. 2014). Like ABC transporters, the purpose of the substrate binding protein is to bind substrate with high affinity and specificity and present it to the membrane spanning domains of the transporter for transport across the membrane (Doeven et al. 2004; Mulligan et al. 2009). The substrate binding protein is found either free in the periplasm of Gram-negative bacteria or anchored to the cytoplasmic membrane in Gram-positive bacteria and archaea (Kelly and Thomas 2001). In contrast, the full TRAP transporter systems are not well characterised, owing to a lack of molecular data describing their structure. The large membrane spanning subunit is predicted to form the membrane translocation pathway through which the substrate passes (Mulligan et al. 2012). The small membrane spanning subunit, which is predicted to contain four transmembrane helices, has been shown to be essential for transport, but its function remains unknown. It also remains unclear whether TRAP sialic acid transporters are either general proton- or general sodium ion-dependent, since there is only one example (below), which is sodium ion-dependent.
TRAP type sialic acid transporters are present in the Pasteurellaceae and Vibrionaceae families (Almagro-Moreno and Boyd 2009). The first sialic acid TRAP transporter system to be characterised in vitro was from H. influenzae (Mulligan et al. 2009), where the small and large membrane spanning subunits are fused. This work demonstrates that, for this TRAP, the substrate binding protein (SiaP) is required for transport and that an electrochemical sodium gradient drives transport. Interestingly, the substrate binding protein is also necessary for reverse transport, suggesting that the interaction is important for changing the conformation of the channel from the open-inward state to the outward-open state. More recently, the same group has determined that the V. cholerae sialic acid TRAP, which is not fused, has a stoichiometry of one large membrane spanning subunit to one small membrane spanning subunit (Mulligan et al. 2012).
Sugar proton symporters for sialic acid transport
The sugar proton symporter from E. coli, NanT, was the very first sialic acid transporter to be discovered (Vimr and Troy 1985). NanT type sialic acid transporters are present in a range of human pathogens within Enterobacteriaceae and Bacteroidetes (Almagro-Moreno and Boyd 2009). NanT is a secondary transporter, meaning that it utilises an electrochemical gradient to facilitate the energetically unfavourable transport of sialic acid against its concentration gradient. NanT is a secondary transporter belonging to the major facilitator family, which utilises the proton electrochemical gradient (pH gradient) to facilitate the unfavourable co-transport of sialic acid against its concentration gradient (Martinez et al. 1995; Vimr et al. 2004; Severi et al. 2010). The stoichiometry of protons required for the transport of sialic acid by NanT is yet to be confirmed.
Most major facilitator superfamily transporters consist of 12 transmembrane helices, placing both the N- and C-termini on the cytoplasmic side of the membrane (Huang et al. 2003). The transmembrane helices are split into six-helical bundles that form amino- and carboxyl-domains connected by an extended central loop within the cytoplasm (Maiden et al. 1987). A number of crystal structures have been solved for other members of the major facilitator superfamily, the first of which includes the E. coli lactose permease sugar proton symporter (Abramson et al. 2003) and the glycerol-3-phosphate inorganic phosphate antiporter (Huang et al. 2003). The mechanism by which these transporters are proposed to move molecules across a membrane is described as the “rocker switch” mechanism (Abramson et al. 2003; Huang et al. 2003). In this mechanism, the transporter alternates between an inward- and outward-facing conformation, where the substrate binding site is only accessible on one side of the membrane at any given time (Huang et al. 2003). More recently, crystal structures of several sugar porter subfamilies within the major facilitator superfamily have been determined, including E. coli D-xylose proton symporter and three eukaryotic glucose transporters (Sun et al. 2012; Deng et al. 2014, 2015; Nomura et al. 2015). On the cytoplasmic side of the structures, instead of an extended central loop, a unique intracellular four-helix domain has been discovered, located between the N- and C-domains. This intracellular four-helix domain has been proposed to be a structural characteristic feature for the sugar porter subfamily in general and is involved in the intracellular gating of the transporters (Sun et al. 2012; Deng and Yan 2016).
Our understanding of sugar proton symporters for sialic acid transport at a molecular level is limited to E. coli NanT (Martinez et al. 1995). Based on the protein sequence, the E. coli NanT sialic acid transporter is predicted to consist of 14 transmembrane helices (Martinez et al. 1995), as opposed to the usual 12 transmembrane helices found in the major facilitator superfamily. Twelve of the transmembrane helices that make up NanT contain residues that are conserved within the major facilitator superfamily (Vimr et al. 2004). However, the two centrally located helices are absent in other secondary transporters (Martinez et al. 1995; Vimr et al. 2004). One of these is predicted to form an amphipathic helix, which may be important for the specificity of this protein towards sialic acid (Vimr et al. 2004). Although E. coli NanT only shares 16.7% identity with XylE, several sugar porter subfamily signature motifs have been identified, including the highly conserved ‘GRR’ motif between TM2 and TM3 and the ‘EXXXXXXRG(N)’ motif between TM4 and TM5 (Martinez et al. 1995). These centrally located membrane helices may form an intracellular four-helix domain, as seen in other sugar transporters.
Sodium solute symporters (SSS) for sialic acid transport
More recently, a novel type of sialic acid transporter (SiaT) has been discovered from S. enterica serovar Typhimurium linked to genes encoding enzymes involved in sialic acid processing and utilisation (Severi et al. 2010). SiaT type sialic acid transporters are present in various other organisms and clinically important human bacterial pathogens, including Staphylococcus aureus (Olson et al. 2013), Photobacterium profundum (Almagro-Moreno and Boyd 2009), C. perfringens (Walters et al. 1999) and C. difficile (Ng et al. 2013). The genes encoding SiaT transporters are widespread among both Gram-positive and -negative species of bacteria, whereas other types of sialic acid transporters are less widespread (Severi et al. 2010).
SiaT belongs to the sodium solute symporter (SSS) family of secondary transporters that co-transport sodium ions with sugars, amino acids, inorganic ions or vitamins (Wright et al. 2004). SSS transporters depend on a sodium gradient for the movement of their respective solutes across the cytoplasmic membrane. This has been demonstrated for the S. enterica serovar Typhimurium SiaT, as its functionality is dependent upon the presence of sodium ions (Severi et al. 2010). It has long been established that cells maintain a low intracellular concentration of sodium ions by actively pumping them out of the cell. In combination with the negative potential of the membrane, this provides the driving force for the transport of solutes into the cell (Schultz and Curran 1970; Faham et al. 2008).
To date, the only representative of the SSS family for which the crystal structure has been solved is the sodium galactose transporter (SGLT) from V. parahaemolyticus (Faham et al. 2008). The structure contains 14 transmembrane helices, and the core domain is comprised of two inverted repeats, which each contain five transmembrane helices; the architecture of this core domain resembles that of other sodium symporters, such as the sodium leucine symporter (LeuT) (Yamashita et al. 2005), the sodium benzyl-hydantoin symporter (Mhp1) (Weyand et al. 2008) and the sodium glycine-betaine/proline-betaine symporter (BetP) (Ressl et al. 2009). Despite the structural similarity, these transporters do not share any sequence homology with one another or with the SSS family (Abramson and Wright 2009).
Similar to the mechanism described for the major facilitator superfamily transporters, sodium symporters also share an alternating access mechanism, where the transporter switches between an inward- and an outward-facing conformation for transport (Faham et al. 2008). In the case of sodium symporters, the sodium motive force drives these conformational changes. The stoichiometry of sodium ions required for transport by SSS transporters varies among family members. For some, two sodium ions are required for transport of their respective solute (Eskandari et al. 1997; Mackenzie et al. 1998), whereas others require only one sodium ion (Turk et al. 2000). The stoichiometry of sodium required for the transport of sialic acid by SiaT is not yet known.
Distribution of the cytoplasmic membrane sialic acid transporters
Whilst most bacterial pathogens possess only one type of sialic acid transporter, there are a few exceptions that are predicted to have two types from different families. For example, P. profundum has a SiaT and a TRAP type sialic acid transporter, S. pneumoniae has a SiaT and a putative ABC sialic acid transporter, and S. enterica serovar Typhimurium has a SiaT and a NanT sialic acid transporter, both of which are linked to operons containing genes involved in sialic acid utilisation (Almagro-Moreno and Boyd 2009; Severi et al. 2010). It is not understood why these organisms produce more than one type of transporter, but it may allow the import of a larger range of sialic acid derivatives that are known in biological contexts.
Concluding remarks
It is an exciting time in the emerging field of sialic acid transporters, reflected in the growing number of studies that identify and begin to characterise the molecular mechanisms that underlie the function. Evidence is mounting to support the view that inhibition of these transport systems is a valid pathway to develop new antibiotics, which elevates the need for the structural and functional data to aid the design of inhibitors. However, many, if not most, transporters are poorly defined at a structural and functional level. A lack of molecular structures that detail how these proteins mediate the transport of sialic acid into bacterial cells means there are many unresolved questions. Given the range of different sialic acids, how is specificity achieved? Across the transporter superfamilies known to transport sialic acid in bacteria, what is the mechanism of transport? How is transport driven? In addition to being interesting questions in and of themselves, the answers may open the way to the development new antibiotics.
References
Abramson J, Wright EM (2009) Structure and function of Na(+)-symporters with inverted repeats. Curr Opin Struct Biol 19:425–432
Abramson J, Smirnova I, Kasho V, Verner G, Kaback HR, Iwata S (2003) Structure and mechanism of the lactose permease of Escherichia coli. Science 301:610–615
Almagro-Moreno S, Boyd EF (2009) Insights into the evolution of sialic acid catabolism among bacteria. BMC Evol Biol 9:118
Bouchet V, Hood DW, Li J et al (2003) Host-derived sialic acid is incorporated into Haemophilus influenzae lipopolysaccharide and is a major virulence factor in experimental otitis media. Proc Natl Acad Sci U S A 100:8898–8903
Caing-Carlsson R, Goyal P, Sharma A et al (2017) Crystal structure of N-acetylmannosamine kinase from Fusobacterium nucleatum. Acta Crystallogr F Struct Biol Commun 73:356–362
Chang D-E, Smalley DJ, Tucker DL et al (2004) Carbon nutrition of Escherichia coli in the mouse intestine. Proc Natl Acad Sci U S A 101:7427–7432
Condemine G, Berrier C, Plumbridge J, Ghazi A (2005) Function and expression of an N-acetylneuraminic acid-inducible outer membrane channel in Escherichia coli. J Bacteriol 187:1959–1965
Deng D, Yan N (2016) GLUT, SGLT, and SWEET: structural and mechanistic investigations of the glucose transporters. Protein Sci 25:546–558
Deng D, Xu C, Sun P et al (2014) Crystal structure of the human glucose transporter GLUT1. Nature 510:121–125
Deng D, Sun P, Yan C et al (2015) Molecular basis of ligand recognition and transport by glucose transporters. Nature 526:391–396
Doeven MK, Abele R, Tampé R, Poolman B (2004) The binding specificity of OppA determines the selectivity of the oligopeptide ATP-binding cassette transporter. J Biol Chem 279:32301–32307
Eitinger T, Rodionov DA, Grote M, Schneider E (2011) Canonical and ECF-type ATP-binding cassette importers in prokaryotes: diversity in modular organization and cellular functions. FEMS Microbiol Rev 35:3–67
Eskandari S, Loo DD, Dai G, Levy O, Wright EM, Carrasco N (1997) Thyroid Na+/I− symporter. Mechanism, stoichiometry, and specificity. J Biol Chem 272:27230–27238
Faham S, Watanabe A, Besserer GM et al (2008) The crystal structure of a sodium galactose transporter reveals mechanistic insights into Na+/sugar symport. Science 321:810–814
Forward JA, Behrendt MC, Wyborn NR, Cross R, Kelly DJ (1997) TRAP transporters: a new family of periplasmic solute transport systems encoded by the dctPQM genes of Rhodobacter capsulatus and by homologs in diverse gram-negative bacteria. J Bacteriol 179:5482–5493
Galdiero S, Falanga A, Cantisani M et al (2012) Microbe–host interactions: structure and role of Gram-negative bacterial porins. Curr Protein Pept Sci 13:843–854
Gangi Setty T, Cho C, Govindappa S, Apicella MA, Ramaswamy S (2014) Bacterial periplasmic sialic acid-binding proteins exhibit a conserved binding site. Acta Crystallogr D Struct Biol 70:1801–1811
Gruteser N, Marin K, Krämer R, Thomas GH (2012) Sialic acid utilization by the soil bacterium Corynebacterium glutamicum. FEMS Microbiol Lett 336:131–138
Higgins CF (1992) ABC transporters: from microorganisms to man. Annu Rev Cell Biol 8:67–113
Hohl M, Briand C, Grütter MG, Seeger MA (2012) Crystal structure of a heterodimeric ABC transporter in its inward-facing conformation. Nat Struct Mol Biol 19:395–402
Holder JW, Ulrich JC, DeBono AC et al (2011) Comparative and functional genomics of Rhodococcus opacus PD630 for biofuels development. PLoS Genet 7:e1002219
Huang Y, Lemieux MJ, Song J, Auer M, Wang DN (2003) Structure and mechanism of the glycerol-3-phosphate transporter from Escherichia coli. Science 301:616–620
Huang Y-L, Chassard C, Hausmann M, Von Itzstein M, Hennet T (2015) Sialic acid catabolism drives intestinal inflammation and microbial dysbiosis in mice. Nat Commun 6:8141
Jeong HG, Oh MH, Kim BS, Lee MY, Han HJ, Choi SH (2009) The capability of catabolic utilization of N-acetylneuraminic acid, a sialic acid, is essential for Vibrio vulnificus pathogenesis. Infect Immun 77:3209–3217
Johnston JW, Zaleski A, Allen S et al (2007) Regulation of sialic acid transport and catabolism in Haemophilus influenzae. Mol Microbiol 66:26–39
Johnston JW, Coussens NP, Allen S et al (2008) Characterization of the N-acetyl-5-neuraminic acid-binding site of the extracytoplasmic solute receptor (SiaP) of nontypeable Haemophilus influenzae strain 2019. J Biol Chem 283:855–865
Jones PM, George AM (2004) The ABC transporter structure and mechanism: perspectives on recent research. Cell Mol Life Sci 61:682–699
Kelly DJ, Thomas GH (2001) The tripartite ATP-independent periplasmic (TRAP) transporters of bacteria and archaea. FEMS Microbiol Rev 25:405–424
Lewis AL, Lewis WG (2012) Host sialoglycans and bacterial sialidases: a mucosal perspective. Cell Microbiol 14:1174–1182
Locher KP, Lee AT, Rees DC (2002) The E. coli BtuCD structure: a framework for ABC transporter architecture and mechanism. Science 296:1091–1098
Mackenzie B, Loo DD, Wright EM (1998) Relationships between Na+/glucose cotransporter (SGLT1) currents and fluxes. J Membr Biol 162:101–106
Maiden MC, Davis EO, Baldwin SA, Moore DC, Henderson PJ (1987) Mammalian and bacterial sugar transport proteins are homologous. Nature 325:641–643
Marion C, Aten AE, Woodiga SA, King SJ (2011a) Identification of an ATPase, MsmK, which energizes multiple carbohydrate ABC transporters in Streptococcus pneumoniae. Infect Immun 79:4193–4200
Marion C, Burnaugh AM, Woodiga SA, King SJ (2011b) Sialic acid transport contributes to pneumococcal colonization. Infect Immun 79:1262–1269
Martinez J, Steenbergen S, Vimr E (1995) Derived structure of the putative sialic acid transporter from Escherichia coli predicts a novel sugar permease domain. J Bacteriol 177:6005–6010
Mulligan C, Geertsma ER, Severi E, Kelly DJ, Poolman B, Thomas GH (2009) The substrate-binding protein imposes directionality on an electrochemical sodium gradient-driven TRAP transporter. Proc Natl Acad Sci U S A 106:1778–1783
Mulligan C, Fischer M, Thomas GH (2011) Tripartite ATP-independent periplasmic (TRAP) transporters in bacteria and archaea. FEMS Microbiol Rev 35:68–86
Mulligan C, Leech AP, Kelly DJ, Thomas GH (2012) The membrane proteins SiaQ and SiaM form an essential stoichiometric complex in the sialic acid tripartite ATP-independent periplasmic (TRAP) transporter SiaPQM (VC1777–1779) from Vibrio cholerae. J Biol Chem 287:3598–3608
Ng KM, Ferreyra JA, Higginbottom SK et al (2013) Microbiota-liberated host sugars facilitate post-antibiotic expansion of enteric pathogens. Nature 502:96–99
Nomura N, Verdon G, Kang HJ et al (2015) Structure and mechanism of the mammalian fructose transporter GLUT5. Nature 526:397–401
North RA, Kessans SA, Atkinson SC et al (2013) Cloning, expression, purification, crystallization and preliminary X-ray diffraction studies of N-acetylneuraminate lyase from methicillin-resistant Staphylococcus aureus. Acta Crystallogr F Struct Biol Commun 69:306–312
North RA, Kessans SA, Griffin MDW et al (2014a) Cloning, expression, purification, crystallization and preliminary X-ray diffraction analysis of N-acetylmannosamine-6-phosphate 2-epimerase from methicillin-resistant Staphylococcus aureus. Acta Crystallogr F Struct Biol Commun 70:650–655
North RA, Seizova S, Stampfli A et al (2014b) Cloning, expression, purification, crystallization and preliminary X-ray diffraction analysis of N-acetylmannosamine kinase from methicillin-resistant Staphylococcus aureus. Acta Crystallogr F Struct Biol Commun 70:643–649
North RA, Watson AJA, Pearce FG et al (2016) Structure and inhibition of N-acetylneuraminate lyase from methicillin-resistant Staphylococcus aureus. FEBS Lett 590:4414–4428
Oldham ML, Davidson AL, Chen J (2008) Structural insights into ABC transporter mechanism. Curr Opin Struct Biol 18:726–733
Olson ME, King JM, Yahr TL, Horswill AR (2013) Sialic acid catabolism in Staphylococcus aureus. J Bacteriol 195:1779–1788
Pezzicoli A, Ruggiero P, Amerighi F, Telford JL, Soriani M (2012) Exogenous sialic acid transport contributes to group B streptococcus infection of mucosal surfaces. J Infect Dis 206:924–931
Phansopa C, Roy S, Rafferty JB et al (2014) Structural and functional characterization of NanU, a novel high-affinity sialic acid-inducible binding protein of oral and gut-dwelling Bacteroidetes species. Biochem J 458:499–511
Poolman B, Konings WN (1993) Secondary solute transport in bacteria. Biochim Biophys Acta 1183:5–39
Post DMB, Mungur R, Gibson BW, Munson RS (2005) Identification of a novel sialic acid transporter in Haemophilus ducreyi. Infect Immun 73:6727–6735
Ressl S, Terwisscha van Scheltinga AC, Vonrhein C, Ott V, Ziegler C (2009) Molecular basis of transport and regulation in the Na(+)/betaine symporter BetP. Nature 458:47–52
Robbe-Masselot C, Maes E, Rousset M, Michalski JC, Capon C (2009) Glycosylation of human fetal mucins: a similar repertoire of O-glycans along the intestinal tract. Glycoconj J 26:397–413
Saurin W, Hofnung M, Dassa E (1999) Getting in or out: early segregation between importers and exporters in the evolution of ATP-binding cassette (ABC) transporters. J Mol Evol 48:22–41
Schultz SG, Curran PF (1970) Coupled transport of sodium and organic solutes. Physiol Rev 50:637–718
Severi E, Randle G, Kivlin P et al (2005) Sialic acid transport in Haemophilus influenzae is essential for lipopolysaccharide sialylation and serum resistance and is dependent on a novel tripartite ATP-independent periplasmic transporter. Mol Microbiol 58:1173–1185
Severi E, Hood DW, Thomas GH (2007) Sialic acid utilization by bacterial pathogens. Microbiology 153:2817–2822
Severi E, Müller A, Potts JR et al (2008) Sialic acid mutarotation is catalyzed by the Escherichia coli beta-propeller protein YjhT. J Biol Chem 283:4841–4849
Severi E, Hosie AHF, Hawkhead JA, Thomas GH (2010) Characterization of a novel sialic acid transporter of the sodium solute symporter (SSS) family and in vivo comparison with known bacterial sialic acid transporters. FEMS Microbiol Lett 304:47–54
Shintre CA, Pike ACW, Li Q et al (2013) Structures of ABCB10, a human ATP-binding cassette transporter in apo- and nucleotide-bound states. Proc Natl Acad Sci U S A 110:9710–9715
Sillanaukee P, Pönniö M, Jääskeläinen IP (1999) Occurrence of sialic acids in healthy humans and different disorders. Eur J Clin Investig 29:413–425
Steenbergen SM, Jirik JL, Vimr ER (2009) YjhS (NanS) is required for Escherichia coli to grow on 9-O-acetylated N-acetylneuraminic acid. J Bacteriol 191:7134–7139
Sun L, Zeng X, Yan C et al (2012) Crystal structure of a bacterial homologue of glucose transporters GLUT1–4. Nature 490:361–366
Turk E, Kim O, le Coutre J et al (2000) Molecular characterization of Vibrio parahaemolyticus vSGLT: a model for sodium-coupled sugar cotransporters. J Biol Chem 275:25711–25716
Vimr ER, Troy FA (1985) Identification of an inducible catabolic system for sialic acids (nan) in Escherichia coli. J Bacteriol 164:845–853
Vimr ER, Kalivoda KA, Deszo EL, Steenbergen SM (2004) Diversity of microbial sialic acid metabolism. Microbiol Mol Biol Rev 68:132–153
Walters DM, Stirewalt VL, Melville SB (1999) Cloning, sequence, and transcriptional regulation of the operon encoding a putative N-acetylmannosamine-6-phosphate epimerase (nanE) and sialic acid lyase (nanA) in Clostridium perfringens. J Bacteriol 181:4526–4532
Weyand S, Shimamura T, Yajima S et al (2008) Structure and molecular mechanism of a nucleobase–cation–symport-1 family transporter. Science 322:709–713
Wirth C, Condemine G, Boiteux C, Bernèche S, Schirmer T, Peneff CM (2009) NanC crystal structure, a model for outer-membrane channels of the acidic sugar-specific KdgM porin family. J Mol Biol 394:718–731
Woo J-S, Zeltina A, Goetz BA, Locher KP (2012) X-ray structure of the Yersinia pestis heme transporter HmuUV. Nat Struct Mol Biol 19:1310–1315
Wright EM, Loo DDF, Hirayama BA, Turk E (2004) Surprising versatility of Na+-glucose cotransporters: SLC5. Physiology (Bethesda) 19:370–376
Yamashita A, Singh SK, Kawate T, Jin Y, Gouaux E (2005) Crystal structure of a bacterial homologue of Na+/cl-dependent neurotransmitter transporters. Nature 437:215–223
Acknowledgements
R.C.J.D. and R.A.N. acknowledge the following for funding support, in part: (1) the New Zealand Royal Society Marsden Fund (15-UOC032) and (2) the Biomolecular Interaction Centre, University of Canterbury. The project has received funding from the European Union’s Seventh Framework Programme for research, technological development and demonstration under grant agreement no. 608743 (to R.F.). This work was also supported by grants from the Swedish Research Council (2011-5790 to R.F.), the Swedish Research Council Formas (2010-1759 to R.F. and 221-2013-730 to W.Y.W.), the Swedish Governmental Agency for Innovation Systems (VINNOVA) (2013-04655 and 2017-00180 to R.F.), Carl Tryggers Stiftelse för Vetenskaplig Forskning (11:147 to R.F.), EMBO (1163-2014 to P.G. and 584-2014 to R.A.N.) and the Centre for Antibiotic Resistance Research (CARe) at the University of Gothenburg (to R.F.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Rachel A. North declares that she has no conflict of interest. Christopher R. Horne declares that he has no conflict of interest. James S. Davies declares that he has no conflict of interest. Daniela M. Remus declares that she has no conflict of interest. Andrew C. Muscroft-Taylor declares that he has no conflict of interest. Parveen Goyal declares that he has no conflict of interest. Weixiao Yuan Wahlgren declares that she has no conflict of interest. S. Ramaswamy declares that he has no conflict of interest. Rosmarie Friemann declares that she has no conflict of interest. Renwick C. J. Dobson declares that he has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
This article is part of a Special Issue on ‘Biomolecules to Bio-nanomachines - Fumio Arisaka 70th Birthday’ edited by Damien Hall, Junichi Takagi and Haruki Nakamura
Rights and permissions
About this article
Cite this article
North, R.A., Horne, C.R., Davies, J.S. et al. “Just a spoonful of sugar...”: import of sialic acid across bacterial cell membranes. Biophys Rev 10, 219–227 (2018). https://doi.org/10.1007/s12551-017-0343-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-017-0343-x