Abstract
p-Nitrophenol (PNP), used primarily for manufacturing pesticides and dyes, has been recognized as a priority environmental pollutant. It is therefore important to reduce the input of this toxicant into the environment and to establish approaches for its removal from the contaminated sites. PNP monooxygenase, a novel enzyme from Gram-positive bacteria like Arthrobacter sp. and Bacillus sp., that comprises two components, a flavoprotein reductase and an oxygenase, catalyzes the initial two sequential monooxygenations to convert PNP to trihydroxybenzene. Accurate and reliable prediction of this enzyme–substrate interactions and binding affinity are of vital importance in understanding these catalytic mechanisms of the two sequential reactions. As crystal structure of the enzyme has not yet been published, we built a homology model for PNP monooxygenase using crystallized chlorophenol 4-monooxygenase from Burkholderia cepacia AC1100 (3HWC) as the template. The model was assessed for its reliability using PROCHECK, ERRAT and ProSA. Molecular docking of the physiological substrates, PNP and 4-nitrocatechol (4-NC), was carried out using Glide v5.7 implemented in Maestro v9.2, and the binding energies were calculated to substantiate the prediction. Docking complexes formed by molecular level interactions of PNP monooxygenase-PNP/4-NC without or with the cofactors, FAD and NADH, showed good correlation with the established experimental evidence that the two-component PNP monooxygenase catalyzes both the hydroxylation of PNP and the oxidative release of nitrite from 4-NC in B. sphaericus JS905. Furthermore, molecular dynamics simulations performed for docking complexes using Desmond v3.0 showed stable nature of the interactions as well.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
With the rapid thrust in industrial and agricultural activity, the recent times have witnessed vast quantities of soil and groundwater resources becoming contaminated with hazardous chemicals. p-Nitrophenol (PNP) is probably the most important contaminant among the mononitrophenols since 27 % of its use in industry is for pesticide manufacture and 13 % is for synthesis of dye components [1]. PNP is also found as a major metabolite of microbial degradation of organophosphorous insecticides, parathion and methyl parathion [2] and is thus an important environmental pollutant [3, 4] posing even odor problems to water bodies [5, 6]. PNP is, therefore, one of the eleven phenolic compounds listed as priority pollutants by the US Environmental Protection Agency [7].
Two alternative pathways for aerobic bacterial degradation that convert PNP to maleylacetate have been elucidated [8]. The first pathway that results in the formation of hydroquinone from PNP, probably via 1,4-benzoquinone, with concomitant release of nitrite [9], is more common in Gram-negative isolates. In the second catabolic pathway, Bacillus sphaericus JS905 [10] hydroxylates PNP to produce 4-nitrocatechol (4-NC), and subsequent oxidative removal of the nitro group from 4-NC as nitrite yields 1,2,4-trihydroxybenzene (THB) as shown in Fig. 1. PNP monooxygenase, a novel enzyme from B. sphaericus JS905 that belongs to a two-component flavin-diffusible monooxygenase family, consisting of two components—a flavoprotein reductase and an oxygenase, catalyzes the first two sequential reactions in the degradation of PNP to THB via 4-NC. The enzyme uses loosely bound FAD as the redox chromophore and is NADH dependent. Genetic evidence that the oxygenase component of the enzyme hydroxylates PNP as well as 4-NC following two sequential monooxygenations has been provided for Arthrobacter sp. strain JS443 [11] and Pseudomonas sp. 1-7 [12]. The nitro substituent from 4-NC is directly removed from the aromatic ring in the form of nitrite.
Understanding the microbial metabolism of nitroaromatic compounds will assist in management options to minimize their persistence in the environment, and one needs to have specific information on various components involved in catalysis that explains the overall function of the system. Our present study concerning analysis of functional sites in PNP monooxygenase relies on the biochemical information provided for B. sphaericus JS905 and the genetic information reported for Arthrobacter sp. strain JS443, since the proposed pathways for PNP degradation in both the bacterial strains are the same involving two sequential monooxygenation reactions. Experimental evidence also implicates four conserved residues (Arg100, Gln158, Arg161 and Thr193) for the substrate binding with oxygenase component of the enzyme [13]. The objective of the present study was primarily to verify whether the same amino acid residues at the catalytic site of the enzyme are involved in the two sequential monooxygenations. Since molecular docking study using 3D structure of an enzyme is one way to predict interaction between the substrate and the active site, we also report the 3D model of PNP monooxygenase and prediction of interaction sites for the substrates, PNP and 4-NC, through molecular modeling, docking and molecular dynamics (MD) simulations.
2 Methodology
2.1 Homology Modeling of PNP Monooxygenase
The protein sequence of PNP monooxygenase from Arthrobacter sp. strain JS443 (accession number ABL75143.1) was retrieved from the NCBI protein database (http://www.ncbi.nlm.nih.gov/protein). BLASTp [14, 15] search for PNP monooxygenase was performed against the protein databank [16] to find a structural template for homology modeling. Chlorophenol 4-monooxygenase of Burkholderia cepacia AC1100 (3HWC) was selected as the structural template and aligned with PNP monooxygenase using ClustalX 1.83 [17]. Modeller9v7 [18, 19] that performs comparative modeling based on satisfaction of spatial restraints was used to build 3D structure of PNP monooxygenase. Twenty 3D structures of PNP monooxygenase were generated, and the structure with the lowest discrete optimized protein energy (DOPE) score was chosen for further analysis [20]. The stereochemical excellence of the selected homology model was confirmed by PROCHECK analysis [21]. The structure of each residue in the model was ensured for their authenticity by WHATCHECK [22], ERRAT [23] and ProSA [24].
2.2 Molecular Docking
As PNP monooxygenase catalyzes the two sequential monooxygenations in both B. sphaericus JS905 [10] and Arthrobacter sp. strain JS443 [11], we performed docking studies for PNP monooxygenase from strain JS443 with the established physiological substrates (PNP, 4-NC) and the cofactors (FAD and NADH) for the enzyme in order to gain insights into the most probable binding interactions.
Each atom of the protein and substrate or cofactor needs to be fixed before molecular docking for any potential aberrations, if any, and for accurate prediction of interactions with better binding affinity. The 3D model of PNP monooxygenase was preprocessed with the protein preparation workflow in the Maestro v9.2 [25]. Hydrogens were added to all the atoms of PNP monooxygenase for subsequent minimization using OPLS 2005 force field by converging heavy atoms with maximum root-mean-square deviation (RMSD) of 0.30 Å. Minimization was performed restraining the heavy atoms with the hydrogen torsion parameters turned off, to allow free rotation of the hydrogen atoms. The structures of PNP, 4-NC, FAD and NADH (Fig. 2) were imported to Maestro v9.2. LigPrep module along with Epik was used to prepare multiple 3D conformations of the four ligand molecules in a pH range of 7 ± 2.
Molecular docking was carried out using Glide v5.7 applying extra precision (XP) method [26, 27] that generates 10000 poses for each substrate during docking and reports the best pose based on the energy term Emodel. Residues of PNP monooxygenase specific for substrates and cofactors were selected to generate a grid of \(20\times 20\times 20\) Å using Glide. The best poses of each ligand were further ranked based on XP Gscore. Lower XP Gscore for a ligand indicates better binding affinity toward the protein. The cutoff XP Gscore parameter for XP docking was set to 0.0 kcal/mol, a constraint set to discard ligands with positive XP Gscore from the final docking output. Binding affinity of 4-NC and PNP with PNP monooxygenase was estimated using Prime/MM-GBSA analysis.
2.3 Molecular Dynamics (MD) Simulations
MD simulations were performed for docking complexes (PNP monooxygenase+PNP+FAD+NADH and PNP monooxygenase+4-NC+FAD+NADH) to evaluate the stability and conformational changes and to gain insights into the natural dynamics on different timescales in solution. Simulations were carried out using Desmond v3.0 [28, 29], implemented in Maestro v9.2 graphical user interface. The system was embedded with simple point charge (SPC) water model and neutralized by replacing solvent molecules with Na\(^{+}\) ions. The final system was simulated through a multistep protocol devised in Maestro v9.2. In brief, the full system was minimized with maximum 2000 interactions of a hybrid of the steepest descent and the limited-memory Broyden–Fletcher–Goldfarb–Shanno (LBFGS) algorithms, with a convergence threshold of 50.0 kcal/mol/Å\(^{2}\) followed by a similar unrestrained minimization with a convergence threshold of 5.0 kcal/mol/Å\(^{2}\).
The minimized system was relaxed with four subsequent short-span simulations: (a) 12-ps simulation in NVT ensemble (temperature 10 K) restraining nonhydrogen solute atoms, (b) 12-ps simulation in the NPT ensemble (temperature 10 K) restraining nonhydrogen solute atoms, (c) 24-ps simulation in the NPT ensemble restrained with solute nonhydrogen atoms (temperature 300 K) and (d) 24-ps simulation in the NPT ensemble (temperature 300 K) with no restraints. The temperatures and pressures in these initial simulations were controlled using Berendsen thermostats and barostats, respectively. These initial minimization and simulations were performed to relax the model before implementing a longer simulation time. The relaxed system was simulated for a simulation time of 5 ns with a time step of 2 fs, NPT ensemble using a Nose–Hoover thermostat at 300 K and Martyna–Tobias–Klein barostat at 1.01325 bar pressure. The simulated systems were analyzed for stability of the docking complex. Energy fluctuations and RMSD of the complexes in each trajectory were analyzed with respect to simulation time. The root-mean-square fluctuations (RMSF) of backbone and side chains atoms of PNP monooxygenase were analyzed for each residue. The docking complex was analyzed and monitored for consistency in hydrogen bonding interactions.
3 Results and Discussion
3.1 3D Model of PNP Monooxygenase
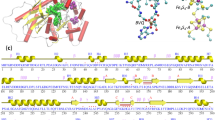
The overall sequence identity between PNP monooxygenase and the template chlorophenol 4-monooxygenase of B. cepacia AC1100 was 45 % with query coverage of 99 %. The sequence alignment for these two sequences enabled us to identify sporadically spread conserved residues all over the sequence (Fig. 3). Conserved regions or positions indicate residues supposedly under stronger evolutionary constraints and thus might be more important for the protein to fulfill its function. Moreover, residues that are specifically conserved in subfamilies point to sequence changes that occurred during the divergence of a common ancestor, and they imply functional changes or the acquisition of modified specificity [30].
Twenty PNP monooxygenase homology models were constructed based on the template structure. All models were assigned a predicted Genetic algorithm 341 (GA341) score and DOPE score. GA341 score of \(\ge\)0.7 suggests that fold assignments in the homology model are correct. GA341 scores for the 20 predicted models were 1.0. Therefore, fold assignments in the homology models were declared accurate. Selection of the best model from the twenty was done based on DOPE score. Measure of conformational energy is represented as DOPE score in homology modeling. Lower DOPE score represents relatively more stable 3D conformation [31]. Thus, the 20th model with the lowest DOPE score (\(-\)54799.56 kcal/mol) was selected for further structural validation.
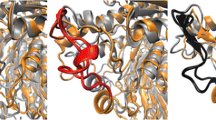
The overall geometric and stereochemical quality of the final modeled structure of PNP monooxygenase was analyzed using PROCHECK, which showed that 92.9 % residues lie in the most favored region and 6.1 % in additionally allowed regions (Fig. 4a). Evaluation of PNP monooxygenase model with ProSA-web revealed the Z-score value as \(-\)8.56, and the overall residue energy of PNP monooxygenase model was largely negative except for some peaks at the end region (Fig. 4b, c). The residue energies of active site and allosteric site regions of the protein were highly negative. The overall quality factor checked from the ERRAT graph was 85.09, indicating the acceptable protein environment (Fig. 4d). The structural superimposition of the model of PNP monooxygenase with the template structure showed backbone RMSD of 0.66 Å. The low backbone RMSD value represents that the predicted structure is reliable. The validated model was submitted to protein model database [32] with ID number PM0077668 (Fig. 5).
3.2 Docking of Substrates into the Active Site of PNP Monooxygenase
Molecular docking helps in determining binding affinity, binding orientations and various bonded and nonbonded enzyme–substrates and enzyme–cofactors interactions in various enzymatic reactions. Docking analysis was implemented for PNP monooxygenase with its two physiological substrates, PNP and 4-NC, and the cofactors, FAD and NADH, to corroborate with the experimental findings of monooxygenation reactions.
Earlier experimental findings showed that active site residues, Arg100 and Thr193, are important for substrate (PNP or 4-NC) binding, while allosteric site residues such as Gln158 and Arg161 are specific for cofactor (FAD or NADH) interaction [13]. Multiple sequence alignment of PNP monooxygenase with homologous proteins, chlorophenol 4-monooxygenase and hydroxyphenylacetate hydoxylase, revealed that residues of the active site and allosteric site are conserved.
A 20\(\,\times\,\)20\(\,\times\,\)20 Å grid was created around centroid of the active site of PNP monooxygenase for PNP and 4-NC docking. Both the substrates, PNP (complex 1) and 4-NC (complex 3), were docked with PNP monooxygenase at the active site with XP Gscore of \(-\)3.434 and \(-\)3.828 kcal/mol, respectively. The (O) atom of hydroxyl group of PNP formed hydrogen bond (h-bond) with residue Thr193, and pi-cation interacted with Arg100 of PNP monooxygenase. The hydrogen atom of one hydroxyl group of 4-NC showed one hydrogen bond with Thr193 of PNP monooxygenase and Arg100 involved in pi-cation interaction with 4-NC (Table 1; Supplementary Fig. 1a, d). Similar binding orientations were observed in both the substrate–PNP monooxygenase complexes.
Further, a docking grid was generated centering on allosteric site of the PNP–PNP monooxygenase complex (complex 1) and 4-NC–PNP monooxygenase complex (complex 3). The grids were generated to determine the probable site of interactions of PNP, 4-NC, FAD and NADH with PNP monooxygenase. Docking interactions of PNP (complex 2) (Fig. 6a) and 4-NC (complex 4) (Fig. 6b) with PNP monooxygenase in the presence of FAD and NADH remained same as that of complex 1 and complex 2, respectively (Table 1; Fig. 6a, b; Supplementary Fig. 1a, d). Binding affinity of NADH and FAD was expressed in terms of XP Gscore in complex 2 and complex 4 (Table 1). NADH was observed to have more binding affinity toward PNP monooxygenase compared to FAD in both the complexes. This result corroborates well with the experimental evidence that FAD is loosely bound to PNP monooxygenase compared to NADH [10]. The docking interactions of FAD and NADH in complex 2 revealed that FAD is interacting with Gln158 and Arg161 through hydrogen bonds while NADH is also forming hydrogen bond with Arg161, and good van der Waals interaction with Gln158 of PNP monooxygenase (Fig. 6a; Supplementary Fig. 1a–d). Similar interactions were also observed in complex 4. FAD was observed to form intermolecular hydrogen bond with 4-NC and good van der Waals interaction with Gln158, while NADH was involved in intermolecular hydrogen bond formation with Arg161 and van der Waals interaction with Gln158 (Fig. 6b; Supplementary Fig. 1d–f). Interaction between FAD and NADH through one intermolecular hydrogen bond was observed in complex 2 (Fig. 6a), while that of complex 4 did not reveal any hydrogen bond (Fig. 6b). Prime/MM-GBSA calculations for PNP showed \(\Delta\)G score of \(-\)28.79 kcal/mol and that of 4-NC showed \(\Delta\)G score of \(-\)27.35 kcal/mol.
3.3 MD Simulation Studies of Docked Complexes
The molecular interactions of PNP monooxygenase with PNP, FAD and NADH, and with 4-NC, FAD and NADH (Fig. 6a, b) corroborated well with the experimental evidences reported in the literature for the substrate and cofactor [13]. Therefore, 5-ns molecular dynamics simulations (MD) were performed for complex 2 (Fig. 7a) and complex 4 (Fig. 7b) to understand the stability of the interactions during PNP degradation in closer-to-natural environmental condition. The interactions obtained through MD simulations are more convincing than docking complexes. Initially, conformational stability of complex 2 and complex 4 was evaluated. The energy plots of complex 2 and complex 4 showed that the energy of the respective systems were stable throughout the simulations (Fig. 8a, b). Analysis of the RMSD plot for protein backbone, cofactors (FAD, NADH) and the substrates (PNP, 4-NC) for complex 2 (Fig. 9a) and complex 4 (Fig. 9b) showed that the RMSD was relatively consistent throughout the MD simulations period. During 5-ns MD simulations of both the complexes, 4-NC showed lower structural rearrangement with PNP monooxygenase compared to PNP. Similarly, FAD showed less structural rearrangement when compared to NADH. Lower RMSD of docking complex during MD simulations represents higher conformational stability of the docking interactions. The root-mean-square fluctuations (RMSF) of a given residue in the MD trajectory were calculated for protein backbone and all side chain atoms of each residue (Fig. 10a, b). RMSF for most of the residues of both the complexes were within the limit of 2.5 Å. The atomic fluctuations of the active site and allosteric site residue were consistent, and backbone residues were also consistent with small conformational peak regions. Analysis of energy plots, RMSD plots and RMSF plots for both the complexes revealed that the two docking complexes were stable during 5-ns MD simulation. Once conformational stability of the docking complexes during 5-ns MD simulations was established, interactions of PNP, 4-NC, FAD and NADH with PNP monooxygenase were analyzed in complex 2 and complex 4.
In molecular docking of complex 2 and complex 4 with PNP and 4-NC, hydroxyl groups were observed to form hydrogen bond with Thr193. Both these substrates were also observed to interact with Arg100 through pi-cation interaction (Supplementary Fig. 1). Hydrogen bond monitoring during MD simulations for substrate–enzyme complexes, PNP and 4-NC showed intermolecular hydrogen bond between the substrates and Thr193 in some trajectories (Supplementary Figs. 3a, b, 4a, b). The presence of hydrogen bond in less number of trajectories for Arg100 is quite evident for its preference for pi-cation interaction in the docking complex, but Thr193 is mainly involved with water-mediated intermolecular hydrogen bond with the substrates in some of trajectories.
In complex 2, Arg161 of PNP monooxygenase interacted with FAD and NADH through hydrogen bond. Gln158 interacted with FAD through hydrogen bond and with NADH through van der Waals interaction. Hydrogen bond monitoring showed that the interaction of FAD and NADH with Arg161 is maintained in all trajectories during MD simulations (Supplementary Fig. 3d, f). MD simulations also revealed that both the cofactors (FAD and NADH) are forming hydrogen bonds with amino group of Arg161 but with different atoms. Suitably, Gln158 showed hydrogen bond with FAD in first \(\sim\)500 ps and \(\sim\)4200 to \(\sim\)4700 ps (Supplementary Fig. 3e), and no hydrogen bond was observed between Gln158 and NADH during 5-ns MD simulations (Supplementary Fig. 3c). The hydrogen bond between FAD and NADH observed in molecular docking was also reproduced (Fig. 11a).
In complex 4, Arg161 was observed to interact through hydrogen bond with NADH in first \(\sim\)200 ps and with FAD in \(\sim\)4500 to \(\sim\)5000 ps (Supplementary Fig. 4d, f). Gln158 was observed to form hydrogen bond in very few trajectories (Supplementary Fig. 4c). The result coincided favorably with the docking results observed for FAD and NADH interactions in complex 4 (Supplementary Fig. 1). Additionally, hydrogen bond interaction between FAD and NADH was identified in complex 4 in all trajectories during MD simulations (Fig. 11b). The final trajectory of 5-ns MD simulations revealed that PNP interacted with residues Phe154, Val155, Ile191 of PNP monooxygenase through a water bridge by SPC4299. The NO\(_{2}\) group of PNP interacted with FAD through intermolecular water bridge formed by three water molecules, from PNP to SPC12001-SPC9446-SPC4359 to FAD. Similarly, NO\(_{2}\) group of PNP was connected with NADH through water bridge with SPC12001-SPC9446-SPC3736 (Fig. 7a; Supplementary Fig. 2). An intermolecular hydrogen bond was also observed between FAD and NADH in complex 2. Water-mediated interactions and intermolecular hydrogen bonds were observed among PNP monooxygenase, PNP, FAD and NADH in all 1042 trajectories. The two cofactors FAD and NADH showed more than one intermolecular hydrogen bonds among themselves (Fig. 11a) in all trajectories. Similar analysis of complex 4 after 5-ns MD simulations revealed FAD interaction with PNP monooxygenase through intermolecular hydrogen bond with residues Asp156, Arg161, Trp207 and Arg208. Two hydrogen bonds were observed between FAD and NADH. NADH was connected to 4-NC through hydrogen bond (Fig. 7b; Supplementary Fig. 2). The intermolecular hydrogen bond monitoring between FAD and NADH showed two hydrogen bonds in most of the trajectories (Fig. 11b).
The analysis of docking complex thus predicted potential path of PNP monooxygenation. Initially, PNP monooxygenase induces oxygenation through Thr193 and Arg100 to form 4-NC. Further, reduction of 4-NC is induced by NADH through FAD to form THB. The binding orientations of ligand molecules predicted through MD simulations showed better correlation to their biologically active states. Our present results clearly corroborate the earlier observations that the two-component monooxygenase catalyzes both the hydroxylation of PNP and the oxidative release of nitrite from 4-NC in B. sphaericus JS905 [10].
4 Conclusions
In the present study, we attempted to create an in silico model for PNP monooxygenase by aiming at the physiological substrates, PNP and 4-NC, along with the cofactors, FAD and NADH. We explored residues Arg100, Gln158 and Thr193 as key catalytic residues which play a major role in controlling the orientation of the nitro group at the active site, and Arg161, Phe154, Val155, Ile191, Trp207 and Arg208 as cofactor-binding residues. Binding orientations of the substrates, PNP and 4-NC, and the cofactors, FAD and NADH, provided a basis for analyzing novel mechanism of PNP monooxygenation reactions in silico. Thus, the modeled 3D structure of PNP monooxygenase showed consistent docking results of enzyme–substrates interactions which correlated well with the earlier experimental results that the enzyme catalyzes two sequential monooxygenation reactions to yield THB with the concomitant release of nitro group as nitrite.
References
Markle RA, Fentiman AF, Steadman JR, Mayer RA (1980) Potentially toxic and hazardous substances in the industrial organic chemicals and organic dyes in pigment industries. National Technical Information Service, Springfield, VA. EPA600/2-80056
Barik S, Sethunathan N (1978) Metabolism of \(p\)-nitrophenol in flooded soils. J Environ Qual 7:349–352
Ramakrishnan B, Megharaj M, Venkateswarlu K, Naidu R, Sethunathan N (2010) The impacts of environmental pollutants on microalgae and cyanobacteria. Crit Rev Environ Sci Technol 40:699–821
Ramakrishnan B, Megharaj M, Venkateswarlu K, Sethunathan N, Naidu R (2011) Mixtures of environmental pollutants: effects on microorganisms and their activities. Rev Environ Contam Toxicol 211:63–120
Bruhn C, Lenke H, Hooper DJ (1987) Nitrosubstituted aromatic compounds as nitrogen source for bacteria. Appl Environ Microbiol 7:208–210
Jones SH, Alexander M (1986) Kinetics of mineralization of phenol in lake water. Appl Environ Microbiol 51:891–897
Keith LH (1980) EPA priority pollutants: where they come from, where they are going? AIChE Symp Ser 77:209–249
Spain JC (1995) Biodegradation of nitroaromatic compounds. Annu Rev Microbiol 49:523–555
Spain JC, Gibson DT (1991) Pathway for biodegradation for \(p\)-nitrophenol in Moraxella sp. Appl Environ Microbiol 57:812–819
Kadiyala V, Spain JC (1998) A two-component monooxygenase catalyzes both the hydroxylation of \(p\)-nitrophenol and the oxidative release of nitrite from 4-nitrocatechol in Bacillus sphaericus JS905. Appl Environ Microbiol 64:2479–2484
Perry LL, Zylstra GJ (2007) Cloning of a gene cluster involved in the catabolism of \(p\)-nitrophenol by Arthrobacter sp. strain JS443 and characterization of the \(p\)-nitrophenol monooxygenase. J Bacteriol 189:7563–7572
Zhang S, Sun W, Xu L, Zheng X, Chu X, Tian J, Wu N, Fan Y (2012) Identification of the para-nitrophenol catabolic pathway, and characterization of three enzymes involved in the hydroquinone pathway, in Pseudomonas sp. 1–7. BMC Microbiol 12:27
Liu PP, Zhang JJ, Zhou NY (2010) Characterization of mutagenesis of a two-component monooxygenase involved in para-nitrophenol degradation by an Arthrobacter strain. Int Biodeterior Biodegrad 64:293–299
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search algorithms. Nucleic Acids Res 50:3389–3402
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucl Acids Res 28:235–242
Chenna R, Sugawana H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003) Multiple sequence alignment with the clustal series of programs. Nucl Acid Res 31:3497–3500
Sali AL, Overington JP (1994) Derivation of rules for comparative protein modeling from a database of protein structure alignments. Protein Sci 3:1582–1596
Sali AL, Potterton F, van Vlijmen H, Karplus M (1995) Evaluation of comparative protein modeling by MODELLER. Proteins 23:318–326
Umamaheswari A, Pradhan D, Hemanthkumar M (2010) Virtual screening for potential inhibitors of homology modeled Leptospira interrogans MurD ligase. J Chem Biol 3:175–187
Laskowski RA, Rullmann JA, MacArthur MW, Kaptein R, Thrornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8:477–486
Hooft R, Vriend WWG, Sander C, Abola EE (1996) Errors in protein structure. Nature 38:272
Colovos C, Yeates TO (1993) Verification of protein crystal structures: patterns of non-bonded atomic interactions. Protein Sci 2:1511–1519
Tomii K, Hirokawa T, Motono C (2005) Protein structure prediction using a variety of profile libraries and 3D verification. Proteins 61:114–121
Maestro v9.0 (2009) Schrodinger, LLC, Portland, OR
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47:1739–1749
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49:6177–6196
Priyadarshini V, Pradhan D, Munikumar M, Umamaheswari A, Rajasekhar D Srinivasa, Rao PVLN (2011) Docking and molecular dynamic simulations of Legionella pneumophila MurB reductase for potential inhibitor design. Biochem Anal Biochem. doi:10.4172/2161-1009.1000101
Sandeep S, Priyadarshini V, Pradhan D, Munikumar M, Umamaheswari A (2012) Docking and molecular dynamics simulations studies of human protein kinase catalytic subunit alpha with antagonist. J Clin Sci Res 1:15–23
Lopez-Romero P, Gomez MJ, Gomez-Puertas P, Valencia A (2004) Prediction of functional sites in proteins by evolutionary methods. In: Kamp RM, Calvette JJ, Choli-Papadopoulou T (eds) Principles and practice: methods in proteome and protein analysis. Springer, Berlin, pp 319-340
Umamaheswari A, Muni Kumar M, Pradhan D, Hemanth Kumar M (2011) Docking studies towards exploring antiviral compounds against envelope protein of yellow fever virus. Interdiscip Sci Comput Life Sci 3:64–77
Castrignano T, De Meo PD, Cozzetto D, Talamo IG, Tramontano A (2006) The PMDB protein model database. Nucl Acids Res 34:306–309
Acknowledgments
KM thanks the University Grants Commission, New Delhi, India, for the financial assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kallubai, M., Amineni, U., Mallavarapu, M. et al. In Silico Approach to Support that p-Nitrophenol Monooxygenase from Arthrobacter sp. Strain JS443 Catalyzes the Initial Two Sequential Monooxygenations. Interdiscip Sci Comput Life Sci 7, 157–167 (2015). https://doi.org/10.1007/s12539-015-0018-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-015-0018-x